1. Introduction
Epilepsy is a chronic neurological disorder whose only known cure is surgery to resect the brain’s seizure onset zone (SOZ, which is often not entirely or exactly removed; the part of the brain actually removed during surgery is known as the resection zone—RZ). However, such surgery requires determination of the RZ and SOZ via a lengthy, inconvenient period of invasive iEEG (intracranial EEG) monitoring. This is because neurologists and neurosurgeons must wait for an ictal event (e.g., a seizure) in order to accurately identify the iEEG electrodes that report abnormal activity and plan for resection surgery. Armed with iEEG recordings of multiple ictal events, physicians inspect these data by eye, which is itself an arduous process. Instead, we look to use interictal iEEG recordings as a basis for surgical planning. If interictal (i.e., everyday activity not from a seizure) iEEG data alone could be used to determine the SOZ, then the SOZ determination phase could be theoretically reduced from at least a week to a matter of hours, which is the time it takes to implant electrodes, run the algorithm, and take out electrodes. It is important to note that this reduction in resection planning time is theoretical and dependent on maintaining a rigorous planning process. In any case, since the current standard of care involves inspection of EEG data streams by the naked eye, this project aims to develop an algorithm that standardizes this process from case to case, further saving physician time and boosting accuracy of SOZ prediction.
The problem of the algorithmic seizure onset zone detection using only interictal data has been approached from a wide variety of angles, apart from GC. Perhaps the most developed is the use of interictal high-frequency oscillations (80–500 Hz; HFOs) derived from iEEG and scalp EEG recordings. HFOs have demonstrated potential in their ability to serve as a biomarker for the seizure onset zone [
1,
2,
3,
4]. Localized HFOs have also been detected from ictal sEEG recordings, supporting the case for the accuracy of HFOs in SOZ detection [
5]. However, despite recent advances in algorithmic HFO detection, the standard protocol remains physician determination of HFO through visual inspection of EEG data [
3,
6]. Furthermore, the HFO methods require processing data in the frequency domain, which is more computationally intensive than the more recent GC time-domain method [
1]. Because of our motivations to cut down on the time needed for SOZ determination, eliminating computational steps and automating the entire process are important for the clinically viability of any SOZ determination methods. Thus, this study builds upon the GC approach, which require neither physician inspection of EEG data nor time-intensive frequency domain processing.
Others have applied variants of the Granger Causality algorithm or graph algorithms that we study to seizure onset detection. Whereas we look at pairs of nodes in our analysis, studies have looked at triplets to determine whether inclusion of a third node enhances the information shared between two other nodes in the same network, called synergistic effects [
7]. These have had some success, finding that synergistic effects in the immediate pre-ictal period correspond to nodes in the SOZ [
8]. Combining this triplet approach with our pairwise one is a potential future area of study. Still others have applied frequency-domain GC in analysis of pre-ictal data and ranked eigenvector centrality in analysis of ictal data to determine the SOZ [
9,
10]. However, none of these studies investigated the relationship between baseline interictal data and ictal data to predict the SOZ.
Still others have adopted network modeling techniques for interictal prediction of the SOZ. These studies have also utilized EEG recordings of interictal patient brain activity [
11,
12]. Demonstrating the difficulty of developing SOZ identification tools that generalize across a larger cohort of patients, one such study found success in identifying robust chimera state markers in interictal recordings for one patient out of a cohort of 15 patients [
11]. We present similarly generalizable SOZ identification algorithms for all epilepsy cases with a much higher success rates. However, due to the diverse nature of epilepsy onset in different regions of the brain and from differing etiologies, identifying a specific subset of clinical cases for which a SOZ identification tool is most useful will likely improve the future usefulness of the tools analyzed both in this study and others. Another study, also based on interictal EEG recordings, interestingly develops a predictive tool to simulate results of resection of any node in a brain’s network [
12]. Their study differs from ours in that a significant portion of patients (7 of 16) did not have successful surgical outcomes [
12]. Without a definitive source of truth (i.e., neurologist-identified SOZ via the current standard of care), assessing the accuracy of a novel SOZ identification tool becomes necessarily speculative. Thus, another contribution of our study is a robust cohort of 24 patients who underwent resection surgery. All 24 had at least a worthwhile improvement (Engel class III or above), and 22 were free of disabling seizures (Engel class I). Nevertheless, in the cited study, even amongst the nine cases with successful surgical outcomes, the predicted SOZ inconsistently matched the actual SOZ [
12], mirroring the difficulties we encountered in our own study. Larger patient cohorts created through a combination of data across studies could prove fruitful for homing in on subsets of patients for which each SOZ/RZ prediction tool offers accurate predictions. We have included data on demographic information, clinical background, and seizure localization for the cohort of patients in this study in
Tables S1 and S2.
Specifically, we study electrocorticography (ECoG) recordings, a form of intracranial EEG (iEEG) monitoring where a skull flap is removed and a grid of electrodes is placed directly onto the brain’s surface. Our population of 25 patients had an average of 102.56 grid electrodes (often referred to as “nodes”) implanted for invasive monitoring. The neurologist-determined SOZ contained an average of 19.62 nodes and the RZ contained an average of 21.88 nodes. One of the 25 patients did not undergo resection surgery after invasive monitoring, resulting in 24 patients with an actual RZ. Investigations of other iEEG recording types (including stereotactic EEG, or sEEG) as they pertain to epilepsy surgery will be the likely subject of later studies by our group.
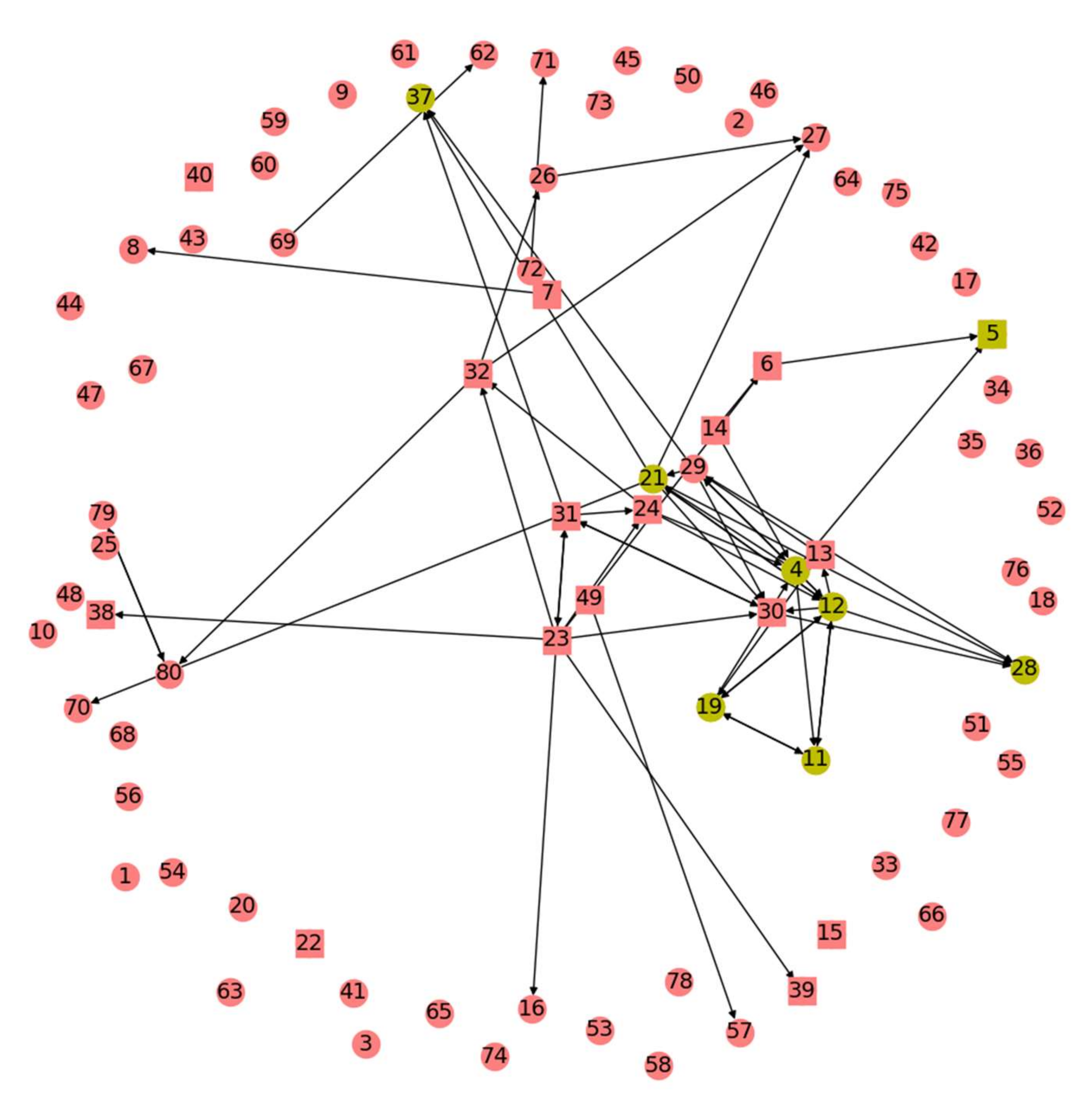
We build off the previous result that time domain Granger Causality (GC) analysis of baseline interictal iEEG data indicates which electrodes seem to be more influential for activity at a network of electrodes [
1]. The retrospective study found that brain regions surrounding these interictally causal electrodes matched the regions chosen for resection through traditional analysis of ictal data with an aggregate probability much smaller than chance (
p < 10
−20) [
1]. GC analyses assign a metric for every pair of electrodes to measure the degree to which activity at one electrode dictates activity at another. This creates a graph of the patient’s brain, also known as a GC map. From the GC map, summation of all the GC values originating at each electrode results in computation of a total GC outdegree metric for that electrode. Nodes are then ranked by their total GC outdegree and the ranks of nodes belonging to the actual SOZ and RZ are summed to create a rank order sum metric. Via comparison of this sum to a random distribution of rank order sums of the same size subset from the same starting number of ranks, it is shown that total GC outdegree results in a statistically significant rank order sum of actual SOZ and RZ nodes with an aggregate probability across all 25 patients much lower than predicted by chance [
1]. However, because the total GC outdegree metric was previously applied without evaluation of alternatives, we start with the GC map and aim to create a more precise algorithm that informs selection of a RZ/SOZ. For ease of comparison, we also use the rank order sum method to evaluate algorithms’ predictive capability. It is important to point out that the Granger statistical approach demonstrates that a particular signal in one channel statistically follows a signal in a different channel. This does not in fact prove that one event is causal, but the term ‘Granger Causality’ is widely used as the name of the statistic. We will keep the name for clarity with this caveat, understanding that the current study begins with the connectivity data derived and reported earlier [
1].
This study comprises three angles from which the above problem is tackled. These were motivated by numerous reasons, chief among them a desire to explore methods that built upon previous GC techniques without adding significant algorithm runtime as well as attempts to naturally mimic brain signals traveling through the network of the GC map. First is a Monte Carlo sampling approach that visits nodes with probabilities weighted by the GC map. Second is an application of Google’s PageRank algorithm where nodes take the place of webpages and are assigned a metric of importance relative to other nodes. Third are centrality algorithms that aim to tease out the subset of nodes most important for transmission of information through the brain’s network. Variants of each algorithm are developed, fine-tuning their application to the problem of RZ/SOZ determination and to the goal of revealing connectivity patterns in an epileptic patient’s brain.
3. Results
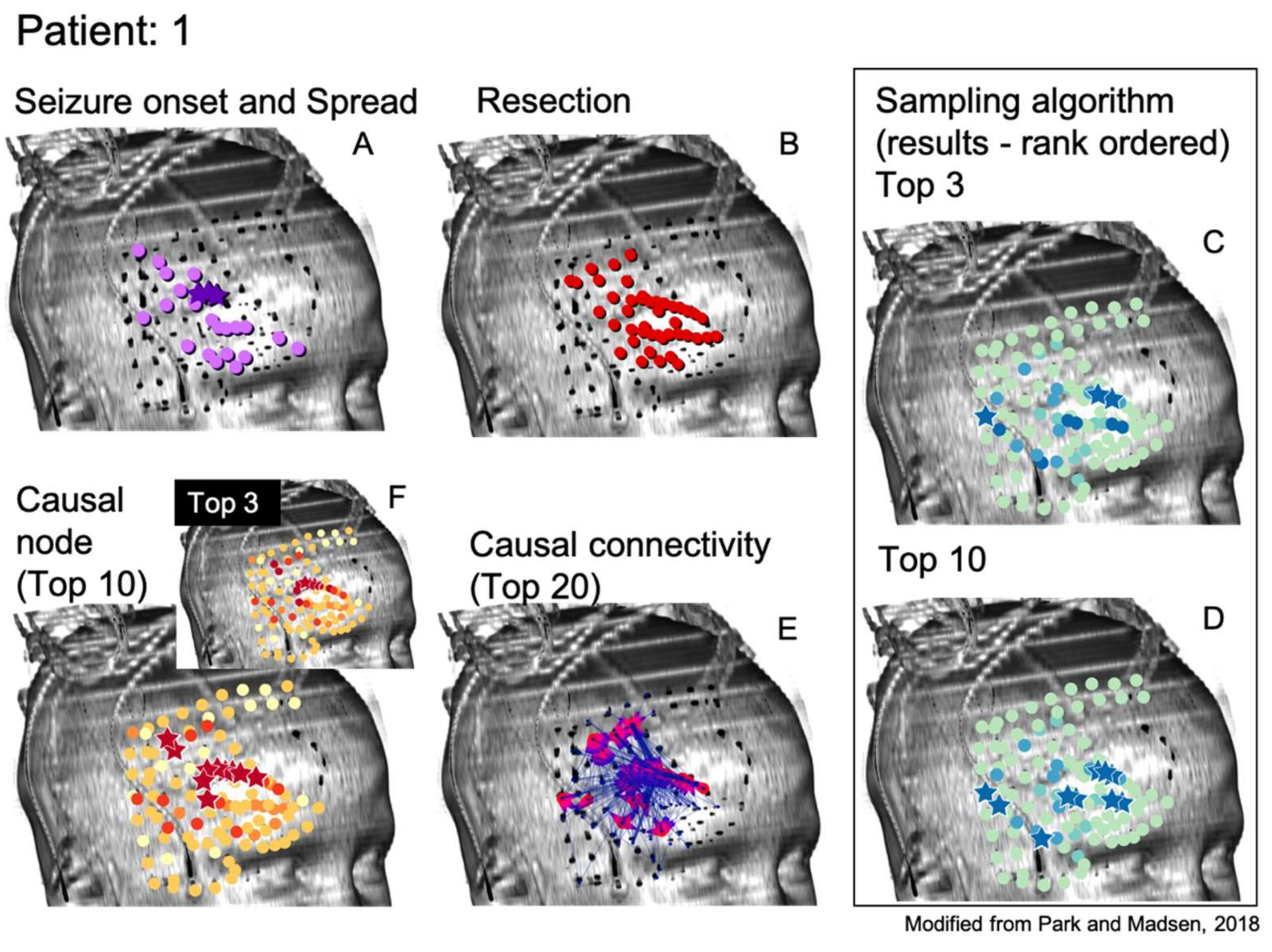
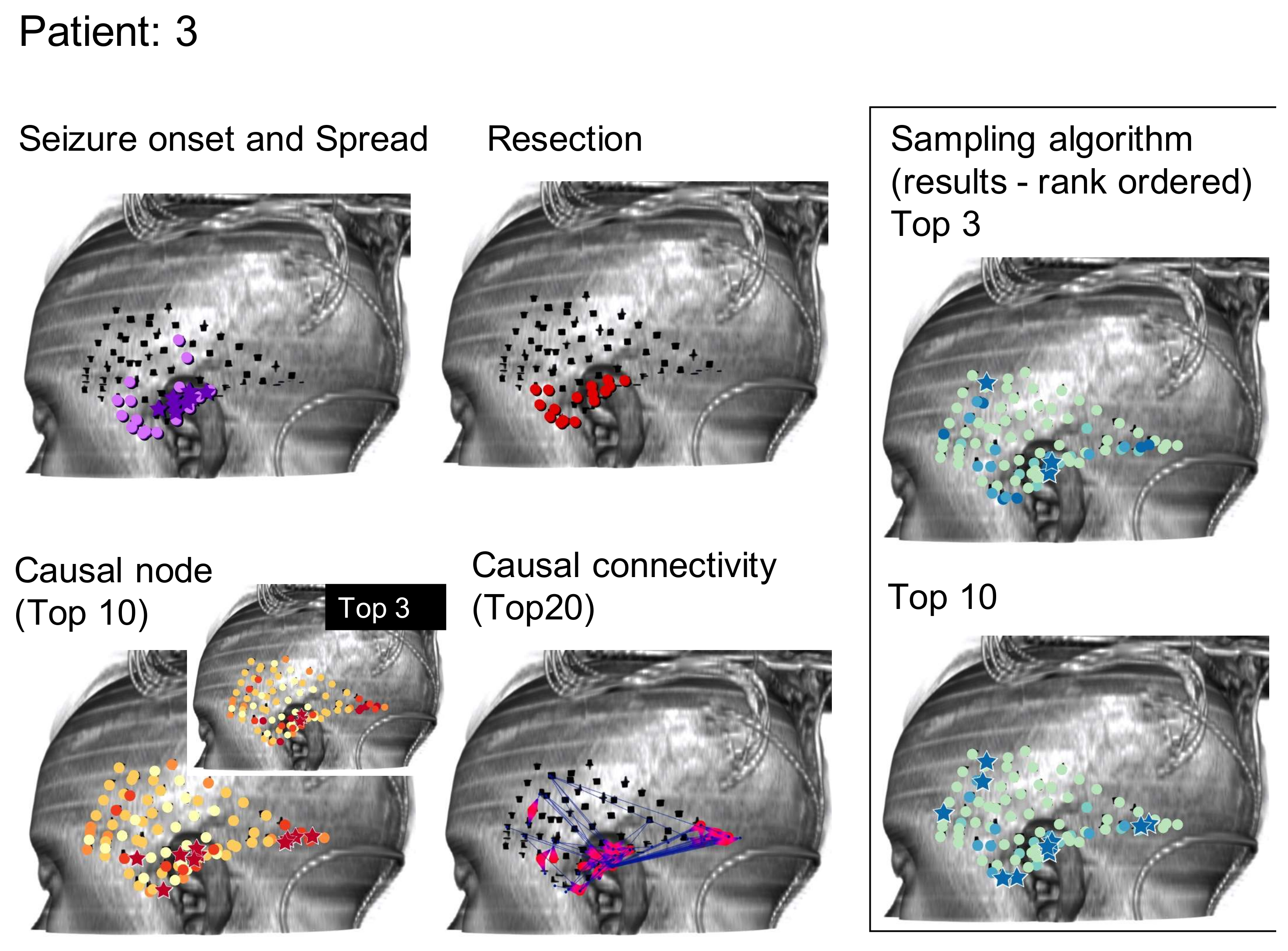
A summary evaluation of all graph algorithms tested via the rank order sum method for RZ is provided in
Figure 5. Displayed are the number of patients, out of the 24 retrospectively tested, that exhibited a significant rank order sum for the RZ (one patient did not undergo resection surgery after completing invasive monitoring). Each algorithm is run at least five times on all patients in the sample and the average of all runs for an algorithm is shown. Certain runs of the Monte Carlo Sampling algorithm variant that keeps track of the visitation interval achieve statistically significant rank order sums for 16 out of 24 patients’ RZ, representing an improvement of three patients from methods in the literature [
1]. As seen in
Figure 5, in general, sampling algorithms utilizing more than one token achieved a larger number of patients with statistically significant rank order sums than sampling with a single token, suggesting that tracking the convergence of multiple tokens eliminates much of the noise found in random sampling. However, as can be seen from the SE depicted by error bars in
Figure 5, there is significant variation between runs in most variants of the Monte Carlo Sampling algorithm, likely limiting its overall usefulness for surgery planning.
PageRank offers consistency of results from run to run, yet does not improve from the literature on the number of patients with a statistically significant rank order sum. Nevertheless, the fact that such a widespread algorithm used for other means in society can also be applied to the problem of seizure onset prediction opens the possibility for exploration of other graph algorithms originally invented for other purposes.
Combining the best of PageRank and Monte Carlo Sampling, indegree and outdegree centrality combined consistency of results with an improvement on the number of patients exhibiting statistically significant rank order sums. Notably, the 13 patients with statistically significant RZ rank order sums from Park and Madsen, 2018, do not completely overlap with the corresponding set of patients for graph algorithms tested in this study. Of the 16 significant patients from the most successful Monte Carlo sampling run, 10 patients are also part of the 13 from the total GC outdegree method from the literature. However, of the 16 significant patients from indegree centrality, 12 are also part of the 13 from total GC outdegree. Nevertheless, the lack of complete overlap suggests a combination of these methods could yield greater RZ and SOZ predication accuracy.
There was minimal improvement in SOZ rank order sums (13.6 patients on average versus 13 for total GC outdegree), likely limiting the immediate clinical potential of any of the algorithms tested. Nevertheless, important findings are derived from comparison of the graph algorithms’ results across their many variants. One such finding is displayed in
Figure 6 and discussed further below, where algorithm variants “reversing” edges to identify the trigger nodes of neural activity instead of the nodes exhibiting that activity itself almost universally perform better than their “forward” counterpart algorithms. For further details, results from each trial of centrality, PageRank, and total GC outdegree algorithms are reported in
Table S3.
4. Discussion
This study is the first to the authors’ knowledge that applies graph algorithms to Granger Causality analyses to holistically consider baseline interictal EEG data for RZ/SOZ detection, using approaches which can be influenced by the topology of nodes’ connections and not just the sum of their GC outdegrees. By demonstrating the comparable utility of these algorithms to those from the literature, we hope to motivate future refinement of other algorithm variants that take this holistic lens. Especially when different graph algorithms yield statistically significant rank order sums for different subsets of patients, further exploration of these algorithms has the potential to exhibit predictive capabilities for patients not yet covered by any previous algorithms.
Other contributions of this study to algorithmic seizure onset and resection zone identification via EEG data are the patterns it reveals in seizure network topography across patients. Namely, the effects of edge direction on algorithm performance reveal that the brain region influential in triggering neural activity better correlates to the SOZ as opposed to the brain regions exhibiting triggered neural activity itself. This observation holds true for Monte Carlo sampling, centrality, and PageRank algorithms, as seen through their tested variants. Uphill sampling, outdegree centrality, and reversed edges PageRank, all of which can be said to have “reversed edges”, give a higher rank to the root cause of neural activity rather than the regions exhibiting activity itself. These patterns are summarized in
Figure 6, where uphill/outdegree edge directions universally correspond to equal or better patient SOZ identification outcomes. This result is intuitive, given that the goal of surgery might be expected to be removal of high generators, rather than receivers, of causative signals. This result is also in line with previous studies based on the total GC outdegree method, which gives a higher rank to nodes that have greater total outward edge weights, likely indicating such nodes are more influential in the generation of activity in a seizure network [
1].
Additionally, we find that stochasticity in many of the algorithm variants in this paper impacts patient results to the point where predictions about which nodes are to be included in the SOZ and RZ are no longer reliable. This point is proven by the success of sampling algorithms with multiple tokens and the requirement that numerous tokens have to simultaneously be present in order to record a visitation. Such an algorithm eliminates the effects of randomness and keys future algorithm variants to be developed to do the same.
Finally, all of the algorithms tested in this study reveal that seizure activity truly is localized to a specific brain region. The relative success of recording nodes’ length of intervals between sampling as opposed to nodes’ overall number of visitations shows this, as sampling intervals seek to tease out cyclical brain activity. This is corroborated by the success of indegree and outdegree centrality compared with betweenness and harmonic centrality. The latter two identify patterns in a node’s role in the context of the entire brain network, because they look at shortest paths between all pairs of nodes. On the other hand, indegree and outdegree zero in on the localized role of a node in relation to only its neighbors. The success of the indegree and outdegree approaches exemplifies the need to focus on localized areas of the brain network instead of approaching all nodes together.
All of these insights were achievable only with information told by the edge weights contained in GC maps, data which were previously discarded. As data becomes available, future work will involve a greater number of patients than the 25 included in this study, in order to better determine the applicability of GC to a wider patient population. These additional patients will include those with sEEG implants as well as more ECoG cases similar to the 25 already studied. Future directions of this work include identifying the interictal time window when GC map-based SOZ and RZ predictions are most accurate. Because we obtain 60 different GC maps selected at random for each patient, these maps can be tagged with time elapsed since previous ictal activity and time to next ictal activity. With this data, an optimal window during which predictions most resemble the actual SOZ and RZ can be identified.
5. Conclusions
In this retrospective study, we develop and test new algorithms based on the entire network as revealed by GC, rather than simply each node’s outdegree. Within our cohort using these approaches, we find informative trends. Firstly, the matching of and improvement on algorithms in the literature by this study’s graph algorithms helps to solidify the relevance of GC maps to epilepsy surgery planning. Secondly, uphill sampling, outdegree centrality, and reversed edges PageRank, all of which can be said to have “reversed edges”, give a higher rank to the root cause of neural activity rather than the regions exhibiting activity itself. Thirdly, stochasticity in some algorithms impacts results such that predictions about nodes to include in the SOZ/RZ are no longer reliable. As such, variants that eliminate stochasticity such as sampling with multiple tokens greatly increase the usefulness of these highly random algorithms. Fourthly, the success of recording nodes’ sampling intervals as opposed to nodes’ overall number of visitations, as well as the success of indegree and outdegree over betweenness and harmonic centrality, reveal that seizure activity tends to localize to a specific brain region. Future work will involve more patients, including sEEG patients, to fine-tune the applicability of GC-based graph algorithms to a wider patient population.