Investigating PRDM8 DNA Methylation in Peripheral Tissues in Borderline Personality Disorder: Association with Symptom Severity but Not Adverse Childhood Experiences
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Cohort
2.2. Measures
2.2.1. Questionnaires
2.2.2. PRDM8 DNAm Analysis in Saliva and Whole Blood
2.3. Data Analyses
2.3.1. Demographic and Clinical Data
2.3.2. Statistical Analyses
3. Results
3.1. Demographic and Clinical Information
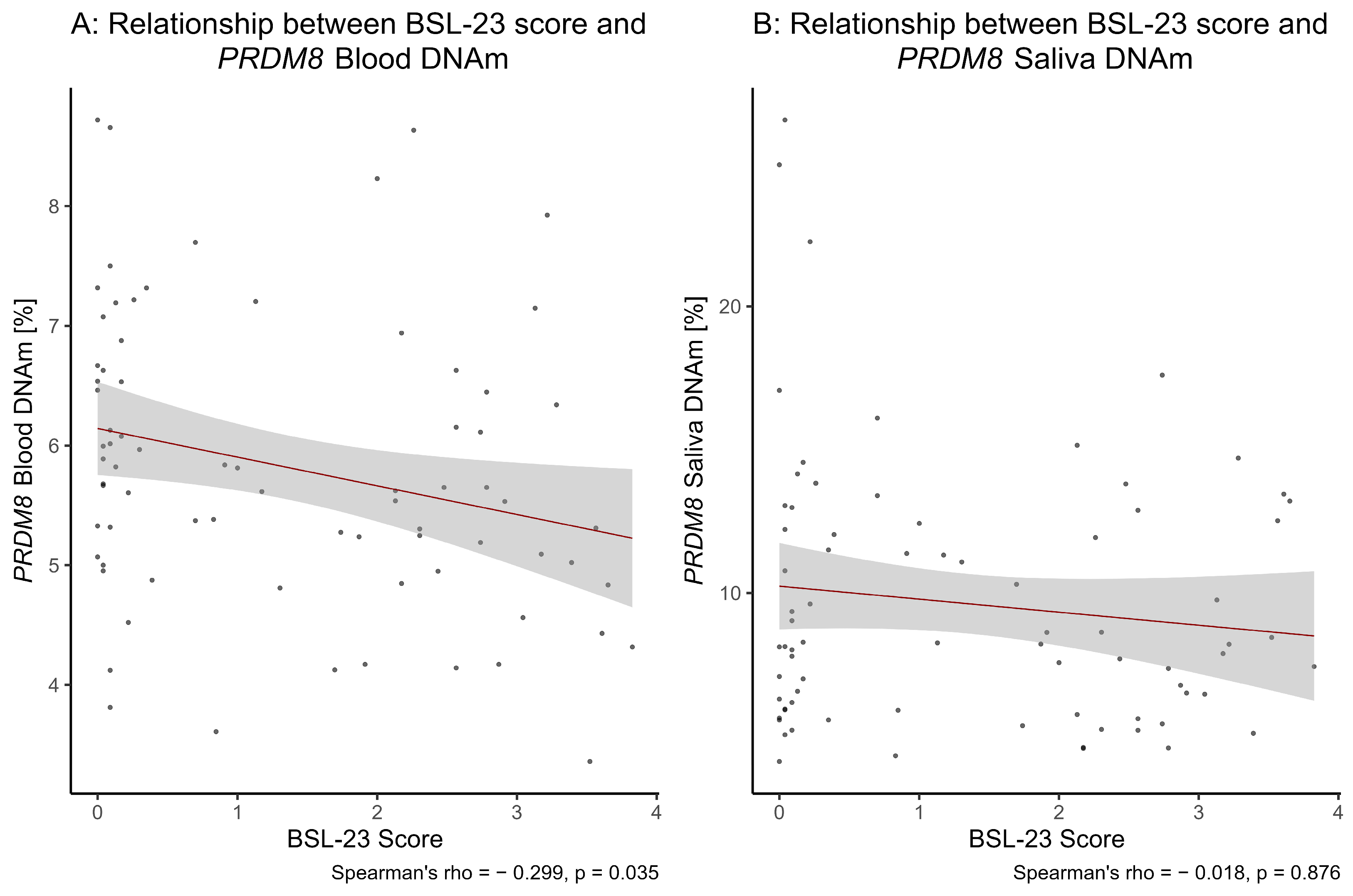
3.2. Severity of Borderline Symptomatology
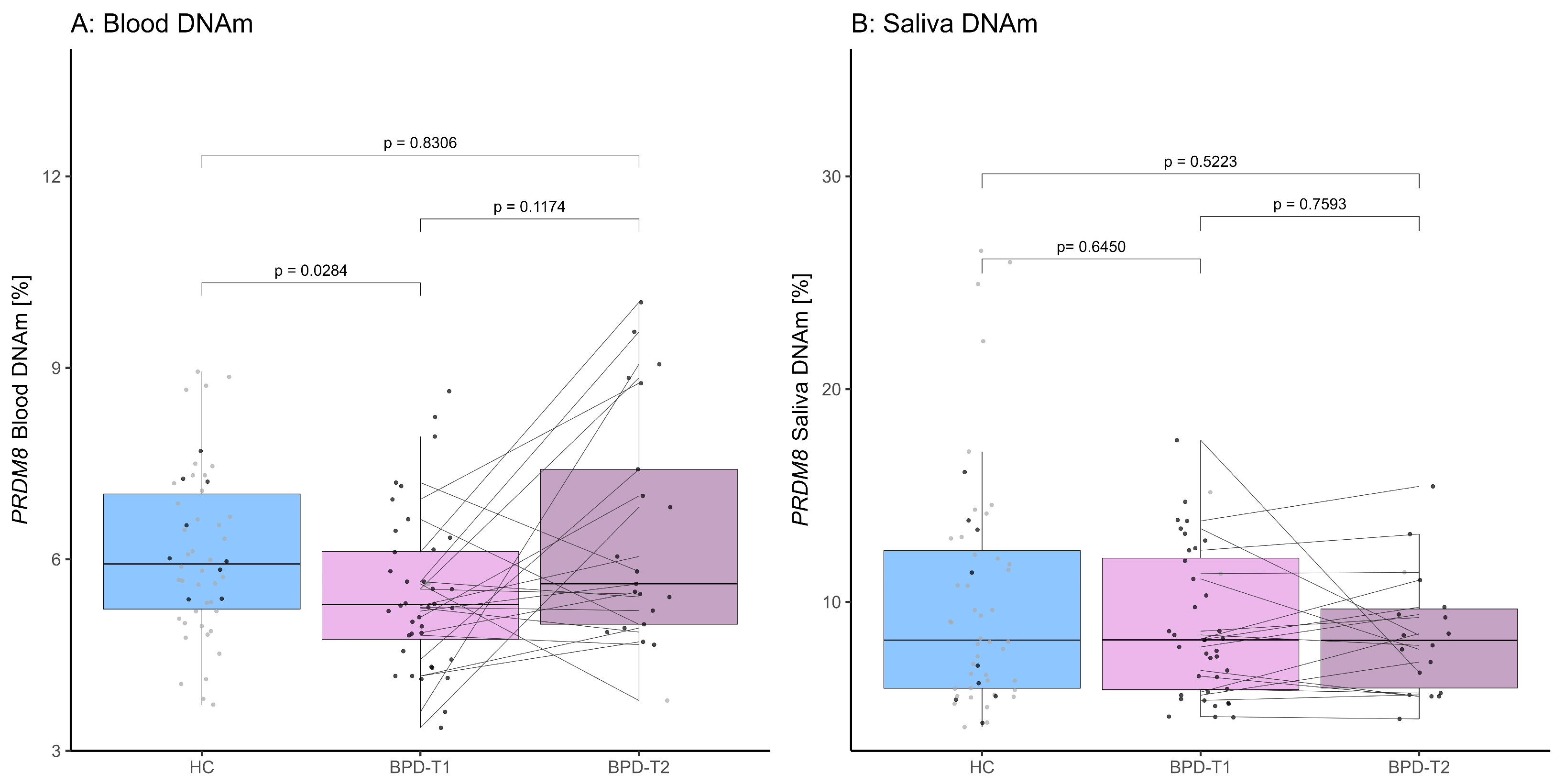
3.3. DNAm Levels of PRDM8 in BPD Patients and HC Individuals
3.4. PRDM8 DNAm Levels in the Context of ACE
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BPD | Borderline Personality Disorder |
| ACE | Adverse Childhood Experience(s) |
| PRDM8 | PR domain zinc finger protein 8 (gene) |
| DBT | Dialectic Behavioral Therapy |
| DNAm | DNA Methylation |
| H3K9 | Lysine 9 of histone H3 |
| HC | Healthy Control |
| T1 | Pre-therapy |
| T2 | Post-therapy |
| IPDE | International Personality Disorder Examination |
| DSM-IV | Diagnostic and Statistical Manual of Mental Disorders, fourth edition |
| CTQ | Childhood Trauma Questionnaire |
| BSL-23 | Borderline Symptom List |
| EDTA | Ethylenediaminetetraacetic (tube) |
| IQR | Interquartile range |
References
- American Psychiatric Association. Association, Diagnostic and Statistical Manual of Mental Disorders: DSM-IV; American Psychiatric Association: Washington, DC, USA, 1994; Volume 4. [Google Scholar]
- Biskin, R.S. The Lifetime Course of Borderline Personality Disorder. Can. J. Psychiatry 2015, 60, 303–308. [Google Scholar] [CrossRef]
- Yen, S.; Shea, M.T.; Battle, C.L.; Johnson, D.M.; Zlotnick, C.; Dolan-Sewell, R.; Skodol, A.E.; Grilo, C.M.; Gunderson, J.G.; Sanislow, C.A.; et al. Traumatic exposure and posttraumatic stress disorder in borderline, schizotypal, avoidant, and obsessive-compulsive personality disorders: Fingings from the collaborative longitudinal personality disorders study. J. Nerv. Ment. Dis. 2002, 190, 510–518. [Google Scholar] [CrossRef]
- Grant, B.F.; Chou, S.P.; Goldstein, R.B.; Huang, B.; Stinson, F.S.; Saha, T.D.; Smith, S.M.; Dawson, D.A.; Pulay, A.J.; Pickering, R.P. Prevalence, correlates, disability, and comorbidity of DSM-IV borderline personality disorder: Results from the Wave 2 National Epidemiologic Survey on Alcohol and Related Conditions. J. Clin. Psychiatry 2008, 69, 533. [Google Scholar] [CrossRef] [PubMed]
- Skodol, A.E.; Gunderson, J.G.; Pfohl, B.; Widiger, T.A.; Livesley, W.J.; Siever, L.J. The borderline diagnosis I: Psychopathology, comorbidity, and personaltity structure. Biol. Psychiatry 2002, 51, 936–950. [Google Scholar] [CrossRef] [PubMed]
- Gescher, D.M.; Kahl, K.G.; Hillemacher, T.; Frieling, H.; Kuhn, J.; Frodl, T. Epigenetics in personality disorders: Today’s insights. Front. Psychiatry 2018, 9, 579. [Google Scholar] [CrossRef]
- Lieb, K.; Zanarini, M.C.; Schmahl, C.; Linehan, M.M.; Bohus, M. Borderline personality disorder. Lancet 2004, 364, 453–461. [Google Scholar] [CrossRef]
- Gunderson, J.G.; Herpertz, S.C.; Skodol, A.E.; Torgersen, S.; Zanarini, M.C. Borderline personality disorder. Nat. Rev. Dis. Primers 2018, 4, 18029. [Google Scholar] [CrossRef]
- Keverne, J.; Binder, E.B. A Review of epigenetics in psychiatry: Focus on environmental risk factors. Med. Genet. 2020, 32, 57–64. [Google Scholar] [CrossRef]
- Dammann, G.; Teschler, S.; Haag, T.; Altmüller, F.; Tuczek, F.; Dammann, R.H. Increased DNA methylation of neuropsychiatric genes occurs in borderline personality disorder. Epigenetics 2011, 6, 1454–1462. [Google Scholar] [CrossRef]
- Arranz, M.J.; Gallego-Fabrega, C.; Martín-Blanco, A.; Soler, J.; Elices, M.; Dominguez-Clavé, E.; Salazar, J.; Vega, D.; Briones-Buixassa, L.; Pascual, J.C. A genome-wide methylation study reveals X chromosome and childhood trauma methylation alterations associated with borderline personality disorder. Transl. Psychiatry 2021, 11, 5. [Google Scholar] [CrossRef]
- Yang, Z.; Zhang, S.; Ouyang, L.; Liao, A.; He, Y.; Li, Z.; Chen, X. DNA methylation and bipolar disorder. J. Psychiatry Brain Sci. 2023, 8, e230012. [Google Scholar] [CrossRef]
- Dall’Aglio, L.; Rijlaarsdam, J.; Mulder, R.H.; Neumann, A.; Felix, J.F.; Kok, R.; Bakermans-Kranenburg, M.J.; van Ijzendoorn, M.H.; Tiemeier, H.; Cecil, C.A.M. Epigenome-wide associations between observed maternal sensitivity and offspring DNA methylation: A population-based prospective study in children. Psychol. Med. 2022, 52, 2481–2491. [Google Scholar] [CrossRef] [PubMed]
- Knoblich, N.; Gundel, F.; Brückmann, C.; Becker-Sadzio, J.; Frischholz, C.; Nieratschker, V. DNA methylation of APBA3 and MCF2 in borderline personality disorder: Potential biomarkers for response to psychotherapy. Eur. Neuropsychopharmacol. 2018, 28, 252–263. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.; Knoblich, N.; Wallisch, A.; Glowacz, K.; Becker-Sadzio, J.; Gundel, F.; Brückmann, C.; Nieratschker, V. Increased BDNF methylation in saliva, but not blood, of patients with borderline personality disorder. Clin. Epigenetics 2018, 10, 109. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.; Banet, N.; Wallisch, A.; Glowacz, K.; Becker-Sadzio, J.; Gundel, F.; Nieratschker, V. Differential COMT DNA methylation in patients with borderline personality disorder: Genotype matters. Eur. Neuropsychopharmacol. 2019, 29, 1295–1300. [Google Scholar] [CrossRef]
- Meaney, M.J.; Szyf, M. Environmental programming of stress responses through DNA methylation: Life at the interface between a dynamic environment and a fixed genome. Dialogues Clin. Neurosci. 2005, 7, 103–123. [Google Scholar] [CrossRef]
- Szyf, M. The early-life social environment and DNA methylation. Clin. Genet. 2012, 81, 341–349. [Google Scholar] [CrossRef]
- Gladish, N.; Merrill, S.; Kobor, M.S. Childhood trauma and epigenetics: State of the science and future. Curr. Environ. Health Rep. 2022, 9, 661–672. [Google Scholar] [CrossRef]
- Mainali, P.; Rai, T.; Rutkofsky, I.H. From child abuse to developing borderline personality disorder into adulthood: Exploring the neuromorphological and epigenetic pathway. Cureus 2020, 12, e9474. [Google Scholar] [CrossRef]
- Estric, C.; Calati, R.; Lopez-Castroman, J. Adverse childhood experiences and neurocognition in borderline personality disorder: A call-to-action perspective review. Harv. Rev. Psychiatry 2022, 30, 248–260. [Google Scholar] [CrossRef]
- Porter, C.; Palmier-Claus, J.; Branitsky, A.; Mansell, W.; Warwick, H.; Varese, F. Childhood adversity and borderline personality disorder: A meta-analysis. Acta Psychiatr. Scand. 2020, 141, 6–20. [Google Scholar] [CrossRef]
- Wiegand, A.; Kreifelts, B.; Munk, M.H.J.; Geiselhart, N.; Ramadori, K.E.; MacIsaac, J.L.; Fallgatter, A.J.; Kobor, M.S.; Nieratschker, V. DNA methylation differences associated with social anxiety disorder and early life adversity. Transl. Psychiatry 2021, 11, 104. [Google Scholar] [CrossRef]
- Edelmann, S.; Balaji, J.; Pasche, S.; Wiegand, A.; Nieratschker, V. DNA Methylation of PXDN is Associated with Early-Life Adversity in Adult Mental Disorders. Biomolecules 2024, 14, 976. [Google Scholar] [CrossRef]
- Megala, J.; Sivakumar, D.; Jha, D.; Kundu, S.; Arora, K.; Gayathri, V. Epigenetic modifications due to childhood trauma causative of potential mental and physical disorders. Int. J. Nutr. Pharmacol. Neurol. Dis. 2021, 11, 41–49. [Google Scholar] [CrossRef]
- Short, A.K.; Weber, R.; Kamei, N.; Wilcox Thai, C.; Arora, H.; Mortazavi, A.; Stern, H.S.; Glynn, L.; Baram, T.Z. Individual longitudinal changes in DNA-methylome identify signatures of early-life adversity and correlate with later outcome. Neurobiol. Stress 2024, 31, 100652. [Google Scholar] [CrossRef]
- Edelmann, S.; Wiegand, A.; Hentrich, T.; Pasche, S.; Schulze-Hentrich, J.M.; Munk, M.H.J.; Fallgatter, A.J.; Kreifelts, B.; Nieratschker, V. Blood transcriptome analysis suggests an indirect molecular association of early life adversities and adult social anxiety disorder by immune-related signal transduction. Front. Psychiatry 2023, 14, 1125553. [Google Scholar] [CrossRef]
- Rivera, L.M.; Uwizeye, G.; Stolrow, H.; Christensen, B.; Rutherford, J.; Thayer, Z. Prenatal exposure to genocide and subsequent adverse childhood events are associated with DNA methylation of SLC6A4, BDNF, and PRDM8 in early adulthood in Rwanda. Sci. Rep. 2024, 14, 27879. [Google Scholar] [CrossRef]
- Musanabaganwa, C.; Wani, A.H.; Donglasan, J.; Fatumo, S.; Jansen, S.; Mutabaruka, J.; Rutembesa, E.; Uwineza, A.; Hermans, E.J.; Roozendaal, B.; et al. Leukocyte methylomic imprints of exposure to the genocide against the Tutsi in Rwanda: A pilot epigenome-wide analysis. Epigenomics 2022, 14, 11–25. [Google Scholar] [CrossRef]
- Eom, G.H.; Kim, K.; Kim, S.-M.; Kee, H.J.; Kim, J.-Y.; Jin, H.M.; Kim, J.-R.; Kim, J.H.; Choe, N.; Kim, K.-B.; et al. Histone methyltransferase PRDM8 regulates mouse testis steroidogenesis. Biochem. Biophys. Res. Commun. 2009, 388, 131–136. [Google Scholar] [CrossRef]
- Ross, S.E.; McCord, A.E.; Jung, C.; Atan, D.; Mok, S.I.; Hemberg, M.; Kim, T.-K.; Salogiannis, J.; Hu, L.; Cohen, S.; et al. Bhlhb5 and Prdm8 form a repressor complex involved in neuronal circuit assembly. Neuron 2012, 73, 292–303. [Google Scholar] [CrossRef]
- Orouji, E.; Peitsch, W.K.; Orouji, A.; Houben, R.; Utikal, J. Unique role of histone methyltransferase PRDM8 in the tumorigenesis of virus-negative merkel cell carcinoma. Cancers 2020, 12, 1057. [Google Scholar] [CrossRef]
- Komai, T.; Iwanari, H.; Mochizuki, Y.; Hamakubo, T.; Shinkai, Y. Expression of the mouse PR domain protein Prdm8 in the developing central nervous system. Gene Expr. Patterns 2009, 9, 503–514. [Google Scholar] [CrossRef]
- Kinameri, E.; Inoue, T.; Aruga, J.; Imayoshi, I.; Kageyama, R.; Shimogori, T.; Moore, A.W. Prdm proto-oncogene transcription factor family expression and interaction with the Notch-Hes pathway in mouse neurogenesis. PLoS ONE 2008, 3, e3859. [Google Scholar] [CrossRef]
- Inoue, M.; Iwai, R.; Yamanishi, E.; Yamagata, K.; Komabayashi-Suzuki, M.; Honda, A.; Komai, T.; Miyachi, H.; Kitano, S.; Watanabe, C.; et al. Deletion of Prdm8 impairs development of upper-layer neocortical neurons. Genes Cells 2015, 20, 758–770. [Google Scholar] [CrossRef]
- Tu, W.J.; Hardy, K.; Sutton, C.R.; McCuaig, R.; Li, J.; Dunn, J.; Tan, A.; Brezar, V.; Morris, M.; Denyer, G.; et al. Priming of transcriptional memory responses via the chromatin accessibility landscape in T cells. Sci. Rep. 2017, 7, 44825. [Google Scholar] [CrossRef]
- Davarzani, A.; Shahrokhi, A.; Hashemi, S.S.; Ghasemi, A.; Habibi Kavashkohei, M.R.; Farboodi, N.; Lang, A.E.; Ghiasi, M.; Rohani, M.; Alavi, A. The second family affected with a PRDM8-related disease. Neurol. Sci. 2022, 43, 3847–3855. [Google Scholar] [CrossRef]
- Turnbull, J.; Girard, J.-M.; Lohi, H.; Chan, E.M.; Wang, P.; Tiberia, E.; Omer, S.; Ahmed, M.; Bennett, C.; Chakrabarty, A.; et al. Early-onset Lafora body disease. Brain 2012, 135, 2684–2698. [Google Scholar] [CrossRef]
- Linehan, M. Cognitive-Behavioral Treatment of Borderline Personality Disorder; Guilford Press: New York, NY, USA, 1993. [Google Scholar]
- Panos, P.T.; Jackson, J.W.; Hasan, O.; Panos, A. Meta-analysis and systematic review assessing the efficacy of dialectical behavior therapy (DBT). Res. Soc. Work. Pract. 2014, 24, 213–223. [Google Scholar] [CrossRef]
- Hernandez-Bustamante, M.; Cjuno, J.; Hernández, R.M.; Ponce-Meza, J.C. Efficacy of dialectical behavior therapy in the treatment of borderline personality disorder: A systematic review of randomized controlled trials. Iran. J. Psychiatry 2024, 19, 119. [Google Scholar] [CrossRef]
- VI confirman den Bosch, L.M.C.; Koeter, M.W.J.; Stijnen, T.; Verheul, R.; van den Brink, W. Sustained efficacy of dialectical behaviour therapy for borderline personality disorder. Behav. Res. Ther. 2005, 43, 1231–1241. [Google Scholar] [CrossRef]
- Bohus, M.; Haaf, B.; Simms, T.; Limberger, M.F.; Schmahl, C.; Unckel, C.; Lieb, K.; Linehan, M.M. Effectiveness of inpatient dialectical behavioral therapy for borderline personality disorder: A controlled trial. Behav. Res. Ther. 2004, 42, 487–499. [Google Scholar] [CrossRef]
- Perroud, N.; Salzmann, A.; Prada, P.; Nicastro, R.; Hoeppli, M.-E.; Furrer, S.; Ardu, S.; Krejci, I.; Karege, F.; Malafosse, A. Response to psychotherapy in borderline personality disorder and methylation status of the BDNF gene. Transl. Psychiatry 2013, 3, e207. [Google Scholar] [CrossRef]
- Quevedo, Y.; Booij, L.; Herrera, L.; Hernández, C.; Jiménez, J.P. Potential epigenetic mechanisms in psychotherapy: A pilot study on DNA methylation and mentalization change in borderline personality disorder. Front. Hum. Neurosci. 2022, 16, 955005. [Google Scholar] [CrossRef]
- Loranger, A.W.; Janca, A.; Sartorius, N. Assessment and Diagnosis of Personality Disorders: The ICD-10 International Personality Disorder Examination (IPDE); Cambridge University Press: Cambridge, UK, 1997. [Google Scholar]
- Bernstein, D.P.; Stein, J.A.; Newcomb, M.D.; Walker, E.; Pogge, D.; Ahluvalia, T.; Stokes, J.; Handelsman, L.; Medrano, M.; Desmond, D.; et al. Development and validation of a brief screening version of the Childhood Trauma Questionnaire. Child Abus. Negl. 2003, 27, 169–190. [Google Scholar] [CrossRef]
- Wingenfeld, K.; Spitzer, C.; Mensebach, C.; Grabe, H.J.; Hill, A.; Gast, U.; Schlosser, N.; Höpp, H.; Beblo, T.; Driessen, M. The German version of the Childhood Trauma Questionnaire (CTQ): Preliminary psychometric properties. Psychother. Psychosom. Med. Psychol. 2010, 60, 442–450. [Google Scholar] [CrossRef]
- Wolf, M.; Limberger, M.F.; Kleindienst, N.; Stieglitz, R.D.; Domsalla, M.; Philipsen, A.; Steil, R.; Bohus, M. Short version of the borderline symptom list (BSL-23): Development and psychometric evaluation. Psychother. Psychosom. Med. Psychol. 2009, 59, 321–324. [Google Scholar] [CrossRef]
- Lapaire, O.; Johnson, K.L.; Bianchi, D.W. Method for extraction of high-quantity and-quality cell-free DNA from amniotic fluid. Prenat. Diagn. 2008, 444, 303–309. [Google Scholar]
- Tagliaferro, S.S.; Zejnelagic, A.; Farrugia, R.; Bezzina Wettinger, S. Comparison of DNA extraction methods for samples from old blood collections. Biotechniques 2021, 70, 243–250. [Google Scholar] [CrossRef]
- Leontiou, C.A.; Hadjidaniel, M.D.; Mina, P.; Antoniou, P.; Ioannides, M.; Patsalis, P.C. Bisulfite conversion of DNA: Performance comparison of different kits and methylation quantitation of epigenetic biomarkers that have the potential to be used in non-invasive prenatal testing. PLoS ONE 2015, 10, e0135058. [Google Scholar] [CrossRef]
- Rotondo, J.C.; Borghi, A.; Selvatici, R.; Magri, E.; Bianchini, E.; Montinari, E.; Corazza, M.; Virgili, A.; Tognon, M.; Martini, F. Hypermethylation-induced inactivation of the IRF6 gene as a possible early event in progression of vulvar squamous cell carcinoma associated with lichen sclerosus. JAMA Dermatol. 2016, 152, 928–933. [Google Scholar] [CrossRef]
- Kampmann, M.-L.; Fleckhaus, J.; Børsting, C.; Jurtikova, H.; Piters, A.; Papin, J.; Gauthier, Q.; Ghemrawi, M.; Doutremepuich, C.; McCord, B.; et al. Collaborative exercise: Analysis of age estimation using a QIAGEN protocol and the PyroMark Q48 platform. Forensic Sci. Res. 2024, 9, owad055. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Braun, P.R.; Han, S.; Hing, B.; Nagahama, Y.; Gaul, L.N.; Heinzman, J.T.; Grossbach, A.J.; Close, L.; Dlouhy, B.J.; Howard, M.A., III; et al. Genome-wide DNA methylation comparison between live human brain and peripheral tissues within individuals. Transl. Psychiatry 2019, 9, 47. [Google Scholar] [CrossRef]
- Bakulski, K.M.; Halladay, A.; Hu, V.W.; Mill, J.; Fallin, M.D. Epigenetic research in neuropsychiatric disorders: The “tissue issue”. Curr. Behav. Neurosci. Rep. 2016, 3, 264–274. [Google Scholar] [CrossRef]
- Kumsta, R. The role of epigenetics for understanding mental health difficulties and its implications for psychotherapy research. Psychol. Psychother. Theory Res. Pract. 2019, 92, 190–207. [Google Scholar] [CrossRef]
- Schiele, M.A.; Gottschalk, M.G.; Domschke, K. The applied implications of epigenetics in anxiety, affective and stress-related disorders-A review and synthesis on psychosocial stress, psychotherapy and prevention. Clin. Psychol. Rev. 2020, 77, 101830. [Google Scholar] [CrossRef] [PubMed]
- Lak, M.; Shakiba, S.; Dolatshahi, B.; Saatchi, M.; Shahrbaf, M.; Jafarpour, A. The prevalence of suicide ideation, suicide attempt and suicide in borderline personality disorder patients: A systematic review and meta-analysis. Gen. Hosp. Psychiatry 2025, 95, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Ng, F.Y.; Bourke, M.E.; Grenyer, B.F. Recovery from borderline personality disorder: A systematic review of the perspectives of consumers, clinicians, family and carers. PLoS ONE 2016, 11, e0160515. [Google Scholar] [CrossRef]
- Alameda, L.; Trotta, G.; Quigley, H.; Rodriguez, V.; Gadelrab, R.; Dwir, D.; Dempster, E.; Wong, C.C.Y.; Di Forti, M. Can epigenetics shine a light on the biological pathways underlying major mental disorders? Psychol. Med. 2022, 52, 1645–1665. [Google Scholar] [CrossRef] [PubMed]
- Hohenauer, T.; Moore, A.W. The Prdm family: Expanding roles in stem cells and development. Development 2012, 139, 2267–2282. [Google Scholar] [CrossRef]
- Weidner, C.I.; Lin, Q.; Birkhofer, C.; Gerstenmaier, U.; Kaifie, A.; Kirschner, M.; Bruns, H.; Balabanov, S.; Trummer, A.; Stockklausner, C.; et al. DNA methylation in PRDM8 is indicative for dyskeratosis congenita. Oncotarget 2016, 7, 10765. [Google Scholar] [CrossRef]
- Marrocco, J.; Gray, J.D.; Kogan, J.F.; Einhorn, N.R.; O’Cinneide, E.M.; Rubin, T.G.; Carroll, T.S.; Schmidt, E.F.; McEwen, B.S. Early life stress restricts translational reactivity in CA3 neurons associated with altered stress responses in adulthood. Front. Behav. Neurosci. 2019, 13, 157. [Google Scholar] [CrossRef]
- Nagy, C.; Torres-Platas, S.G.; Mechawar, N.; Turecki, G. Repression of astrocytic connexins in cortical and subcortical brain regions and prefrontal enrichment of H3K9me3 in depression and suicide. Int. J. Neuropsychopharmacol. 2017, 20, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Cypris, O.; Eipel, M.; Franzen, J.; Rösseler, C.; Tharmapalan, V.; Kuo, C.-C.; Vieri, M.; Nikolić, M.; Kirschner, M.; Brümmendorf, T.H.; et al. PRDM8 reveals aberrant DNA methylation in aging syndromes and is relevant for hematopoietic and neuronal differentiation. Clin. Epigenetics 2020, 12, 125. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhang, J.; Huang, S.; He, X. Genome-wide analysis reveals that exon methylation facilitates its selective usage in the human transcriptome. Brief. Bioinform. 2018, 19, 754–764. [Google Scholar] [CrossRef] [PubMed]
- Shayevitch, R.; Askayo, D.; Keydar, I.; Ast, G. The importance of DNA methylation of exons on alternative splicing. RNA 2018, 24, 1351–1362. [Google Scholar] [CrossRef]
- Jones, P.A. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 2012, 13, 484–492. [Google Scholar] [CrossRef]
- Rijlaarsdam, J.; Pappa, I.; Walton, E.; Bakermans-Kranenburg, M.J.; Mileva-Seitz, V.R.; Rippe, R.C.A.; Roza, S.J.; Jaddoe, V.W.V.; Verhulst, F.C.; Felix, J.F.; et al. An epigenome-wide association meta-analysis of prenatal maternal stress in neonates: A model approach for replication. Epigenetics 2016, 11, 140–149. [Google Scholar] [CrossRef]
| Variable | HC | BPD | p (W) |
|---|---|---|---|
| n = 53 | n = 40 | ||
| Age [years] median ± IQR | 26.00 ± 9.00 | 28.00 ± 10.25 | 0.1266 (1257.00) |
| Sex female [%] | 46 [86.79] | 33 [82.50] | 0.5751 * |
| ACE high [%] | 10 [18.87] | 38 [95.00] | < 0.0001 * |
| Emotional abuse score median ± IQR | 7.00 ± 3.00 | 17.00 ± 7.25 | <0.0001 (1898.00) |
| Emotional neglect score median ± IQR | 8.00 ± 4.00 | 17.00 ± 8.25 | <0.0001 (1830.00) |
| Physical abuse score median ± IQR | 5.00 ± 1.00 | 9.50 ± 8.50 | <0.0001 (1713.50) |
| Physical neglect score median ± IQR | 5.00 ± 2.00 | 9.50 ± 6.25 | <0.0001 (1717.50) |
| Sexual abuse score median ± IQR | 5.00 ± 0.00 | 10.00 ± 8.50 | <0.0001 (1658.50) |
| CTQ total score median ± IQR | 32.00 ± 9.00 | 59.00 ± 25.25 | <0.0001 (1996.50) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bender, A.; Bertele, L.; Musaoglu, M.N.; Pasche, S.; Edelmann, S.; Nieratschker, V. Investigating PRDM8 DNA Methylation in Peripheral Tissues in Borderline Personality Disorder: Association with Symptom Severity but Not Adverse Childhood Experiences. Brain Sci. 2025, 15, 950. https://doi.org/10.3390/brainsci15090950
Bender A, Bertele L, Musaoglu MN, Pasche S, Edelmann S, Nieratschker V. Investigating PRDM8 DNA Methylation in Peripheral Tissues in Borderline Personality Disorder: Association with Symptom Severity but Not Adverse Childhood Experiences. Brain Sciences. 2025; 15(9):950. https://doi.org/10.3390/brainsci15090950
Chicago/Turabian StyleBender, Annika, Laila Bertele, Mirac Nur Musaoglu, Sarah Pasche, Susanne Edelmann, and Vanessa Nieratschker. 2025. "Investigating PRDM8 DNA Methylation in Peripheral Tissues in Borderline Personality Disorder: Association with Symptom Severity but Not Adverse Childhood Experiences" Brain Sciences 15, no. 9: 950. https://doi.org/10.3390/brainsci15090950
APA StyleBender, A., Bertele, L., Musaoglu, M. N., Pasche, S., Edelmann, S., & Nieratschker, V. (2025). Investigating PRDM8 DNA Methylation in Peripheral Tissues in Borderline Personality Disorder: Association with Symptom Severity but Not Adverse Childhood Experiences. Brain Sciences, 15(9), 950. https://doi.org/10.3390/brainsci15090950







