3.2. Artificial Physarum polycephalum Colony
In this subsection, to overcome the shortcomings of algorithm search, a novel artificial
Physarum polycephalum colony is designed on the basis of natural Physarum in the literature [
17,
18,
47].
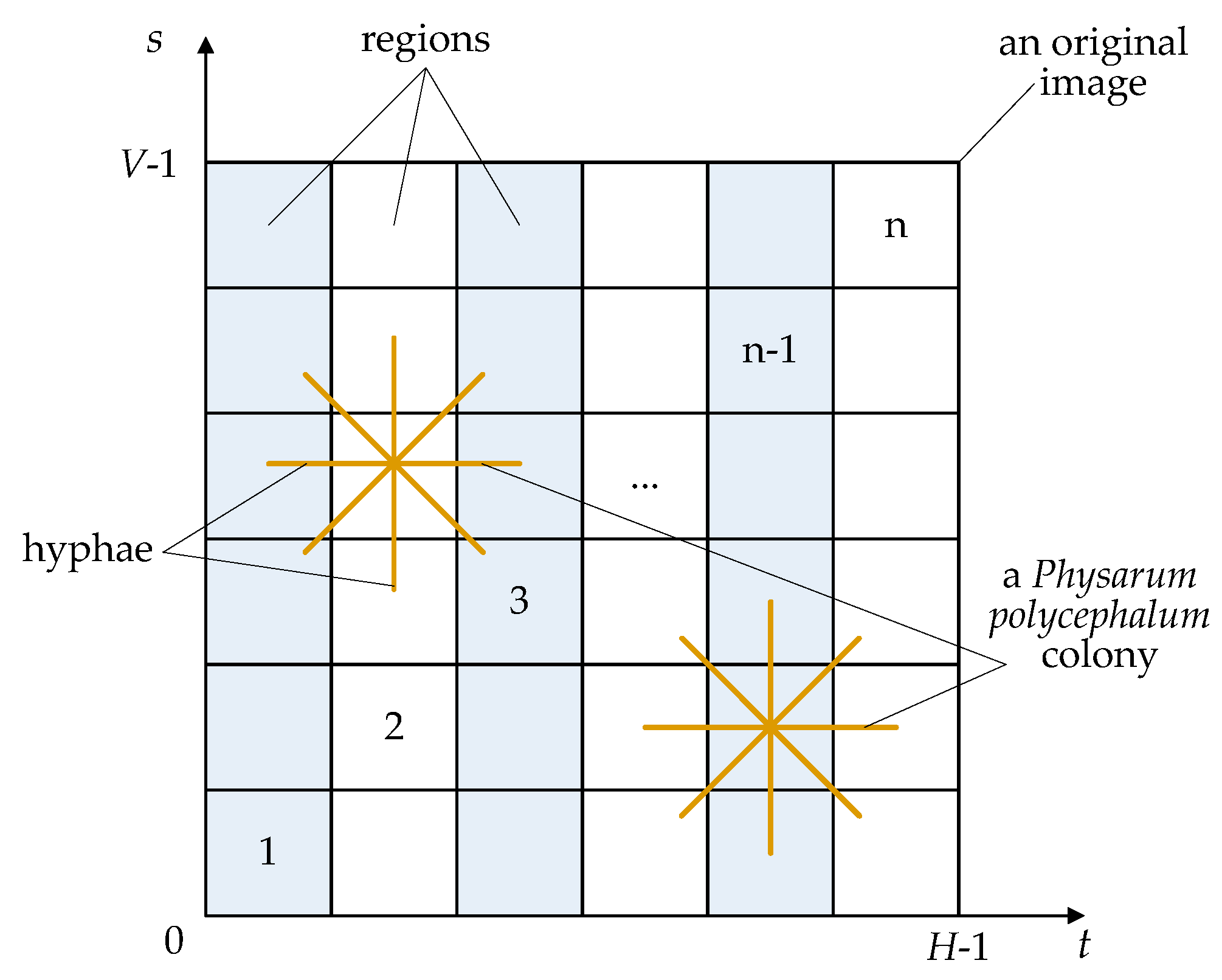
Figure 1 shows an artificial
Physarum polycephalum colony on a 2-D segmentation image. In our artificial
Physarum polycephalum colony algorithm, there are three main components, i.e., an original image, an artificial
Physarum polycephalum colony, and a fitness function.
First, the original image in
Figure 1 is not only the living area of an artificial
Physarum polycephalum colony but also the solution space of the threshold image segmentation or other complex problems. In an original image, there are a lot of pixels with different gray levels. An artificial
Physarum polycephalum colony can search for feasible solutions on an original image or solution space. Each feasible solution is a segmentation model of the original image. The artificial
Physarum polycephalum colony should search for feasible solutions as the food sources for its living and the feasible solutions are randomly distributed in the solution space. The food sources can be absorbed by the artificial
Physarum polycephalum colony and changed into nutrients flowing in its body.
Second, several
Physarum polycephalum constitute an artificial
Physarum polycephalum colony in
Figure 1, where an artificial
Physarum polycephalum employs many filiform hyphae to randomly search for feasible solutions. The artificial
Physarum polycephalum colony algorithm counts the number of hyphae as the population size
S. The artificial
Physarum polycephalum colony employs three learning mechanisms, i.e., self-learning, social learning, and free learning. Hence, different hyphae can learn from each other and many hyphae can cross to share the environmental information. The artificial
Physarum polycephalum finding the food sources and obtaining more nutrients will survive; otherwise, it will be absorbed and disappear. The leaning mechanism and absorbing behavior can effectively simulate the swarm behavior of a real
Physarum polycephalum colony.
Third, the artificial
Physarum polycephalum colony utilizes a fitness function to evaluate its living state, which is not shown in
Figure 1. The threshold
t and region threshold
s segment the original image into several parts, and the fitness depends on the nutrients from the food sources. The artificial
Physarum polycephalum colony does not know the positions of the feasible solutions from the beginning, but it can judge the feasible solution by a fitness function. If an artificial
Physarum polycephalum or a hyphae finds a solution, it will obtain a high fitness value. The fitness function can be designed according to the problems to be solved, such as threshold image segmentation. Different artificial
Physarum polycephalum can cross and learn the nutrient information to share the fitness information.
Based on the three basic components, the artificial Physarum polycephalum colony algorithm uses two main operations to search for the optimal solutions, i.e., expansion and contraction.
In the expansion operation, the artificial Physarum polycephalum colony employs three learning mechanisms to produce more hyphae, i.e., self-learning, social-learning, and free-learning. At the initialization, the artificial Physarum polycephalum colony produces a population size S of filiform hyphae, where each filiform hyphae is a randomly generated feasible solution. In expansion, the filiform hyphae with a population size of S remain unchanged by self-learning, and all these artificial Physarum polycephalum hyphae can learn from each other’s experience to produce a new generation of hyphae with a population size of S by social-learning. At the same time, the artificial Physarum polycephalum colony will search the whole solution space by free learning to randomly generate several new hyphae completely unrelated to existing hyphae. Now, the population size of hyphae in expansion operation is greater than twice the original state 2S.
In contraction operation, the artificial Physarum polycephalum colony calculates the fitness of all hyphae according to the fitness function and selects the top S best hyphae. Other hyphae with low fitness will be absorbed and disappear. Now, the population size of hyphae in contraction operation restores to the initial value S. The contraction operations can help the artificial Physarum polycephalum colony converge to the optimal solution and eliminate hyphae with low fitness. After multiple expansion and contraction iterations and fitness comparisons, the artificial Physarum polycephalum colony can find the optimal solution to the threshold image segmentation.
3.3. A Fitness Function for Entropy Threshold
This subsection provides the fitness function for the artificial
Physarum polycephalum colony to implement threshold image segmentation. To explore the average information content of different pixels in image segmentation, the fitness of the proposed threshold image segmentation is based on the maximum entropy model proposed by Kapur [
13,
40] and maximizes the total entropy of the segmented target and background. The Kapur’s entropy can help us obtain the largest amount of segmentation information on an original image.
In different threshold models, the 1-D segmentation method is the fastest in segmentation speed since it does not consider the gray values of different pixels on an original image. The 3-D segmentation is the lowest in segmentation speed since it considers 3-D gray information. The 2-D segmentation method is between 1-D and 3-D in image segmentation speed since it considers the gray information of each pixel and its neighborhood but not the 3-D gray information. Here, 2-D segmentation is employed since it can obtain balanced performance, as shown in
Figure 1. A set of thresholds
t = {
} segments the target and background into several regions, and the other regions are mainly image noise.
The fitness function should instruct the artificial Physarum polycephalum colony to be efficient and accurate in searching for the best threshold () to obtain the best segmentation quality. A good fitness function should make full use of the gray information of the original image and suppress the noise from target and background areas.
Here, the Kapur’s entropy
can be calculated for the set of thresholds (
) as follows [
13,
40]:
where
indicates Kapur’s entropy in all areas. Hence, Kapur’s entropy in each area can be computed as
where the parameters
w0,
w1, …,
wn indicate the possibility for each segmentation cluster, and
pi indicates the possibility of the occurrence of pixels with a gray value
i. Now, Kapur’s entropy can be obtained for each cluster and instruct the artificial
Physarum polycephalum colony to evaluate the characteristics of clusters. There is
The combination maximization function fit is the optimal threshold group. The artificial Physarum polycephalum colony uses the fitness function to select the optimal solution. The higher the fitness function, the better the quality is achieved.
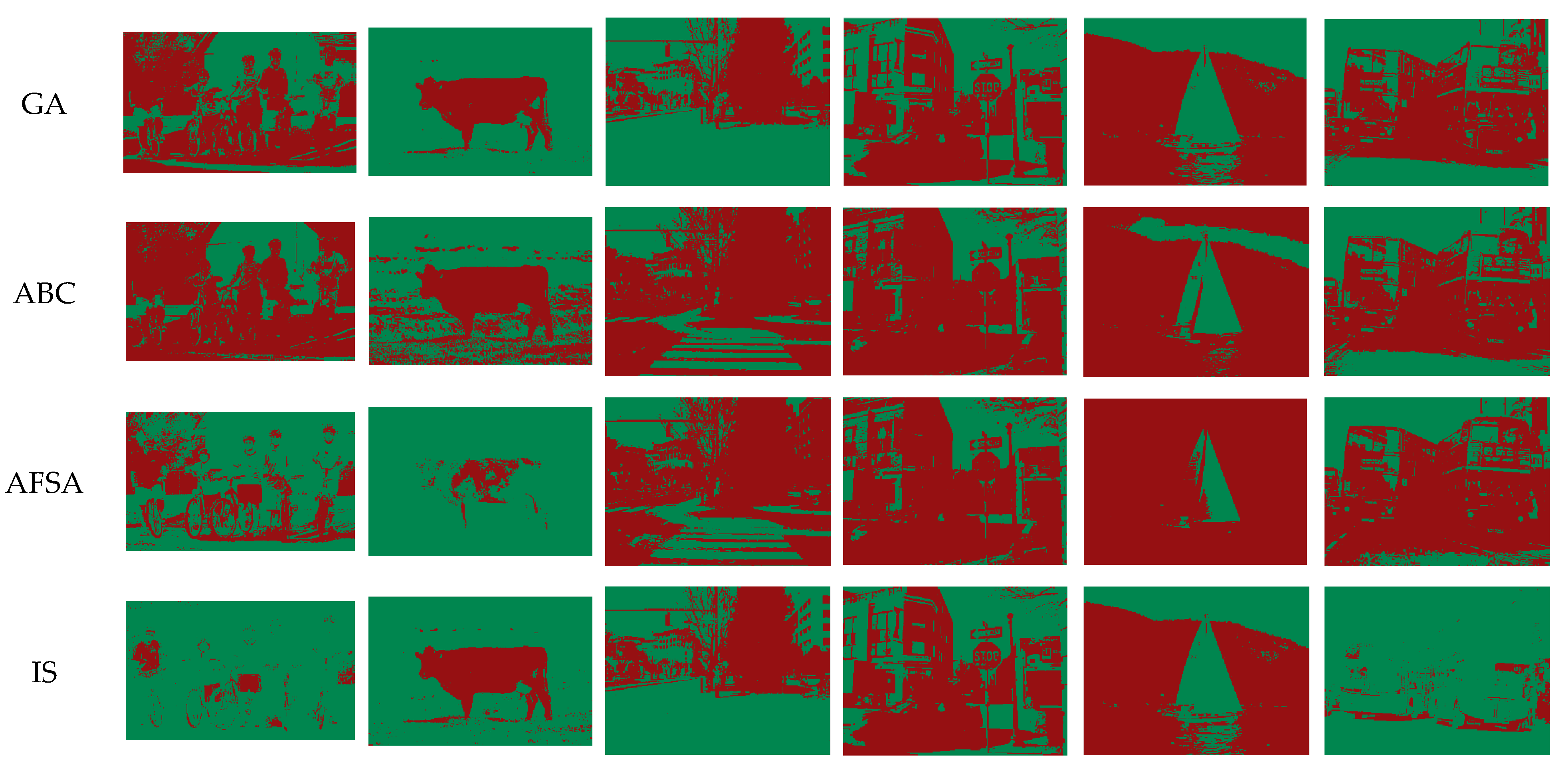
3.4. Performance Measurement
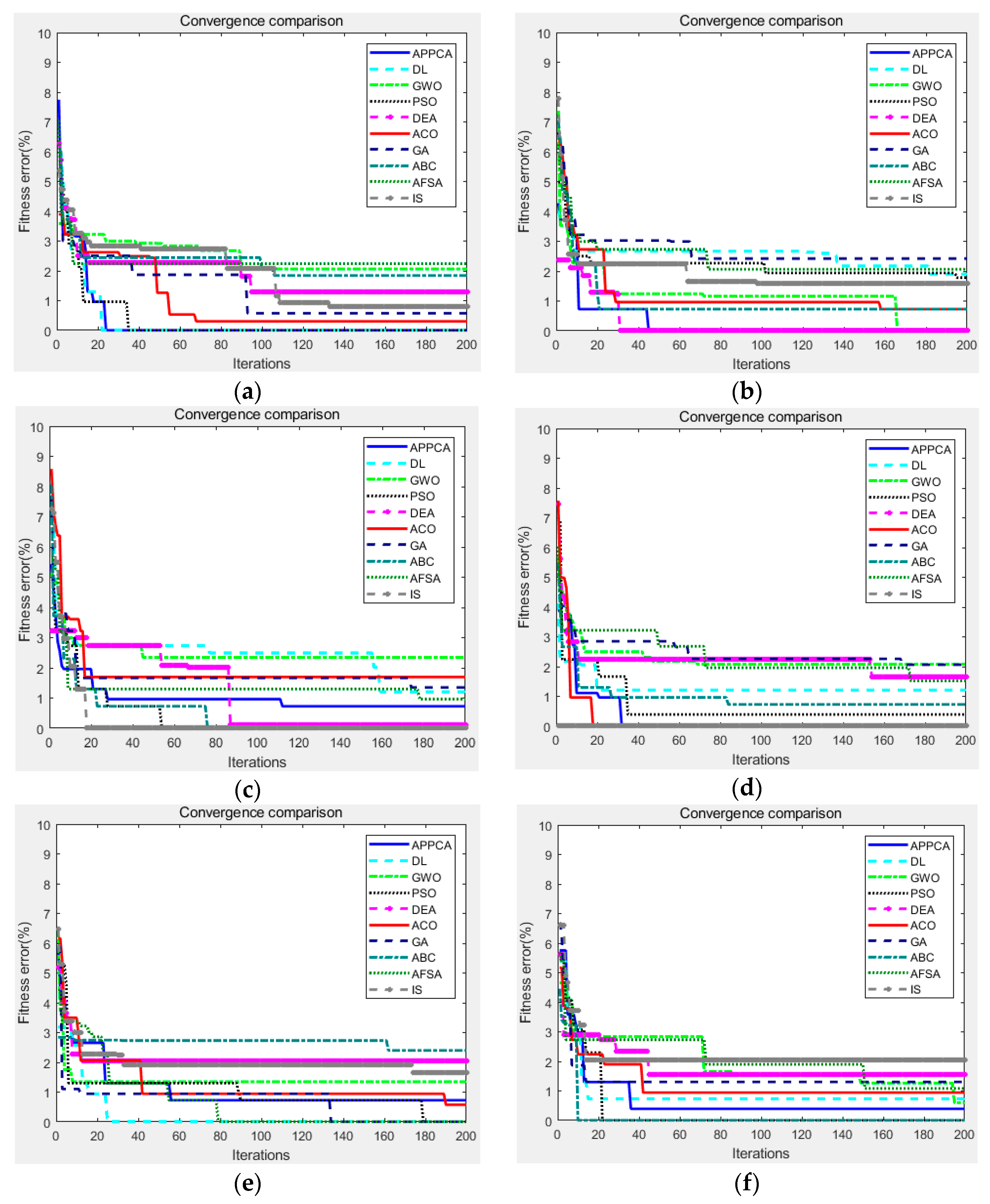
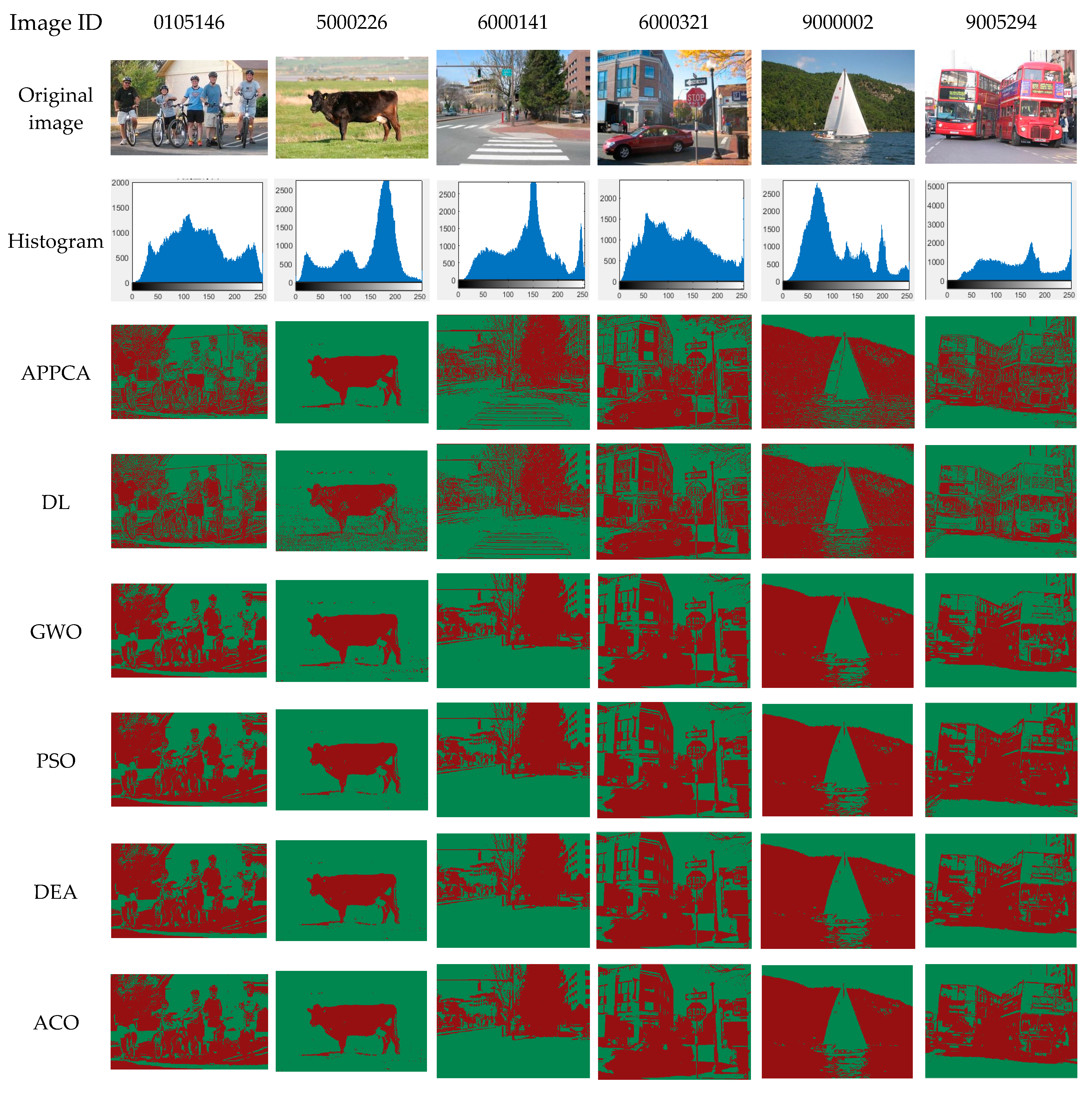
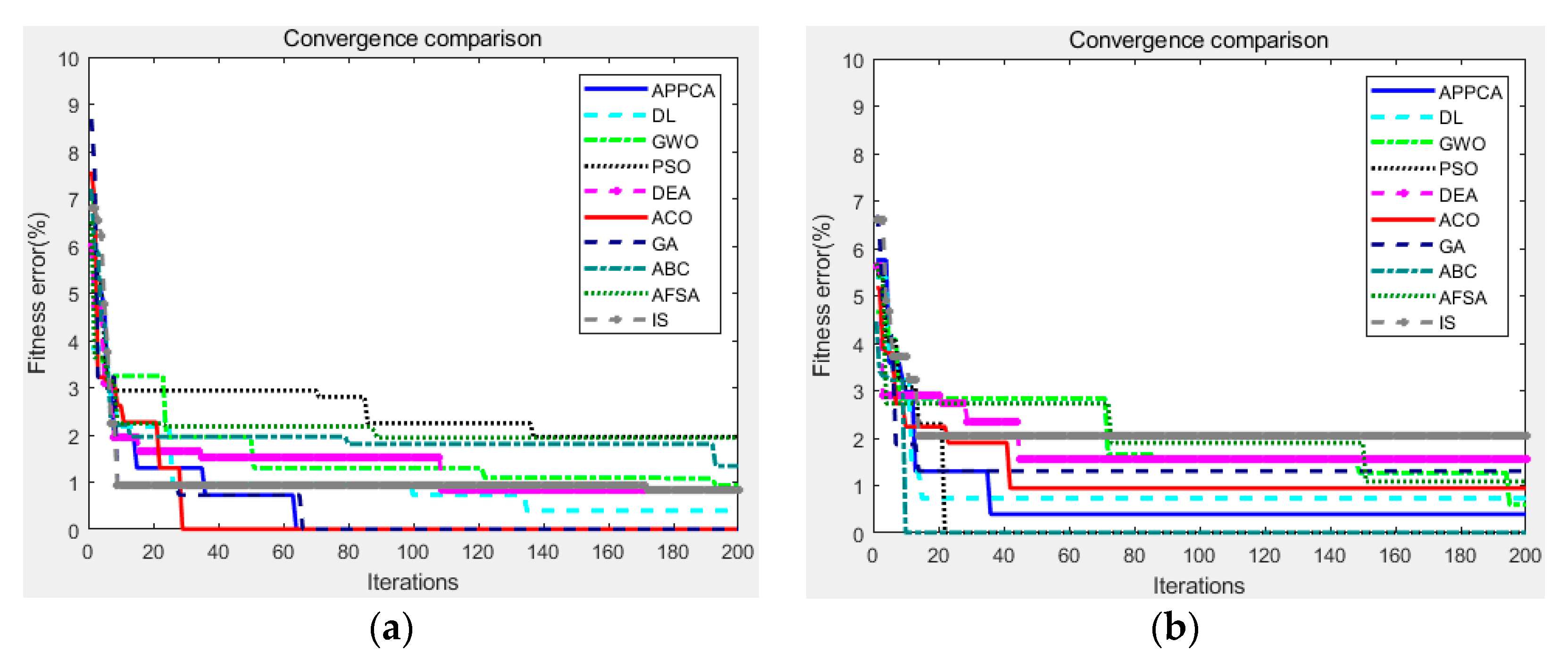
This subsection describes the evaluation indicators for the performance measurement of threshold image segmentation. These indicators can measure the segmentation performance from different aspects based on the original image, where the original image is a reference and the segmentation models from the same original image are subjected to the image noise. In the benchmark tests of
Section 5, the indicators can help us compare the performance difference between the proposed APPC algorithm and the traditional artificial intelligence ones.
First, the mean squared error (
MSE) is a popular index to measure the difference between two images by computing the average sum of the squares between two images. The
MSE can evaluate the total difference of all pixels on two whole images, where a lower
MSE value means less difference between two images. The
MSE can be written as follows:
where
H represents the image size in the horizontal direction,
V represents the image size in the vertical direction,
aij is a pixel on the segmented image
A with a pair of coordinates (
i,
j), and
bij is a pixel on the benchmark image
B with a pair of coordinates (
i,
j). The drawback of
MSE is that the value often has the same weight in different grayscale. For example, a high
MSE can be obtained from a noisy background on a well-recognizable image.
Second, the peak signal-to-noise ratio (
PSNR) is an important indicator of the quality of image segmentation. It calculates the ratio of the effective signal to the noise signal on an image, that is, the ratio of the segmented image to the benchmark image. According to the mean square deviation in Equation (6), the
PSNR can be defined for two images with the same size:
where
MSE is the mean squared error in Equation (6), and
L is the grayscale level of the image. A high
PSNR often means high quality, and a low
PSNR often means low quality. A high
PSNR also indicates good image recognition.
Third, the correlation coefficient (
CorCoe) is also a popular indicator to measure the linear correlation between two images. The
CorCoe can compare the intensity difference of each pixel between two images and the average intensity difference between two whole images. The
CorCoe can be calculated as follows:
where
and
indicate the intensity value on each pixel with a pair of coordinates (
i,
j) for two images A and B, respectively.
and
indicate the arithmetic mean of intensity value for two images
A and
B, respectively.
H represents the image size in the horizontal direction, and
V represents the image size in the vertical direction. The
CorCoe metric often takes values in [−1, 1], where 1 represents the highest match between two images, but −1 represents the lowest match between two images.
Fourth, Jaccard’s index (
Jaccard), also called Intersection-Over-Union, IoU, is another popular indicator to measure the segmentation quality. The
Jaccard metric calculates the size of the intersection divided by the union size of each set of pixels to measure the similarity between finite sets of pixels. The calculation equation can be obtained as follows:
where
A is the segmented image with additive noise and
B is the benchmark image. The numerator
calculates the overlap area of images
A and
B, and the denominator
describes the fusion area of images
A and
B.
Jaccard index ranges from 0 to 1, where a value of 1 indicates a perfect overlap or the greatest similarity, but a value of 0 indicates no spatial overlap or an absolute mismatch.
Fifth, the Sørensen Dice coefficient (
Dice) is similar to Jaccard’s coefficient but is the double overlap region divided by the total number of pixels in two images. The
Dice and
Jaccard are positively correlated and are easy to reach a consensus assessment. For example, they both may determine model
A is better than model
B in segmentation performance, or vice versa. The calculation equation is as follows:
where
A is the segmented image with additive noise and
B is the benchmark image. The numerator
calculates the overlap area of images
A and
B, and the denominator
describes the fusion area of images
A and
B.
Dice coefficient also ranges from 0 to 1, where a value of 1 indicates a perfect overlap or the greatest similarity, but a value of 0 indicates no spatial overlap or an absolute mismatch.
Sixth, the Structural Similarity Index (SSIM) calculates a multiplicative combination of three image components, i.e., brightness, contrast, and texture. The SSIM can describe the scene structure information on an image.
For a gray image
A, the average gray level
can be calculated to estimate the brightness, so the brightness can be written as follows:
where
H represents the image size in the horizontal direction,
V represents the image size in the vertical direction, and
aij is a pixel on the segmented image
A with a pair of coordinates (
i,
j).
The measurement system removes the average grayscale value
from the image. For a digital image
A, standard deviation
can be used as the contrast estimation value.
According to the brightness
in Equation (11) and the contrast
in Equation (12), the texture can be expressed as their function. Therefore, the
SSIM can be calculated as follows:
where the average gray level
and
estimates the brightness, and the standard deviation
and
evaluates the contrast. The constant
is used to avoid instability of the evaluation system when the brightness approaches 0, often
.
L is the grayscale level of the image, i.e.,
L = 255 for 8-bit grayscale images.
is a positive constant far less than 1. The constant
is used to avoid instability of the evaluation system when the contrast approaches 0, often
.
L is the grayscale level of the image, and
is a positive constant far less than 1.
In the experimental results of
Section 5, the performance measurement indicators above are employed for objective evaluation and algorithm comparison, which can objectively verify the proposed APPC algorithm and compare the segmentation performance.
3.5. Algorithm Flow of APPCA
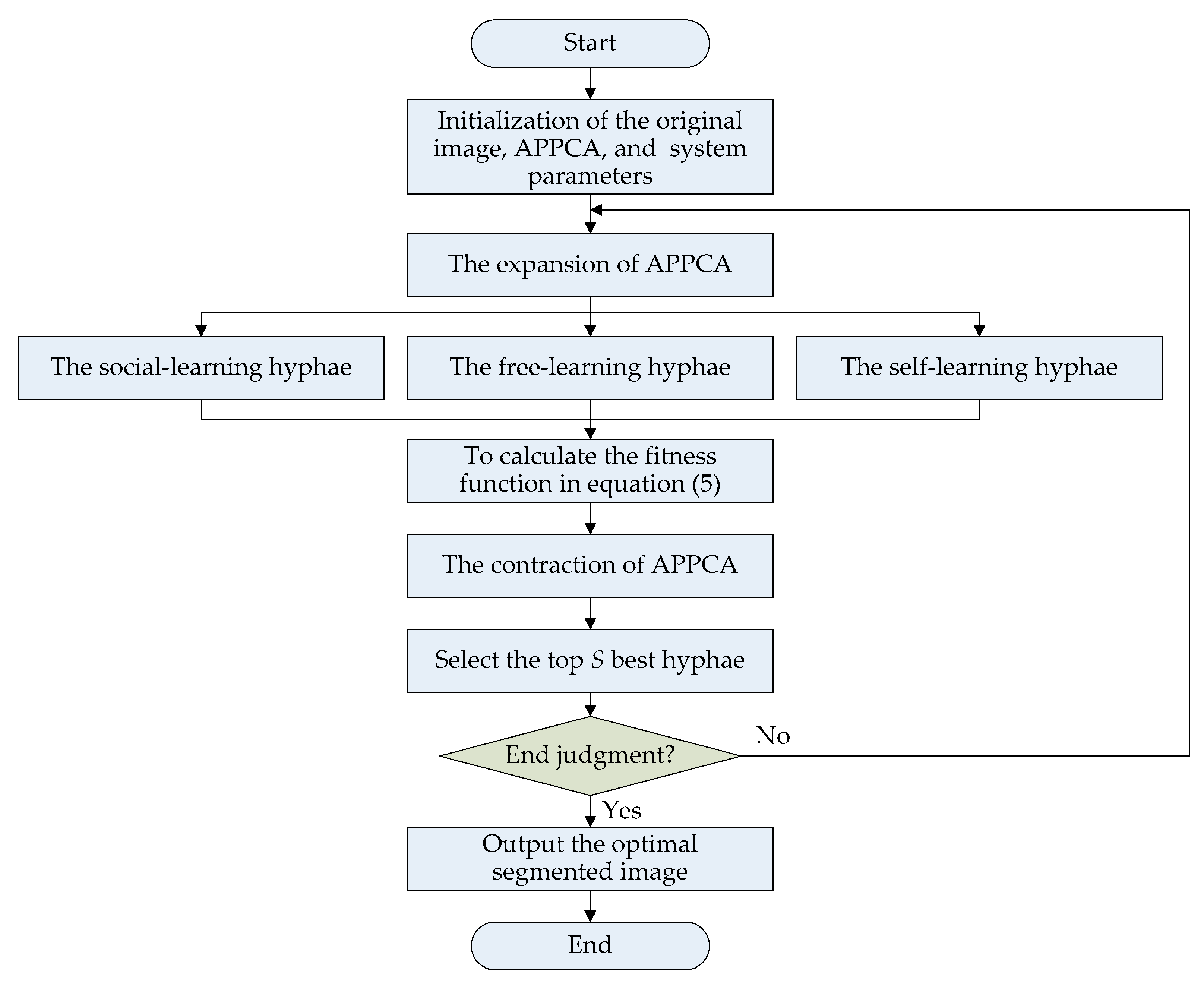
This subsection provides the algorithm flow of APPCA. To solve the threshold image segmentation problem, all
Physarum polycephalum will continuously search for the optimal solutions by iterative computation according to the algorithm flow, as shown in
Figure 2. The whole algorithm flow includes four main stages, namely initialization, expansion, contraction, and end judgment.
In the initialization stage, there are three main parts, i.e., the original image, the artificial Physarum polycephalum colony, and the system simulation parameters.
In the expansion stage, multiple Physarum polycephalum will search for the optimal solutions according to the fitness in Equation (5). The Physarum polycephalum colony selects the best solutions to the fitness function of Kapur’s entropy. The Physarum polycephalum colony will learn the nutrient information to expand and contract with self-learning, social learning, and free learning to implement efficient image segmentation. Therefore, the population size increases.
In the contraction stage, the Physarum polycephalum colony will evaluate the fitness of hyphae and select the best ones for the next iterations. The optimal hyphae with high fitness will be left and the population size will decrease. After many iterations of expanding and contracting, it is possible to obtain an optimal segmented image. For all Physarum polycephalum, it is not compulsory to search for all solutions. It is sufficient to select the optimal hyphae through a sufficient number of iterative calculations.
At the end of iteration learning, the error varying becomes less and less, and then the solution can be output. A predefined error threshold eth, the number of iterations Ite_max, or a maximum calculation time limit can be used as an end judgment before the optimal solutions are generated.