Abstract
Traditional She medicine is part of China’s cultural heritage and has become remarkably popular worldwide. The Shi-Liang tea is made from the processed leaves of Chimonanthus salicifolius S. Y. Hu and Chimonanthus zhejiangensis M. C. Liu. To ensure the safety and efficacy of Shi-Liang tea, we used DNA barcoding based on the internal transcribed spacer 2 (ITS2) of nuclear ribosomal DNA to distinguish the original plant sources of Shi-Liang tea from closely related species. All 71 ITS2 sequences were aligned by Clustal-W, and genetic distances were computed using MEGA 6.0 according to the Kimura 2-parameter model. The results indicated that the sequence lengths of ITS2 regions of the original plants of Shi-Liang tea and closely related species ranged from 256 bp to 260 bp. Interspecific genetic distances ranged from 0 to 0.078. The neighbor-joining (NJ) tree showed that the original plants of Shi-Liang tea species can be easily differentiated from closely related species. Distinct molecular differences were found between the secondary structures of ITS2 sequences from Shi-Liang tea and closely related species. The results in the present investigation suggested that the ITS2 could be an effective DNA marker to identify the original plants of Shi-Liang tea and their closely related species. These DNA barcodes can greatly benefit the supervision of the commercial circulation of She medicine.
1. Introduction
Traditional She medicine is an important part of folk medicine with a long history. The She medicine has attracted much attention from the Han and She ethnic groups in Southeastern China, especially in the Zhejiang and Fujian provinces. This traditional Chinese medicine was gradually developed to prevent and treat diseases under particular historical conditions and special geographical environments when the She minority group struggled with diseases to survive and reproduce [1,2,3]. Shi-Liang tea, which is one of the most widely used She medicines, is made from the leaves of two Chimonanthus species in the Calycanthaceae family: Chimonanthus salicifolius S. Y. Hu and Chimonanthus zhejiangensis M. C. Liu [4]. The Calycanthaceae family includes ten species that belong to three genera and seven species of two genera distributed in China. Chimonanthus is a Chinese endemic genus and mainly distributed in the Zhejiang, Fujian, and Jiangxi provinces. Chimonanthus plants have also been introduced to Japan, North Korea, Europe, and North America, among others. Shi-Liang tea was indexed as a commonly used type of She medicine in the 2005 edition of the Chinese traditional medicine processing standard of Zhejiang [5].
Previous studies showed that Shi-Liang tea mainly contains volatile oils and flavonoids, among others. The tea can be used to treat diseases such as the common cold or influenza, deficiency of food stagnation, abdominal pain, and acid regurgitation [6,7,8,9,10,11]. Shi-Liang tea exhibits antibacterial, anti-inflammatory, antipyretic, analgesic, antitussive, and expectorant effects, such as remarkable effects against strep throat and bronchitis and certain curative effects on high blood pressure [6,7,8,9,10,11]. Chemical differences and biodiversity are known among the original plants used in Shi-Liang tea and several other species in the Calycanthaceae family, such as Chimonanthus nitens Oliv, Chimonanthus praecox (L.) Link, Chimonanthus campanulatus R. H. Chang et C. S. Ding, Calycanthus floridus L., and Calycanthus chinensis Cheng et S. Y. Chang [6,7,8,9,10,11,12,13,14]. The classification of family Calycanthaceae has long been controversial because of the similar morphologies among Calycanthaceae species [6,15,16]. Therefore, several studies have investigated pharmacology [6,17], chemical composition [6,7,8,9,10,11,12,13,14], and classification [6,15,16] of the original plant sources of Shi-Liang tea and closely related species. Shi-Liang tea is made from processed leaves, which are difficult to authenticate based on traditional morphology. The original plant sources of Shi-Liang tea are impossible to distinguish from their closely related species by mere visual inspection. The misuse and abuse of folk medicine could pose a risk to consumer safety and hinder the long-term development of traditional Chinese medicine. Therefore, the accurate identification of Shi-Liang tea species and closely related species is urgently needed.
With the development of molecular biology and bioinformatics, DNA barcoding has recently become a hotspot for identifying species and biodiversity research [18,19,20,21,22]. This biological method uses the short and universal segments of DNA to rapidly and accurately identify species. DNA barcoding has become an important approach to remedy the limitations of morphology-based identification systems. The candidate barcodes, psbA-trnH, matK, rbcL, rpoC1, ycf5, ITS, and ITS2, were evaluated according to their potential in species identification. The ITS2 DNA barcode has been reported to be the most suitable region to identify medicinal plants based on the analytical results from more than 6600 medicinal plants and their closely related species [22,23]. Moreover, the secondary structure of ITS2 is highly conserved throughout the eukaryotes and correlated with taxonomic classification. The application of secondary structures not only improves the stability of resulting trees, but more importantly also increases the accuracy of phylogenetic estimations [24,25,26]. The two-dimensional DNA barcode is one of the most appropriate symbologies for DNA sequences information. This barcode has the largest coding capacity and relatively high compression ratio. The two-dimensional DNA is also similar to the barcode scanning used in supermarkets to distinguish different kinds of goods [27]. In this study, DNA barcoding technology was used to identify Shi-Liang tea (C. salicifolius and C. zhejiangensis) and five other species in the family Calycanthaceae using ITS2 to ensure the accuracy and safety of Calycanthaceae in traditional folk medicine practice. Moreover, further study of other She medicine by identifying these species in family Calycanthaceae will provide technical support for the research and development of She medicine. This study will provide a frontier on folk medicine application in China.
2. Materials and Methods
2.1. Materials
Thirty-six samples from seven species in the Calycanthaceae family, namely, 18 C. salicifolius, 10 C. zhejiangensis, 1 C. praecox, 1 C. nitens, 2 C. campanulatus, 1 Calycanthus floridus, and 3 Calycanthus chinensis, were collected from the Zhejiang, Shaanxi, Guangxi, Hunan, Jiangxi, and Anhui provinces. All the samples were authenticated by Kejun Cheng and deposited at the Institute of Medicinal Plant Development, Chinese Academy of Medical Sciences & Peking Union Medical College (Beijing, China). Moreover, another 35 sequences of Calycanthaceae were downloaded from GenBank. The details of the plant materials are listed in Table 1 and the GenBank accession numbers are shown in Table S1.

Table 1.
Sample detail information of Shi-Liang tea and its closely related species.
2.2. DNA Extraction, Amplification and Sequencing
Approximately 20 mg of the silicon-dried leaves were rubbed for 2 min at 30 times/s in a Fast Prep bead mill (Retsch MM400, Haan, Germany). The total genomic DNA (gDNA) was isolated from the crushed materials using the Plant Genomic DNA Kit (Tiangen Biotech Co., Ltd., Beijing, China) according to the manufacturer’s instructions and stored at −20 °C. The ITS2 sequences from each plant were amplified in a reaction containing 2 μL of gDNA (approximately 30 ng), 12.5 μL of 2× Taq Mix Master, and 1 μL each of the 2.5 μM upstream (S2F: 5′-ATGCGATACTTGGTGTGAAT-3′) and downstream (S3R: 5′-GACGCTTCTCCAGACTACAAT-3′) primers, with ddH2O to a volume of 25 μL. The PCR program consisted of an initial denaturation step at 94 °C for 5 min, followed by 40 cycles of 94 °C for 30 s, 56 °C for 30 s, and 72 °C for 45 s, with a final extension step at 72 °C for 10 min [28]. The amplified products were separated on 1.2% agarose gels and sequenced after gel extraction. All 71 ITS2 sequences were aligned with Clustal-W, and genetic distances were computed with MEGA 6.0 [28] according to the Kimura 2-parameter (K2P) model. A neighbor-joining (NJ) tree was built, and the bootstrap test (1000 replications) was applied to determine the reliability of each branch, and Ceratophyllum echinatum from Calycanthaceae were used as the outgroup. The secondary structures of the ITS2 sequences were predicted according to the ITS2 database by Schultz et al. [24,29,30]. The two-dimensional DNA barcode images were obtained by encoding the Latin names of the Shi-Liang tea species and closely related species, as well as the ITS2 sequences, using the coding program and the open source PHP QR Code. Mobile terminals (such as Android and iPhone devices) can be used as QR code scanners to read the information. Finally, the information was made available online (http://www.tcmbarcode.cn) to identify and analyze ITS2 sequences [27,31].
3. Results and Discussion
3.1. ITS2 Sequence Characteristics of Shi-Liang Tea Species and Closely Related Species
The total gDNA was isolated from the crushed materials. The PCR amplification efficiency of ITS2 was 100%, and 36 ITS2 sequences were successfully obtained. The 38 ITS2 sequences for Shi-Liang tea included 23 sequences of C. salicifolius and 15 sequences of C. zhejiangensis. The ITS2 sequences of C. salicifolius ranged from 256 bp to 258 bp, while those of C. zhejiangensis were 256 bp in length. The average GC contents of the C. salicifolius and C. zhejiangensis sequences were 71.2% and 71.1%, respectively (Table 2). The intra-specific K2P distances of the ITS2 sequences ranged from 0 to 0.008 in C. salicifolius and from 0 to 0.004 in C. zhejiangensis (Table 3) and the average inter-specific distance among Shi-Liang tea species and its closely related species is 0.047. The minimum inter-specific K2P distances of the ITS2 sequences is the one between C. zhejiangensis and C. nitens which ranged from 0.004 to 0.008 and the maximum one is between C. nitens and Ca. chinensis. All 71 ITS2 sequences were 256 bp to 260 bp in length, with 36 nucleotide variation sites, including 20 parsim-informative sites. The variable site in the Shi-Liang tea species and closely related species are displayed in Figure S1.

Table 2.
Sequence length and average GC contents of Shi-Liang tea and its closely related species.

Table 3.
Intra- and inter-specific K2P distance of Shi-Liang tea and its closely related species.
3.2. Clustering Analysis of Shi-Liang Tea Sources and Closely Related Species
The NJ tree is a method for constructing phylogenetic trees using either DNA sequences or amino acid sequences. It takes a distance matrix as input and specifies the distance between each pair of taxa [31]. In this study, the NJ tree was constructed by using all 71 ITS2 sequences of Shi-Liang tea species and closely related species and the outgroup to study their phylogenetic relationships (Figure 1). It is obvious that the ITS2 sequences of Shi-Liang tea species (C. salicifolius, C. zhejiangensis) formed one clade, which could be successfully distinguished from the closely related species: C. praecox, C. nitens, C. campanulatus, Ca. floridus, and Ca. chinensis, which formed one clade, respectively. Therefore, Shi-Liang tea sources could be successfully distinguished from closely related species C. praecox, C. nitens, C. campanulatus, Ca. floridus, and Ca. chinensis by DNA barcoding based on the ITS2 sequence, but C. salicifolius and C. zhejiangensis cannot be distinguished from each other by ITS2 DNA barcoding. The two commercial samples, CH-17 and CH-18, formed one clade with C. salicifolius in the NJ tree, which showed that they were identified as the labeled species.
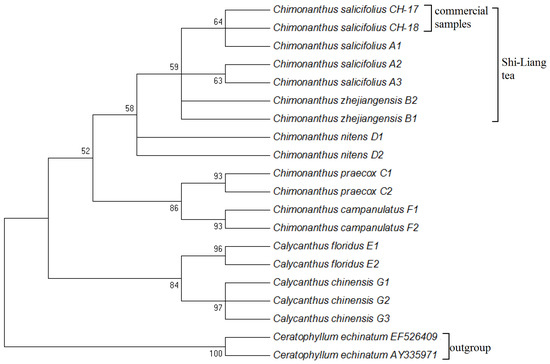
Figure 1.
The NJ-tree of Shi-Liang tea and its closely related species.
3.3. Comparison of ITS2 Secondary Structures of Shi-Liang Tea Species and Closely Related Species
Previous studies showed that the secondary structures of ITS2 are conserved and can provide useful biological information for alignment [24,25,32]. These structures can be considered as molecular morphological characteristics to identify species. We compared the secondary structures of Shi-Liang tea species and closely related species. The secondary structures of 71 ITS2 sequences had four similar helices: Helices I, II, III, and IV. The secondary structures of species in Chimonanthus were relatively more conserved in Helices III and IV, but the positions, sizes, and numbers of loops in Helices I and II were different. By contrast, the secondary structures between Ca. floridus and Ca. chinensis were highly conserved in Helices II, III, and IV, but had fewer differences in Helix I. These results show that these helices can be used as molecular characteristics to identify species [25,26] (Figure 2). Therefore, Shi-Liang tea species could be clearly distinguished from closely related species based on their secondary structures.
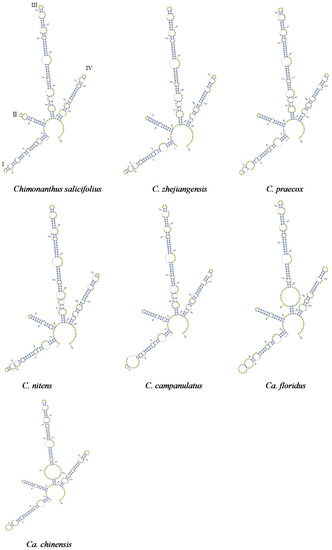
Figure 2.
The ITS2 secondary structure of Shi-Liang tea and its closely related species.
3.4. Two-Dimensional DNA Barcodes to Monitor Market Circulation and Promote the Modernization of Shi-Liang Tea Species and Closely Related Species
The two-dimensional DNA barcode is the most appropriate tool for DNA sequences, with the largest coding capacity and relatively high compression ratio [27]. In this study, ITS2 sequences of Shi-Liang tea species and closely related species were converted into colored barcode images based on our software code (Figure 3) [31,33]. This system consisted of the DNA barcode sequences and the cross-platform conversion of DNA barcode information. The application of two-dimensional DNA barcodes is beneficial to effectively monitor the quality and regulate the market circulation of Shi-Liang tea.

Figure 3.
ITS2 sequences barcode and two-dimensional DNA barcode image of Shi-Liang tea and its closely related species (■ A■ T■ C■ G).
The image list in the order of C. salicifolius, C. zhejiangensis, C. praecox, C. nitens, C. campanulatus, Calycanthus floridus, and Ca. chinensis.
4. Discussion
4.1. Application of ITS2 Sequences in Identification of She Medicine
DNA markers have become convenient tools for species identification and molecular systematics as a result of the advances in molecular technology [34,35,36]. However, ITS/ITS2 regions as universal barcodes present several problems and limitations, such as ITS/ITS2 showed poor PCR success in non-angiosperms [36]. Moreover, this parameter is quite variable in numerous phylogenetic studies. The amplification segment length of ITS/ITS2 was more than 1100 bp in some species and long poly-G, poly-C, and poly-A in ITS2/ITS made it difficult to sequence and sequence analysis [36]. Additionally, the nuclear gene itself has many copies and numerous variations within sequences, hindering it as a common barcode [37]. While Song et al. [38] found that the mutation of ITS2 in a genome is frequent but major variants of ITS2 are conserved and predominant. The usage of the major variants alone is sufficient for phylogeny construction and species determination in most cases. ITS2 is one of the most important core markers in molecular systematics and evolution [23]. Thus, this trait has shown great potential as a barcode to identify plants by providing the highest rates of accurate identification at the species and genus levels [23,26,30]. Compared with other candidate DNA barcodes, namely, psbA-trnH, matK, rbcL, rpoC1, ycf5, and ITS, the ITS2 sequence represents the most suitable region for DNA barcoding applications to classify medicinal plant species [22]. Most medicinal plants such as Cornus officinalis, Menispermum dauricum, Rhodiola crenulata, and Selaginella tamariscina could be identified effectively based on ITS2 barcode [19,20,35,39]. In the present study, the ITS2 region was used to distinguish Shi-Liang tea species from closely related species. The results showed that the ITS2 barcoding regions could be easily amplified with sufficient discrimination as a candidate marker to identify Shi-Liang tea.
4.2. DNA Barcoding Identification of Herbal Tea
DNA barcoding identification can identify counterfeit species with highly similar biological traits that cause great difficulty in morphological identification. This technology can accurately and rapidly identify several species or even dozens of families based on one or a few selected gene segments. This technology has the advantages of simple operation and good repeatability of results. The method can be universally applied to build a unified database and evaluation platform, as well as to promote and standardize plant classification. DNA sequences have been extensively used in systematic taxonomic research for species identification [40,41]. DNA barcoding technology has been increasingly applied in authenticating herbal tea [42,43]. Stoeckle et al. [42] used rbcL or matK barcodes to test the ingredients of 146 commercial tea products and showed that a third of the identified DNA was not found on herbal tea labels. Li et al. [43] systematically collected 177 samples of raw materials and established DNA barcodes for the identified botanical sources of Chinese “cooling” beverages. In this study, the CH-17 and CH-18 are commercial tea samples and high-quality DNA in both of them were extracted from these samples and the NJ tree and BLAST result based on similarity showed that they were identified as the labeled species.
China is the “hometown” of tea and the “birthplace” of tea culture. Chinese tea culture has a long history, and this culture has spread worldwide. Tea is edible and can be used as an antidote. Long-term tea-drinking can benefit health and prolong life-span. An ancient saying has stated that “tea is the essence of heaven and earth, and is the foundation of life” [44,45,46]. Shi-Liang tea is a health drink used by the She minority group for hundreds of years. The importance of standardization and quality control of Shi-Liang tea has increased because of its rapidly growing popularity and trade. To ensure the use of correct materials in tea production, this study demonstrated the use of DNA barcodes to identify the source species of Shi-Liang tea (C. salicifolius and C. zhejiangensis) and other Calycanthaceae species using the ITS2 sequence. The results of sequence alignment demonstrated the significant differences in ITS2 sequences between the original species of Shi-Liang tea and closely related species. The nucleotide variation sites among these sequences reached up to 37. The NJ tree constructed based on the ITS2 sequences demonstrated that the Shi-Liang tea species (C. salicifolius and C. zhejiangensis) formed one clade, which can be successfully distinguished from closely related species.
4.3. Application of the Secondary Structure of ITS2 Sequences in Traditional Chinese Medicine Identification
The ITS2 region of nuclear DNA is a powerful marker for DNA barcoding because of the abundant variation in primary sequences and secondary structures. The application of secondary structures of ITS2 sequences to reconstruct phylogenies could improve the stability of obtained trees and increase the accuracy of phylogenetic estimations [24,25,26]. Schultz et al. [26] built an ITS2 database web server, which facilitates the detection and prediction of secondary structure of ITS2 sequences. Zhao et al. [47] compared the secondary structures of the ITS2 sequences of Acanthopanacis cortex and its adulterants, providing another dimensionality to species identification. Thus, we focused on the divergence of primary sequences of ITS2 and the use of variations in the secondary structures to identify the She medicine species. By comparing the secondary structure of Shi-Liang tea species and closely related species, the results show that the secondary structures of all analyzed species had four similar helices: Helices I, II, III, and IV. Major differences among these helices could more distinctly distinguish the original plants of Shi-Liang tea species from closely related species than primary sequences.
4.4. Application and Significance of Two-Dimensional DNA Barcodes
Two-dimensional DNA barcodes have the largest coding capacity with a relatively high compression ratio. These barcodes can be divided into multiple data areas and can be easily scanned. The information is stored in multiple QR code symbols that can be reconstructed as a single data symbol to allow for high error tolerance. The use of a two-dimensional DNA barcode as the standard DNA barcode presentation format will promote the practical applications of DNA barcodes. Xin et al. [33] combined QR codes and DNA barcodes to construct a two-dimensional DNA barcode system to regulate the circulation of TCM. Liu et al. [27] compared different types of one-dimensional and two-dimensional barcodes for five DNA barcode markers, namely, ITS2, rbcL, matK, psbA-trnH, and CO1, and developed a web server for further use of QR code-based DNA barcodes. In this study, we obtained a two-dimensional DNA barcode for Shi-Liang tea, which can be used commercially to monitor the circulation of She medicine. The application of two-dimensional DNA barcode technology promotes the effective monitoring of quality and regulation of the commercial circulation of Shi-Liang tea.
5. Conclusions
ITS2 sequences were examined to identify and distinguish the Shi-Liang tea species from closely related species. The results showed that the ITS2 regions can be used as DNA barcodes, providing a new technique to promote the development of She medicine. DNA barcoding based on the ITS2 sequence could differentiate medical plants in the Calycanthaceae family and successfully classify other She medicines. Our research supports the development of traditional She medicine and its applications, especially in terms of quality control and management.
Supplementary Materials
The following are available online at http://www.mdpi.com/2076-3417/7/3/195/s1, Table S1: The accession numbers of Shi-Liang Tea and its closely related species, Figure S1: The sequence alignment of the Shi-Liang tea species and closely related species. The figure of sequence alignment is the site number.
Acknowledgments
This work was supported by funds from the Key Projects in the National Science and Technology Pillar Program (No.2011BAI07B08), the National Natural Science Foundation of China (No. 81303305), the Zhejiang Major Science & Technology Project of New Agricultural Varieties (No. 2016C02058), the Lishui Research Fund (No. LS20140010), and the Postdoctoral Research Fund of Zhejiang Province. The authors are grateful to Shilin Chen (Institute of Chinese Materia Medica, China Academy of Chinese Medical Sciences, Beijing, China) for his valuable guidance and Xinhe Pan (Lishui Institute of Forestry, Lishui, Zhejiang, China) and Shujuan Li (Xi’an botanical garden, Xi’an, China) for their assistance in the samples collections.
Author Contributions
H.Y. and K.C. conceived the experiments, S.M., Q.L., and J.F. designed the experiments, S.M. and H.Z. performed the experiments, S.M., W.C., and C.J. analyzed the data, S.M., H.Y., and K.C. wrote the paper.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Lei, H.; Li, J.; Zheng, S.; Fan, L.; Li, S.; Cheng, W.; Hua, J.; Yu, Y.; Dai, D.; Xie, W. Resources and Application of She’s Nationality Wild Medicinal Plants. Chin. J. Chin. Mater. Med. 2014, 39, 3180–3183. [Google Scholar]
- Lei, H.; Tao, Y.; Yan, L. Recent Progress in Research of She Nationality Medicine. J. Med. Pharm. Chin. Min. 2006, 12, 8–10. [Google Scholar]
- Lei, H.; Li, S. Chinese She Nationality Medicine; China Press of Traditional Chinese Medicine: Beijing, China, 2008. [Google Scholar]
- Wang, L.; Yan, L.; Yang, T.; Zhou, J. HPLC Determination of Rutin Quercein in Traditional She Medicine Diet Cool Tea. Chin. Arch. Tradit. Chin. Med. 2014, 32, 2916–2918. [Google Scholar]
- Zhejiang Province Food and Drug Administration. Zhejiang Province Processing Standard of Traditional Chinese Medicine; Zhejiang Science and Technology Press: Zhejiang, China, 2005; pp. 310–311. [Google Scholar]
- Xiao, K.; Liu, Y. Classification, chemical composition and pharmacological activity research progress of Chimonanthus. Res. Pract. Chin. Med. 2003, 17, 59–61. [Google Scholar]
- Ting, O.; Mai, X. Analysis of the Chemical Contituents of Essential Oil from Chimonanthus zhejiangensis by GC-MS. J. Chin. Med. Mater. 2010, 33, 385–386. [Google Scholar]
- Zhang, Y.; Hua, J.; Wang, X.; Cheng, W.; Lei, H.; Cheng, K.; Yu, P. Chemical constituents of chloroform fraction from leaves of Chimonanthus salicifolius. China. J. Chin. Mater. Med. 2013, 38, 2661–2664. [Google Scholar]
- Cui, X.; Li, M.; Zhou, B.; Li, J.; Cheng, D.; Zheng, P. Simultaneous Determination of Four Active Compounds in Chimonanthus nitens by HPLC. Chin. J. Exp. Tradit. Med. Formulae 2013, 19, 85–87. [Google Scholar]
- Mao, J.; Wang, W.; Yu, L.; Cheng, K. Study on Methodolgy of Content Determination of Three Flavonoids in Traditional She Medicine Shi Liang Cha. China Pharm. 2014, 17, 1808–1810. [Google Scholar]
- Yang, C.; Cheng, W.; Mei, J.; Yi, Y.; Cheng, J.; Ying, G.; Wang, H. Extraction and purification of total coumarins from Chimonanthus salicifolius Hu with ultrasonic assisted extraction. J. Zhejiang Univ. Technol. 2012, 40, 247–249. [Google Scholar]
- Ma, G.; Yang, G.; Xiong, J.; Cheng, K.; Hu, J. Salicifoxazines A and B, new cytotoxic tetrahydro-1,2-oxazine-containing tryptamine-derived alkaloids from the leaves of Chimonanthus salicifolius. Tetrahedron Lett. 2015, 56, 4071–4075. [Google Scholar] [CrossRef]
- Xu, J.; Cheng, K. Studies on the Alkaloids of the Calycanthaceae and Their Syntheses. Molecules 2015, 20, 6715–6738. [Google Scholar] [CrossRef] [PubMed]
- Lv, J.; Zhang, L.; Chu, X.; Zhou, J. Chemical composition, antioxidant and antimicrobial activity of the extracts of the flowers of the Chinese plant Chimonanthus praecox. Nat. Prod. Res. 2012, 6, 1363–1367. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Zhao, M.; Lan, T.; He, S. Research on China’s Chimonanthus plant taxonomy and Germplasm Resources. J. Beijing For. Univ. 1992, 14, 112–118. [Google Scholar]
- Zhu, Y.; Huang, Q. Studies on the identification of plants of Chimonanthus genus in China. J. Shanghai Med. Univ. 1987, 14, 451–455. [Google Scholar]
- Liu, Z.; Xi, J.; Schröder, S.; Wang, W.; Xie, T.; Wang, Z.; Bao, S.; Fei, J. Chimonanthus nitens var. salicifolius Aqueous Extract Protects against 5-Fluorouracil Induced Gastrointestinal Mucositis in a Mouse Model. Evid. Based Complement. Altern. Med. 2013, 4, 789263. [Google Scholar]
- Li, M.; Cao, H.; But, P.P.; Shaw, P.C. Identification of herbal medicinal materials using DNA barcodes. J. Syst. Evol. 2011, 49, 271–283. [Google Scholar] [CrossRef]
- Yang, P.; Li, X.; Zhou, H.; Hu, H.; Zhang, H.; Sun, W.; Wang, Y.; Yao, H. Molecular Identification of Chinese Materia Medica and Its Adulterants Using ITS2 and psbA-trnH Barcodes: A Case Study on Rhizoma Menispermi. Chin. Med. 2014, 5, 190–198. [Google Scholar] [CrossRef]
- Hou, D.; Song, J.; Yao, H.; Han, J.; Pang, X.; Shi, L.; Wang, X.; Chen, S. Molecular identification of Corni fructus and its adulterants by ITS/ITS2 sequences. Chin. J. Nat. Med. 2013, 11, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Pang, X.; Song, J.; Shi, L.; Yao, H.; Han, J.; Leon, C. A renaissance in herbal medicine identification: From morphology to DNA. Biotechnol. Adv. 2014, 32, 1237–1244. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Yao, H.; Han, J.; Liu, C.; Song, J.; Shi, L.; Zhu, Y.; Ma, X.; Gao, T.; Pang, X.; et al. Validation of the ITS2 regional a novel DNA barcode for identifying medicinal plant species. PLoS ONE 2010, 5, e8613. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Song, J.; Liu, C.; Luo, K.; Han, J.; Li, Y.; Pang, X.; Xu, H.; Zhu, Y.; Xiao, P.; et al. Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE 2010, 5, e13102. [Google Scholar] [CrossRef] [PubMed]
- Koetschan, C.; Förster, F.; Keller, A.; Schleicher, T.; Ruderisch, B.; Schwarz, R.; Müller, T.; Wolf, M.; Schultz, J. The ITS2 Database III-sequences and structures for phylogeny. Nucleic Acids Res. 2010, 38, 275–279. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.; Schleicher, T.; Schultz, J.; Müller, T.; Dandekar, T.; Wolf, M. 5.8S-28S rRNA interaction and HMM-based ITS2 annotation. Gene 2009, 430, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.; Maisel, S.; Gerlach, D.; Müller, T.; Wolf, M. A common core of secondary structure of the internal transcribed spacer 2 (ITS2) throughout the Eukaryota. RNA 2005, 11, 361–364. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Shi, L.; Xu, X.; Li, H.; Xing, H.; Liang, D.; Jiang, K.; Pang, X.; Song, J.; Chen, S. DNA Barcode Goes Two-Dimensions: DNA QR Code Web Server. PLoS ONE 2012, 7, e35146. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Selig, C.; Wolf, M.; Müller, T.; Dandekar, T.; Schultz, J. Homology modelling RNA structure for molecular systematics. Nucleic Acids Res. 2008, 36, D377–D380. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.; Muller, T.; Achtziger, M.; Seibel, P.N.; Dandekar, T.; Wolf, M. The internal transcribed spacer 2 database-a web server for (not only) low level phylogenetic analyses. Nucleic Acids Res. 2006, 34, W704–W707. [Google Scholar] [CrossRef] [PubMed]
- Chen, S. Standard DNA Barcodes of Chinese Materia Medica in Chinese Pharmacopoeia; Science Press: Beijing, China, 2015. [Google Scholar]
- Coleman, A. Pan-eukaryote ITS2 homologies revealed by RNA secondary structure. Nucleic Acids Res. 2007, 35, 3322–3329. [Google Scholar] [CrossRef] [PubMed]
- Xin, T.; Li, X.; Yao, H.; Han, J.; Song, J.; Chen, S. A two-dimensional DNA barcode system for circulation regulation of traditional Chinese medicine. Sci. Sin. Vitae 2015, 45, 695–702. [Google Scholar]
- Yao, H.; Song, J.; Ma, X.; Liu, C.; Li, Y.; Xu, H.; Han, J.; Duan, L.; Chen, S. Identification of Dendrobium species by a candidate DNA barcode sequence: The chloroplast psbA-trnH intergenic region. Planta Med. 2009, 75, 667–669. [Google Scholar] [CrossRef] [PubMed]
- Xin, T.; Li, X.; Yao, H.; Lin, Y.; Ma, X.; Cheng, R.; Song, J.; Ni, L.; Fan, C.; Chen, S. Survey of commercial Rhodiola products revealed species diversity and potential safety issues. Sci. Rep. 2015, 5, 8337. [Google Scholar] [CrossRef] [PubMed]
- Sass, C.; Little, D.; Stevenson, D.; Specht, C. DNA barcoding in the cycadales: Testing the potential of proposed barcoding markers for species identification of cycads. PLoS ONE 2007, 2, e1154. [Google Scholar] [CrossRef] [PubMed]
- Kress, W.; Erickson, D. A two-locus global DNA barcode for land plants: The coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE 2007, 2, e508. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Shi, L.; Li, D.; Sun, Y.; Niu, Y.; Chen, Z.; Luo, H.; Pang, X.; Sun, Z.; Liu, C.; Lv, A.; Deng, Y.; Larson-Rabin, Z.; Wilkinson, M.; Chen, S. Extensive pyrosequencing reveals frequent intra-genomic variations of internal transcribed spacer regions of nuclear ribosomal DNA. PLoS ONE 2012, 7, e43971. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Song, J.; Gao, Y.; Sun, Q.; Yao, H.; Wu, Q.; Chao, J.; Zhou, J.; Xue, W.; Duan, J. Application of the ITS2 region for barcoding medicinal plants of Selaginellaceae in Pteridophyta. PLoS ONE 2013, 8, e67818. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y. The Study of Tea Culture in Contemporary China and the Future Direction of Chinese. Jiangxi Soc. Sci. 2005, 7, 7–18. [Google Scholar]
- Chen, S.; Guo, B.; Zhang, G.; Yan, Z.; Luo, G.; Sun, S.; Wu, H.; Huang, L.; Pang, X.; Chen, J. Advances of studies on new technology and method for identifying traditional Chinese medicinal materials. Chin. J. Chin. Mater. Med. 2012, 37, 1043–1055. [Google Scholar]
- Stoeckle, M.Y.; Gamble, C.C.; Kirpekar, R.; Young, G.; Ahmed, S.; Little, D.P. Commercial Teas Highlight Plant DNA Barcode Identification Successes and Obstacles. Sci. Rep. 2011, 1, 42. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wong, K.; Chan, W.; Li, J.; But, P.P.; Cao, H. Establishment of DNA barcodes for the identification of the botanical sources of the Chinese ‘cooling’ beverage. Food Control. 2012, 25, 758–766. [Google Scholar] [CrossRef]
- Lei, H.; Li, S.; Li, J. Study of Traditional She’s Medicine Characteristics and Its Application. Chin. Arch. Tradit. Chin. Med. 2013, 31, 1911–1913. [Google Scholar]
- Zhang, Y. Research on Chinese Tea Culture of Ming Dynasty from Historical Perspective of Health Promotion; China Academy of Chinese Medical Sciences: Beijing, China, 2012. [Google Scholar]
- Huang, X. Studies on the Booming of Tea Culture and Its Effects in Modern Time; Zhejiang University: Hangzhou, China, 2003. [Google Scholar]
- Zhao, S.; Chen, X.; Song, J.; Pang, X.; Chen, S. Internal transcribed spacer 2 barcode: A good tool for identifying Acanthopanacis cortex. Front. Plant Sci. 2015, 6, 840. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).