Using Level-Based Multiple Reasoning in a Web-Based Intelligent System for the Diagnosis of Farmed Fish Diseases
Abstract
:1. Introduction
- A novel knowledge acquisition and representation method.
- Introduction of a level-based diagnostic process for farmed fish diseases.
- Introduction of a novel integration of reasoning approaches for disease diagnosis.
- Integration of an image recognition system in the diagnosis process.
2. Background Knowledge
2.1. Rule-Based Representation and Reasoning
then h
then fish-disease is ceratothoa
2.2. Rules with Certainty Factors
then fish-disease is ceratothoa (0.7)
2.3. Probabilistic Reasoning with Bayes Theorem
then h (hypothesis)
2.4. Case-Based Reasoning
2.5. Ontology-Based Representation and Reasoning
- Concepts (domain entities organized in hierarchies, regarded as classes)
- Relationships (between concepts)
- Properties (attributes of the concepts that define them)
- Axioms or Rules (logical statements that hold true in the domain)
- Individuals (specific entities, regarded as instances of classes)
- Class hierarchy reasoning
- Concept subsumption
- Property restriction inference
- Consistency checking
- Rule-based reasoning
- Logic-based reasoning
3. Related Work
4. Materials and Methods
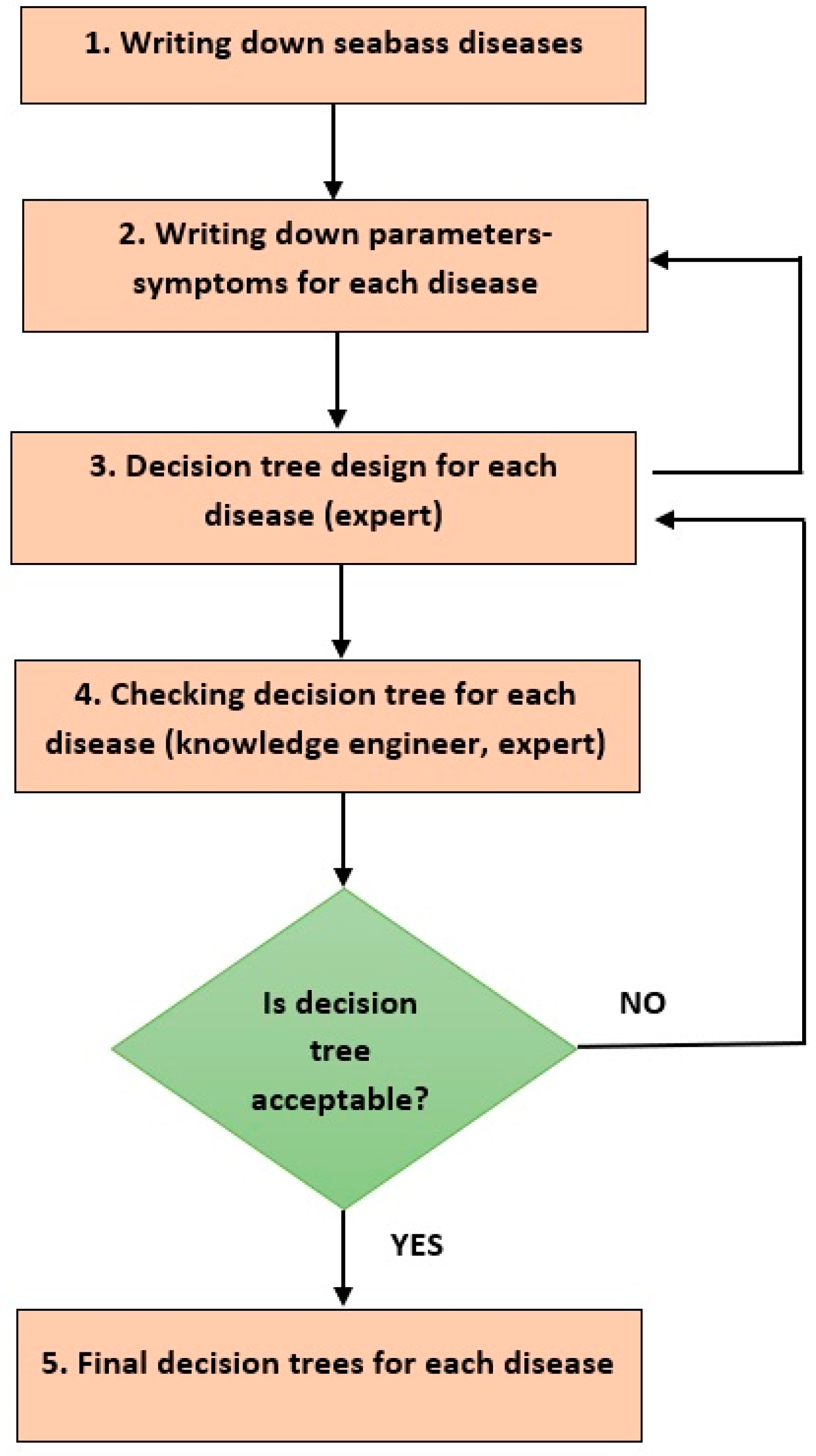
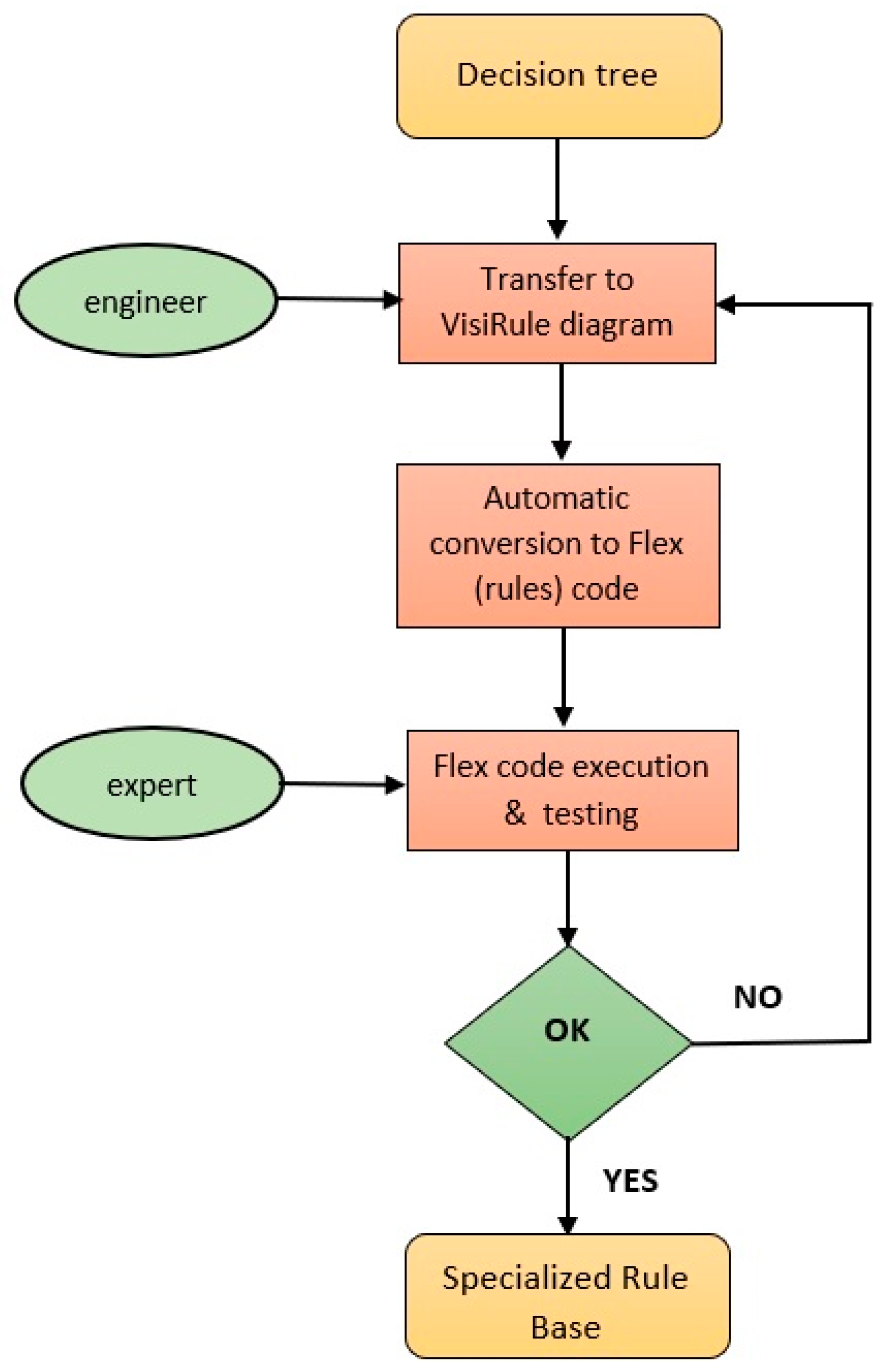
4.1. Knowledge Acquisition and Representation
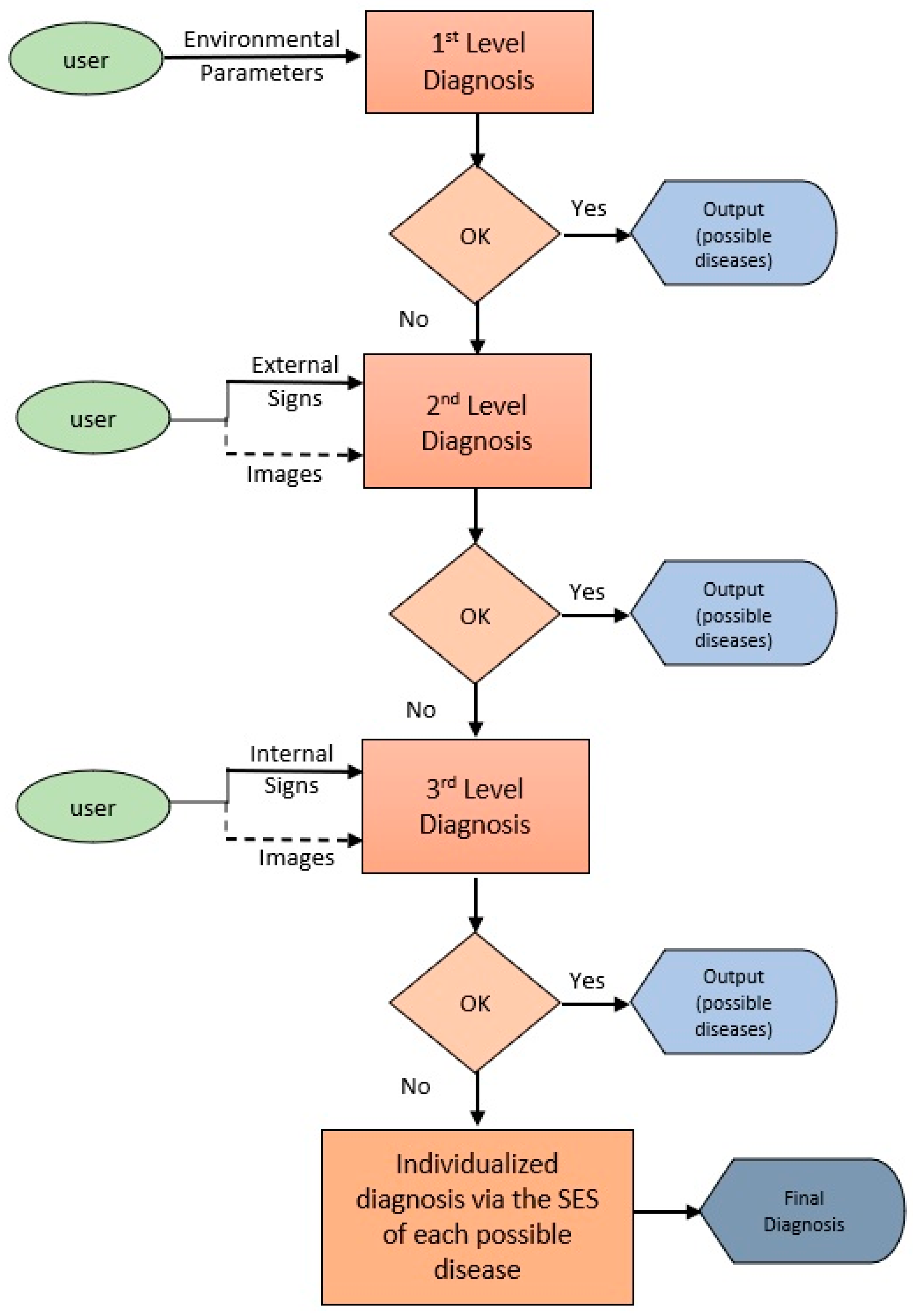
4.2. Diagnostic Process
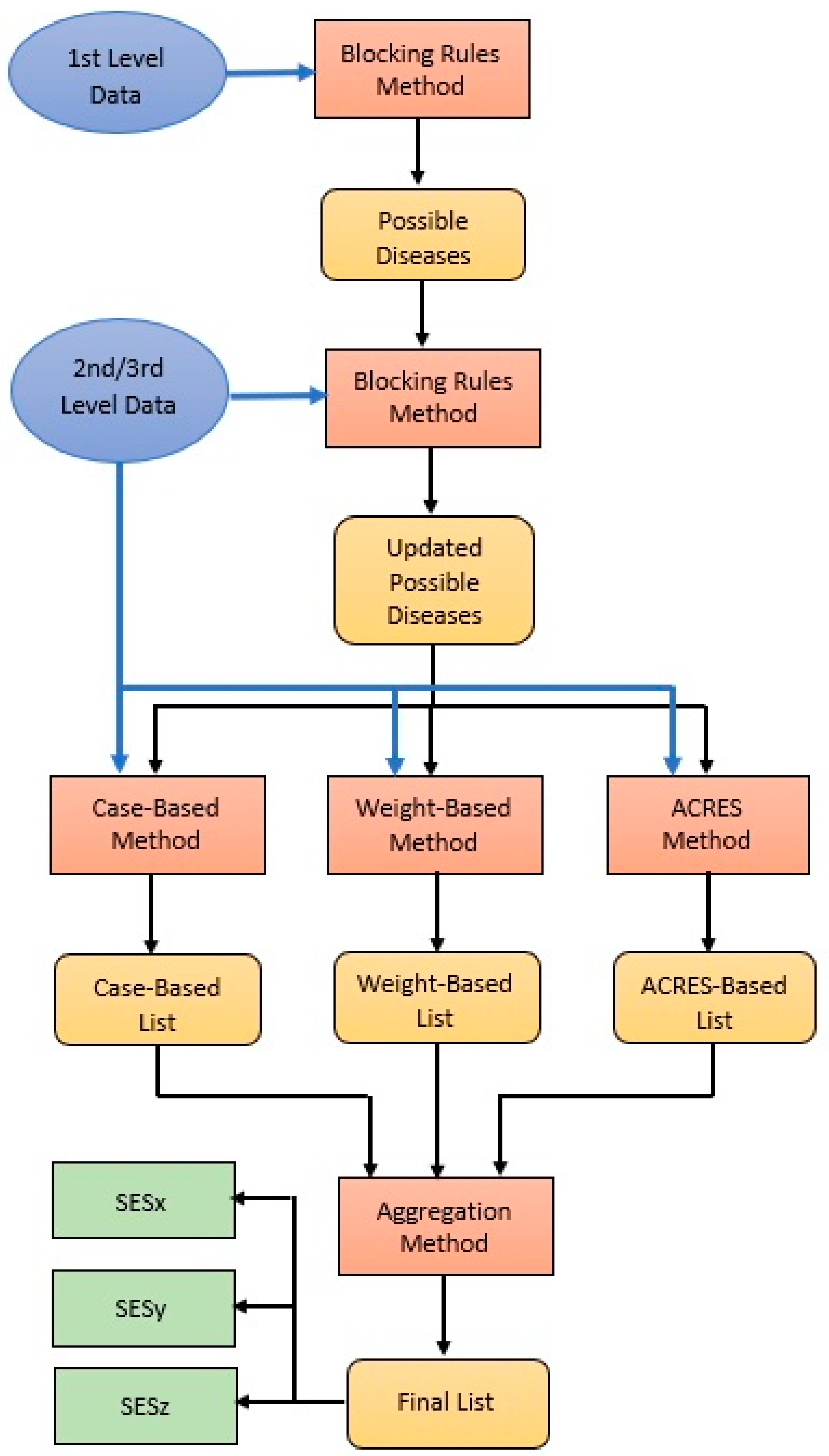
4.3. Diagnostic Methods
4.3.1. Blocking Rules
4.3.2. Case Based Ordering (CBO)
- a: number of common elements in the two lists
- b: number of elements present only in the user list (user provided symptoms)
- c: number of items present only in the disease list (disease symptoms)
kulcz2 = 0, if ((a = b = 0 ≠ c) or (a = c = 0 ≠ b))
kulcz2 = [(a/(a + b)) + (a/(a + c))], otherwise
4.3.3. Method of Weights (WM)
4.3.4. The ACRES Method
4.3.5. Aggregation Method (AGM)
- The most common of the three first ranked diseases in the three lists gets first in the final list. If all the first three are different, majority voting is applied to the three first ranked plus the three second ranked, and so on.
- To determine the second disease in the final list, we apply majority voting to the first three and the second three diseases in the lists after removing the occurrences of the disease selected as first in the final list in the previous step in the same way.
- This continues until the third most probable disease on the final list is determined.
4.3.6. Overall Diagnostic Process
| Algorithm 1: Overall Diagnostic Process |
| input: L = [d1, d2, …, dn]: all diseases list, BRx: blocking rules of level x ϵ {1,2,3}, x = 1: level counter, exit = false output: L = [dp1, dp2, …, dpm]: ordered possible diseases
|
4.3.7. Specialized Expert Systems Based Diagnosis
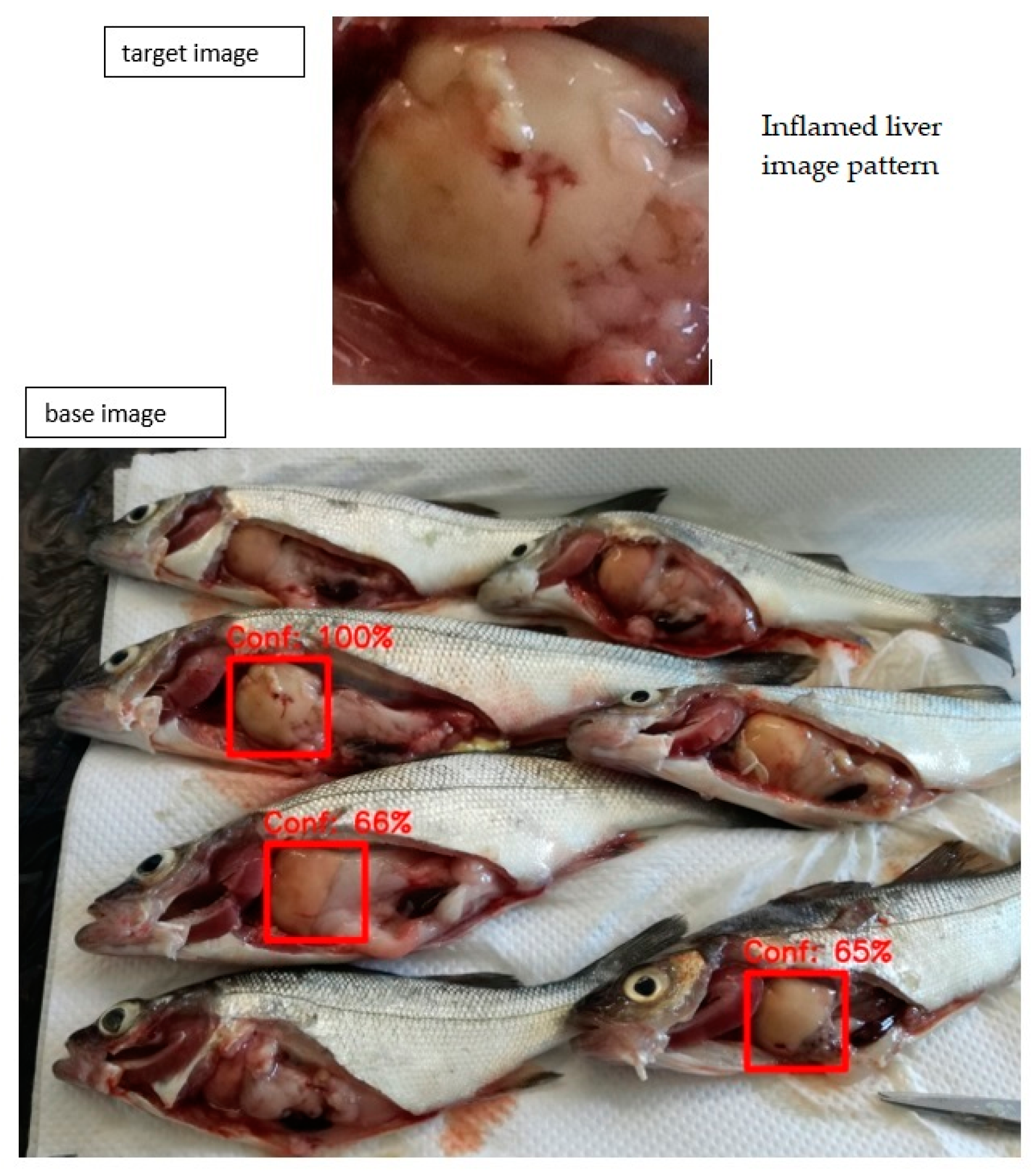
4.4. Image Recognition System (IRS)
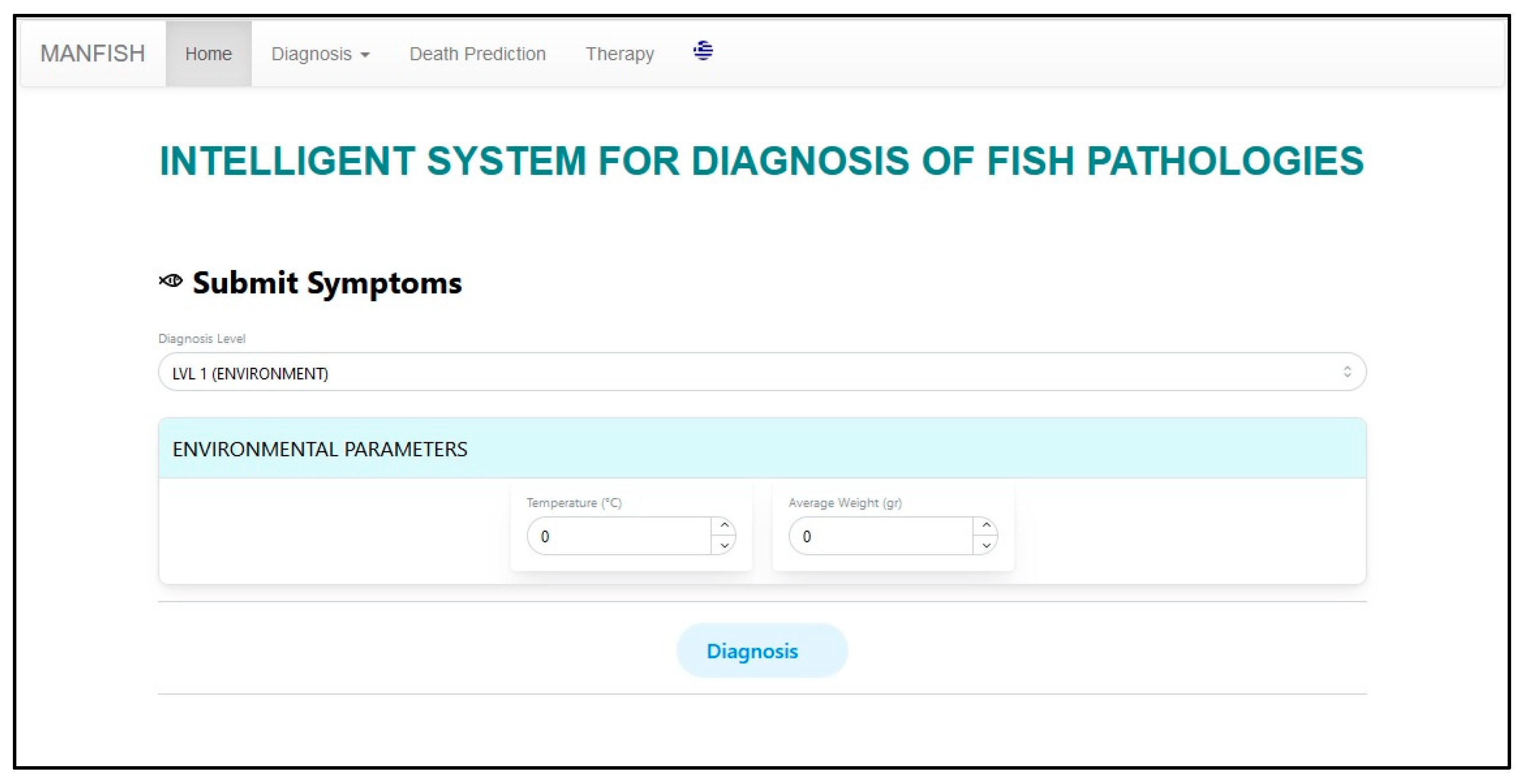
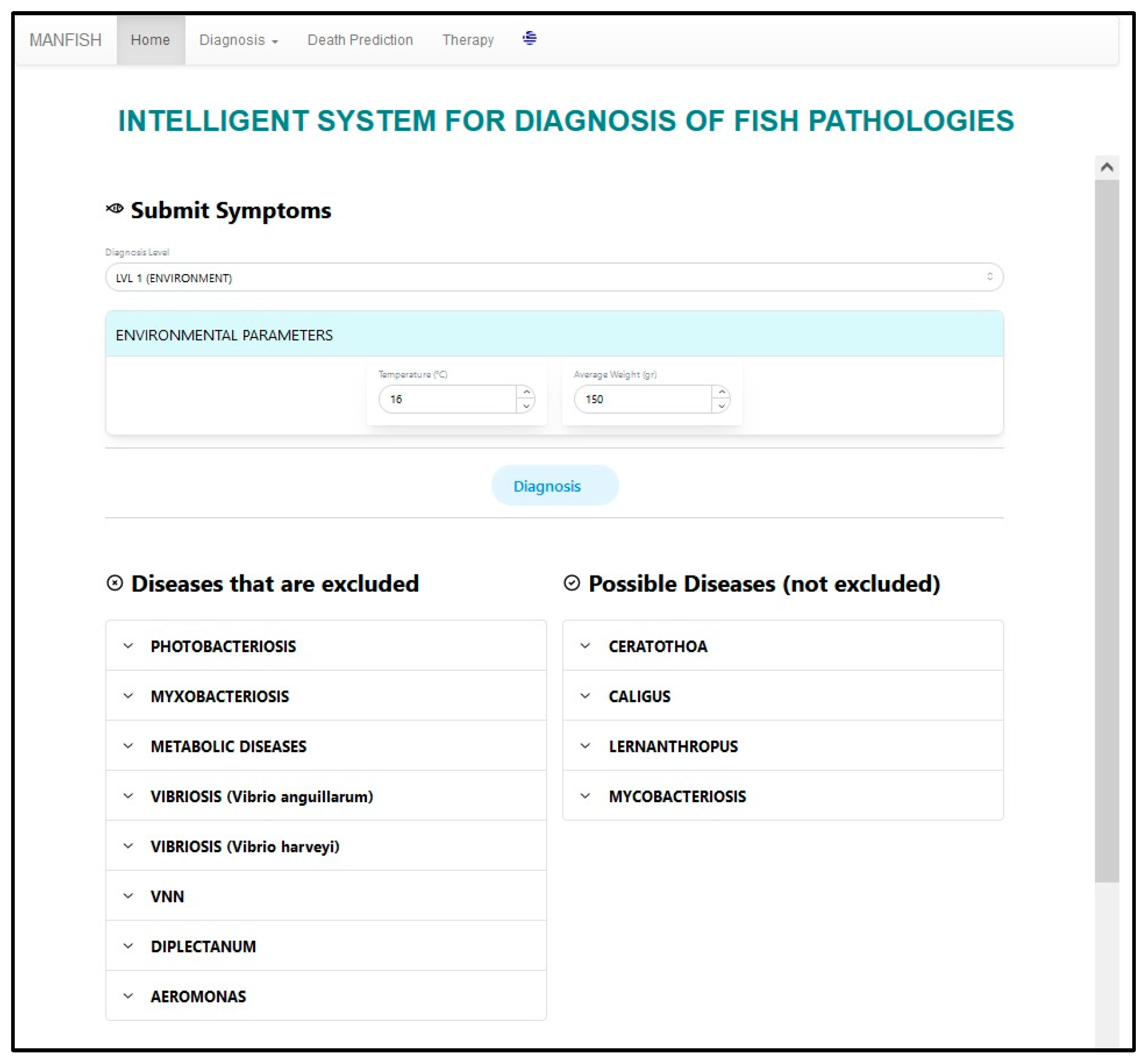
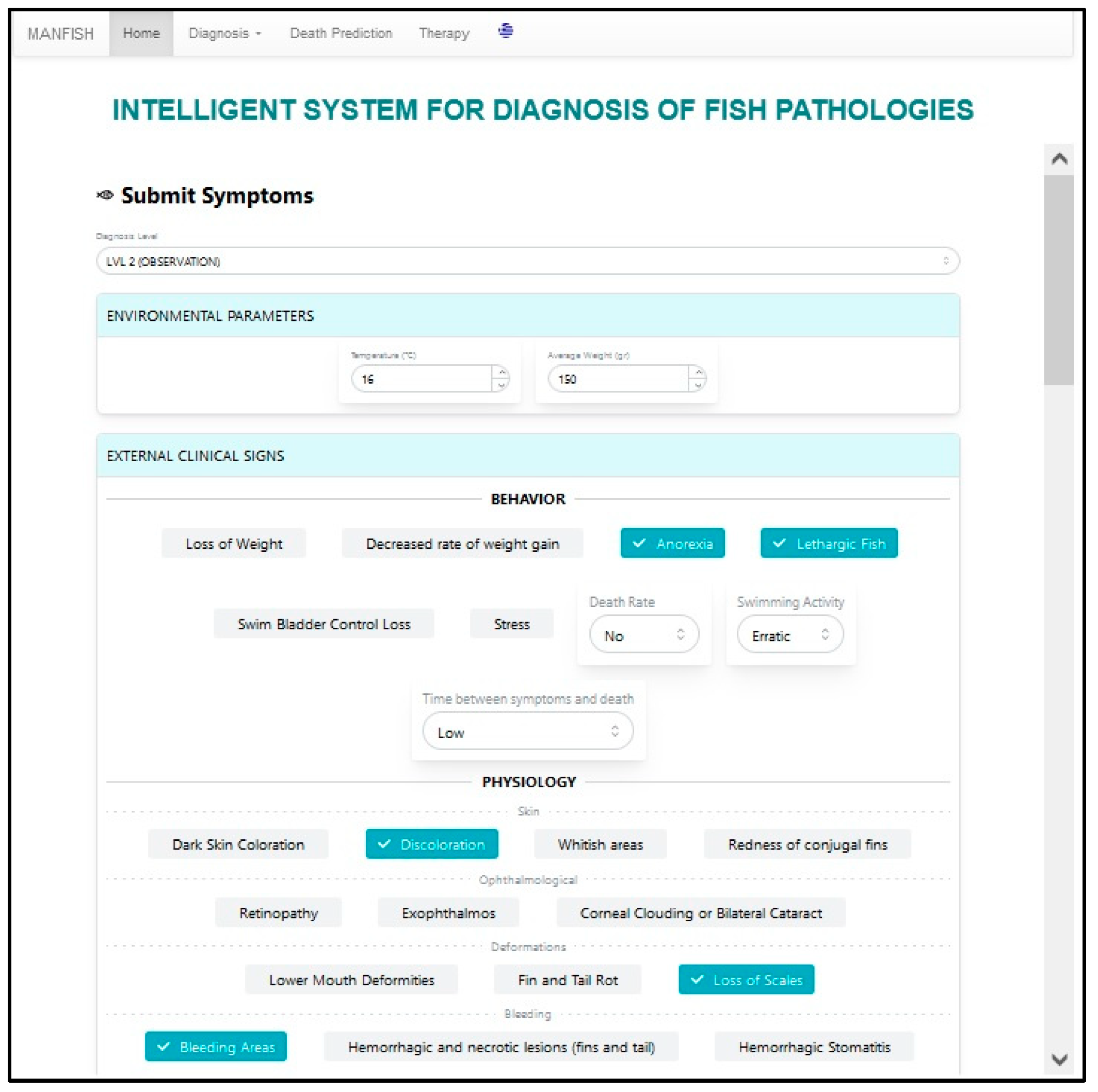
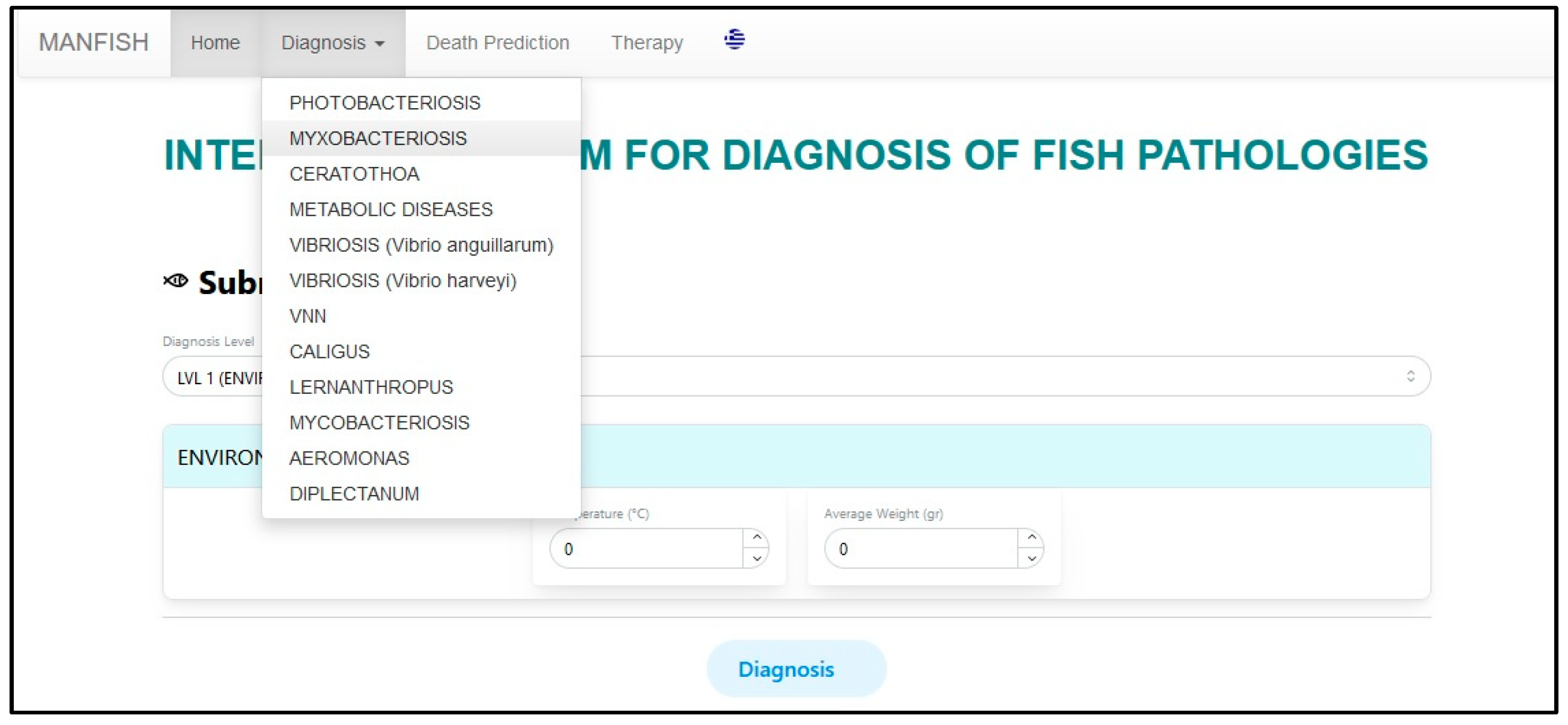
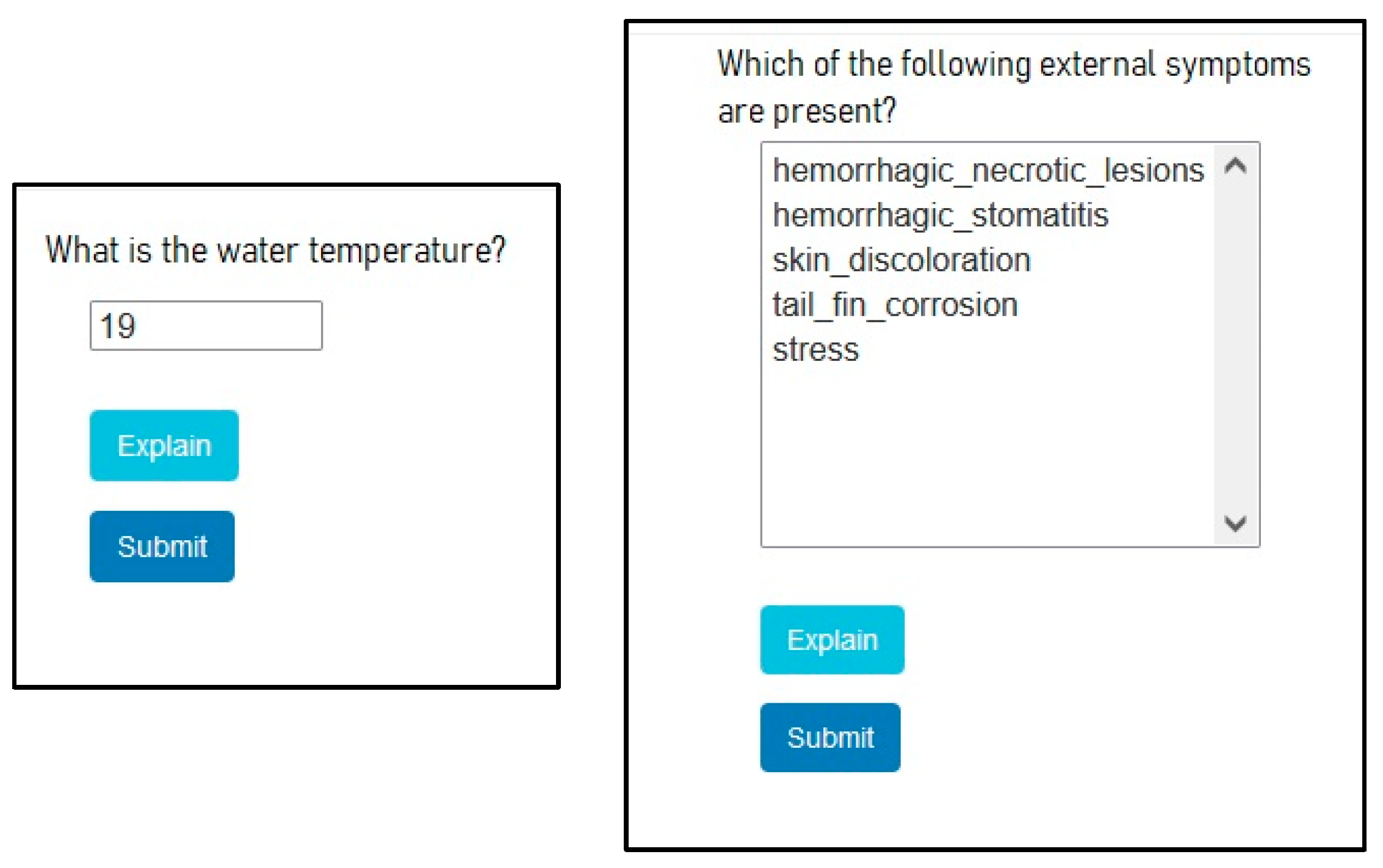
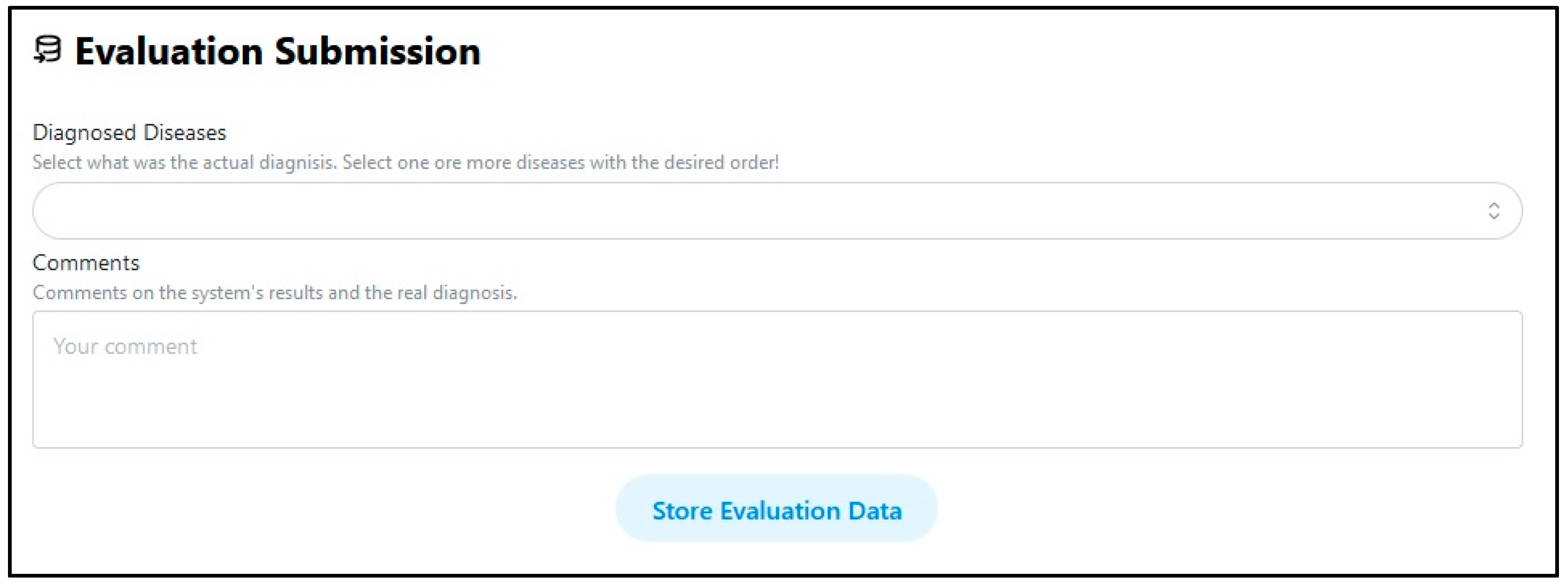
5. User Interface
6. System Evaluation
6.1. Expert Based Tuning and Evaluation
6.2. Artificial Data Based Evaluation
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Maldonado-Miranda, J.J.; Castillo-Perez, L.J.; Ponce-Hernandez, A.; Carranza-Alvarez, C. Chapter 19—Summary of economic losses due to bacterial pathogens in aquaculture industry. In Bacterial Fish Diseases; Academic Press: Cambridge, MA, USA, 2022; pp. 39–417. [Google Scholar]
- Sánchez, J.L.F.; Le Breton, A.; Brun, E.; Vendramin, N.; Spiliopoulos, G.; Furones, D.; Basurco, B. Assessing the economic impact of diseases in Mediterranean grow-out farms culturing European sea bass. Aquaculture 2022, 547, 737530. [Google Scholar] [CrossRef]
- Zrncic, S. (Ed.) Diagnostic Manual for the Main Pathogens in European Seabass and Gilthead Seabream Aquaculture; Options Méditerranéennes, Series B: Studies and Research, No. 75; CIHEAM: Zaragoza, Spain, 2020; p. 172. [Google Scholar]
- Li, D.; Li, X.; Wang, Q.; Hao, Y. Advanced Techniques for the Intelligent Diagnosis of Fish Diseases: A Review. Animals 2022, 12, 2938. [Google Scholar] [CrossRef]
- Grosan, C.; Abraham, A. Intelligent Systems-A Modern Approach; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar]
- Richter, M.M.; Weber, R.O. Case Based Reasoning-A Textbook; Springer: Berlin/Heidelberg, Germany; New York, NY, USA; Dordrecht, The Netherlands; London, UK, 2013. [Google Scholar]
- Chatzilygeroudis, K.; Perikos, I.; Hatzilygeroudis, I. Machine Learning Basics. In Intelligent Computing for Interactive System Design: Statistics, Digital Signal Processing, and Machine Learning in Practice; Eslambolchilar, P., Komninos, A., Dunlop, M., Eds.; ACM: New York, NY, USA, 2021; pp. 143–193. [Google Scholar] [CrossRef]
- Aggarwal, C.C. Neural Networks and Deep Learning: A Textbook; Springer International Publishing AG: Berlin/Heidelberg, Germany, 2018. [Google Scholar]
- Zhao, S.; Zhang, S.; Liu, J.; Wang, H.; Li, D.; Zhao, R. Application of machine learning in intelligent fish aquaculture: A Review. Aquaculture 2021, 540, 736724. [Google Scholar] [CrossRef]
- Ahmed, S.; Aurpa, T.T.; Azad, A.K. Fish Disease Detection Using Image Based Machine Learning Technique in Aquaculture. J. King Saud Univ. Comput. Inf. Sci. 2022, 34, 5170–5182. [Google Scholar] [CrossRef]
- Sun, M.; Yang, X.; Xie, Y. Deep Learning in Aquaculture: A Review. J. Comput. 2020, 31, 294–319. [Google Scholar]
- Hatzilygeroudis, I.; Dimitropoulos, K.; Kovas, K.; Theodorou, J.A. Expert Systems for Farmed Fish Disease Diagnosis: An Overview and a Proposal. J. Mar. Sci. Eng. 2023, 11, 1084. [Google Scholar] [CrossRef]
- Negnevitsky, M. Artificial Intelligence-A Guide to Intelligent Systems, 3rd ed.; Pearsons Higher Education: River Street Hoboken, NJ, USA, 2010. [Google Scholar]
- Shortliffe, E.H.; Buchanan, B.G. A model of inexact reasoning in medicine. Math. Biosci. 1975, 23, 351–379. [Google Scholar] [CrossRef]
- Sumartono, I.; Arisandi, D.; Siahaan, A.P.U.; Aan, M. Expert System of Catfish Disease Determinant Using Certainty Factor Method. Int. J. Recent Trends Eng. Res. 2017, 3, 202–209. [Google Scholar] [CrossRef]
- Aamodt, A.; Plaza, E. Case-based reasoning: Foundational issues, methodological variations and system approaches. Artif. Intell. Commun. 1994, 7, 39–59. [Google Scholar] [CrossRef]
- Kolodner, J.L. Case-Based Reasoning; Morgan Kaufmann: San Mateo, CA, USA, 1993. [Google Scholar]
- Keet, C.M. An Introduction to Ontology Engineering. 2020 College Publications. Open Book. ISBN: 978-1-84890-295-4. Available online: https://people.cs.uct.ac.za/~mkeet/files/OEbook.pdf (accessed on 18 November 2023).
- Qin, H.; Xiao, J.; Gao, X.; Wang, H. Horse-Expert: An aided expert system for diagnosing horse diseases. Pol. J. Veter-Sci. 2016, 19, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Li, D. Aquatic Animal Disease Diagnosis System Based on Android. In Proceedings of the 9th IFIP WG 5.14 International Conference on Computer and Computing Technologies in Agriculture (CCTA), Beijing, China, 27–30 September 2016; pp. 115–124. [Google Scholar]
- Munirah, M.Y.; Suriawati, S.; Teresa, P.P. Design and Development of Online Dog Diseases Diagnosing System. Int. J. Inf. Educ. Technol. 2016, 6, 913–916. [Google Scholar] [CrossRef]
- Gebre-Amanuel, E.K.; Taddesse, F.G.; Assalif, A.T. Web Based Expert System for Diagnosis of Cattle Disease. In Proceedings of the 10th International Conference on Management of Digital EcoSystems (MEDES’18), Tokyo, Japan, 25–28 September 2018; ACM: New York, NY, USA, 2018; p. 8. [Google Scholar] [CrossRef]
- Fahrozi, W.; Harahap, C.B.; Syahputra, A.; Pane, R. Expert System of Diagnosing Koi’s Fish Disease by Certainty Factor Method. In Proceedings of the 2018 6th International Conference on Cyber and IT Service Management (CITSM), Parapat, Indonesia, 7–9 August 2018; pp. 1–5. [Google Scholar] [CrossRef]
- Alarcón-Salvatierra, A.; Bazán-Vera, W.; Samaniego-Cobo, T.; Anchundia, S.M.; Alarcón-Salvatierra, P. SE-DiagEnf: An Ontology-Based Expert System for Cattle Disease Diagnosis. In Technologies and Innovation. CITI 2018. Communications in Computer and Information Science; Valencia-García, R., Alcaraz-Mármol, G., Del Cioppo-Morstadt, J., Vera-Lucio, N., Bucaram-Leverone, M., Eds.; Springer Nature: Cham, Switzerland, 2018; Volume 883, pp. 70–81. [Google Scholar] [CrossRef]
- Sihotang, H.T.; Riandari, F.; Simanjorang, R.M.; Simangunsong, A.; Hasugian, P.S. Expert System for Diagnosis Chicken Disease using Bayes Theorem. J. Phys. Conf. Ser. 2019, 1230, 012066. [Google Scholar] [CrossRef]
- Tomatala, M.F.; Arundaa, R.; Damodalag, H. Fish Disease Diagnosis using Case-based Reasoning with Euclidean Distance. In Proceedings of the 4th International Conference of Vocational Higher Education (ICVHE 2019)—Empowering Human Capital Towards Sustainable 4.0 Industry, Manado, Indonesia, 3–4 September 2019; SCITEPRESS-Science and Technology Publications, LDA: Setúbal, Portugal, 2021; pp. 215–221. [Google Scholar]
- Mardiyanto, F.F.; Satria, F. Expert System for Diagnosis Diseases in Betta Fish Based on Android. Int. J. Artif. Intell. Robot. Technol. IJAIRTec 2021, 1, 35–44. [Google Scholar]
- Riyanto, E.D.; Prasetyo, E.; Zainal, R.F.; Rubaningtyas, R.; Setyatama, F.; Herulambang, W.; Alim, S.; Tias, R.F. Design of Expert System Diagnosis of Catfish Disease with Forward Chaining Method. J. Electr. Eng. Comput. Sci. 2022, 7, 1215–1222. [Google Scholar] [CrossRef]
- Cha, S.-H. Comprehensive Survey on Distance/Similarity Measures between Probability Density Functions. Int. J. Math. Models Methods Appl. Sci. 2007, 1, 300–307. [Google Scholar]
- Widians, J.A.; Puspitasari, N.; Febriansyah, A. Disease Diagnosis System using Certainty Factor. In Proceedings of the 6th International Conference on Electrical, Electronics and Information Engineering (ICEEIE 2019), Denpasar, Indonesia, 3–4 October 2019; pp. 303–308. [Google Scholar]
- Saputri, A.E.; Sevani, N.; Saputra, F.; Sali, R.K. Using Certainty Factor Method to Handle Uncertain Condition in Hepatitis Diagnosis. ComTech Comput. Math. Eng. Appl. 2020, 11, 1–10. [Google Scholar] [CrossRef]
- Muntiari, N.R.; Hanif, K.H. Application of The Certainty Factor Method for Diagnosing Osteoarthritis using The Python Programming Language. J. Adv. Health Inf. Res. JAHIR 2023, 1, 21–27. [Google Scholar] [CrossRef]
- Hatzilygeroudis, I.; Kovas, K. A Tool for Automatic Creation of Rule-Based Expert Systems with CFs. In IFIP Advances in Information and Communication Technology, Volume 339, Artificial Intelligence Applications and Innovations (AIAI-10); Springer: Berlin/Heidelberg, Germany, 2010; pp. 195–202. [Google Scholar]
- Konstantinopoulou, G.; Kovas, K.; Hatzilygeroudis, I.; Prentzas, J. An Approach using Certainty Factor Rules for Aphasia Diagnosis. In Proceedings of the 10th International Conference on Information, Intelligence, Systems and Applications (IISA 2019), Patras, Greece, 15–17 July 2019. [Google Scholar]
- Brunelli, R. Template Matching Techniques in Computer Vision: Theory and Practice; Wiley: Hoboken, NJ, USA, 2009; ISBN 978-0-470-51706-2. [Google Scholar]




| No | Disease | Category |
|---|---|---|
| 1 | Aeromonas disease | Bacterial |
| 2 | Mycobacteriosis | Bacterial |
| 3 | Myxobacteriosis | Bacterial |
| 4 | Photobacteriosis | Bacterial |
| 5 | Vibriosis (Vibrio anguillarum) | Bacterial |
| 6 | Vibriosis (Vibrio harveyi) | Bacterial |
| 7 | Caligus | Parasitic |
| 8 | Ceratothoa | Parasitic |
| 9 | Diplectanum | Parasitic |
| 10 | Lernanthropus | Parasitic |
| 11 | VNN | Viral |
| 12 | Metabolic Diseases | Metabolic |
| Parameter/Symptom | Type (Values) |
|---|---|
| Temperature | numeric |
| Lethargic fish | boolean |
| Anorexia | boolean |
| Hemorrhagic stomatitis | boolean |
| Skin color darkening | boolean |
| Skin discoloration | boolean |
| Skin ulcers | Categorical (no, small, mild, large) |
| Hemorrhagic and necrotic skin changes in fins | boolean |
| Hemorrhagic skin changes in tail | boolean |
| Necrotic skin changes in fins | boolean |
| Necrotic skin changes in tail | boolean |
| Corrosion of tail | boolean |
| Corrosion of fins | boolean |
| Stress | boolean |
| (Sub)Category | Parameter | Type |
|---|---|---|
| Environmental | ||
| Temperature | numeric | |
| Average weight | numeric | |
| External Clinical Signs (Symptoms) | ||
| Behavioral | Anorexia | boolean |
| Lethargic fish | boolean | |
| Weight loss | boolean | |
| Reduced weight increase rate | boolean | |
| Swimming Bladder Control Loss | boolean | |
| Stress | boolean | |
| Mortality | Categorical (zero, massive, nonmassive) | |
| Swimming Behavior | Categorical (normal, alien, slow, fast) | |
| Time between symptoms and deaths | Categorical (normal, small, large) | |
| Physiological | Skin color darkening | boolean |
| Skin discoloration | boolean | |
| Whitish areas | boolean | |
| Conjugal fins redness | boolean | |
| Retinopathy | boolean | |
| Exophthalmos | boolean | |
| Corneal Clouding | boolean | |
| Mouth lower jaw deformity | boolean | |
| Corrosion-Necrosis of Tail and Fins | boolean | |
| Scales Loss | boolean | |
| Bleeding areas | boolean | |
| Hemorrhagic and necrotic lesions | boolean | |
| Hemorrhagic stomatitis | boolean | |
| Skin ulcers | Categorical (no, small, mild, large) | |
| Fins and Tail ulcers | boolean | |
| Scoliosis or Lordosis or Hypercalcification | boolean | |
| Release of Mucus Fecal Casts | boolean | |
| Internal Clinical Signs (Symptoms) | ||
| Gills | Excess gill mucous | boolean |
| Gill discoloring | boolean | |
| Anemic gills | boolean | |
| Local changes | boolean | |
| INTERNAL Organs | Pseudoenteritis | boolean |
| Splenomegaly | boolean | |
| Granulomatosis | boolean | |
| Inflamed liver | boolean | |
| Symptoms | Photo-Bacteriosis | Myxo-Bacteriosis | Myco-Bacteriosis | Ceratothoa |
|---|---|---|---|---|
| Weight loss | X | |||
| Mortality (deaths) | nomass, mass | |||
| Anorexia | X | X | X | |
| Lethargic fish | X | |||
| Swimming Behavior | normal, alien | |||
| Color Darkening | X | X | ||
| Skin discoloration | X | |||
| Mouth Lower Jaw Deformity | X | |||
| Fins and tail corrosion | X | |||
| Scales Loss | X | |||
| Stomatitis | X | |||
| Skin ulcers | X | X | ||
| Gill mucous | X | |||
| Anemic gills | X | |||
| Granulomatosis | X |
| Laboratory Parameters |
|---|
| Microbiological |
| GRAM Stain |
| Bacterial Culture |
| Viral Culture |
| Antibiogram |
| Urinary adhesion |
| Blood test |
| Biochemical API test |
| Microscopic |
| Gills microscopic examination |
| Histopathological examination |
| Molecular |
| PCR test |
| Chemical |
| Chemical examination |
| Method | Accuracy | Top-3 Accuracy | Average Rank |
|---|---|---|---|
| CBO | 0.511 | 0.919 | 1.825 |
| WBM | 0.435 | 0.897 | 1.982 |
| ACRES | 0.622 | 0.909 | 1.743 |
| AGM | 0.658 | 0.940 | 1.569 |
| Disease | Accuracy | Top-3 Accuracy | Average Rank |
|---|---|---|---|
| pasteridiasi | 0.322 | 0.762 | 2.399 |
| mixovaktiridiasi | 0.673 | 0.940 | 1.547 |
| metavolika | 0.800 | 0.960 | 1.327 |
| don-vibrio-anguil | 0.707 | 0.967 | 1.473 |
| don-vibrio-harv | 0.660 | 0.947 | 1.580 |
| egkefalopatheia | 0.700 | 0.973 | 1.407 |
| ceratothoa | 0.720 | 1.000 | 1.333 |
| galicus | 0.520 | 0.933 | 1.773 |
| diplectanum | 0.520 | 0.840 | 2.007 |
| lernantropus | 0.627 | 0.973 | 1.493 |
| mycobacteriosis | 0.773 | 0.987 | 1.353 |
| aeromonada | 0.860 | 0.993 | 1.180 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kovas, K.; Hatzilygeroudis, I.; Dimitropoulos, K.; Spiliopoulos, G.; Poulos, K.; Abatzidou, E.; Aravanis, T.; Ilias, A.; Kanlis, G.; Theodorou, J.A. Using Level-Based Multiple Reasoning in a Web-Based Intelligent System for the Diagnosis of Farmed Fish Diseases. Appl. Sci. 2023, 13, 13059. https://doi.org/10.3390/app132413059
Kovas K, Hatzilygeroudis I, Dimitropoulos K, Spiliopoulos G, Poulos K, Abatzidou E, Aravanis T, Ilias A, Kanlis G, Theodorou JA. Using Level-Based Multiple Reasoning in a Web-Based Intelligent System for the Diagnosis of Farmed Fish Diseases. Applied Sciences. 2023; 13(24):13059. https://doi.org/10.3390/app132413059
Chicago/Turabian StyleKovas, Konstantinos, Ioannis Hatzilygeroudis, Konstantinos Dimitropoulos, Georgios Spiliopoulos, Konstantinos Poulos, Evi Abatzidou, Theofanis Aravanis, Aristeidis Ilias, Grigorios Kanlis, and John A. Theodorou. 2023. "Using Level-Based Multiple Reasoning in a Web-Based Intelligent System for the Diagnosis of Farmed Fish Diseases" Applied Sciences 13, no. 24: 13059. https://doi.org/10.3390/app132413059
APA StyleKovas, K., Hatzilygeroudis, I., Dimitropoulos, K., Spiliopoulos, G., Poulos, K., Abatzidou, E., Aravanis, T., Ilias, A., Kanlis, G., & Theodorou, J. A. (2023). Using Level-Based Multiple Reasoning in a Web-Based Intelligent System for the Diagnosis of Farmed Fish Diseases. Applied Sciences, 13(24), 13059. https://doi.org/10.3390/app132413059









