NLRP12 Inflammasome Expression in the Rat Brain in Response to LPS during Morphine Tolerance
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Morphine and LPS Administration
2.3. RNA Isolation and Preparation of cDNA
2.4. Real-Time PCR Array
2.5. PCR Array Data Analysis
2.6. LINCS Analysis
3. Results
3.1. Expression Profile of Inflammasome-Related Genes Following an LPS Challenge, with and without Morphine Tolerance
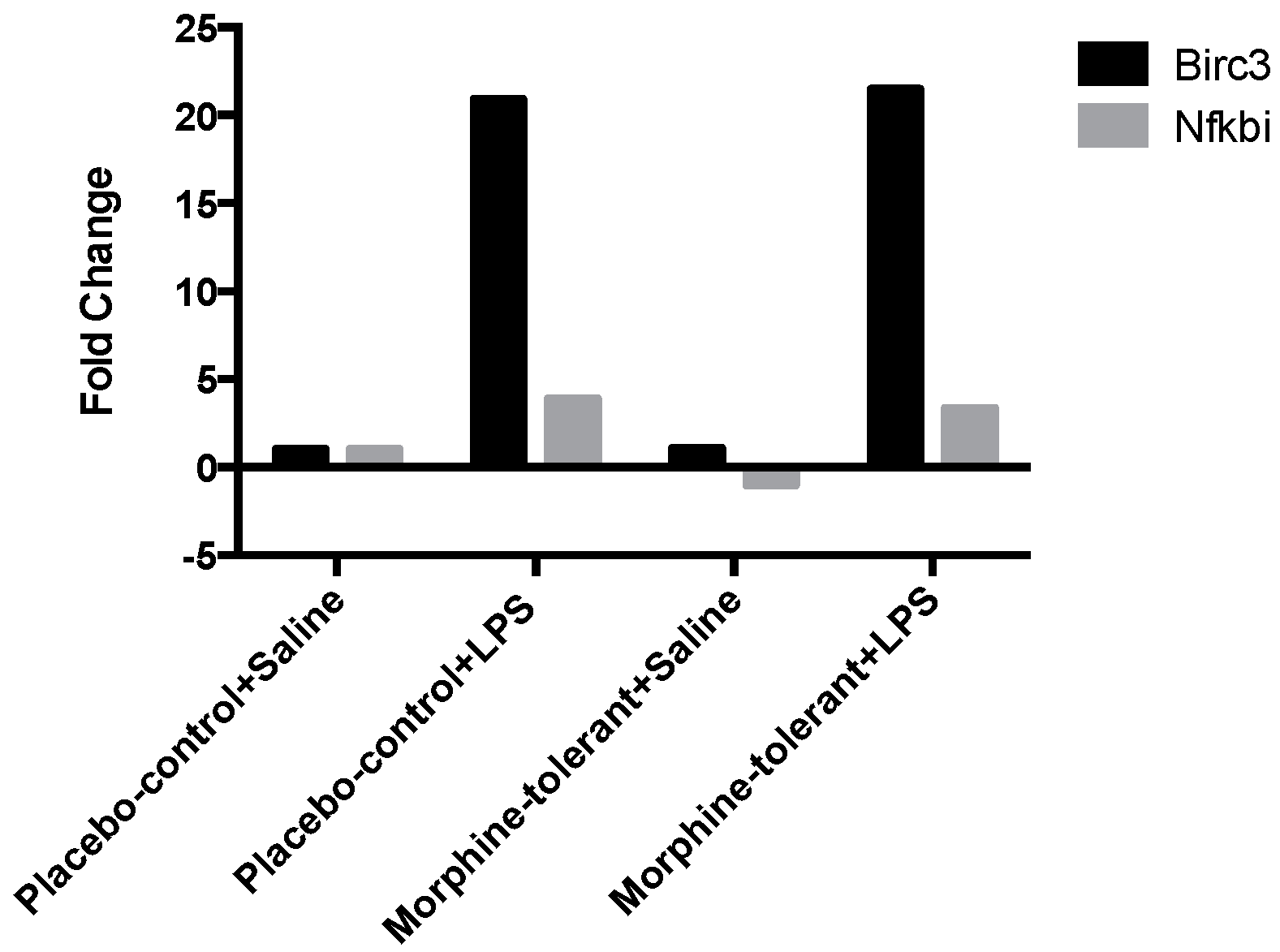
3.2. Expression Profile of Inflammasome-Related Downstream Signaling Genes Following an LPS Challenge, With and Without Morphine Tolerance
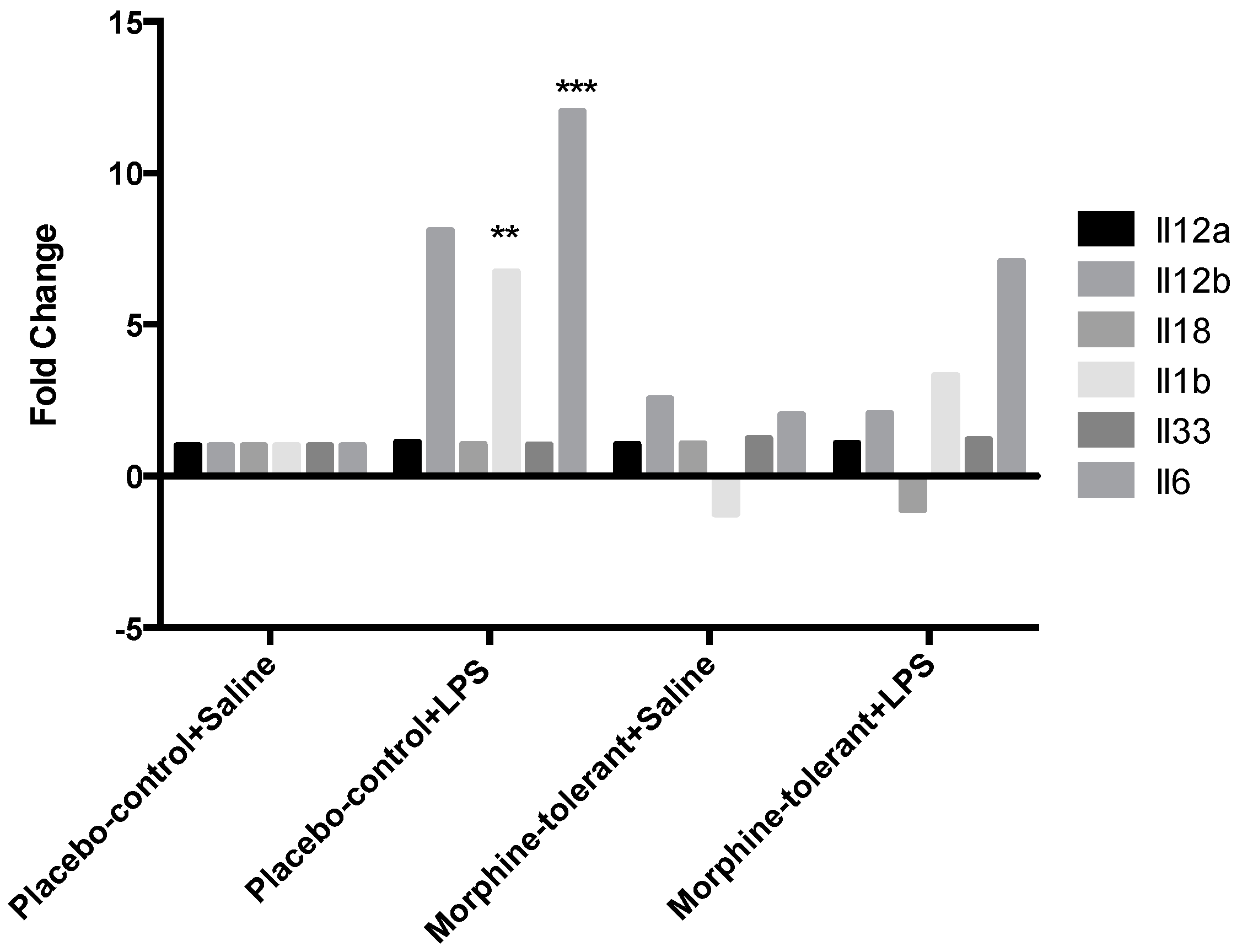
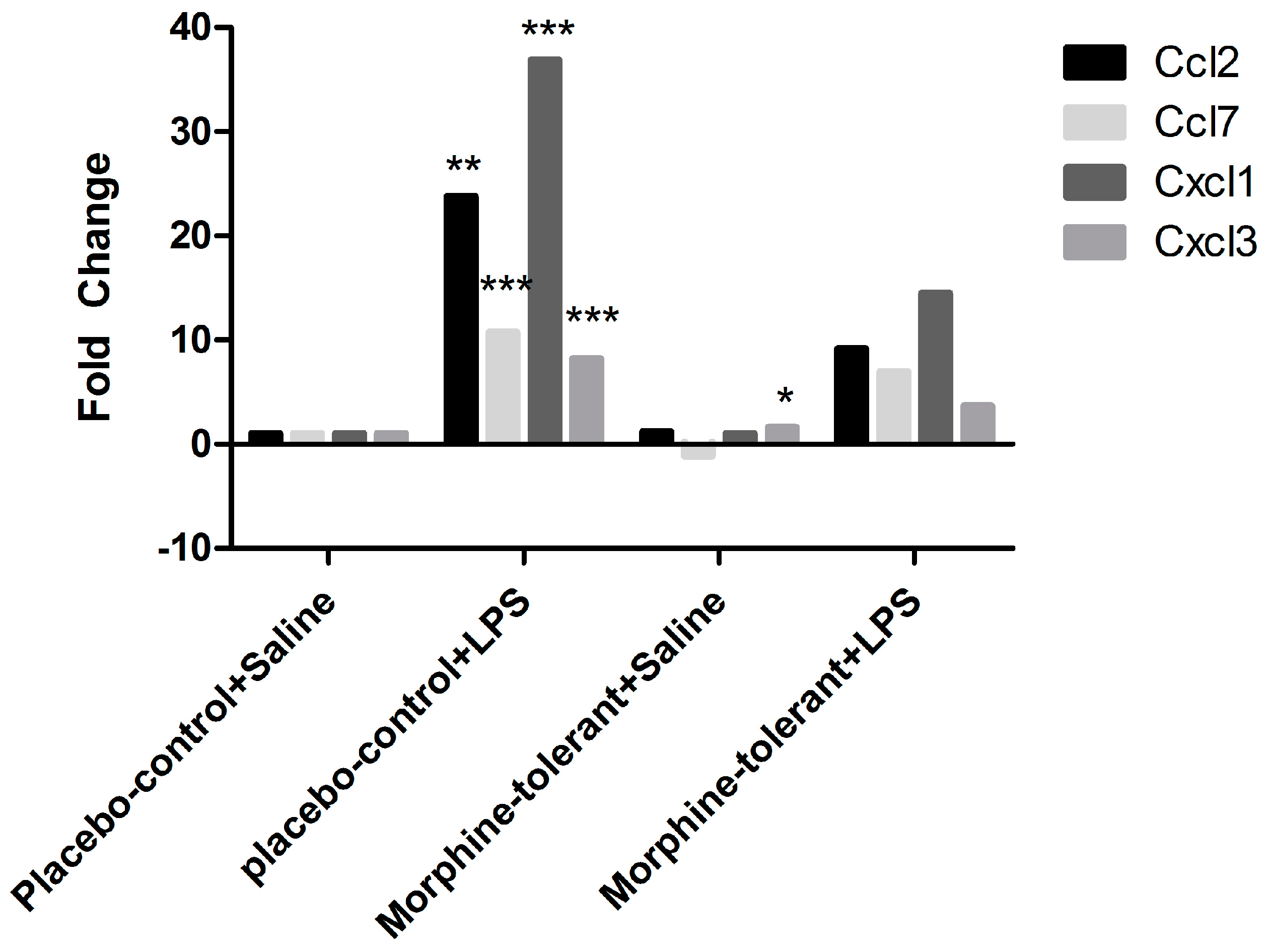
3.3. Expression Profile of Inflammasome-Related Chemokine and Cytokine Genes after an LPS Challenge, with and without Morphine Tolerance
3.4. LINCS Analysis of the Differentially Expressed Genes
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A
| Gene Symbol | Full Name | mRNA Entry |
|---|---|---|
| Card6 | caspase recruitment domain family, member 6 | NM_001106413.1 |
| Casp1 | caspase 1 | NM_012762.2 |
| Casp12 | caspase 12 | NM_130422.1 |
| Casp8 | caspase 8 | NM_022277.1 |
| Naip2 | NLR family, apoptosis inhibitory protein 6 | XM_008760697.2 |
| Nlrp12 | NLR family, pyrin domain containing 12 | NM_001169142.1 |
| Nlrp5 | NLR family, pyrin domain containing 5 | NM_001107474.1 |
| Nlrc4 | NLR family, CARD domain containing 4 | NM_001309432.1 |
| Nlrp1a | NLR family, pyrin domain containing 1A | NM_001145755.2 |
| Nlrp3 | NLR family, pyrin domain containing 3 | NM_001191642.1 |
| Nlrp6 | NLR family, pyrin domain containing 6 | NM_134375.3 |
| Nlrx1 | NLR family member X1 | NM_001025010.1 |
| Nod2 | nucleotide-binding oligomerization domain containing 2 | NM_001106172.1 |
| Pycard | PYD and CARD domain containing | NM_172322.1 |
| Bcl2 | BCL2, apoptosis regulator | NM_016993.1 |
| Bcl2l1 | BCL2 like 1 | NM_001033670.1 |
| Birc2 | baculoviral IAP repeat-containing 2 | NM_021752.2 |
| Birc3 | baculoviral IAP repeat-containing 3 | NM_023987.3 |
| Cflar | CASP8 and FADD-like apoptosis regulator | NM_001033864.2 |
| Chuk | conserved helix-loop-helix ubiquitous kinase | NM_001107588.1 |
| Ciita | class II, major histocompatibility complex, transactivator | NM_001270803.1 |
| Ctsb | cathepsin B | NM_022597.2 |
| Fadd | Fas associated via death domain | NM_152937.2 |
| Hsp90aa1 | heat shock protein 90, alpha (cytosolic), class A member 1 | NM_175761.2 |
| Hsp90ab1 | heat shock protein 90 alpha family class B member 1 | NM_001004082.3 |
| Ikbkb | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta | NM_053355.2 |
| Ikbkg | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma | NM_199103.1 |
| Irak1 | interleukin-1 receptor-associated kinase 1 | NM_001127555.1 |
| Map3k7 | mitogen activated protein kinase kinase kinase 7 | NM_001107920.2 |
| Map3k7ip1 | TGF-beta activated kinase 1/MAP3K7 binding protein 1 | NM_001109976.2 |
| Map3k7ip2 | TGF-beta activated kinase 1/MAP3K7 binding protein 2 | NM_001012062.1 |
| Mapk1 | mitogen activated protein kinase 1 | NM_053842.2 |
| Mapk11 | mitogen-activated protein kinase 11 | NM_001109532.2 |
| Mapk12 | mitogen-activated protein kinase 12 | NM_021746.1 |
| Mapk13 | mitogen activated protein kinase 13 | NM_019231.2 |
| Mapk14 | mitogen activated protein kinase 14 | NM_031020.2 |
| Mapk3 | mitogen activated protein kinase 3 | NM_017347.2 |
| Mapk8 | mitogen-activated protein kinase 8 | NM_053829.2 |
| Mapk9 | mitogen-activated protein kinase 9 | NM_001270544.1 |
| Mefv | Mediterranean fever | NM_031634.1 |
| Myd88 | myeloid differentiation primary response 88 | NM_198130.1 |
| Nfkb1 | nuclear factor kappa B subunit 1 | NM_001276711.1 |
| Nfkbia | NFKB inhibitor alpha | NM_001105720.2 |
| Nfkbib | NFKB inhibitor beta | NM_030867.2 |
| P2rx7 | purinergic receptor P2X 7 | NM_019256.1 |
| Panx1 | Pannexin 1 | NM_001270548.1 |
| Pea15a | phosphoprotein enriched in astrocytes 15 | NM_001013231.1 |
| Pstpip1 | proline-serine-threonine phosphatase-interacting protein 1 | NM_001106824.2 |
| Ptgs2 | prostaglandin-endoperoxide synthase 2 | NM_017232.3 |
| Rage | MOK protein kinase | NM_001010965.1 |
| Rela | RELA proto-oncogene, NF-kB subunit | NM_199267.2 |
| Ripk2 | receptor-interacting serine-threonine kinase 2 | NM_001191865.1 |
| Sugt1 | SGT1 homolog, MIS12 kinetochore complex assembly cochaperone | NM_001013051.1 |
| Tirap | TIR domain containing adaptor protein | XM_017596001.1 |
| Hsp90b1 | heat shock protein 90 beta family member 1 | NM_001012197.2 |
| Traf6 | TNF receptor associated factor 6 | NM_001107754.2 |
| Txnip | thioredoxin interacting protein | NM_001008767.1 |
| Xiap | E3 ubiquitin-protein ligase XIAP | NM_022231.2 |
| Ccl11 | C-C motif chemokine ligand 11 | NM_019205.1 |
| Ccl12 | chemokine (C-C motif) ligand 12 | NM_001105822.1 |
| Ccl2 | C-C motif chemokine ligand 2 | NM_031530.1 |
| Ccl5 | C-C motif chemokine ligand 5 | NM_031116.3 |
| Ccl7 | C-C motif chemokine ligand 7 | NM_001007612.1 |
| Cxcl1 | C-X-C motif chemokine ligand 1 | NM_030845.1 |
| Cxcl3 | C-X-C motif chemokine ligand 3 | NM_138522.1 |
| Cd40lg | CD40 ligand | NM_053353.1 |
| Ifnb1 | interferon beta 1 | NM_019127.1 |
| Ifng | interferon gamma | NM_138880.2 |
| Il12a | interleukin 12A | NM_053390.1 |
| Il12b | interleukin 12B | NM_022611.1 |
| Il18 | interleukin 18 | NM_019165.1 |
| Il1b | interleukin 1 beta | NM_031512.2 |
| Il33 | interleukin 33 | NM_001014166.1 |
| Il6 | interleukin 6 | NM_012589.2 |
| Irf1 | interferon regulatory factor 1 | NM_012591.1 |
| Irf2 | interferon regulatory factor 2 | NM_001047086.1 |
| Irf3 | interferon regulatory factor 3 | NM_001006969.1 |
| Irf4 | interferon regulatory factor 4 | NM_001106108.1 |
| Irf5 | interferon regulatory factor 5 | NM_001106586.1 |
| Irf6 | interferon regulatory factor 6 | NM_001108859.1 |
| Tnfsf11 | tumor necrosis factor superfamily member 11 | NM_057149.1 |
| Tnfsf14 | tumor necrosis factor superfamily member 14 | NM_001191803.1 |
| Tnfsf4 | tumor necrosis factor superfamily member 4 | NM_053552.1 |
| Gene Symbol | Full name |
|---|---|
| VPS28 | VPS28, ESCRT-I subunit |
| PROCR | protein C receptor |
| CHMP2A | charged multivesicular body protein 2A |
| MB | myoglobin |
| ZNF768 | zinc finger protein 768 |
| RBPJ | recombination signal binding protein for immunoglobulin kappa J region |
| WARS2 | tryptophanyl tRNA synthetase 2 (mitochondrial) |
| TBX2 | T-box 2 |
| MRPS2 | mitochondrial ribosomal protein S2 |
| MAP3K14 | mitogen-activated protein kinase kinase kinase 14 |
| SMARCE1 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 |
| AHRR | aryl-hydrocarbon receptor repressor |
| GPX7 | glutathione peroxidase 7 |
| ATP5F1 | ATP synthase, H+ transporting, mitochondrial Fo complex subunit B1 |
| CALR | calreticulin |
| GPR110 | G protein-coupled receptor 110 |
| ELF4 | E74 like ETS transcription factor 4 |
| FGFR1 | fibroblast growth factor receptor 1 |
| F7 | coagulation factor VII |
| AHR | aryl hydrocarbon receptor |
| UBE2L6 | ubiquitin-conjugating enzyme E2L 6 |
| PAFAH1B3 | platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 |
| JUNB | JunB proto-oncogene, AP-1 transcription factor subunit |
| RYK | receptor-like tyrosine kinase |
| ARG1 | arginase 1 |
| ZNF324 | zinc finger protein 324 |
| ATP5D | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit |
References
- Ossipov, M.H.; Lai, J.; King, T.; Vanderah, T.W.; Malan, T.P., Jr.; Hruby, V.J.; Porreca, F. Antinociceptive and nociceptive actions of opioids. J. Neurobiol. 2004, 61, 126–148. [Google Scholar] [CrossRef] [PubMed]
- Bonnet, M.P.; Beloeil, H.; Benhamou, D.; Mazoit, J.X.; Asehnoune, K. The μ opioid receptor mediates morphine-induced tumor necrosis factor and interleukin-6 inhibition in toll-like receptor 2-stimulated monocytes. Anesth. Analg. 2008, 106, 1142–1149. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, M.R.; Coats, B.D.; Lewis, S.S.; Zhang, Y.; Sprunger, D.B.; Rezvani, N.; Baker, E.M.; Jekich, B.M.; Wieseler, J.L.; Somogyi, A.A.; et al. Proinflammatory cytokines oppose opioid-induced acute and chronic analgesia. Brain Behav. Immun. 2008, 22, 1178–1189. [Google Scholar] [CrossRef] [PubMed]
- Mohan, S.; Davis, R.L.; DeSilva, U.; Stevens, C.W. Dual regulation of mu opioid receptors in SK-N-SH neuroblastoma cells by morphine and interleukin-1beta: Evidence for opioid-immune crosstalk. J. Neuroimmunol. 2010, 227, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Ocasio, F.M.; Jiang, Y.; House, S.D.; Chang, S.L. Chronic morphine accelerates the progression of lipopolysaccharide-induced sepsis to septic shock. J. Neuroimmunol. 2004, 149, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Flores, L.R.; Wahl, S.M.; Bayer, B.M. Mechanisms of morphine-induced immunosuppression: Effect of acute morphine administration on lymphocyte trafficking. J. Pharmacol. Exp. Ther. 1995, 272, 1246–1251. [Google Scholar] [PubMed]
- Gomez-Flores, R.; Weber, R.J. Inhibition of interleukin-2 production and downregulation of IL-2 and transferrin receptors on rat splenic lymphocytes following pag morphine administration: A role in natural killer and T cell suppression. J. Interferon Cytokine Res. 1999, 19, 625–630. [Google Scholar] [CrossRef] [PubMed]
- Patel, N.A.; Romero, A.A.; Zadina, J.E.; Chang, S.L. Chronic exposure to morphine attenuates expression of interleukin-1 beta in the rat hippocampus. Brain Res. 1996, 712, 340–344. [Google Scholar] [CrossRef]
- Roy, S.; Charboneau, R.G.; Barke, R.A. Morphine synergizes with lipopolysaccharide in a chronic endotoxemia model. J. Neuroimmunol. 1999, 95, 107–114. [Google Scholar] [CrossRef]
- Hilburger, M.E.; Adler, M.W.; Truant, A.L.; Meissler, J.J., Jr.; Satishchandran, V.; Rogers, T.J.; Eisenstein, T.K. Morphine induces sepsis in mice. J. Infect. Dis. 1997, 176, 183–188. [Google Scholar] [CrossRef] [PubMed]
- Martinon, F.; Burns, K.; Tschopp, J. The inflammasome: A molecular platform triggering activation of inflammatory caspases and processing of proil-beta. Mol. Cell 2002, 10, 417–426. [Google Scholar] [CrossRef]
- Guarda, G.; Zenger, M.; Yazdi, A.S.; Schroder, K.; Ferrero, I.; Menu, P.; Tardivel, A.; Mattmann, C.; Tschopp, J. Differential expression of NLRP3 among hematopoietic cells. J. Immunol. 2011, 186, 2529–2534. [Google Scholar] [CrossRef] [PubMed]
- Martinon, F.; Mayor, A.; Tschopp, J. The inflammasomes: Guardians of the body. Annu. Rev. Immunol. 2009, 27, 229–265. [Google Scholar] [CrossRef] [PubMed]
- Strowig, T.; Henao-Mejia, J.; Elinav, E.; Flavell, R. Inflammasomes in health and disease. Nature 2012, 481, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Allan, S.M.; Tyrrell, P.J.; Rothwell, N.J. Interleukin-1 and neuronal injury. Nat. Rev. Immunol. 2005, 5, 629–640. [Google Scholar] [CrossRef] [PubMed]
- Allen, I.C.; Wilson, J.E.; Schneider, M.; Lich, J.D.; Roberts, R.A.; Arthur, J.C.; Woodford, R.M.; Davis, B.K.; Uronis, J.M.; Herfarth, H.H.; et al. NLRP12 suppresses colon inflammation and tumorigenesis through the negative regulation of noncanonical NF-kappab signaling. Immunity 2012, 36, 742–754. [Google Scholar] [CrossRef] [PubMed]
- Williams, K.L.; Lich, J.D.; Duncan, J.A.; Reed, W.; Rallabhandi, P.; Moore, C.; Kurtz, S.; Coffield, V.M.; Accavitti-Loper, M.A.; Su, L.; et al. The caterpiller protein monarch-1 is an antagonist of toll-like receptor-, tumor necrosis factor alpha-, and mycobacterium tuberculosis-induced pro-inflammatory signals. J. Biol. Chem. 2005, 280, 39914–39924. [Google Scholar] [CrossRef] [PubMed]
- Lich, J.D.; Williams, K.L.; Moore, C.B.; Arthur, J.C.; Davis, B.K.; Taxman, D.J.; Ting, J.P. Monarch-1 suppresses non-canonical NF-kappab activation and P52-dependent chemokine expression in monocytes. J. Immunol. 2007, 178, 1256–1260. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Health (NIH) Library of Network-Based Cellular Signatures (LINCS) Program. Available online: http://www.lincsproject.org/ (accessed on 11 June 2016).
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.P.; Subramanian, A.; Ross, K.N.; et al. The connectivity map: Using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.L.; Moldow, R.L.; House, S.D.; Zadina, J.E. Morphine affects the brain-immune axis by modulating an interleukin-1 beta dependent pathway. Adv. Exp. Med. Biol. 1996, 402, 35–42. [Google Scholar] [PubMed]
- Graf, J.A.; Patel, J.A.; Chang, S.L. Chronic exposure to morphine, but not ethanol, attenuates the expression of interleukin-1 beta converting enzyme in rat spleen. Immunol. Lett. 1997, 58, 153–157. [Google Scholar] [CrossRef]
- Zadina, J.E.; Kastin, A.J.; Harrison, L.M.; Ge, L.J.; Chang, S.L. Opiate receptor changes after chronic exposure to agonists and antagonists. Ann. N. Y. Acad. Sci. 1995, 757, 353–361. [Google Scholar] [CrossRef] [PubMed]
- House, S.D.; Mao, X.; Wu, G.; Espinelli, D.; Li, W.X.; Chang, S.L. Chronic morphine potentiates the inflammatory response by disrupting interleukin-1beta modulation of the hypothalamic-pituitary-adrenal axis. J. Neuroimmunol. 2001, 118, 277–285. [Google Scholar] [CrossRef]
- RT2 Profiler PCR Array Date Analysis version 3.5. Available online: http://pcrdataanalysis.sabiosciences.com/pcr/arrayanalysis.php (accessed 22 January 2017).
- Ma, J.; Malladi, S.; Beck, A.H. Systematic analysis of sex-linked molecular alterations and therapies in cancer. Sci. Rep. 2016, 6, 19119. [Google Scholar] [CrossRef] [PubMed]
- Siavelis, J.C.; Bourdakou, M.M.; Athanasiadis, E.I.; Spyrou, G.M.; Nikita, K.S. Bioinformatics methods in drug repurposing for Alzheimer’s disease. Brief. Bioinform. 2016, 17, 322–335. [Google Scholar] [CrossRef] [PubMed]
- Allen, I.C.; Lich, J.D.; Arthur, J.C.; Jania, C.M.; Roberts, R.A.; Callaway, J.B.; Tilley, S.L.; Ting, J.P. Characterization of NLRP12 during the development of allergic airway disease in mice. PLoS ONE 2012, 7, e30612. [Google Scholar] [CrossRef] [PubMed]
- Dostert, C.; Guarda, G.; Romero, J.F.; Menu, P.; Gross, O.; Tardivel, A.; Suva, M.L.; Stehle, J.C.; Kopf, M.; Stamenkovic, I.; et al. Malarial hemozoin is a NALP3 inflammasome activating danger signal. PLoS ONE 2009, 4, e6510. [Google Scholar] [CrossRef] [PubMed]
- Zaki, M.H.; Vogel, P.; Malireddi, R.K.; Body-Malapel, M.; Anand, P.K.; Bertin, J.; Green, D.R.; Lamkanfi, M.; Kanneganti, T.D. The nod-like receptor NLRP12 attenuates colon inflammation and tumorigenesis. Cancer Cell 2011, 20, 649–660. [Google Scholar] [CrossRef] [PubMed]
- Beug, S.T.; Cheung, H.H.; LaCasse, E.C.; Korneluk, R.G. Modulation of immune signalling by inhibitors of apoptosis. Trends Immunol. 2012, 33, 535–545. [Google Scholar] [CrossRef] [PubMed]
- Scherer, D.C.; Brockman, J.A.; Chen, Z.; Maniatis, T.; Ballard, D.W. Signal-induced degradation of I kappa B alpha requires site-specific ubiquitination. Proc. Natl. Acad. Sci. USA 1995, 92, 11259–11263. [Google Scholar] [CrossRef] [PubMed]
- Homji, N.F.; Mao, X.; Langsdorf, E.F.; Chang, S.L. Endotoxin-induced cytokine and chemokine expression in the HIV-1 transgenic rat. J. Neuroinflamm. 2012, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Del Fresno, C.; Garcia-Rio, F.; Gomez-Pina, V.; Soares-Schanoski, A.; Fernandez-Ruiz, I.; Jurado, T.; Kajiji, T.; Shu, C.; Marin, E.; Gutierrez del Arroyo, A.; et al. Potent phagocytic activity with impaired antigen presentation identifying lipopolysaccharide-tolerant human monocytes: Demonstration in isolated monocytes from cystic fibrosis patients. J. Immunol. 2009, 182, 6494–6507. [Google Scholar] [CrossRef] [PubMed]
- Fang, H.; Pengal, R.A.; Cao, X.; Ganesan, L.P.; Wewers, M.D.; Marsh, C.B.; Tridandapani, S. Lipopolysaccharide-induced macrophage inflammatory response is regulated by ship. J. Immunol. 2004, 173, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Martich, G.D.; Boujoukos, A.J.; Suffredini, A.F. Response of man to endotoxin. Immunobiology 1993, 187, 403–416. [Google Scholar] [CrossRef]
- Porter, K.J.; Gonipeta, B.; Parvataneni, S.; Appledorn, D.M.; Patial, S.; Sharma, D.; Gangur, V.; Amalfitano, A.; Parameswaran, N. Regulation of lipopolysaccharide-induced inflammatory response and endotoxemia by beta-arrestins. J. Cell Physiol. 2010, 225, 406–416. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.; Sarkar, S.; Chang, S.L. Involvement of the NLRP3 inflammasome in the modulation of an LPS-induced inflammatory response during morphine tolerance drug and alcohol dependence. Drug Alcohol Depend. 2013, 132, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Limiroli, E.; Gaspani, L.; Panerai, A.E.; Sacerdote, P. Differential morphine tolerance development in the modulation of macrophage cytokine production in mice. J. Leukoc. Biol. 2002, 72, 43–48. [Google Scholar] [PubMed]

| morphine-tolerant + saline/placebo control + saline | placebo control + LPS/placebo control + saline | morphine-tolerant + LPS/morphine-tolerant + saline | ||||
|---|---|---|---|---|---|---|
| Gene | Fold Change | p-value * | Fold Change ** | p-value * | Fold Change ** | p-value * |
| Card6 | −1.2347 | 0.078926 | −1.5292 | 0.004159 | −1.714 | 0.017499 |
| Casp1 | 1.1505 | 0.299299 | 1.2132 | 0.276593 | 1.2764 | 0.02969 |
| Casp12 | 1.0972 | 0.462347 | 1.5422 | 0.003614 | 1.4249 | 0.092085 |
| Casp8 | −1.0402 | 0.612597 | −1.1363 | 0.391056 | −1.2652 | 0.139999 |
| Naip2 | −1.4483 | 0.067329 | −1.1031 | 0.619799 | −1.2316 | 0.25758 |
| Nlrp12 | −1.6504 | 0.674422 | −7.31 | 0.002731 | −4.5124 | 0.025372 |
| Nlrp5 | 1.494 | 0.054876 | 1.274 | 0.20786 | 1.2819 | 0.027352 |
| Nlrc4 | 1.0028 | 0.915792 | 1.1349 | 0.159712 | 1.1003 | 0.445719 |
| Nlrp1a | −1.0898 | 0.470844 | −1.2027 | 0.001147 | −1.3308 | 0.1379 |
| Nlrp3 | 1.3773 | 0.134647 | 1.5114 | 0.054565 | 1.3956 | 0.02407 |
| Nlrp6 | 1.205 | 0.420019 | −1.6225 | 0.358997 | 1.0293 | 0.784547 |
| Nlrx1 | −1.1574 | 0.24832 | −1.1494 | 0.191026 | 1.039 | 0.679959 |
| Nod2 | −1.6257 | 0.650152 | 1.0901 | 0.811946 | 1.2826 | 0.50395 |
| Pycard | −1.0575 | 0.610143 | −1.0285 | 0.797363 | −1.2363 | 0.030115 |
 : 6–10 fold decrease;
: 6–10 fold decrease;  : 2–5 fold decrease;
: 2–5 fold decrease;  : <2 fold;
: <2 fold;  : 2–5 fold increase;
: 2–5 fold increase;  : 6–10 fold increase;
: 6–10 fold increase;  : 11–30 fold increase;
: 11–30 fold increase;  : 31–50 fold increase.
: 31–50 fold increase.| morphine-tolerant + saline/placebo control + saline | placebo control + LPS/placebo control + saline | morphine-tolerant + LPS/morphine-tolerant + saline | ||||
|---|---|---|---|---|---|---|
| Gene | Fold Change | p-value * | Fold Change ** | p-value * | Fold Change ** | p-value * |
| Bcl2 | 1.0469 | 0.863233 | 1.0586 | 0.845254 | 1.2398 | 0.339246 |
| Bcl2l1 | −1.1256 | 0.01111 | 1.0572 | 0.476643 | −1.0291 | 0.671184 |
| Birc2 | 1.0509 | 0.586337 | −1.0281 | 0.622487 | 1.0699 | 0.531417 |
| Birc3 | 1.0648 | 0.587513 | 20.9356 | 0.011065 | 21.5025 | 0.001873 |
| Cflar | 1.0226 | 0.697346 | 1.3021 | 0.004748 | 1.2155 | 0.140978 |
| Chuk | 1.1867 | 0.119108 | −1.0233 | 0.754891 | 1.1593 | 0.073058 |
| Ciita | −1.0617 | 0.750879 | 1.0619 | 0.873363 | 1.4678 | 0.39646 |
| Ctsb | −1.0009 | 0.978499 | −1.0656 | 0.617885 | −1.1046 | 0.493204 |
| Fadd | −1.1261 | 0.258884 | 1.1235 | 0.252486 | 1.0306 | 0.71174 |
| Hsp90aa1 | 1.2774 | 0.114349 | 1.0391 | 0.743375 | 1.189 | 0.149024 |
| Hsp90ab1 | 1.0752 | 0.376876 | −1.0968 | 0.305427 | 1.1579 | 0.18515 |
| Ikbkb | 1.0718 | 0.550623 | −1.0011 | 0.962992 | 1.1799 | 0.201955 |
| Ikbkg | 1.0924 | 0.324904 | −1.0402 | 0.492725 | 1.1419 | 0.079202 |
| Irak1 | 1.1549 | 0.062648 | 1.0287 | 0.755395 | 1.2213 | 0.242955 |
| Map3k7 | −1.2495 | 0.002404 | −1.1008 | 0.434596 | −1.2497 | 0.114056 |
| Map3k7ip1 | −1.0335 | 0.664499 | −1.1151 | 0.369226 | 1.02 | 0.864431 |
| Map3k7ip2 | −1.2869 | 0.000867 | −1.1169 | 0.456355 | −1.3427 | 0.007785 |
| Mapk1 | −1.0527 | 0.468581 | −1.0887 | 0.159097 | 1.0316 | 0.724595 |
| Mapk11 | −1.0673 | 0.530192 | −1.0706 | 0.578016 | −1.0633 | 0.571223 |
| Mapk12 | 1.1424 | 0.420684 | 1.1889 | 0.372287 | −1.0255 | 0.780819 |
| Mapk13 | −1.1629 | 0.924422 | −1.5423 | 0.355826 | 1.0504 | 0.651078 |
| Mapk14 | 1.0018 | 0.940725 | −1.0318 | 0.81795 | 1.0335 | 0.709962 |
| Mapk3 | −1.1294 | 0.224192 | −1.121 | 0.372958 | −1.0495 | 0.76638 |
| Mapk8 | 1.0383 | 0.883566 | −1.0215 | 0.798267 | 1.0037 | 0.911563 |
| Mapk9 | −1.1252 | 0.03886 | −1.0075 | 0.87718 | −1.0805 | 0.288977 |
| Mefv | 1.2572 | 0.278807 | 1.2582 | 0.463496 | 1.9192 | 0.028283 |
| Myd88 | 1.0211 | 0.791137 | 1.0103 | 0.917416 | 1.2185 | 0.03802 |
| Nfkb1 | 1.1098 | 0.279116 | 1.2919 | 0.089881 | 1.3323 | 0.033357 |
| Nfkbia | −1.0294 | 0.835021 | 3.885 | 0.001215 | 3.3744 | 0.001348 |
| Nfkbib | −1.1473 | 0.330475 | −1.0153 | 0.762037 | −1.1167 | 0.643504 |
| P2rx7 | −1.1586 | 0.428492 | 1.0742 | 0.625549 | 1.0578 | 0.661519 |
| Panx1 | −1.076 | 0.275575 | 1.1455 | 0.406863 | 1.1101 | 0.334734 |
| Pea15a | −1.0943 | 0.291531 | −1.043 | 0.625202 | −1.0802 | 0.492135 |
| Pstpip1 | −1.1041 | 0.527175 | −1.1018 | 0.680263 | −1.0232 | 0.87342 |
| Ptgs2 | −1.08 | 0.499364 | 1.5126 | 0.000571 | 1.1762 | 0.323717 |
| Rage | 1.0729 | 0.478448 | −1.1273 | 0.17423 | 1.1854 | 0.16574 |
| Rela | 1.0171 | 0.804744 | 1.1801 | 0.417806 | 1.2504 | 0.106174 |
| Ripk2 | −1.1057 | 0.237762 | 1.2913 | 0.028006 | 1.1199 | 0.389288 |
| Sugt1 | 1.0063 | 0.918273 | −1.1023 | 0.519396 | −1.1264 | 0.106262 |
| Tirap | −1.3098 | 0.094434 | −1.0971 | 0.535721 | 1.0809 | 0.374146 |
| Hsp90b1 | 1.0475 | 0.680071 | −1.0334 | 0.69801 | −1.0877 | 0.314466 |
| Traf6 | −1.0344 | 0.915204 | −1.0171 | 0.943304 | 1.0988 | 0.437323 |
| Txnip | −1.0761 | 0.70951 | −1.0718 | 0.699702 | −1.0187 | 0.789181 |
| Xiap | 1.0572 | 0.503462 | −1.0558 | 0.471524 | 1.0089 | 0.93573 |
 : 6–10 fold decrease;
: 6–10 fold decrease;  : 2–5 fold decrease;
: 2–5 fold decrease;  : <2 fold;
: <2 fold;  : 2–5 fold increase;
: 2–5 fold increase;  : 6–10 fold increase;
: 6–10 fold increase;  : 11–30 fold increase;
: 11–30 fold increase;  : 31–50 fold increase.
: 31–50 fold increase.| morphine-tolerant + saline/placebo control + saline | placebo control + LPS/placebo control + saline | morphine-tolerant + LPS/morphine-tolerant + saline | ||||
|---|---|---|---|---|---|---|
| Gene | Fold Change ** | p-value * | Fold Change ** | p-value * | Fold Change ** | p-value * |
| Ccl11 | −2.4662 | 0.023104 | −2.8639 | 0.16576 | −2.7346 | 0.249229 |
| Ccl12 | −1.4362 | 0.646529 | 2.0214 | 0.219015 | 2.1108 | 0.399119 |
| Ccl2 | 1.1756 | 0.521412 | 23.7845 | 0.001354 | 9.1497 | 0.106064 |
| Ccl5 | −1.105 | 0.472434 | 1.2909 | 0.290174 | −1.0949 | 0.766893 |
| Ccl7 | −1.0841 | 0.688167 | 10.7188 | 0.000259 | 6.9528 | 0.163034 |
| Cxcl1 | 1.0131 | 0.944854 | 36.8609 | 0 | 14.4718 | 0.130008 |
| Cxcl3 | 1.6208 | 0.019673 | 8.1909 | 0.000076 | 3.6477 | 0.170486 |
| Cd40lg | −2.2796 | 0.215753 | 1.1119 | 0.704126 | −2.4054 | 0.273798 |
| Ifnb1 | 1.2519 | 0.936419 | 1.6899 | 0.391302 | 1.5695 | 0.513801 |
| Ifng | 1.5355 | 0.290413 | 1.1698 | 0.767882 | 2.2615 | 0.088346 |
| Il12a | 1.05 | 0.57557 | 1.1178 | 0.33855 | 1.0837 | 0.456191 |
| Il12b | 2.5587 | 0.26828 | 8.1055 | 0.102341 | 2.077 | 0.198782 |
| Il18 | 1.068 | 0.60015 | 1.0484 | 0.707002 | −1.1151 | 0.48359 |
| Il1b | −1.2476 | 0.284501 | 6.7366 | 0.001383 | 3.3136 | 0.089312 |
| Il33 | 1.2342 | 0.054615 | 1.0407 | 0.583823 | 1.1978 | 0.072613 |
| Il6 | 2.023 | 0.45937 | 12.0303 | 0.008058 | 7.0943 | 0.083969 |
| Irf1 | 1.1299 | 0.521156 | 3.4575 | 0.000386 | 3.4104 | 0.01276 |
| Irf2 | −1.1481 | 0.050782 | 1.0027 | 0.916481 | 1.0556 | 0.593809 |
| Irf3 | −1.2824 | 0.024657 | −1.3225 | 0.225481 | −1.2478 | 0.088625 |
| Irf4 | −1.1491 | 0.43182 | −1.0557 | 0.846866 | 1.0263 | 0.858979 |
| Irf5 | −1.1024 | 0.38296 | −1.1468 | 0.376953 | 1.0391 | 0.841777 |
| Irf6 | 1.1055 | 0.413463 | 1.012 | 0.792025 | 1.118 | 0.272222 |
| Tnfsf11 | −1.6077 | 0.800779 | −1.362 | 0.519436 | −1.3266 | 0.965379 |
| Tnfsf14 | −1.396 | 0.243156 | −1.8445 | 0.126355 | −1.4278 | 0.629476 |
| Tnfsf4 | −1.0175 | 0.997398 | −1.0887 | 0.790199 | −1.3422 | 0.420711 |
 : 6–10 fold decrease;
: 6–10 fold decrease;  : 2–5 fold decrease;
: 2–5 fold decrease;  : <2 fold;
: <2 fold;  : 2–5 fold increase;
: 2–5 fold increase;  : 6–10 fold increase;
: 6–10 fold increase;  : 11–30 fold increase;
: 11–30 fold increase;  : 31–50 fold increase.
: 31–50 fold increase.| (A) | Rank | placebo-Saline vs. Placebo-LPS | Placebo-Saline vs. Morphine-Tolerant-Saline | Morphine-Tolerant-Saline vs. Morphine-Tolerant-LPS |
| 1 | VPS28 | SMARCE1 | AHR | |
| 2 | PROCR | AHRR | UBE2L6 | |
| 3 | CHMP2A | GPX7 | PAFAH1B3 | |
| 4 | MB | ATP5F1 | VPS28 | |
| 5 | ZNF768 | CALR | JUNB | |
| 6 | RBPJ | GPR110 | RYK | |
| 7 | WARS2 | CHMP2A | ARG1 | |
| 8 | TBX2 | ELF4 | PROC | |
| 9 | MRPS2 | FGFR1 | ZNF324 | |
| 10 | MAP3K14 | F7 | ATP5D | |
| (B) | Gene Rank | Placebo-Saline vs. Placebo-LPS | Placebo-Saline vs. Morphine-Tolerant-Saline | Morphine-Tolerant-Saline vs. Morphine-Tolerant-LPS |
| VPS28 | 1 | 42 | 4 | |
| PROCR | 2 | 491 | 681 | |
| CHMP2A | 3 | 7 | 177 | |
| SMARCE1-1 | 675 | 1 | 504 | |
| AHRR | 2383 | 2 | 460 | |
| GPX7 | 488 | 3 | 2137 | |
| AHR | 46 | 769 | 1 | |
| UBE2L6 | 89 | 545 | 2 | |
| PAFAH1B3 | 95 | 622 | 3 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, S.L.; Huang, W.; Mao, X.; Sarkar, S. NLRP12 Inflammasome Expression in the Rat Brain in Response to LPS during Morphine Tolerance. Brain Sci. 2017, 7, 14. https://doi.org/10.3390/brainsci7020014
Chang SL, Huang W, Mao X, Sarkar S. NLRP12 Inflammasome Expression in the Rat Brain in Response to LPS during Morphine Tolerance. Brain Sciences. 2017; 7(2):14. https://doi.org/10.3390/brainsci7020014
Chicago/Turabian StyleChang, Sulie L., Wenfei Huang, Xin Mao, and Sabroni Sarkar. 2017. "NLRP12 Inflammasome Expression in the Rat Brain in Response to LPS during Morphine Tolerance" Brain Sciences 7, no. 2: 14. https://doi.org/10.3390/brainsci7020014
APA StyleChang, S. L., Huang, W., Mao, X., & Sarkar, S. (2017). NLRP12 Inflammasome Expression in the Rat Brain in Response to LPS during Morphine Tolerance. Brain Sciences, 7(2), 14. https://doi.org/10.3390/brainsci7020014





