Issue-Based Clustering of Scholarly Articles
Abstract
Featured Application
Abstract
1. Introduction
2. Background
2.1. Citation-Based Similarity Measures
2.2. Content-Based Similarity Measures
3. Development of ICRT
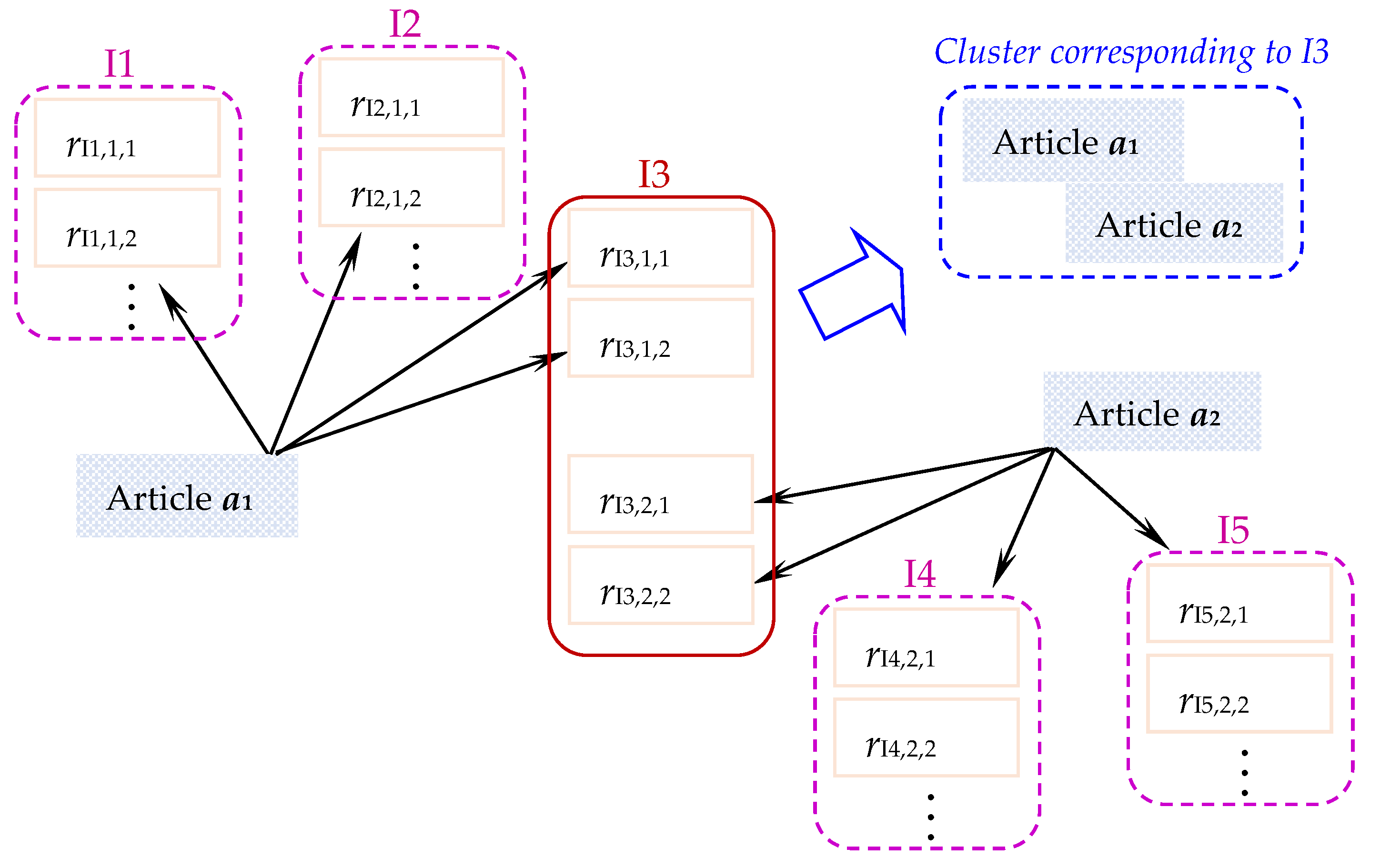
3.1. Estimation of Issue-Based Similarity (IBS)
3.2. Identification of the Set of Issue-Indicative Terms (IIT)
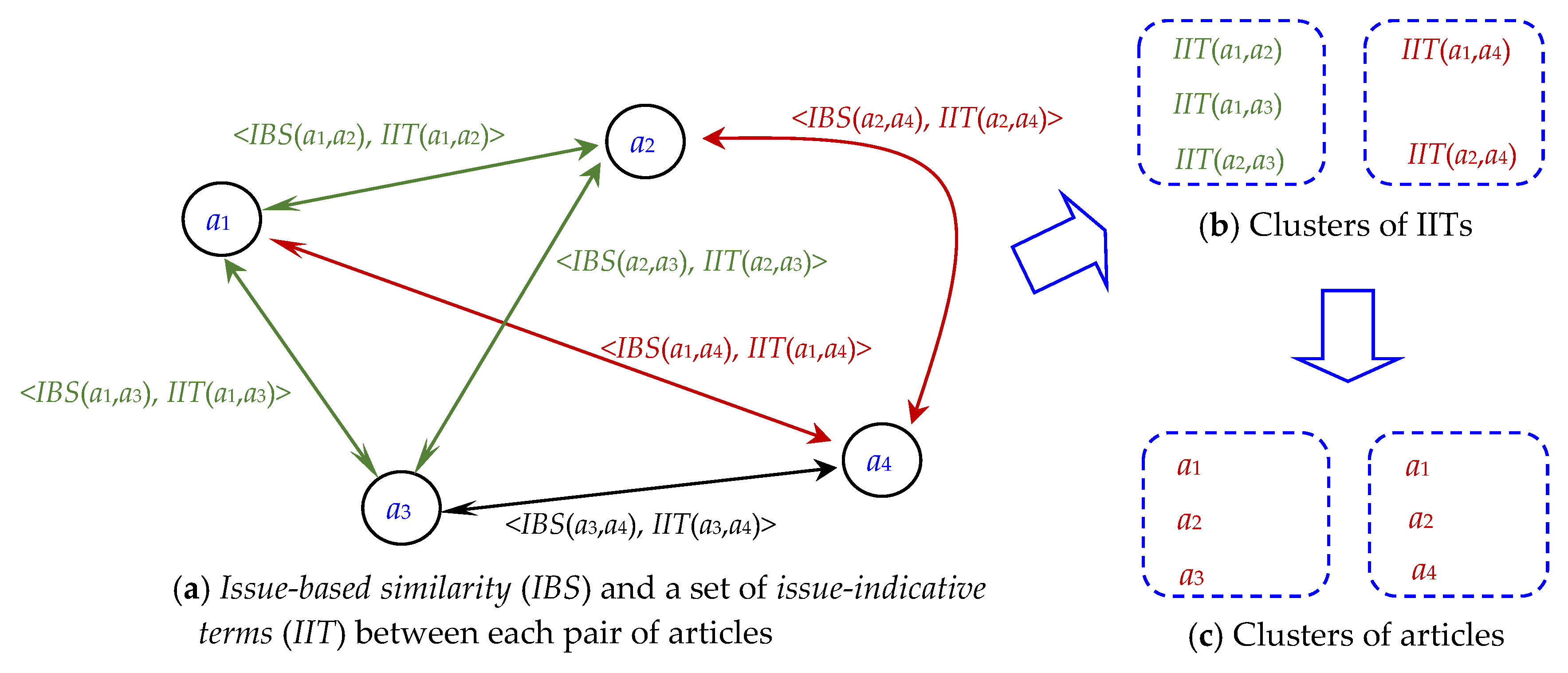
3.3. Clustering of IITs and Involving Articles
4. Experiments
4.1. Experimental Data
4.2. Underlying Systems for Overlapping Clustering
4.3. Evaluation Criteria
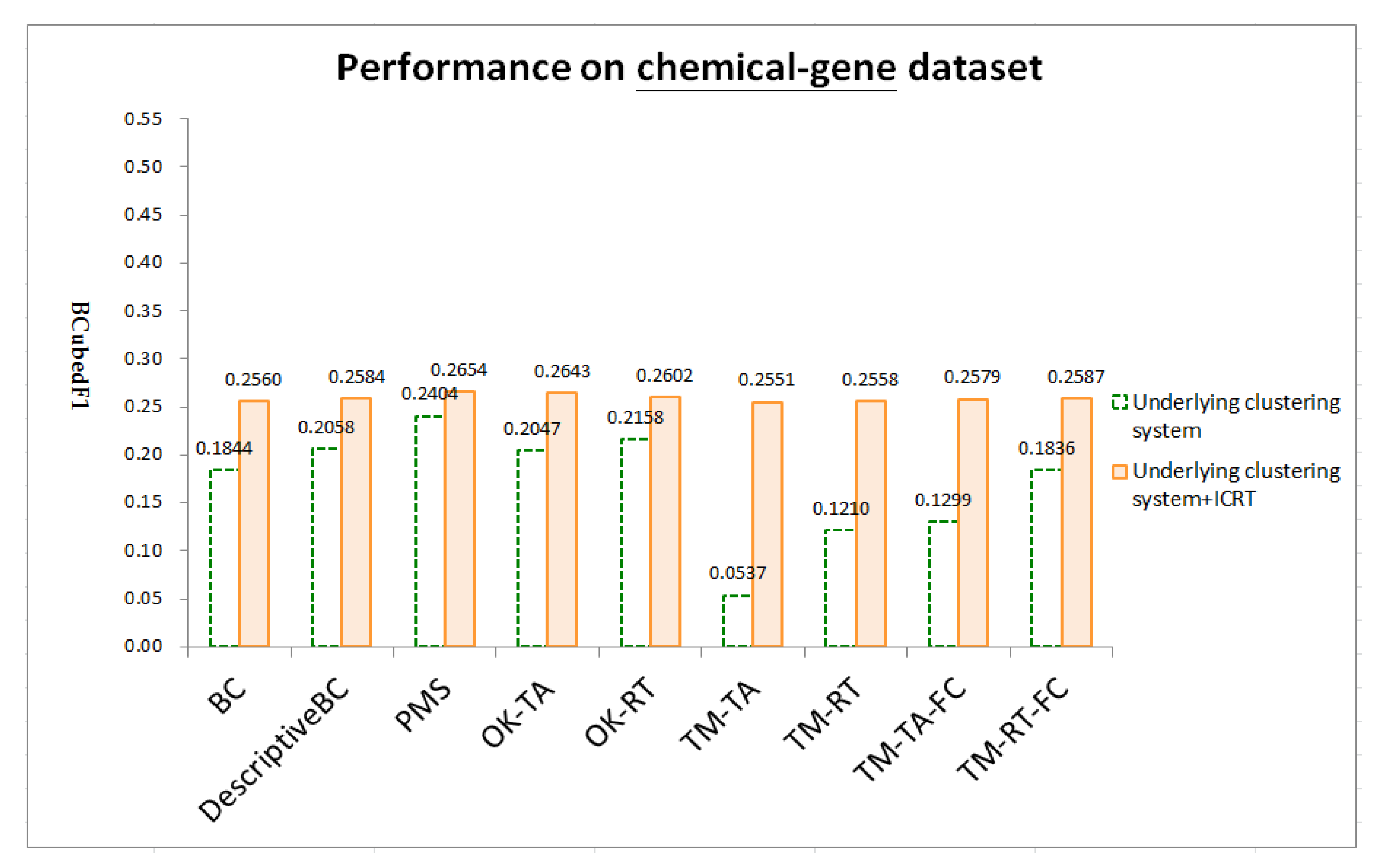
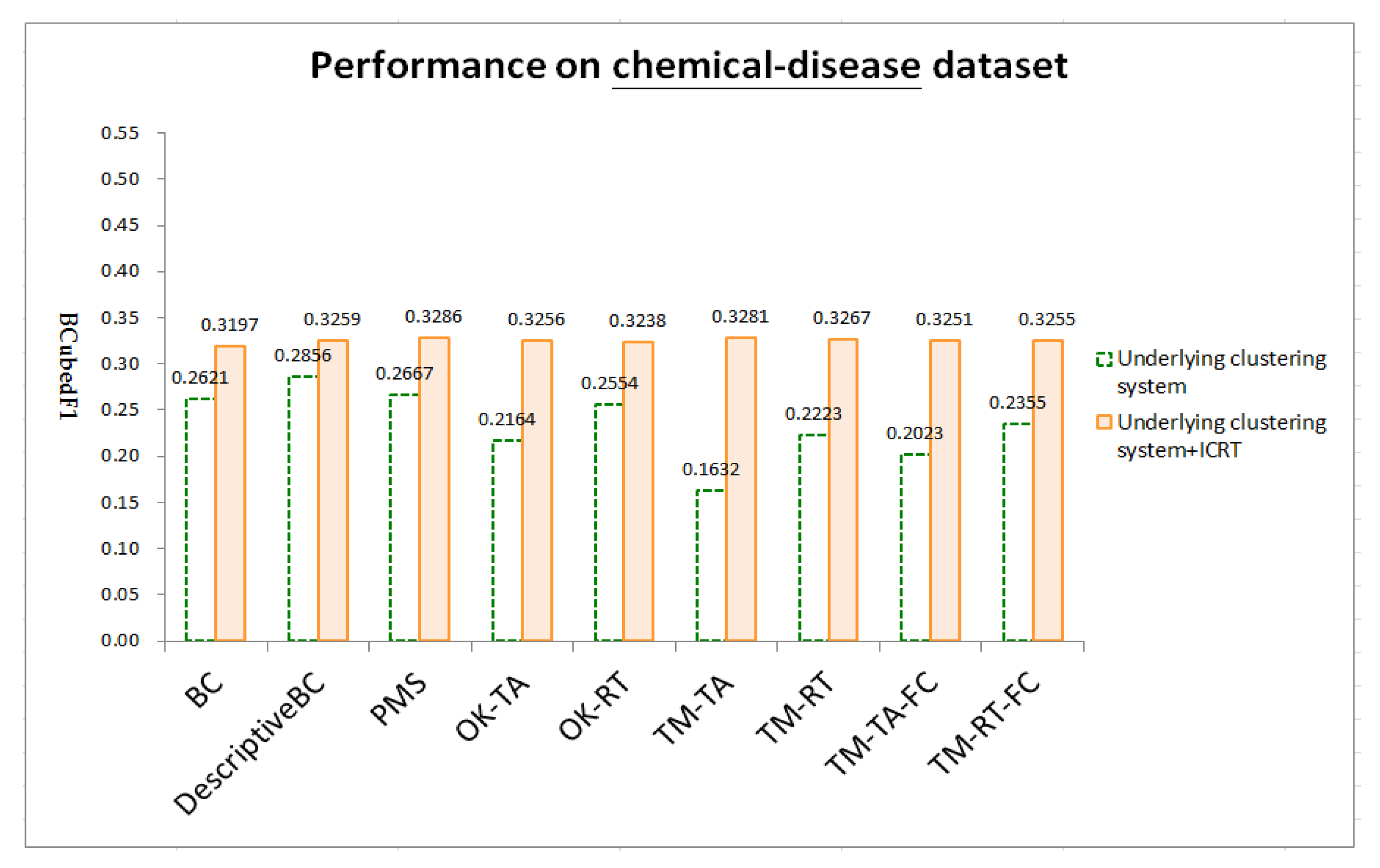
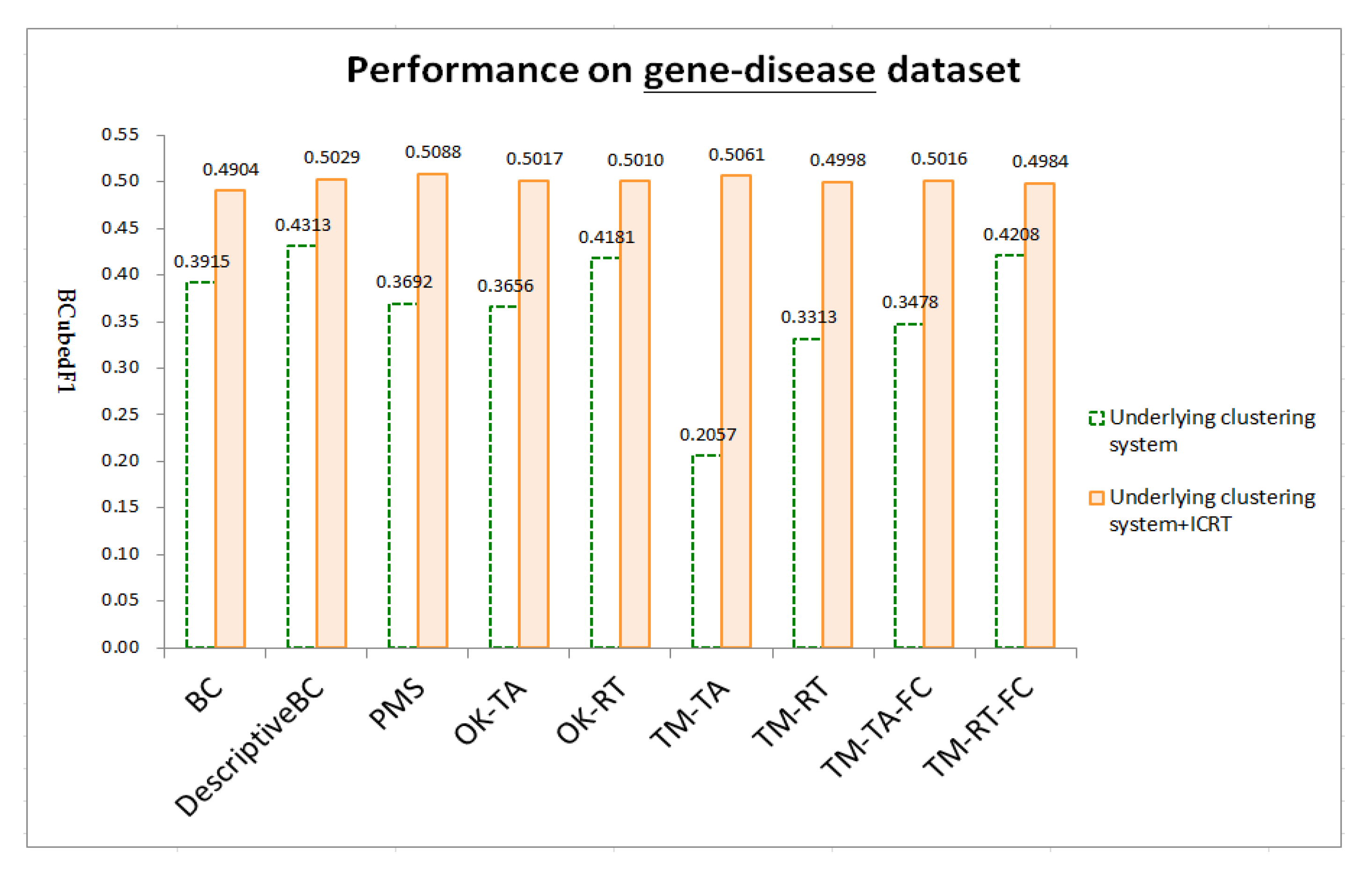
4.4. Results
4.5. A Case Study
5. Conclusions and Future Work
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAQ—When Is Data Updated? Available online: http://ctdbase.org/help/faq/;jsessionid=92111C8A6B218E4B2513C3B0BEE7E63F?p=6422623 (accessed on 11 December 2018).
- Expert Reviewers—Genetics Home Reference—NIH. Available online: https://ghr.nlm.nih.gov/about/expert-reviewers (accessed on 11 December 2018).
- About OMIM. Available online: https://www.omim.org/about (accessed on 11 December 2018).
- Peters, G.; Crespo, F.; Lingras, P.; Weber, R. Soft clustering—Fuzzy and rough approaches and their extensions and derivatives. Int. J. Approx. Reason. 2013, 54, 307–322. [Google Scholar] [CrossRef]
- Bezdek, J.C.; Ehrlich, R.; Full, W. FCM: The Fuzzy c-means Clustering Algorithm. Comput. Geosci. 1984, 10, 191–203. [Google Scholar] [CrossRef]
- Sisodia, D.S.; Verma, S.; Vyas, O.P. A Subtractive Relational Fuzzy C-Medoids Clustering Approach to Cluster Web User Sessions from Web Server Logs. Int. J. Appl. Eng. Res. 2017, 12, 1142–1150. [Google Scholar]
- Krishnapuram, R.; Joshi, A.; Yi, L. A Fuzzy Relative of the k-Medoids Algorithm with Application to Web Document and Snippet Clustering. In Proceedings of the IEEE International Conference on Fuzzy Systems, Seoul, Korea, 22–25 August 1999; pp. 1281–1286. [Google Scholar]
- Šubelj, L.; van Eck, N.J.; Waltman, L. Clustering Scientific Publications Based on Citation Relations: A Systematic Comparison of Different Methods. PLoS ONE 2016, 11, e0154404. [Google Scholar] [CrossRef] [PubMed]
- Small, H.G. Co-citation in the scientific literature: A new measure of relationship between two documents. J. Am. Soc. Inf. Sci. 1973, 24, 265–269. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, Y.; Liu, R.; Zhang, J. Knowledge-transfer analysis based on co-citation clustering. Scientometrics 2013, 3, 859–869. [Google Scholar] [CrossRef]
- Kessler, M.M. Bibliographic coupling between scientific papers. Am. Doc. 1963, 14, 10–25. [Google Scholar] [CrossRef]
- Couto, T.; Cristo, M.; Gonc¸alves, M.A.; Calado, P.; Nivio Ziviani, N.; Moura, E.; Ribeiro-Neto, B. A Comparative Study of Citations and Links in Document Classification. In Proceedings of the 6th ACM/IEEE-CS Joint Conference on Digital Libraries, Chapel Hill, NC, USA, 11–15 June 2006; pp. 75–84. [Google Scholar]
- Boyack, K.W.; Klavans, R. Co-citation analysis, bibliographic coupling, and direct citation: Which citation approach represents the research front most accurately? J. Am. Soc. Inf. Sci. Technol. 2010, 61, 2389–2404. [Google Scholar] [CrossRef]
- Liu, R.-L. Passage-based Bibliographic Coupling: An Inter-Article Similarity Measure for Biomedical Articles. PLoS ONE 2015, 10, e0139245. [Google Scholar] [CrossRef]
- Janssens, F.; Glänzel, W.; De Moor, B. A hybrid mapping of information science. Scientometrics 2008, 75, 607–631. [Google Scholar] [CrossRef]
- Liu, R.-L. A New Bibliographic Coupling Measure with Descriptive Capability. Scientometrics 2017, 110, 915–935. [Google Scholar] [CrossRef]
- Tian, G.; Jing, L. Recommending scientific articles using bi-relational graph-based iterative RWR. In Proceedings of the 7th ACM Conference on Recommender Systems, Hong Kong, China, 12–16 October 2013; pp. 399–402. [Google Scholar]
- Whissell, J.S.; Clarke, C.L.A. Effective Measures for Inter-Document Similarity. In Proceedings of the 22nd ACM International Conference on Conference on Information & Knowledge Management, San Francisco, CA, USA, 17 October–1 November 2013; pp. 1361–1370. [Google Scholar]
- Boyack, K.W.; Newman, D.; Duhon, R.J.; Klavans, R.; Patek, M.; Biberstine, J.R. Clustering More than Two Million Biomedical Publications: Comparing the Accuracies of Nine Text-Based Similarity Approaches. PLoS ONE 2011, 6, e18029. [Google Scholar] [CrossRef] [PubMed]
- Glenisson, P.; Glanzel, W.; Janssens, F.; De Moor, B. Combining full text and bibliometric information in mapping scientific disciplines. Inf. Process. Manag. 2005, 41, 1548–1572. [Google Scholar] [CrossRef]
- Landauer, T.K.; Laham, D.; Derr, M. From paragraph to graph: Latent semantic analysis for information visualization. Proc. Natl. Acad. Sci. USA 2004, 101 (Suppl. 1), 5214–5219. [Google Scholar] [CrossRef] [PubMed]
- Robertson, S.E.; Walker, S.; Beaulieu, M. Okapi at TREC-7: Automatic ad hoc, filtering, VLC and interactive. In Proceedings of the 7th Text REtrieval Conference (TREC 7), Gaithersburg, MD, USA, 1 July 1998; pp. 253–264. [Google Scholar]
- PubMed Help—PubMed Help—NCBI Bookshelf. Available online: https://www.ncbi.nlm.nih.gov/books/NBK3827/#pubmedhelp.Computation_of_Similar_Articl (accessed on 11 December 2018).
- Lin, J.; Wilbur, W.J. PubMed related articles: A probabilistic topic-based model for content similarity. BMC Bioinform. 2007, 8, 423. [Google Scholar] [CrossRef] [PubMed]
- Blei, D.M. Probabilistic topic models. Commun. ACM 2012, 55, 77–84. [Google Scholar] [CrossRef]
- Yau, C.-K.; Porter, A.L.; Newman, N.C.; Suominen, A. Clustering scientific documents with topic modeling. Scientometrics 2014, 100, 767–786. [Google Scholar] [CrossRef]
- Xie, P.; Xing, E.P. Integrating Document Clustering and Topic Modeling. In Proceedings of the Twenty-Ninth Conference on Uncertainty in Artificial Intelligence, Bellevue, WA, USA, 11–15 August 2013; pp. 694–703. [Google Scholar]
- The Comparative Toxicogenomics Database | CTD. Available online: http://ctdbase.org/ (accessed on 11 December 2018).
- Davis, A.P.; Grondin, C.J.; Johnson, R.J.; Sciaky, D.; King, B.L.; McMorran, R. The Comparative Toxicogenomics Database: Update 2017. Nucleic Acids Res. 2017, 45, D972–D978. [Google Scholar] [CrossRef] [PubMed]
- Wiegers, T.C.; Davis, A.P.; Cohen, K.B.; Hirschman, L.; Mattingly, C.J. Text mining and manual curation of chemical-gene-disease networks for the Comparative Toxicogenomics Database (CTD). BMC Bioinform. 2009, 10, 326. [Google Scholar] [CrossRef]
- Home—PMC—NCBI. Available online: https://www.ncbi.nlm.nih.gov/pmc/ (accessed on 11 December 2018).
- [Table, Stopwords]—PubMed Help—NCBI Bookshelf. Available online: https://www.ncbi.nlm.nih.gov/books/NBK3827/table/pubmedhelp.T.stopwords/ (accessed on 13 March 2018).
- MetaMap—A Tool For Recognizing UMLS Concepts in Text. Available online: https://metamap.nlm.nih.gov/ (accessed on 13 March 2018).
- GitHub—Senderle/Topic-Modeling-Tool: A Point-and-Click Tool for Creating and Analyzing Topic Models Produced by MALLET. Available online: https://github.com/senderle/topic-modeling-tool (accessed on 13 March 2018).
- Amigo, E.; Gonzalo, J.; Artiles, J.; Verdejo, F. A comparison of extrinsic clustering evaluation metrics based on formal constraints. Inf. Retr. 2009, 12, 461–486. [Google Scholar] [CrossRef]
- Banerjee, A.; Krumpelman, C.; Ghosh, J.; Basu, S.; Mooney, R.J. Model based overlapping clustering. In Proceedings of the International Conference on Knowledge Discovery and Data Mining, Chicago, IL, USA, 18 August 2005; pp. 532–537. [Google Scholar]
- Lewis, D.D.; Schapire, R.E.; Callan, P.; Papka, R. Training Algorithms for Linear Text Classifiers. In Proceedings of the 19th Annual International ACM SIGIR Conference on Research and Development in Information Retrieval, Zurich, Switzerland, 18–22 August 1996; pp. 298–306. [Google Scholar]
- Liu, R.-L. Context-based Term Frequency Assessment for Text Classification. J. Am. Soc. Inf. Sci. Technol. 2010, 61, 300–309. [Google Scholar]
- Tong, T.; Dinakarpandian, D.; Lee, Y. Literature Clustering using Citation Semantics. In Proceedings of the 42nd Hawaii International Conference on System Sciences, Big Island, HI, USA, 5–8 January 2009. [Google Scholar]
- Janssens, F.; Zhang, L.; De Moor, B.; Glänzel, W. Hybrid clustering for validation and improvement of subject-classification schemes. Inf. Process. Manag. 2009, 45, 683–702. [Google Scholar] [CrossRef]

| Dataset | # Clusters Having X Articles | # Articles Belonging to X Clusters | # Articles Published in Year X | ||||||
|---|---|---|---|---|---|---|---|---|---|
| X = 2 Articles | X = 3 Articles | X ≥ 4 Articles | X = 1 Cluster | X = 2 Clusters | X ≥ 3 Clusters | X ≤ 2005 | X in [2006,2010] | X ≥ 2011 | |
| (1) <chemical, gene> | 6719 (50.49%) | 3140 (23.60%) | 3448 (25.91%) | 2669 (31.59%) | 1614 (19.10%) | 4167 (49.31%) | 1215 (14.38%) | 3233 (38.26%) | 4002 (47.36%) |
| (2) <chemical, disease> | 1338 (54.59%) | 479 (19.54%) | 634 (25.87%) | 3176 (60.80%) | 1235 (23.64%) | 813 (15.56%) | 1840 (35.22%) | 1919 (36.73%) | 1465 (28.05%) |
| (3) <gene, disease> | 414 (72.50%) | 94 (16.46%) | 63 (11.04%) | 902 (78.43%) | 188 (16.35%) | 60 (5.22%) | 170 (14.78%) | 446 (38.78%) | 534 (46.44%) |
| Similarity Measures | Content | Overlapping Clustering | ||
|---|---|---|---|---|
| References | Titles & Abstracts | Fuzzy Clustering | Thresholding | |
| (1) Bibliographic Coupling: BC | √ | √ | √ | |
| (2) DescriptiveBC | √ | √ | √ | |
| (3) PubMed-based similarity: PMS | √ | √ | √ | |
| (4-1) OK: OK-TA | √ | √ | √ | |
| (4-2) OK: OK-RT | √ | √ | √ | |
| (5-1) Topic Modeling: TM-TA | √ | √ | ||
| (5-2) Topic Modeling: TM-RT | √ | √ | ||
| (5-3) Topic Modeling: TM-TA-FC | √ | √ | √ | |
| (5-4) Topic Modeling: TM-RT-FC | √ | √ | √ | |
| Article | Associated Entities (Curated by CTD) | Errors Made by the Systems | |||
|---|---|---|---|---|---|
| TERT | DC | DKC1 | IPF | ||
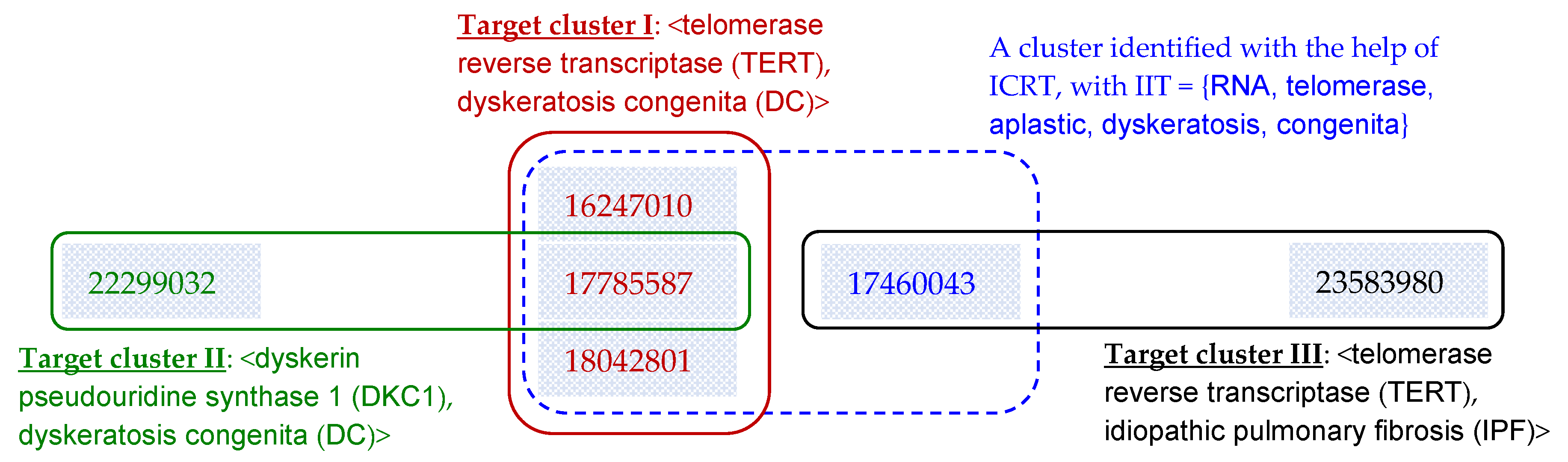
| (1) 17785587 (Telomerase reverse-transcriptase homozygous mutations in autosomal recessive dyskeratosis congenita and Hoyeraal-Hreidarsson syndrome) | √ | √ | √ | OK-TA: <TERT, DC> (Target cluster I in Figure 7) is not identified for this article. | |
| (2) 17460043 (Adult-onset pulmonary fibrosis caused by mutations in telomerase) | √ | √ | All systems (including those enhanced with ICRT): Group this article into <TERT, DC> (Target cluster I in Figure 7). | ||
| (3) 23583980 (Genome-wide association study identifies multiple susceptibility loci for pulmonary fibrosis) | √ | √ | BC and DescriptiveBC: Group this article into <TERT, DC> (Target cluster I in Figure 7). | ||
| (4) 22299032 (Zebrafish models for dyskeratosis congenita reveal critical roles of p53 activation contributing to hematopoietic defects through RNA processing) | √ | √ | All underlying systems except BC and DescriptiveBC: (1) Fail to group this article into <DKC1, DC> (Target cluster II in Figure 7); and/or (2) Group this article into <TERT, DC> (Target cluster I in Figure 7). | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, R.-L.; Hsu, C.-K. Issue-Based Clustering of Scholarly Articles. Appl. Sci. 2018, 8, 2591. https://doi.org/10.3390/app8122591
Liu R-L, Hsu C-K. Issue-Based Clustering of Scholarly Articles. Applied Sciences. 2018; 8(12):2591. https://doi.org/10.3390/app8122591
Chicago/Turabian StyleLiu, Rey-Long, and Chih-Kai Hsu. 2018. "Issue-Based Clustering of Scholarly Articles" Applied Sciences 8, no. 12: 2591. https://doi.org/10.3390/app8122591
APA StyleLiu, R.-L., & Hsu, C.-K. (2018). Issue-Based Clustering of Scholarly Articles. Applied Sciences, 8(12), 2591. https://doi.org/10.3390/app8122591




