Optimized Resolution-Oriented Many-to-One Intensity Standardization Method for Magnetic Resonance Images
Abstract
1. Introduction
2. Methods
2.1. Network Architecture
2.2. Adversarial Loss and Cycle Consistency Loss
2.3. Normalized Mutual Information Loss
2.4. Entire Loss Function
3. Data and Preprocessing
3.1. Dataset
3.2. Advanced Weak-Pair Data Input Strategy
3.3. Optimized Training Strategy with Synchronized Batch Normalization
4. Experiments and Results
4.1. The Resolution Preservation Ability Enhancement Brought by the Optimizations on Methodology
4.2. Other Visual and Numerical Comparison with the Previous Methods
5. Discussion
5.1. Analysis of the Model Stability of the Anomalies and Lesions
5.2. The Fusion of Various Losses during the Training Process
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| MRI | Magnetic Resonance Imaging |
| MR | Magnetic Resonance |
| CycleGAN | Cycle Generative Adversarial Network |
| PDFs | Probability Density Functions |
| NMI | Normalized Mutual Information |
| CARN | Cascading Residual Network |
| LeakyReLU | Leaky Rectified Linear Unit |
| LSGAN | Least Squares Generative Adversarial Networks |
| HGG | High Grade Glioma |
| LGG | Low Grade Glioma |
| MNI | Montreal Neurological Institute |
| PSNR | Peak Signal-to-noise Ratio |
| SSIM | Structural Similarity |
| VIF | Visual Information Fidelity |
| UQI | Universal Quality Index |
| IFC | Image Fidelity Criterion |
| GMSD | Gradient Magnitude Similarity Deviation |
| MSE | Mean Square Error |
| HC | Histogram Correlation |
| AD | Average Disparity |
| mRMR | Minimum-redundancy-maximum-relevance |
| GA | Genetic-algorithm |
| GPUs | Graphics Processing Units |
References
- Bradley, J.E.; Panagiotis, K.; Zeynettin, A.; Timothy, L.K. Machine learning for medical imaging. RadioGraphics 2017, 37, 505–515. [Google Scholar]
- Yu, J.; Shi, Z.; Lian, Y.; Li, Z.; Liu, T.; Gao, Y.; Wang, Y.; Chen, L.; Mao, Y. Noninvasive IDH1 mutation estimation based on a quantitative radiomics approach for grade II glioma. Eur. Radiol. 2017, 27, 3509–3522. [Google Scholar] [CrossRef] [PubMed]
- Feis, R.A.; Bouts, M.J.R.J.; Panman, J.L.; Jiskoot, L.C.; Dopper, E.G.P.; Schouten, T.M.; Vos, F.D.; Grond, J.V.D.; Swieten, J.C.V.; Rombouts, S.A.R.B. Single-subject classification of presymptomatic frontotemporal dementia mutation carriers using multimodal MRI. NeuroImage Clin. 2019, 22, 101718. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, B.N.; Hardie, R.C.; Kebede, T.M.; Sprague, M.J. Optimized feature selection-based clustering approach for computer-aided detection of lung nodules in different modalities. Pattern Anal. Appl. 2019, 22, 559–571. [Google Scholar] [CrossRef]
- Raj, A.; Kuceyeski, A.; Weiner, M. A network diffusion model of disease progression in dementia. Neuron 2012, 73, 1204–1215. [Google Scholar] [CrossRef] [PubMed]
- Shiradkar, R.; Podder, T.K.; Algohary, A.; Viswanath, S.; Ellis, R.J.; Madabhushi, A. Radiomics based targeted radiotherapy planning (Rad-TRaP): A computational framework for prostate cancer treatment planning with MRI. Radiat. Oncol. 2016, 11, 148. [Google Scholar] [CrossRef] [PubMed]
- Ingrisch, M.; Schneider, M.J.; Norenberg, D.; Figueiredo, G.N.D.; Maier-Hein, K.; Suchorska, B.; Schuller, U.; Albert, N.; Bruckmann, H.; Reiser, M.; et al. Radiomic analysis reveals prognostic information in T1-weighted baseline magnetic resonance imaging in patients with glioblastoma. Investig. Radiol. 2017, 52, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, B.N.; Hardie, R.C.; Kebede, T.M. Performance analysis of a computer-aided detection system for lung nodules in CT at different slice thicknesses. J. Med. Imaging 2018, 1, 014504. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Shi, Z.; Wang, Y.; Yu, J.; Chen, L.; Guo, Y.; Zhang, Q.; Mao, Y. Histological grade and type classification of glioma using Magnetic Resonance Imaging. In Proceedings of the 9th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics, Datong, China, 15–17 October 2016; pp. 1808–1813. [Google Scholar]
- Nyul, L.G.; Udupa, J.K.; Zhang, X. New variants of a method of MRI scale standardization. IEEE Trans. Med. Imaging 2000, 19, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Collewet, G.; Strzelecki, M.; Mariette, F. Influence of MRI acquisition protocols and image intensity normalization methods on texture classification. Magn. Reson. Imaging 2004, 22, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Madabhushi, A.; Udupa, J.K. New methods of MR image intensity standardization via generalized scale. Med. Phys. 2006, 33, 3426–3434. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Shi, L.; Luo, Y.; Yang, W.; Li, H.; Liang, P.; Li, K.; Mok, V.C.; Chu, W.C.; Wang, D. Histogram-based normalization technique on human brain magnetic resonance images from different acquisitions. Biomed. Eng. Online 2015, 14, 73. [Google Scholar] [CrossRef] [PubMed]
- Nunzio, G.D.; Cataldo, R.; Carla, A. Robust intensity standardization in brain magnetic resonance images. J. Digit. Imaging 2015, 28, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Jager, F.; Deuerling-Zheng, Y.; Frericks, B.; Wacker, F.; Hornegger, J. A new method for MRI intensity standardization with application to lesion detection in the brain. In Proceedings of the 11th International Fall Workshop Vision, Modeling, and Visualization, Aachen, Germany, 22–24 November 2006; Volume 1010, pp. 269–276. [Google Scholar]
- Jager, F.; Hornegger, J. Nonrigid registration of joint histograms for intensity standardization in magnetic resonance imaging. IEEE Trans. Med. Imaging 2009, 28, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Dzyubachyk, O.; Staring, M.; Reijnierse, M.; Lelieveldt, B.P.; Geest, R.J.V.D. Inter-station intensity standardization for whole-body MR data. Magn. Reson. Med. 2017, 77, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Robitaille, N.; Mouiha, A.; Crepeault, B.; Valdivia, F.; Duchesne, S. Tissue-based MRI intensity standardization: application to multicentric datasets. Int. J. Biomed. Imaging 2012, 2012, 347120. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Pan, J.; Guo, Y.; Yu, J.; Zhang, J.; Geng, D.; Wang, Y. N-D point cloud registration for intensity normalization on magnetic resonance images. In Proceedings of the VipIMAGE 2017, Porto, Portugal, 18–20 October 2017; pp. 121–130. [Google Scholar]
- Gao, Y.; Pan, J.; Guo, Y.; Yu, J.; Zhang, J.; Geng, D.; Wang, Y. Optimised MRI intensity standardisation based on multi-dimensional sub-regional point cloud registration. Comput. Methods Biomech. Biomed. Eng. Imaging Vis. 2019, 7, 1–10. [Google Scholar] [CrossRef]
- Gao, Y.; Liu, Y.; Wang, Y.; Shi, Z.; Yu, J. A universal intensity standardization method based on a many-to-one weak-paired cycle generative adversarial network for magnetic resonance images. IEEE Trans. Med. Imaging 2019, 38, 2059–2069. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.Y.; Park, T.; Isola, P.; Efros, A.A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the 2017 IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2242–2251. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: convolutional networks for biomedical image segmentation. In Proceedings of the 18th International Conference on Medical Image Computing and Computer Assisted Intervention, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar]
- Ahn, N.; Kang, B.; Sohn, K.A. Fast, accurate, and lightweight super-resolution with cascading residual network. In Proceedings of the 2018 European Conference on Computer Vision, Munich, Gemany, 8–14 September 2018. [Google Scholar]
- Wang, C.; Macnaught, G.; Papanastasiou, G.; MacGillivray, T.; Newby, D. Unsupervised learning for cross-domain medical image synthesis uing deformation invariant cycle consistency networks. In Proceedings of the 2018 International Workshop on Simulation and Synthesis in Medical Imaging, Granada, Spain, 16 September 2018; pp. 52–60. [Google Scholar]
- Ji, C.; Yu, J.; Wang, Y.; Chen, L.; Shi, Z.; Mao, Y. Brain tumor segmentation in MR slices using improved GrowCut algorithm. In Proceedings of the 7th International Conference on Graphic and Image Processing, Singapore, 23–25 October 2015. [Google Scholar]
- Kendall, A.; Gal, Y.; Cipolla, R. Multi-task learning using uncertainty to weigh losses for scene geometry and semantics. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Salt Lake City, UT, USA, 18–22 June 2018; pp. 7482–7491. [Google Scholar]
- BenTaieb, A.; Hamarneh, G. Uncertainty driven multi-loss fully convolutional networks for histopathology. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention Workshop on Large-scale Annotation of Biomedical data and Expert Label Synthesis (MICCAI LABELS), Quebec City, QC, Canada, 10–14 September 2017; pp. 155–163. [Google Scholar]
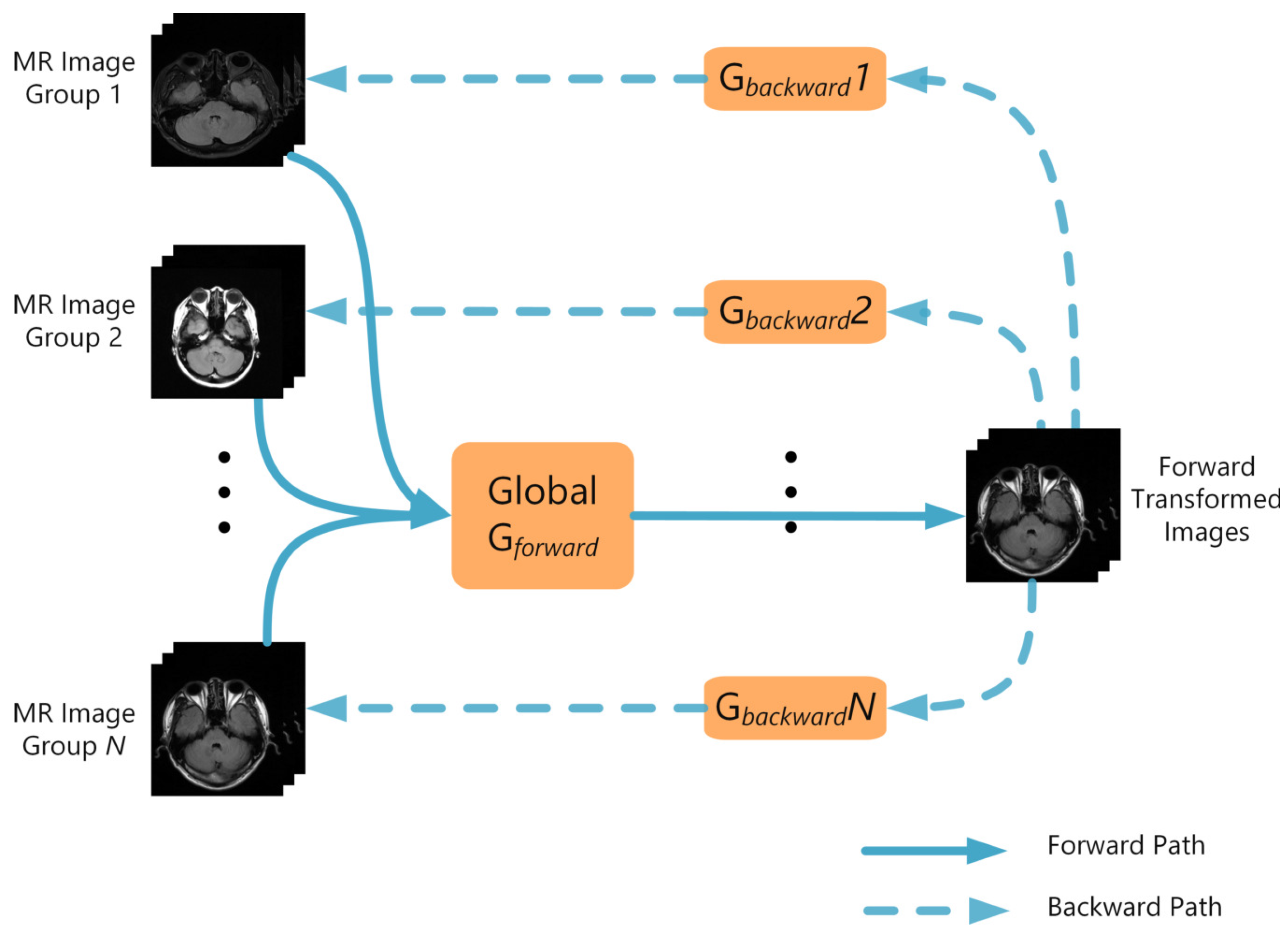



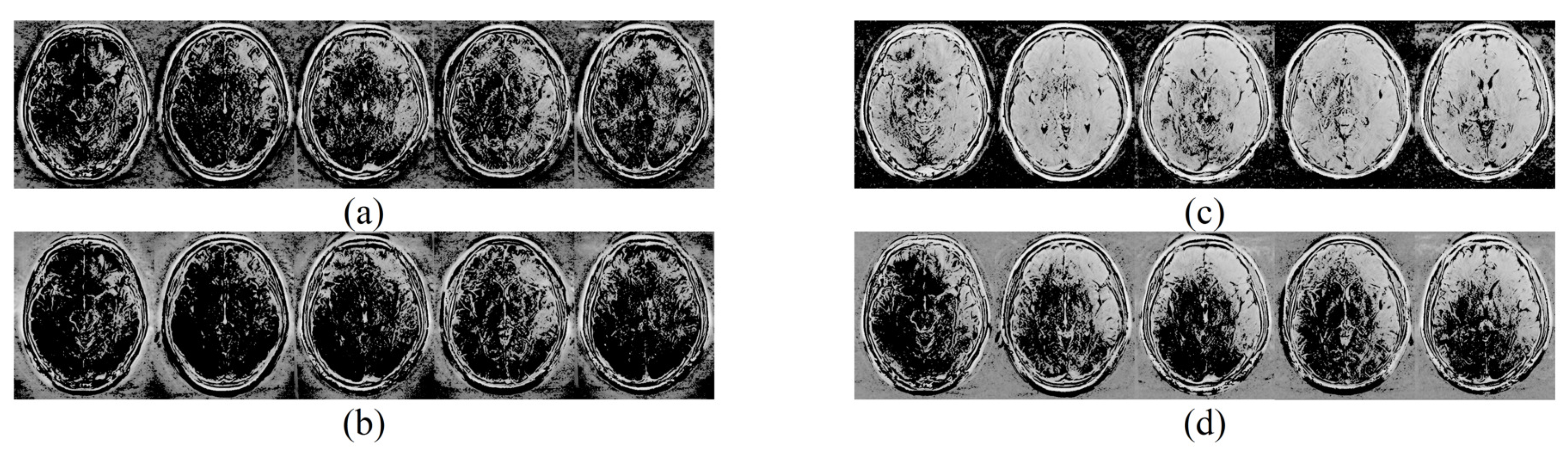
| (a) | |||||
| Image Group | 1 | 2 | 3 | 4 | 5 |
| Image number | 1904 | 1472 | 464 | 1196 | 656 |
| Patient number | 119 | 92 | 29 | 80 | 41 |
| Slice thickness (mm) | 6 | 6 | 6 | 6 | 4 |
| Pulse repetition time/Echo time/Inverse time (ms/ms/ms) | 8000/102/2370 | 9000/102/2500 | 8500/102/2439 | 8800/152/2100 | 8525/142/2100 |
| Imaging frequency (MHz) | 123.1678 | 123.2407 | 123.2463 | 127.7692 | 127.7706 |
| Pixel bandwidth (Hz/pixel) | 287 | 287 | 287 | 195.312 | 195.312 |
| Pixel spacing (mm) | 0.4492 | 0.4492 | 0.4492 | 0.4688 | 0.4688 |
| Receive coil | 12-channel head coil | 12-channel head coil | 12-channel head coil | 8-channel head coil | 8-channel head coil |
| (b) | |||||
| Image Group | 6 | 7 | 8 | 9 | |
| Image number | 924 | 548 | 644 | 384 | |
| Patient number | 46 | 30 | 32 | 20 | |
| Slice thickness (mm) | 5 | 5 | 5.5 | 5 | |
| Pulse repetition time/Echo time/Inverse time (ms/ms/ms) | 8500/90/2439 | 8500/91/2439 | 9000/83/2500 | 9000/128/2500 | |
| Imaging frequency (MHz) | 123.2587 | 123.2622 | 123.2001 | 123.2292 | |
| Pixel bandwidth (Hz/pixel) | 289 | 287 | 201 | 285 | |
| Pixel spacing (mm) | 0.6875 | 0.4297 | 0.8984 | 0.8984 | |
| Receive coil | 12-channel head coil | 12-channel head coil | 12-channel head coil | 32-channel head coil | |
| Image Group | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Pulse sequence | Axial T2W BLADE | T1W FLAIR | T2W FLAIR | Axial T2 PRO- PELLER | T1 FLAIR | T2 FLAIR |
| Slice thickness (mm) | 6 | 6 | 6 | 6 | 6 | 6 |
| Pulse repetition time/Echo time/nverse time (ms/ms/ms) | 3500/95/NA | 2000/17/857 | 8000/102/2370 | 4582/96/NA | 1872/22/720 | 8525/146/2100 |
| Imaging frequency (MHz) | 123.1678 | 123.1678 | 123.1678 | 127.7705 | 127.7705 | 127.7705 |
| Pixel bandwidth (Hz/pixel) | 287 | 287 | 287 | 195.312 | 195.312 | 195.312 |
| Pixel spacing (mm) | 0.8984 | 0.8984 | 0.4492 | 0.4688 | 0.4688 | 0.4688 |
| Group | Train | Test |
|---|---|---|
| Instrument | SIEMENS MAGNETOM Verio 3.0T MRI scanner | GE Discovery MR750 3.0T MRI scanner |
| Patient with HGG | 34 | 17 |
| Patient with LGG | 32 | 11 |
| Slice thickness (mm) | 6 | 4 |
| Pulse repetition time/Echo time/Inverse time (ms/ms/ms) | 8000/102/2370 | 8525/141.9/2100 |
| Imaging frequency (MHz) | 123.1678 | 127.7706 |
| Pixel bandwidth (Hz/pixel) | 287 | 195.312 |
| Transmit coil name | Body | 8HRBRAIN |
| Pixel spacing (mm) | 0.4492 | 0.4688 |
| Evaluation Criteria | PSNR | SSIM | IFC | UQI | VIF |
|---|---|---|---|---|---|
| The proposed method | 37.31 | 0.9663 | 4.164 | 0.6169 | 0.1588 |
| Original universal method | 37.21 | 0.9690 | 3.212 | 0.6138 | 0.1233 |
| Evaluation Criteria | PSNR | SSIM | IFC | UQI | VIF |
|---|---|---|---|---|---|
| The proposed method | 65.51 | 0.9992 | 4.318 | 0.7568 | 0.2413 |
| Original universal method | 65.13 | 0.9989 | 4.294 | 0.7538 | 0.2431 |
| Test Group | Original Images | Standardized Images with the Original Method | Standardized Images with the Proposed Method |
|---|---|---|---|
| Accuracy | 0.7143 | 0.8214 | 0.8571 |
| Sensitivity | 0.6471 | 0.7647 | 0.8824 |
| Specificity | 0.8182 | 0.9091 | 0.8182 |
| Evaluation Criteria | Original Images | The Proposed Method | Original Universal Method | Histogram Matching Method | Joint Histogram Registration Method |
|---|---|---|---|---|---|
| PSNR | 53.75 | 65.51 | 65.13 | 61.39 | 64.27 |
| SSIM | 0.9829 | 0.9992 | 0.9989 | 0.9968 | 0.9988 |
| HC | 0.2724 | 0.9273 | 0.8865 | 0.8049 | 0.8825 |
| GMSD | 0.1328 | 0.1026 | 0.1027 | 0.1291 | 0.1245 |
| MSE | |||||
| AD | 30.8 | 4.505 | 1.093 | 13.54 | 8.467 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, Y.; Wang, Y.; Yu, J. Optimized Resolution-Oriented Many-to-One Intensity Standardization Method for Magnetic Resonance Images. Appl. Sci. 2019, 9, 5531. https://doi.org/10.3390/app9245531
Gao Y, Wang Y, Yu J. Optimized Resolution-Oriented Many-to-One Intensity Standardization Method for Magnetic Resonance Images. Applied Sciences. 2019; 9(24):5531. https://doi.org/10.3390/app9245531
Chicago/Turabian StyleGao, Yuan, Yuanyuan Wang, and Jinhua Yu. 2019. "Optimized Resolution-Oriented Many-to-One Intensity Standardization Method for Magnetic Resonance Images" Applied Sciences 9, no. 24: 5531. https://doi.org/10.3390/app9245531
APA StyleGao, Y., Wang, Y., & Yu, J. (2019). Optimized Resolution-Oriented Many-to-One Intensity Standardization Method for Magnetic Resonance Images. Applied Sciences, 9(24), 5531. https://doi.org/10.3390/app9245531





