In Silico Identification of Six Mushroom-Derived Sterol and Triterpenoid Compounds as Potential P-Glycoprotein Modulators in Multidrug Resistance
Abstract
1. Introduction
2. Materials and Methods
2.1. Molecular Docking
2.1.1. Database of Bioactive Mushroom Compounds
2.1.2. Protein Structure Preparation
2.1.3. Redocking and Validation
2.1.4. Virtual Screening
2.2. Toxicity Prediction
2.3. ADME Analysis
3. Results
3.1. Molecular Docking
3.1.1. Protein Structure and Redocking Validation
3.1.2. Virtual Screening
3.2. Toxicity Prediction
3.3. ADME Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Goebel, J.; Chmielewski, J.; Hrycyna, C.A. The roles of the human ATP-binding cassette transporters P-glycoprotein and ABCG2 in multidrug resistance in cancer and at endogenous sites: Future opportunities for structure-based drug design of inhibitors. Cancer Drug Resist. 2021, 4, 784. [Google Scholar] [CrossRef]
- Park, H.J. Current uses of mushrooms in cancer treatment and their anticancer mechanisms. Int. J. Mol. Sci. 2022, 23, 10502. [Google Scholar] [CrossRef]
- Mora Lagares, L.; Pérez-Castillo, Y.; Minovski, N.; Novič, M. Structure–function relationships in the human P-glycoprotein (ABCB1): Insights from molecular dynamics simulations. Int. J. Mol. Sci. 2021, 23, 362. [Google Scholar] [CrossRef]
- Nanayakkara, A.K.; Follit, C.A.; Chen, G.; Williams, N.S.; Vogel, P.D.; Wise, J.G. Targeted inhibitors of P-glycoprotein increase chemotherapeutic-induced mortality of multidrug resistant tumor cells. Sci. Rep. 2018, 8, 967. [Google Scholar] [CrossRef] [PubMed]
- Waghray, D.; Zhang, Q. Inhibit or evade multidrug resistance P-glycoprotein in cancer treatment: Miniperspective. J. Med. Chem. 2017, 61, 5108–5121. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, S.; Islam, M.J.; Parves, M.R.; Khan, M.A.; Tabussum, L.; Ahmed, S.; Ali, M.A.; Fakayode, S.O.; Halim, M.A. Designing potent inhibitors against the multidrug resistance P-glycoprotein. J. Biomol. Struct. Dyn. 2022, 40, 9403–9415. [Google Scholar] [CrossRef] [PubMed]
- Marques, S.M.; Šupolíková, L.; Molčanová, L.; Šmejkal, K.; Bednar, D.; Slaninová, I. Screening of natural compounds as P-glycoprotein inhibitors against multidrug resistance. Biomedicines 2021, 9, 357. [Google Scholar] [CrossRef]
- Yang, Z.; Cai, Y.; Yang, X.; Li, Y.; Wu, Q.; Yu, Y.; Chen, Z.; Wei, B.; Tian, J.M.; Bao, X.; et al. Novel benzo five-membered heterocycle derivatives as P-glycoprotein inhibitors: Design, synthesis, molecular docking, and anti-multidrug resistance activity. J. Med. Chem. 2023, 66, 5550–5566. [Google Scholar] [CrossRef]
- Afonso de Lima, C.; de Souza Bueno, I.L.; Nunes Siqueira Vasconcelos, S.; Sciani, J.M.; Ruiz, A.L.T.G.; Foglio, M.A.; Carvalho, J.E.d.; Barbarini Longato, G. Reversal of ovarian cancer cell lines multidrug resistance phenotype by the association of apiole with chemotherapies. Pharmaceuticals 2020, 13, 327. [Google Scholar] [CrossRef]
- Calhelha, R.C.; Shiraishi, C.S.; Ribeiro, L.; Carocho, M.; Abreu, R.; Coutinho, P.; Barros, L.; Vaz, J.; Ferreira, I.C. New Trends from Fungi Secondary Metabolism in the Pharmaceutical Industry. In Natural Secondary Metabolites: From Nature, Through Science, to Industry; Springer: Berlin/Heidelberg, Germany, 2023; pp. 823–850. [Google Scholar]
- Wang, L.; Sun, Y. Efflux mechanism and pathway of verapamil pumping by human P-glycoprotein. Arch. Biochem. Biophys. 2020, 696, 108675. [Google Scholar] [CrossRef]
- Dewanjee, S.; Dua, T.K.; Bhattacharjee, N.; Das, A.; Gangopadhyay, M.; Khanra, R.; Joardar, S.; Riaz, M.; De Feo, V.; Zia-Ul-Haq, M. Natural products as alternative choices for P-glycoprotein (P-gp) inhibition. Molecules 2017, 22, 871. [Google Scholar] [CrossRef]
- Nagy, H.; Goda, K.; Fenyvesi, F.; Bacsó, Z.; Szilasi, M.; Kappelmayer, J.; Lustyik, G.; Cianfriglia, M.; Szabó Jr, G. Distinct groups of multidrug resistance modulating agents are distinguished by competition of P-glycoprotein-specific antibodies. Biochem. Biophys. Res. Commun. 2004, 315, 942–949. [Google Scholar] [CrossRef]
- Mollazadeh, S.; Sahebkar, A.; Hadizadeh, F.; Behravan, J.; Arabzadeh, S. Structural and functional aspects of P-glycoprotein and its inhibitors. Life Sci. 2018, 214, 118–123. [Google Scholar] [CrossRef]
- Gou, Y.; Zheng, X.; Li, W.; Deng, H.; Qin, S. Polysaccharides produced by the mushroom trametes robiniophila murr boosts the sensitivity of hepatoma cells to oxaliplatin via the miR-224-5p/ABCB1/P-gp Axis. Integr. Cancer Ther. 2022, 21, 15347354221090221. [Google Scholar] [CrossRef] [PubMed]
- Doğan, H.H.; Kars, M.D.; Özdemir, Ö.; Gündüz, U. Fomes fomentarius and Tricholoma anatolicum (Agaricomycetes) extracts exhibit significant multiple drug-resistant modulation activity in drug-resistant breast cancer cells. Int. J. Med. Mushrooms 2020, 22, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.W.; Li, B.; Lai, E.T.C.; Chen, L.; Huang, J.J.H.; Cheung, A.L.M.; Cheung, P.C.K. Water extract from Pleurotus pulmonarius with antioxidant activity exerts in vivo chemoprophylaxis and chemosensitization for liver cancer. Nutr. Cancer 2014, 66, 989–998. [Google Scholar] [CrossRef]
- Teng, Y.N.; Chang, C.S.; Lee, T.E.; Hung, C.C. Cordycepin re-sensitizes multidrug resistance cancer cells to chemotherapeutic agents through modulating P-glycoprotein expression and ATPase function. J. Funct. Foods 2016, 26, 681–690. [Google Scholar] [CrossRef]
- Fonseca, J.; Vaz, J.A.; Ricardo, S. The potential of mushroom extracts to improve chemotherapy efficacy in cancer cells: A systematic review. Cells 2024, 13, 510. [Google Scholar] [CrossRef]
- Shin, M.J.; Chae, H.J.; Lee, J.W.; Koo, M.H.; Kim, H.J.; Seo, J.B.; YanIllia, S.; Park, S.H.; Lo, H.E.; Kim, S.H.; et al. Lucidumol A, Purified Directly from Ganoderma lucidum, Exhibits Anticancer Effect and Cellular Inflammatory Response in Colorectal Cancer. Evid.-Based Complement. Altern. Med. 2022, 2022, 7404493. [Google Scholar] [CrossRef]
- CFR Ferreira, I.; Vaz, J.A.; Vasconcelos, M.H.; Martins, A. Compounds from wild mushrooms with antitumor potential. Anti-Cancer Agents Med. Chem.-Anti Agents 2010, 10, 424–436. [Google Scholar] [CrossRef]
- Borges, B.F. Preparação e Screening Virtual de uma Biblioteca de Compostos de Baixo Peso Molecular Oriundos de Cogumelos Contra Proteínas da Família BCL-2. Master’s Thesis, Instituto Politecnico de Braganca (Portugal), Bragança, Portugal, 2018. [Google Scholar]
- Shiraishi, C.S.H. Estudo do Potencial Anti-inflamatório de Uma Biblioteca de Compostos Naturais de Cogumelos por Screening Virtual Contra as Enzimas Cox (-1 E-2). Master’s Thesis, Instituto Politecnico de Braganca (Portugal), Bragança, Portugal, 2020. [Google Scholar]
- Li, W.; Zhang, H.; Assaraf, Y.G.; Zhao, K.; Xu, X.; Xie, J.; Yang, D.H.; Chen, Z.S. Overcoming ABC transporter-mediated multidrug resistance: Molecular mechanisms and novel therapeutic drug strategies. Drug Resist. Updat. 2016, 27, 14–29. [Google Scholar] [CrossRef] [PubMed]
- Thomas, H.; Coley, H.M. Overcoming multidrug resistance in cancer: An update on the clinical strategy of inhibiting p-glycoprotein. Cancer Control 2003, 10, 159–165. [Google Scholar] [CrossRef]
- Gagliano, T.; Gentilin, E.; Benfini, K.; Di Pasquale, C.; Tassinari, M.; Falletta, S.; Feo, C.; Tagliati, F.; Uberti, E.d.; Zatelli, M.C. Mitotane enhances doxorubicin cytotoxic activity by inhibiting P-gp in human adrenocortical carcinoma cells. Endocrine 2014, 47, 943–951. [Google Scholar] [CrossRef]
- Consoli, U.; Priebe, W.; Ling, Y.H.; Mahadevia, R.; Griffin, M.; Zhao, S.; Perez-Soler, R.; Andreeff, M. The novel anthracycline annamycin is not affected by P-glycoprotein-related multidrug resistance: Comparison with idarubicin and doxorubicin in HL-60 leukemia cell lines. Blood 1996, 88, 633–644. [Google Scholar] [CrossRef]
- Sander, T.; Freyss, J.; Von Korff, M.; Rufener, C. DataWarrior: An open-source program for chemistry aware data visualization and analysis. J. Chem. Inf. Model. 2015, 55, 460–473. [Google Scholar] [CrossRef]
- Land, H.; Humble, M.S. YASARA: A tool to obtain structural guidance in biocatalytic investigations. In Protein Engineering: Methods and Protocols; Humana Press: New York, NY, USA, 2018; pp. 43–67. [Google Scholar]
- Nosol, K.; Romane, K.; Irobalieva, R.N.; Alam, A.; Kowal, J.; Fujita, N.; Locher, K.P. Cryo-EM structures reveal distinct mechanisms of inhibition of the human multidrug transporter ABCB1. Proc. Natl. Acad. Sci. USA 2020, 117, 26245–26253. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A. Software news and update AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function. Effic. Optim. Multithreading 2009, 31, 455–461. [Google Scholar]
- Bell, E.W.; Zhang, Y. DockRMSD: An open-source tool for atom mapping and RMSD calculation of symmetric molecules through graph isomorphism. J. Cheminform. 2019, 11, 40. [Google Scholar] [CrossRef]
- Schrödinger, LLC. The PyMOL Molecular Graphics System, Version 1.8; Schrödinger, LLC: New York, NY, USA, 2015. [Google Scholar]
- Schöning-Stierand, K.; Diedrich, K.; Ehrt, C.; Flachsenberg, F.; Graef, J.; Sieg, J.; Penner, P.; Poppinga, M.; Ungethüm, A.; Rarey, M. Proteins Plus: A comprehensive collection of web-based molecular modeling tools. Nucleic Acids Res. 2022, 50, W611–W615. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, P.; Kemmler, E.; Dunkel, M.; Preissner, R. ProTox 3.0: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2024, 52, W513–W520. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Ma, J.; Li, M.; Zhang, Y.; Jiang, B.; Zhao, X.; Huai, C.; Shen, L.; Zhang, N.; He, L.; et al. Cytochrome P450 enzymes and drug metabolism in humans. Int. J. Mol. Sci. 2021, 22, 12808. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef]
- Toledo, B.; González-Titos, A.; Hernández-Camarero, P.; Perán, M. A brief review on chemoresistance; targeting cancer stem cells as an alternative approach. Int. J. Mol. Sci. 2023, 24, 4487. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Lei, Y.; Wang, Y.; Lai, J.; Wang, J.; Xia, F. Mechanism of multidrug resistance to chemotherapy mediated by P-glycoprotein. Int. J. Oncol. 2023, 63, 119. [Google Scholar] [CrossRef]
- Emran, T.B.; Shahriar, A.; Mahmud, A.R.; Rahman, T.; Abir, M.H.; Siddiquee, M.F.R.; Ahmed, H.; Rahman, N.; Nainu, F.; Wahyudin, E.; et al. Multidrug resistance in cancer: Understanding molecular mechanisms, immunoprevention and therapeutic approaches. Front. Oncol. 2022, 12, 891652. [Google Scholar] [CrossRef]
- Pinzi, L.; Rastelli, G. Molecular docking: Shifting paradigms in drug discovery. Int. J. Mol. Sci. 2019, 20, 4331. [Google Scholar] [CrossRef]
- Agu, P.; Afiukwa, C.; Orji, O.; Ezeh, E.; Ofoke, I.; Ogbu, C.; Ugwuja, E.; Aja, P. Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Sci. Rep. 2023, 13, 13398. [Google Scholar] [CrossRef]
- Wahono, C.S.; Syaban, M.F.R.; Pratama, M.Z.; Rahman, P.A.; Erwan, N.E. Exploring the potential of phytoconstituents from Phaseolus vulgaris L against CXC motif chemokine receptor 4 (CXCR4): A bioinformatic and molecular dynamic simulations approach. Egypt. J. Med. Hum. Genet. 2024, 25, 52. [Google Scholar] [CrossRef]
- Castro-Alvarez, A.; Costa, A.M.; Vilarrasa, J. The performance of several docking programs at reproducing protein–macrolide-like crystal structures. Molecules 2017, 22, 136. [Google Scholar] [CrossRef]
- Chen, Y.M.; Sung, H.C.; Kuo, Y.H.; Hsu, Y.J.; Huang, C.C.; Liang, H.L. The Effects of Ergosta-7, 9 (11), 22-trien-3β-ol from Antrodia camphorata on the Biochemical Profile and Exercise Performance of Mice. Molecules 2019, 24, 1225. [Google Scholar] [CrossRef]
- Ma, J.Q.; Zhang, Y.J.; Tian, Z.K. Anti-oxidant, anti-inflammatory and anti-fibrosis effects of ganoderic acid A on carbon tetrachloride induced nephrotoxicity by regulating the Trx/TrxR and JAK/ROCK pathway. Chem.-Biol. Interact. 2021, 344, 109529. [Google Scholar] [CrossRef] [PubMed]
- Shao, G.; He, J.; Meng, J.; Ma, A.; Geng, X.; Zhang, S.; Qiu, Z.; Lin, D.; Li, M.; Zhou, H.; et al. Ganoderic acids prevent renal ischemia reperfusion injury by inhibiting inflammation and apoptosis. Int. J. Mol. Sci. 2021, 22, 10229. [Google Scholar] [CrossRef]
- Chen, S.Y.; Chang, C.L.; Chen, T.H.; Chang, Y.W.; Lin, S.B. Colossolactone H, a new Ganoderma triterpenoid exhibits cytotoxicity and potentiates drug efficacy of gefitinib in lung cancer. Fitoterapia 2016, 114, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Marei, H.E.; Althani, A.; Afifi, N.; Hasan, A.; Caceci, T.; Pozzoli, G.; Morrione, A.; Giordano, A.; Cenciarelli, C. p53 signaling in cancer progression and therapy. Cancer Cell Int. 2021, 21, 703. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Kang, N.; Song, E.; Wie, M.; Lee, E.A.; Hwang, S.; Lee, D.; Ra, J.S.; Park, I.B.; Park, J.; et al. ATAD5 promotes replication restart by regulating RAD51 and PCNA in response to replication stress. Nat. Commun. 2019, 10, 5718. [Google Scholar] [CrossRef]
- Wong, I.L.; Wang, B.C.; Yuan, J.; Duan, L.X.; Liu, Z.; Liu, T.; Li, X.M.; Hu, X.; Zhang, X.Y.; Jiang, T.; et al. Potent and nontoxic chemosensitizer of P-glycoprotein-mediated multidrug resistance in cancer: Synthesis and evaluation of methylated epigallocatechin, gallocatechin, and dihydromyricetin derivatives. J. Med. Chem. 2015, 58, 4529–4549. [Google Scholar] [CrossRef]
- Wu, Q.P.; Xie, Y.Z.; Deng, Z.; Li, X.M.; Yang, W.; Jiao, C.W.; Fang, L.; Li, S.Z.; Pan, H.H.; Yee, A.J.; et al. Ergosterol peroxide isolated from Ganoderma lucidum abolishes microRNA miR-378-mediated tumor cells on chemoresistance. PLoS ONE 2012, 7, e44579. [Google Scholar] [CrossRef]
- Li, J.; Cao, L.; Yuan, C.; Jiang, Z.; Cai, H.; Xu, W.; Han, Y.; Chen, L.; Zhang, Q.; Jiang, R.; et al. Ganoderma lucidum extract reverses hepatocellular carcinoma multidrug resistance via inhibiting the function of P-glycoprotein in vitro and in vivo. Ital. J. Food Sci. 2023, 35, 90–98. [Google Scholar] [CrossRef]
- Zhao, Y.Y.; Shen, X.; Chao, X.; Ho, C.C.; Cheng, X.L.; Zhang, Y.; Lin, R.C.; Du, K.J.; Luo, W.J.; Chen, J.Y.; et al. Ergosta-4, 6, 8 (14), 22-tetraen-3-one induces G2/M cell cycle arrest and apoptosis in human hepatocellular carcinoma HepG2 cells. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2011, 1810, 384–390. [Google Scholar] [CrossRef]
- Chang, C.W.; Chen, Y.S.; Chen, C.C.; Chan, I.O.; Chen, C.C.; Sheu, S.J.; Lin, T.w.; Chou, S.H.; Liu, C.J.; Lee, T.C.; et al. Targeting cancer initiating cells by promoting cell differentiation and restoring chemosensitivity via dual inactivation of STAT3 and src activity using an active component of antrodia cinnamomea mycelia. Oncotarget 2016, 7, 73016. [Google Scholar] [CrossRef] [PubMed]
- Amen, Y.M.; Zhu, Q.; Tran, H.B.; Afifi, M.S.; Halim, A.F.; Ashour, A.; Mira, A.; Shimizu, K. Lucidumol C, a new cytotoxic lanostanoid triterpene from Ganoderma lingzhi against human cancer cells. J. Nat. Med. 2016, 70, 661–666. [Google Scholar] [CrossRef] [PubMed]



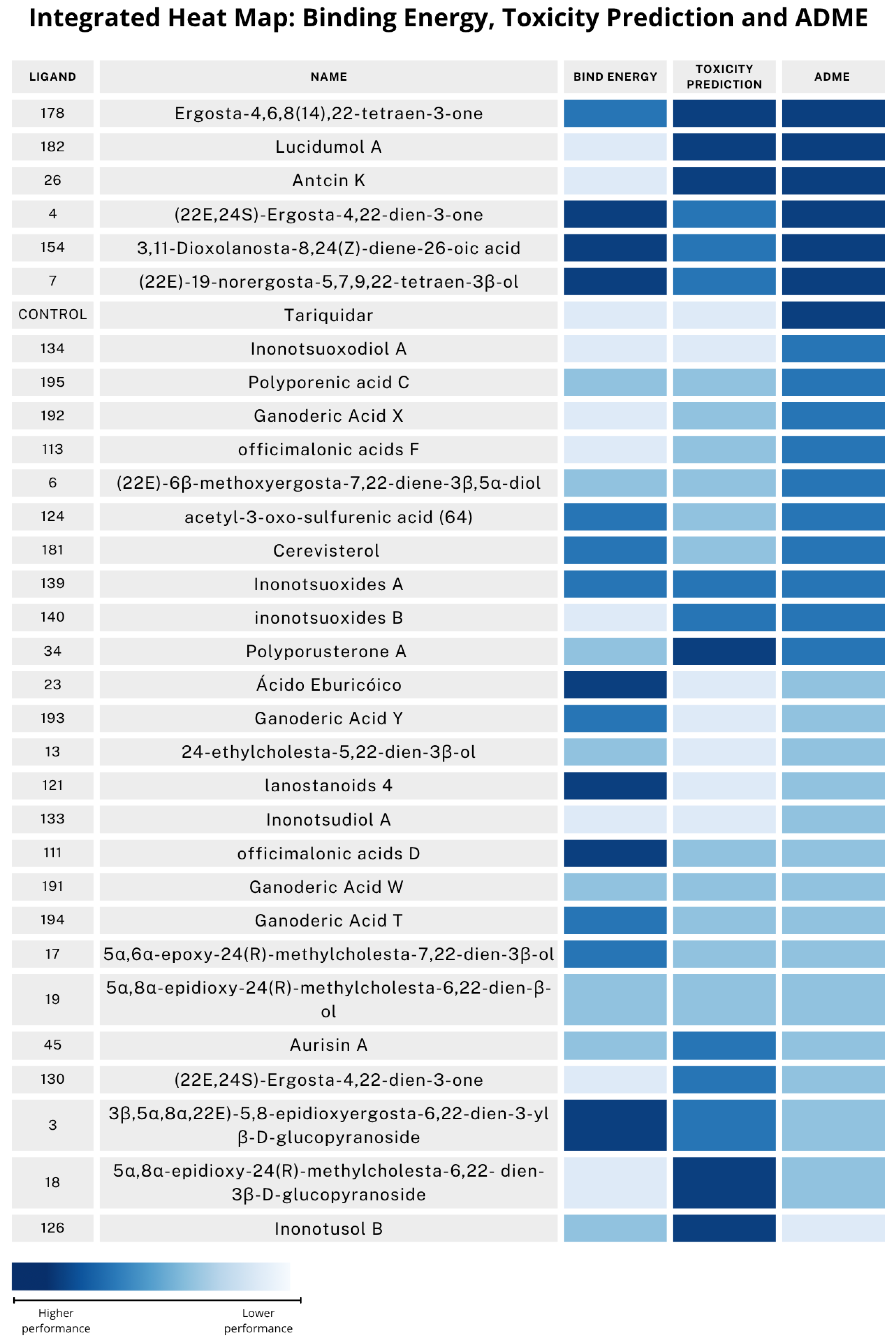
| Compound | Name | Predicted Binding Energy [kcal/mol] | Con.Surf [] |
|---|---|---|---|
| 121 | Lanostanoid 4 | −13.19 | 470.73 |
| 120 | Lanostanoid 3 | −12.67 | 495.39 |
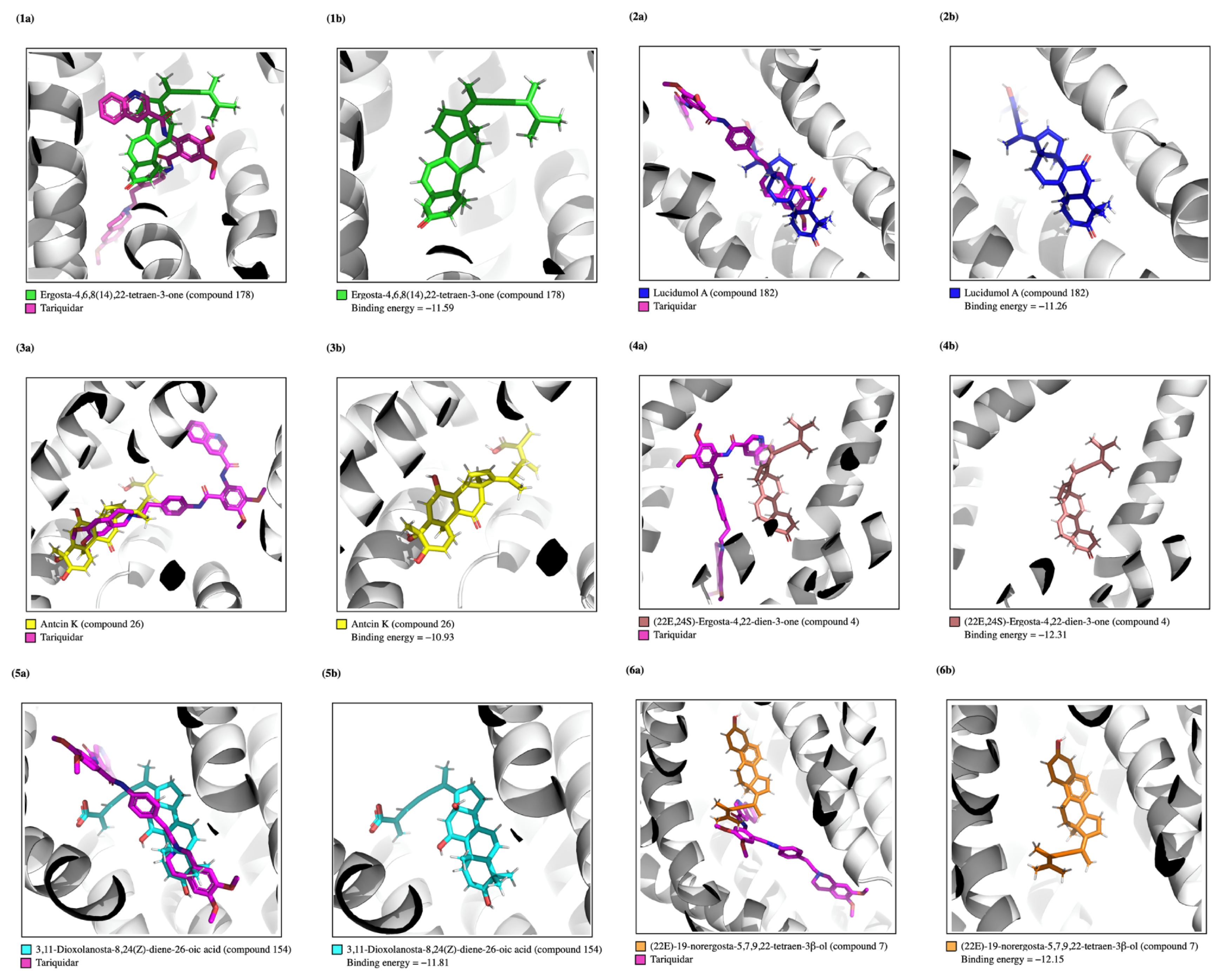
| 4 | (22E,24S)-Ergosta-4,22-dien-3-one | −12.31 | 461.12 |
| 177 | Ergosterol | −12.27 | 462.35 |
| 68 | Inoscavin A | −12.22 | 445.33 |
| 7 | (22E)-19-Norergosta-5,7,9,22-tetraen-3-ol | −12.15 | 384.84 |
| 111 | Officimalonic acids D | −12.09 | 565.94 |
| 1 | +E4:F38(3,5,8,22E,24S)-5,8-Epidioxyergosta-6,9(11),22-trien-3-ol | −12.07 | 448.19 |
| 21 | 9,11-Dehydroergosterol peroxide | −12.04 | 449.10 |
| 10 | (22E)-Ergosta-5,7,22-trien-3-ol | −12.01 | 457.70 |
| 9 | (22E)-5,6-Epoxyergosta-8,14,22-triene-3,7-diol | −11.98 | 447.32 |
| 3 | 3,5,8,22E)-5,8-Epidioxyergosta-6,22-dien-3-yl-D-glucopyranoside | −11.93 | 506.27 |
| 12 | (3,5,22E)-Ergosta-7,22,24(28)-trien-3-ol | −11.92 | 455.67 |
| 23 | Eburicoic acid | −11.91 | 467.92 |
| 32 | Ergosterol peroxide | −11.87 | 465.10 |
| 168 | 3-Oxo-24-methyl-5-lanost-8,25-dien-21-oic acid | −11.86 | 476.27 |
| 171 | 22E-5-Ergost-7,9(11),22-trien-3-ol | −11.84 | 466.39 |
| 154 | 3,11-Dioxolanosta-8,24(Z)-diene-26-oic acid | −11.81 | 444.14 |
| 194 | Ganoderic Acid T | −11.74 | 587.44 |
| 124 | Acetyl-3-oxo-sulfurenic acid (64) | −11.74 | 512.85 |
| 17 | 5,6-Epoxy-24(R)-methylcholesta-7,22-dien-3-ol | −11.73 | 437.11 |
| 190 | Ganoderic Acid H | −11.66 | 503.58 |
| 112 | Officimalonic acids E | −11.66 | 550.17 |
| 48 | Colossolactone H (Colo H) | −11.64 | 469.63 |
| 178 | Ergosta-4,6,8(14),22-tetraen-3-one | −11.59 | 367.21 |
| 114 | Officimalonic acids G | −11.52 | 562.15 |
| 193 | Ganoderic Acid Y | −11.49 | 459.87 |
| 20 | 5,6-Epoxyergosta-8(14)-ene-3,7-diol | −11.48 | 455.29 |
| 139 | Inonotsuoxides A | −11.48 | 438.71 |
| 181 | Cerevisterol | −11.46 | 441.80 |
| 13 | 24-Ethylcholesta-5,22-dien-3-ol | −11.33 | 390.82 |
| 28 | Estelasterol | −11.31 | 408.88 |
| 191 | Ganoderic Acid W | −11.30 | 533.11 |
| 19 | 5,8-Epidioxy-24(R)-methylcholesta-6,22-dien--ol | −11.29 | 426.07 |
| 195 | Polyporenic acid C | −11.28 | 472.30 |
| 182 | Lucidumol A | −11.26 | 507.86 |
| 144 | Chagabusone A | −11.23 | 466.75 |
| 34 | Polyporusterone A | −11.22 | 466.28 |
| 45 | Aurisin A | −11.18 | 429.27 |
| 126 | Inonotusol B | −11.14 | 443.88 |
| 6 | (22E)-6-Methoxyergosta-7,22-diene-3,5-diol | −11.11 | 463.33 |
| 132 | Spiroinonotsuoxodiol | −11.06 | 469.46 |
| 192 | Ganoderic Acid X | −11.04 | 492.63 |
| 27 | Eringiacetal A | −11.00 | 388.48 |
| 189 | Ganoderic acid F | −11.00 | 481.37 |
| 110 | Officimalonic acids C | −10.99 | 554.09 |
| 118 | Lanostanoid 1 (58) | −10.95 | 472.98 |
| 133 | Inonotsudiol A | −10.95 | 455.79 |
| 26 | Antcin K | −10.93 | 496.48 |
| 140 | Inonotsuoxides B | −10.92 | 426.16 |
| 102 | Piptolinic acid E | −10.92 | 487.53 |
| 8 | (22E)-3,5,9-Trihydroxyergosta-7,22-dien-6-one | −10.91 | 410.35 |
| 18 | 5,8-Epidioxy-24(R)-methylcholesta-6,22-dien-3-D-glucopyranoside | −10.88 | 554.78 |
| 130 | Inonotusol F | −10.87 | 451.37 |
| 113 | Officimalonic acids F | −10.87 | 549.20 |
| 11 | (22E)-Ergosta-7,22-dien-3-ol | −10.86 | 425.94 |
| 119 | Lanostanoid 2 | −10.85 | 502.87 |
| 134 | Inonotsuoxodiol A | −10.79 | 478.10 |
| 22 | Dehydroeburicoic acid | −10.79 | 480.87 |
| Control | Tariquidar | −10.78 | 633.80 |
| Compound | Name | Predicted Toxicity Class | Predicted LD50 | Final Score |
|---|---|---|---|---|
| 178 | Ergosta-4,6,8(14),22-tetraen-3-one | 6 | 10,000 mg/kg | 309 |
| 18 | 5,8-Epidioxy-24(R)-methylcholesta-6,22- dien-3-D-glucopyranoside | 6 | 39,800 mg/kg | 309 |
| 34 | Polyporusterone A | 6 | 9000 mg/kg | 307 |
| 182 | Lucidumol A | 6 | 5010 mg/kg | 306 |
| 26 | Antcin K | 6 | 9000 mg/kg | 305 |
| 126 | Inonotusol B | 6 | 9000 mg/kg | 303 |
| 3 | 3,5,8,22E)-5,8-Epidioxyergosta-6,22-dien-3-yl-D-glucopyranoside | 5 | 4000 mg/kg | 264 |
| 4 | (22E,24S)-Ergosta-4,22-dien-3-one | 5 | 2300 mg/kg | 263 |
| 130 | Inonotusol F | 5 | 5000 mg/kg | 263 |
| 154 | 3,11-Dioxolanosta-8,24(Z)-diene-26-oic acid | 5 | 3389 mg/kg | 262 |
| 139 | Inonotsuoxides A | 5 | 3520 mg/kg | 262 |
| 45 | Aurisin A | 5 | 2500 mg/kg | 262 |
| 140 | Inonotsuoxides B | 5 | 3520 mg/kg | 262 |
| 124 | Acetyl-3-oxo-sulfurenic acid (64) | 5 | 5000 mg/kg | 261 |
| 17 | 5,6-Epoxy-24(R)-methylcholesta-7,22-dien-3-ol | 5 | 5000 mg/kg | 261 |
| 181 | Cerevisterol | 5 | 2340 mg/kg | 261 |
| 19 | 5,8-Epidioxy-24(R)-methylcholesta-6,22-dien--ol | 5 | 5000 mg/kg | 261 |
| 6 | (22E)-6-Methoxyergosta-7,22-diene-3,5-diol | 5 | 2340 mg/kg | 260 |
| 194 | Ganoderic Acid T | 5 | 3000 mg/kg | 259 |
| 111 | Officimalonic acids D | 5 | 3267 mg/kg | 258 |
| 191 | Ganoderic Acid W | 5 | 3000 mg/kg | 258 |
| 192 | Ganoderic Acid X | 5 | 5000 mg/kg | 258 |
| 113 | Officimalonic acids F | 5 | 3267 mg/kg | 258 |
| 195 | Polyporenic acid C | 5 | 5000 mg/kg | 257 |
| 121 | Lanostanoid 4 | 4 | 1000 mg/kg | 218 |
| 7 | (22E)-19-Norergosta-5,7,9,22-tetraen-3-ol | 4 | 590 mg/kg | 218 |
| 133 | Inonotsudiol A | 4 | 2000 mg/kg | 218 |
| 134 | Inonotsuoxodiol A | 4 | 1000 mg/kg | 218 |
| 23 | Eburicoic acid | 4 | 1000 mg/kg | 217 |
| 193 | Ganoderic Acid Y | 4 | 1000 mg/kg | 217 |
| 13 | 24-Ethylcholesta-5,22-dien-3-ol | 4 | 890 mg/kg | 217 |
| Control | Tariquidar | 4 | 1570 mg/kg | 217 |
| Compound | Name | P-gp Substrate | GI Absorption50 | BBB Permeant | Lipinski’s Rule of Five | ADME Score |
|---|---|---|---|---|---|---|
| 178 | Ergosta-4,6,8(14),22-tetraen-3-one | No | High | Yes | Yes; 1 violation: | 4 |
| 182 | Lucidumol A | Yes | High | No | Yes; 0 violations | 4 |
| 26 | Antcin K | Yes | High | No | Yes; 0 violations | 4 |
| 4 | (22E,24S)-Ergosta-4,22-dien-3-one | No | High | Yes | Yes; 1 violation: | 4 |
| 154 | 3,11-Dioxolanosta-8,24(Z)-diene-26-oic acid | Yes | High | No | Yes; 0 violations | 4 |
| 7 | (22E)-19-Norergosta-5,7,9,22-tetraen-3-ol | No | High | Yes | Yes; 1 violation: | 4 |
| Control | Tariquidar | No | Low | No | Yes; 1 violation: | 4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fonseca, J.; Shiraishi, C.S.H.; Abreu, R.M.V.; Ricardo, S.; Vaz, J.A. In Silico Identification of Six Mushroom-Derived Sterol and Triterpenoid Compounds as Potential P-Glycoprotein Modulators in Multidrug Resistance. Appl. Sci. 2025, 15, 8772. https://doi.org/10.3390/app15168772
Fonseca J, Shiraishi CSH, Abreu RMV, Ricardo S, Vaz JA. In Silico Identification of Six Mushroom-Derived Sterol and Triterpenoid Compounds as Potential P-Glycoprotein Modulators in Multidrug Resistance. Applied Sciences. 2025; 15(16):8772. https://doi.org/10.3390/app15168772
Chicago/Turabian StyleFonseca, Jéssica, Carlos S. H. Shiraishi, Rui M. V. Abreu, Sara Ricardo, and Josiana A. Vaz. 2025. "In Silico Identification of Six Mushroom-Derived Sterol and Triterpenoid Compounds as Potential P-Glycoprotein Modulators in Multidrug Resistance" Applied Sciences 15, no. 16: 8772. https://doi.org/10.3390/app15168772
APA StyleFonseca, J., Shiraishi, C. S. H., Abreu, R. M. V., Ricardo, S., & Vaz, J. A. (2025). In Silico Identification of Six Mushroom-Derived Sterol and Triterpenoid Compounds as Potential P-Glycoprotein Modulators in Multidrug Resistance. Applied Sciences, 15(16), 8772. https://doi.org/10.3390/app15168772







