Modelling and Predicting the Growth of Escherichia coli and Staphylococcus aureus in Co-Culture with Geotrichum candidum and Lactic Acid Bacteria in Milk
Abstract
1. Introduction
2. Material and Methods
2.1. Microorganisms and Culture Conditions
2.2. Preparation of Inoculum and Experiments
2.3. Quantification of Microorganisms
2.4. Mathematical Models
2.4.1. Modelling the Microbial Interaction in Co-Cultures
- A.
- H-GD model with the competition coefficients
2.4.2. Parameter Determination and Evaluation of Model Performance
3. Results and Discussion
3.1. One-Step Analysis of Competitive Growth
3.2. Model Validation
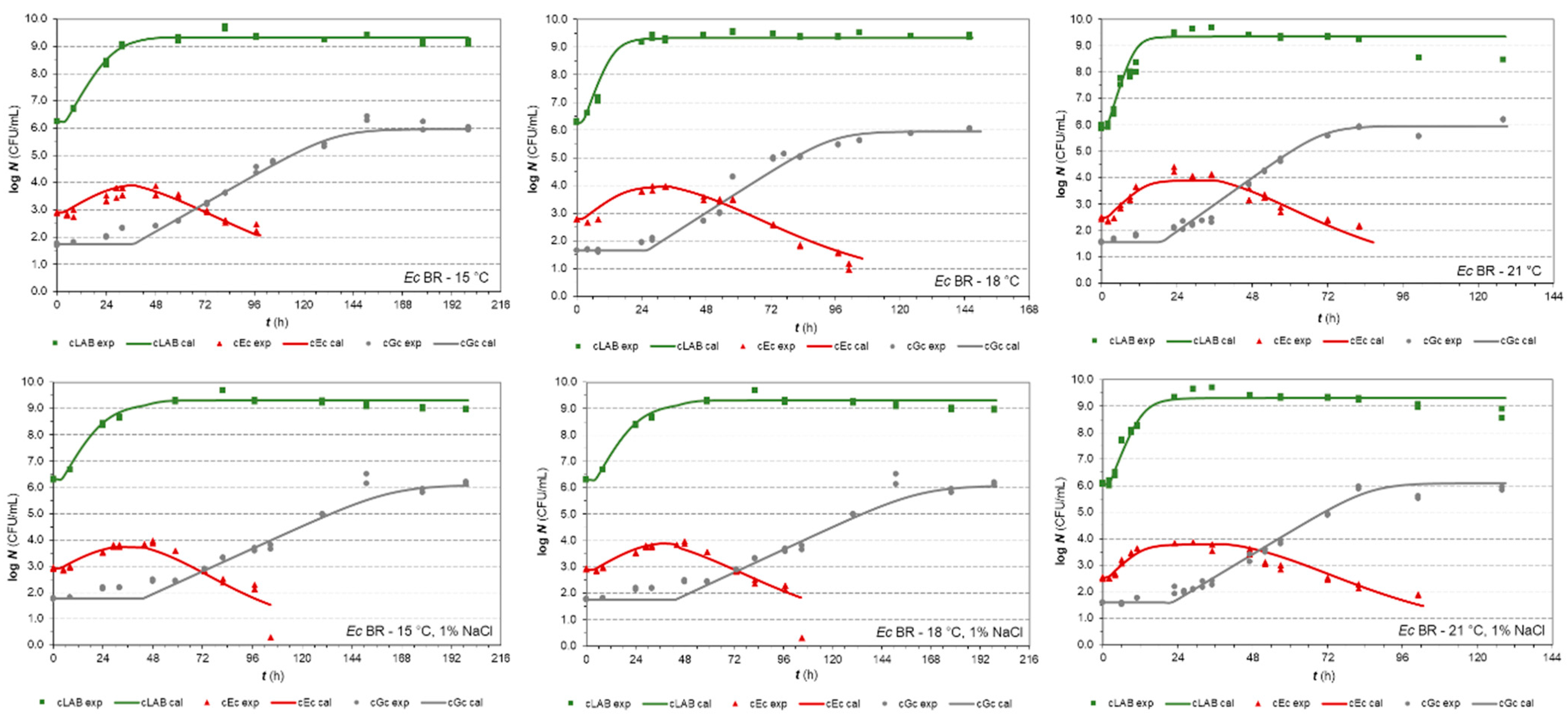
3.2.1. E. coli Isolates in Co-Cultures
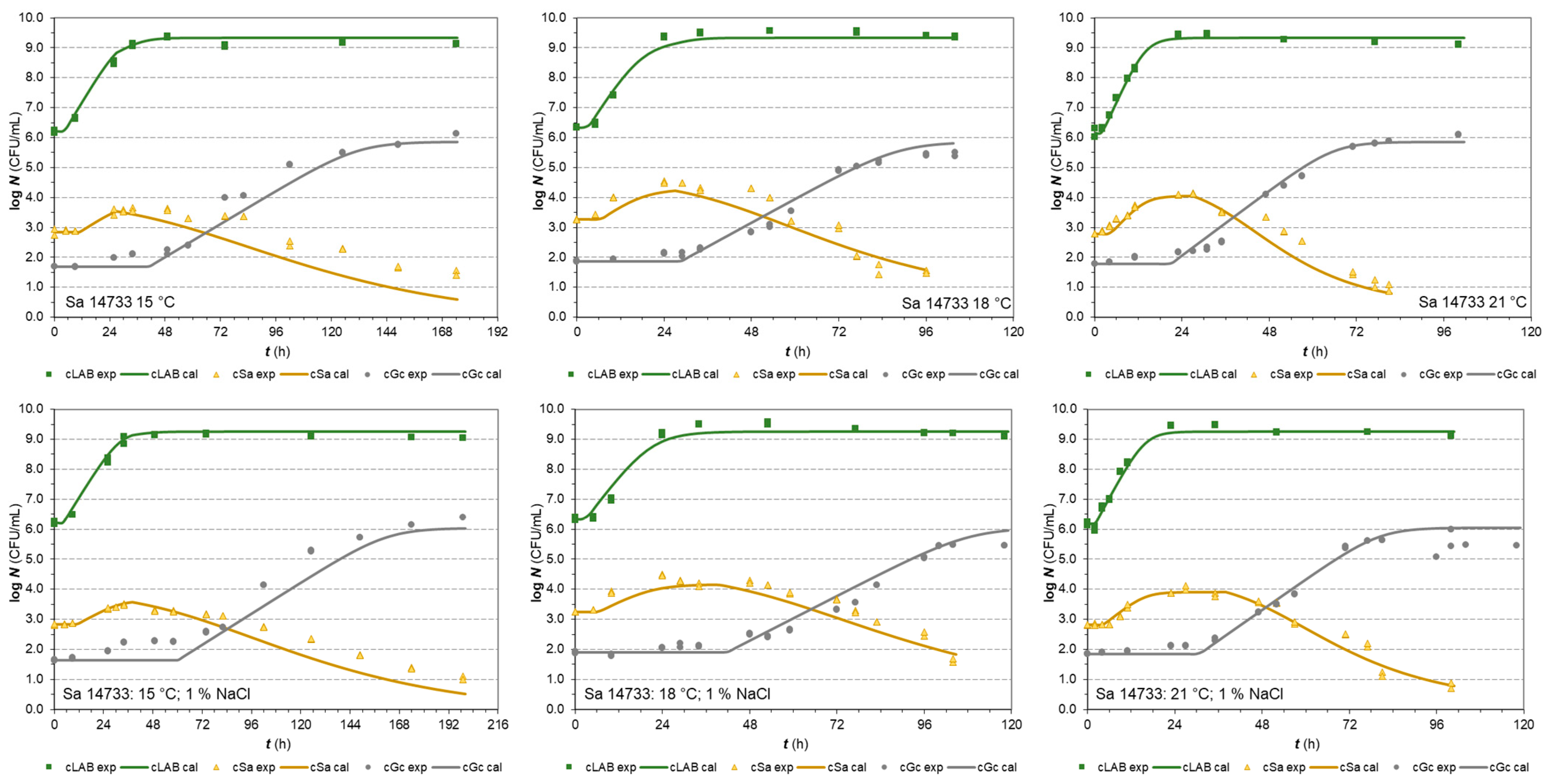
3.2.2. S. aureus Isolates in Co-Cultures
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Whiting, R.; Buchanan, R. A classification of models for predictive microbiology. Food Microbiol. 1993, 10, 175–177. [Google Scholar]
- Jewell, K. Comparison of 1-step and 2-step methods of fitting microbiological models. Int. J. Food Microbiol. 2012, 160, 145–161. [Google Scholar] [CrossRef] [PubMed]
- Huang, L. IPMP Global Fit—A one-step direct data analysis tool for predictive microbiology. Int. J. Food Microbiol. 2017, 262, 38–48. [Google Scholar] [CrossRef]
- Vereecken, K.; Dens, E.; Van Impe, J. Predictive modelling of mixed microbial populations in food products: Evaluation of two-species models. J. Theor. Biol. 2000, 205, 53–72. [Google Scholar] [CrossRef] [PubMed]
- Giménez, B.; Dalgaard, P. Modelling and predicting the simultaneous growth of Listeria monocytogenes and spoilage micro-organisms in cold-smoked salmon. J. Appl. Microbiol. 2004, 96, 96–109. [Google Scholar] [CrossRef]
- Possas, A.; Bonilla-Luque, O.M.; Valero, A. From Cheese-Making to Consumption: Exploring the Microbial Safety of Cheeses through Predictive Microbiology Models. Foods 2021, 10, 355. [Google Scholar] [CrossRef]
- Metz, M.; Sheehan, J.; Feng, P.C.H. Use of indicator bacteria for monitoring sanitary quality of raw milk cheeses—A literature review. Food Microbiol. 2020, 85, 103283. [Google Scholar] [CrossRef]
- Gonzales-Barron, U.; Gonçalves-Tenório, A.; Rodrigues, V.; Cadavez, V. Foodborne pathogens in raw milk and cheese of sheep and goat origin: A meta-analysis approach. Curr. Opin. Food Sci. 2017, 18, 7–13. [Google Scholar] [CrossRef]
- Desmarchelier, P.; Fegan, N. Pathogens in Milk: Escherichia coli. In Encyclopedia of Dairy Sciences, 2nd ed.; Fuquay, J.W., Ed.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 60–66. [Google Scholar]
- Krebs de Souza, C.; Angioletti, B.L.; Hoffmann, T.G.; Bertoli, S.L.; Reiter, M.G. Promoting the appreciation and marketability of artisanal Kochkäse (traditional German cheese): A review. Int. Dairy J. 2022, 126, 105244. [Google Scholar] [CrossRef]
- Piqueras, J.; Chassard, C.; Callon, C.; Rifa, E.; Rifa, S.; Lebecque, A.D. Lactic Starter Dose Shapes S. aureus and STEC O26:H11 Growth, and Bacterial Community Patterns in Raw Milk Uncooked Pressed Cheeses. Microorganisms 2021, 9, 1081. [Google Scholar] [CrossRef]
- Costanzo, N.; Ceniti, C.; Santoro, A.; Clausi, M.T.; Casalinuovo, F. Foodborne Pathogen Assessment in Raw Milk Cheeses. Int. J. Food Sci. 2020, 2020, 3616713. [Google Scholar] [CrossRef] [PubMed]
- Eliskases-Lechner, F.; Guéguen, M.; Panoff, J.M. Geotrichum candidum. In Encyclopedia of Dairy Sciences, 3rd ed.; Mcsweeney, P.L.H., McNamara, J.P., Eds.; Elsevier: Amsterdam, The Netherlands, 2022; Volume 4, pp. 561–569. [Google Scholar]
- Kačániová, M.; Kunová, S.; Štefániková, J.; Felšöciová, S.; Godočíková, L.; Horská, E.; Nagyová, Ľ.; Haščík, P.; Terentjeva, M. Microbiota of the traditional Slovak sheep cheese “Bryndza”. J. Microbiol. Biotechnol. Food Sci. 2019, 9, 482–486. [Google Scholar] [CrossRef]
- Koňuchová, M.; Valík, Ľ. Modelling the Radial Growth of Geotrichum candidum: Effects of Temperature and Water Activity. Microorganisms 2021, 9, 532. [Google Scholar] [CrossRef] [PubMed]
- Marcellino, S.N.; Benson, D.R. The good, the bad, and the ugly: Tales of mold-ripened cheese. In Cheese and Microbes, 1st ed.; Donnelly, C.W., Ed.; ASM Press: Washington, DC, USA, 2014; pp. 95–132. [Google Scholar]
- Fusco, V.; Chieffi, D.; De Angelis, M. Fresh pasta filata cheeses: Composition, role, and evolution of the microbiota in their quality and safety. J. Dairy Sci. 2022, 105, 9347–9366. [Google Scholar] [CrossRef] [PubMed]
- Mayo, B.; Rodríguez, J.; Vázquez, L.; Flórez, A.B. Microbial Interactions within the Cheese Ecosystem and Their Application to Improve Quality and Safety. Foods 2021, 10, 602. [Google Scholar] [CrossRef] [PubMed]
- Ačai, P.; Valík, Ľ.; Medveďová, A. One- and Two-Step Kinetic Data Analysis Applied for Single and Co-Culture Growth of Staphylococcus aureus, Escherichia coli, and Lactic Acid Bacteria in Milk. Appl. Sci. 2021, 11, 8673. [Google Scholar] [CrossRef]
- Lobacz, A.; Kowalik, J.; Zulewska, J. Determination of the survival kinetics of Salmonella spp. on the surface of ripened raw milk cheese during storage at different temperatures. Int. J. Food Sci. Technol. 2020, 55, 610–618. [Google Scholar] [CrossRef]
- Valík, Ľ.; Ačai, P.; Medveďová, A. Application of competitive models in predicting the simultaneous growth of Staphylococcus aureus and lactic acid bacteria in milk. Food Control 2018, 87, 145–152. [Google Scholar] [CrossRef]
- Valík, Ľ.; Görner, F.; Sonneveld, C. Fermentation of ewe’s lump cheese under conditions of artisan production. Breed. Sheep Goats 2004, 24, 23–24. [Google Scholar]
- Palo, V.; Kaláb, M. Slovak sheep cheeses. Milchwissenshaft 1984, 39, 518–521. [Google Scholar]
- Šipošová, P.; Koňuchová, M.; Valík, Ľ.; Medveďová, A. Growth dynamics of lactic acid bacteria and dairy microscopic fungus Geotrichum candidum during their co-cultivation in milk. Food Sci. Technol. Int. 2020, 27, 572–582. [Google Scholar] [CrossRef] [PubMed]
- Ačai, P.; Medveďová, A.; Mančušková, T.; Valík, Ľ. Growth prediction of two bacterial populations in co-culture with lactic acid bacteria. Food Sci. Technol. Int. 2019, 25, 692–700. [Google Scholar] [CrossRef] [PubMed]
- Ačai, P.; Valík, Ľ.; Medveďová, A.; Rosskopf, F. Modelling and predicting the simultaneous growth of Escherichia coli and lactic acid bacteria in milk. Food Sci. Technol. Int. 2016, 22, 475–484. [Google Scholar] [CrossRef]
- EN ISO 15214; Microbiology of Food and Animal Feeding Stuffs. Horizontal Method for the Enumeration of Mesophilic Lactic Acid Bacteria. Colony-Count Technique at 30 °C. International Organization of Standardization (ISO): Geneva, Switzerland, 2005.
- EN ISO 21527-1; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Yeasts and Moulds—Part 1: Colony Count Technique in Products with Water Activity Greater Than 0.95. International Organization of Standardization (ISO): Geneva, Switzerland, 2008.
- National Standard Method F23; Enumeration of Enterobacteriaceae by the Colony Count Technique. Standards Unit, Evaluations and Standards Laboratory, Health Protection Agency: Hong Kong, China, 2005; p. 23.
- EN ISO 6888-1; Microbiology of Food and Animal Feeding Stuffs. Horizontal Method for the Enumeration of Coagulase-Positive Staphylococci (Staphylococcus aureus and Other Species). Part 1: Technique Using Baird-Parker Agar Medium. International Organization of Standardization (ISO): Geneva, Switzerland, 2001.
- Huang, L. Optimization of a new mathematical model for bacterial growth. Food Control 2013, 32, 283–288. [Google Scholar] [CrossRef]
- Ratkowsky, D.A.; Olley, J.; McMeekin, T.A.; Ball, A. Relationship Between Temperature and Growth Rate of Bacterial Cultures. J. Bacteriol. 1982, 149, 1–5. [Google Scholar] [CrossRef]
- Buchanan, R.L. Predictive food microbiology. Trends Food Sci. Technol. 1993, 4, 6–11. [Google Scholar] [CrossRef]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; Van’t Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 1990, 56, 1875–1881. [Google Scholar] [CrossRef]
- Huang, L. Mathematical modeling and validation of growth of Salmonella Enteritidis and background microorganisms in potato salad—One-step kinetic analysis and model development. Food Control 2016, 68, 69–76. [Google Scholar] [CrossRef]
- Medveďová, A.; Kočiš-Kovaľ, M.; Valík, Ľ. Effect of salt and temperature on the growth of Escherichia coli PSII. Acta Aliment. 2021, 50, 180–188. [Google Scholar]
- Baranyi, J.; Pin, C.; Ross, T. Validating and comparing predictive models. Int. J. Food Microbiol. 1999, 48, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Hwang, C.; Liu, Y.; Renye, J.; Jia, Z. Growth competition between lactic acid bacteria and Listeria monocytogenes during simultaneous fermentation and drying of meat sausages—A mathematical modeling. Food Res. Int. 2022, 158, 111553. [Google Scholar] [CrossRef] [PubMed]
- Licitra, G.; Caccamo, M.; Lortal, S. Artisanal Products Made With Raw Milk. In Raw Milk: Balance between Hazards and Benefits; Nero, L.A., De Carvalho, A.F., Eds.; Academic Press: London, UK, 2019; pp. 175–221. [Google Scholar] [CrossRef]
- Šipošová, P.; Lehotová, V.; Valík, Ľ.; Medveďová, A. Microbiological quality assessment of raw milk from a vending machine and of traditional Slovakian pasta filata cheeses. J. Food Nutr. Res. 2020, 59, 272–279. [Google Scholar]
- Lehotová, V.; Antálková, V.; Medveďová, A.; Valík, Ľ. Quantitative Microbiological Analysis of Artisanal Stretched Quantitative Microbiological Analysis of Artisanal Stretched. Appl. Sci. 2021, 11, 2680. [Google Scholar] [CrossRef]
- Bettera, L.; Levante, A.; Bancalari, E.; Bottari, B.; Gatti, M. Lactic acid bacteria in cow raw milk for cheese production: Which and how many? Front. Microbiol. 2023, 13, 1092224. [Google Scholar] [CrossRef]
- Schoustra, S.; van der Zon, C.; Groenenboom, A.; Moonga, H.B.; Shindano, J.; Smid, E.J.; Hazeleger, W. Microbiological safety of traditionally processed fermented foods based on raw milk, the case of Mabisi from Zambia. LWT Food Sci. Technol. 2022, 171, 113997. [Google Scholar] [CrossRef]
- Medveďová, A.; Koňuchová, M.; Kvočiková, K.; Hatalová, I.; Valík, Ľ. Effect of Lactic Acid Bacteria Addition on the Microbiological Safety of Pasta-Filata Types of Cheeses. Front. Microbiol. 2020, 11, 612528. [Google Scholar] [CrossRef]
- Medveďová, A.; Valík, Ľ. Staphylococcus aureus: Characterisation and Quantitative Growth Description in Milk and Artisanal Raw Milk Cheese Production. In Structure and Function of Food Engineering; Eissa, A.A., Ed.; InTech: Rijeka, Croatia, 2012; pp. 71–102. [Google Scholar]
- Dalcanton, F.; Carrasco, E.; Pérez-Rodríguez, F.; Posada-Izquierdo, G.D.; Falcão de Aragão, G.M.; García-Gimeno, R.M. Modeling the Combined Effects of Temperature, pH, and Sodium Chloride and Sodium Lactate Concentrations on the Growth Rate of Lactobacillus plantarum ATCC 8014. J. Food Qual. 2018, 2018, 1726761. [Google Scholar] [CrossRef]
- Medveďová, A.; Šipošová, P.; Mančušková, T.; Valík, Ľ. The effect of salt and temperature on the growth of Fresco culture. Fermentation 2019, 5, 3390. [Google Scholar] [CrossRef]
- Rodríguez-Sánchez, S.; Ramos, I.M.; Rodríguez-Pérez, M.; Poveda, J.M.; Seseña, S.; Palop, M.L. Lactic acid bacteria as biocontrol agents to reduce Staphylococcus aureus growth, enterotoxin production and virulence gene expression. LWT 2022, 170, 114025. [Google Scholar] [CrossRef]
- Sreekumar, O.; Hosono, A. Immediate effect of Lactobacillus acidophilus on the intestinal flora and fecal enzymes of rats and the in vitro inhibition of Escherichia coli in coculture. J. Dairy Sci. 2000, 83, 931–939. [Google Scholar] [CrossRef]
- Medveďová, A.; Liptáková, D.; Hudecová, A.; Valík, Ľ. Quantification of the growth competition of lactic acid bacteria: A case of co-culture with Geotrichum candidum and Staphylococcus aureus. Acta Chim. Slovaca 2008, 1, 192–201. [Google Scholar]
- Aldarf, M.; Fourcade, F.; Amrane, A.; Prigent, Y. Diffusion of lactate and ammonium in relation to growth of Geotrichum candidum at the surface of solid media. Biotechnol. Bioeng. 2004, 87, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Medveďová, A.; Havlíková, A.; Lehotová, V.; Valík, Ľ. Staphylococcus aureus 2064 growth as affected by temperature and reduced water activity. Ital. J. Food Saf. 2019, 8, 188–193. [Google Scholar] [CrossRef] [PubMed]
- Canon, F.; Nidelet, T.; Guédon, E.; Thierry, A.; Gagnaire, V. Understanding the Mechanisms of Positive Microbial Interactions That Benefit Lactic Acid Bacteria Co-cultures. Front. Microbiol. 2020, 11, 2088. [Google Scholar] [CrossRef] [PubMed]
- Šipošová, P.; Koňuchová, M.; Valík, Ľ.; Trebichavská, M.; Medveďová, A. Quantitative Characterization of Geotrichum candidum Growth in Milk. Appl. Sci. 2021, 11, 4619. [Google Scholar] [CrossRef]
- Ross, T. Indices for performance evaluation of predictive models in food microbiology. J. Appl. Microbiol. 1996, 81, 501–508. [Google Scholar] [CrossRef]
| Microorganism | Isolate | Origin |
|---|---|---|
| DVS® Fresco® 1000NG | - | commercial LAB culture; Christian Hansen, Hoersholm, Denmark |
| G. candidum | J | Slovakian traditional cheese “Bryndza” |
| E. coli | Br | Slovakian traditional cheese “Bryndza” |
| PS2 | laboratory-produced pasta-filata cheese from raw cows’ milk | |
| S. aureus | 2064 | Slovakian ewes’ lump cheese |
| 14733 | milk vending machine biofilm |
| Parameters | E. coli (Isolate BR) | E. coli (Isolate PS2) | ||
|---|---|---|---|---|
| In Milk | 1% NaCl in Milk | In Milk | 1% NaCl in Milk | |
| xmax,Lab | 9.34 ± 0.04 | 9.32 ± 0.03 | 9.36 ± 0.04 | 9.33 ± 0.04 |
| xmax,Ec | 4.17 ± 0.16 | 3.95 ± 0.10 | 5.14 ± 0.17 | 5.14 ± 0.10 |
| xmax,Gc | 5.96 ± 0.08 | 6.09 ± 0.10 | 5.72 ± 0.08 | 6.04 ± 0.17 |
| ILE | 1.158 ± 0.093 | 1.254 ± 0.100 | 0.957 ± 0.059 | 0.951 ± 0.054 |
| IEL | 0.526 ± 0.045 | 0.536 ± 0.049 | 0.588 ± 0.042 | 0.513 ± 0.035 |
| kGc | 0.850 ± 0.038 | 0.710 ± 0.025 | 0.931 ± 0.048 | 0.749 ± 0.046 |
| kref | 0.101 ± 0.006 | 0.101 ± 0.006 | 0.133 ± 0.006 | 0.081 ± 0.006 |
| xres,Ec | 0.4 a | 0.42 ± 0.16 | 1.20 ± 0.29 | 0.5 d |
| zEc | 30.67 ± 5.68 | 32.25 d | 6.38 ± 0.70 | 28.21 ± 5.76 |
| bλ,Gc | 0.0109 ± 0.0003 | 0.0101 ± 0.0003 | 0.0096 ± 0.0002 | 0.0085 ± 0.0002 |
| bT,Gc b | 0.0228 b | 0.0228 b | 0.0228 a | 0.0228 a |
| Tmin,Gc b | 0.00 b | 0.00 b | 0.00 a | 0.00 a |
| bλ,Lab c | 0.0343 c | 0.0343 c | 0.0343 b | 0.0343 b |
| bT,Lab c | 0.0384 c | 0.0384 c | 0.0384 b | 0.0384 b |
| Tmin,Lab | 1.11 c | 1.11 c | 1.11 b | 1.11 b |
| bλ,Ec | 0.0493 c | 0.0493 c | 0.0365 ± 0.0045 | 0.0366 ± 0.0044 |
| bT,Ec | 0.0421 c | 0.0421 c | 0.052 c | 0.052 c |
| Tmin,Ec | 4.16 c | 4.16 c | 4.80 c | 4.80 c |
| Parameters | S. aureus (Isolate 2064) | S. aureus (Isolate 14733) | ||
|---|---|---|---|---|
| In Milk | 1% NaCl in Milk | In Milk | 1% NaCl in Milk | |
| xmax,Lab | 9.43 ± 0.03 | 9.40 ± 0.05 | 9.34 ± 0.03 | 9.25 ± 0.03 |
| xmax,Sa | 3.83 ± 0.15 | 4.17 ± 0.11 | 4.43 ± 0.12 | 4.43 ± 0.16 |
| xmax,Gc | 5.65 ± 0.12 | 5.82 ± 0.17 | 5.85 ± 0.11 | 6.04 ± 0.15 |
| ILS | 1.262 ± 0.056 | 1.083 ± 0.057 | 1.064 ± 0.044 | 0.912 ± 0.043 |
| ISL | 0.308 ± 0.144 | 0.174 ± 0.089 | 0.705 ± 0.079 | 0.526 ± 0.054 |
| cLS | - | - | - | - |
| cSL | - | - | - | - |
| kGc | 0.995 ± 0.067 | 0.778 ± 0.058 | 0.906 ± 0.048 | 0.850 ± 0.055 |
| kref | 0.133 ± 0.022 | 0.102 ± 0.007 | 0.107 ± 0.007 | 0.094 ± 0.006 |
| xres,Sa | 1.47 ± 0.13 | 0.3 c | 0.5 c | 0.5 c |
| zSa | 9.46 ± 1.21 | 10.44 ± 0.51 | 11.49 ± 1.18 | 13.79 ± 1.67 |
| bλ,Gc | 0.0092 ± 0.0002 | 0.0086 ± 0.0003 | 0.0104 ± 0.0003 | 0.0086 ± 0.0002 |
| bT,Gc | 0.0228 a | 0.0228 a | 0.0228 a | 0.0228 a |
| Tmin,Gc | 0.00 a | 0.00 a | 0.00 a | 0.00 a |
| bλ,Lab | 0.0343 b | 0.0343 b | 0.0384 b | 0.0384 b |
| bT,Lab | 0.0384 b | 0.0384 b | 1.11 b | 1.11 b |
| Tmin,Lab | 1.11 b | 1.11 b | 0.0302 b | 0.0302 b |
| bλ,Sa | 0.0302 b | 0.0302 b | 0.0409 b | 0.0409 b |
| bT,Sa | 0.0409 b | 0.0409 b | 5.02 b | 5.02 b |
| Tmin,Sa | 5.02 b | 5.02 b | ||
| Indices | E. coli BR | E. coli PS2 | S. aureus 2064 | S. aureus 14733 | ||||
|---|---|---|---|---|---|---|---|---|
| in Milk | 1% NaCl in Milk | in Milk | 1% NaCl in Milk | in Milk | 1% NaCl in Milk | in Milk | 1% NaCl in Milk | |
| SSE | 14.719 | 16.080 | 19.450 | 25.719 | 10.625 | 11.725 | 15.184 | 17.592 |
| R2 | 0.992 | 0.991 | 0.987 | 0.986 | 0.991 | 0.991 | 0.989 | 0.988 |
| p | 10 | 10 | 11 | 11 | 10 | 10 | 10 | 10 |
| RMSE | 0.251 | 0.254 | 0.289 | 0.324 | 0.280 | 0.284 | 0.270 | 0.282 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ačai, P.; Koňuchová, M.; Valík, Ľ. Modelling and Predicting the Growth of Escherichia coli and Staphylococcus aureus in Co-Culture with Geotrichum candidum and Lactic Acid Bacteria in Milk. Appl. Sci. 2023, 13, 8713. https://doi.org/10.3390/app13158713
Ačai P, Koňuchová M, Valík Ľ. Modelling and Predicting the Growth of Escherichia coli and Staphylococcus aureus in Co-Culture with Geotrichum candidum and Lactic Acid Bacteria in Milk. Applied Sciences. 2023; 13(15):8713. https://doi.org/10.3390/app13158713
Chicago/Turabian StyleAčai, Pavel, Martina Koňuchová, and Ľubomír Valík. 2023. "Modelling and Predicting the Growth of Escherichia coli and Staphylococcus aureus in Co-Culture with Geotrichum candidum and Lactic Acid Bacteria in Milk" Applied Sciences 13, no. 15: 8713. https://doi.org/10.3390/app13158713
APA StyleAčai, P., Koňuchová, M., & Valík, Ľ. (2023). Modelling and Predicting the Growth of Escherichia coli and Staphylococcus aureus in Co-Culture with Geotrichum candidum and Lactic Acid Bacteria in Milk. Applied Sciences, 13(15), 8713. https://doi.org/10.3390/app13158713








