Optimization of Discrete Wavelet Transform Feature Representation and Hierarchical Classification of G-Protein Coupled Receptor Using Firefly Algorithm and Particle Swarm Optimization
Abstract
Featured Application
Abstract
1. Introduction
2. G-Protein Coupled Receptor
3. The Procedures of Protein Feature Representation and Classification
3.1. Data Collection
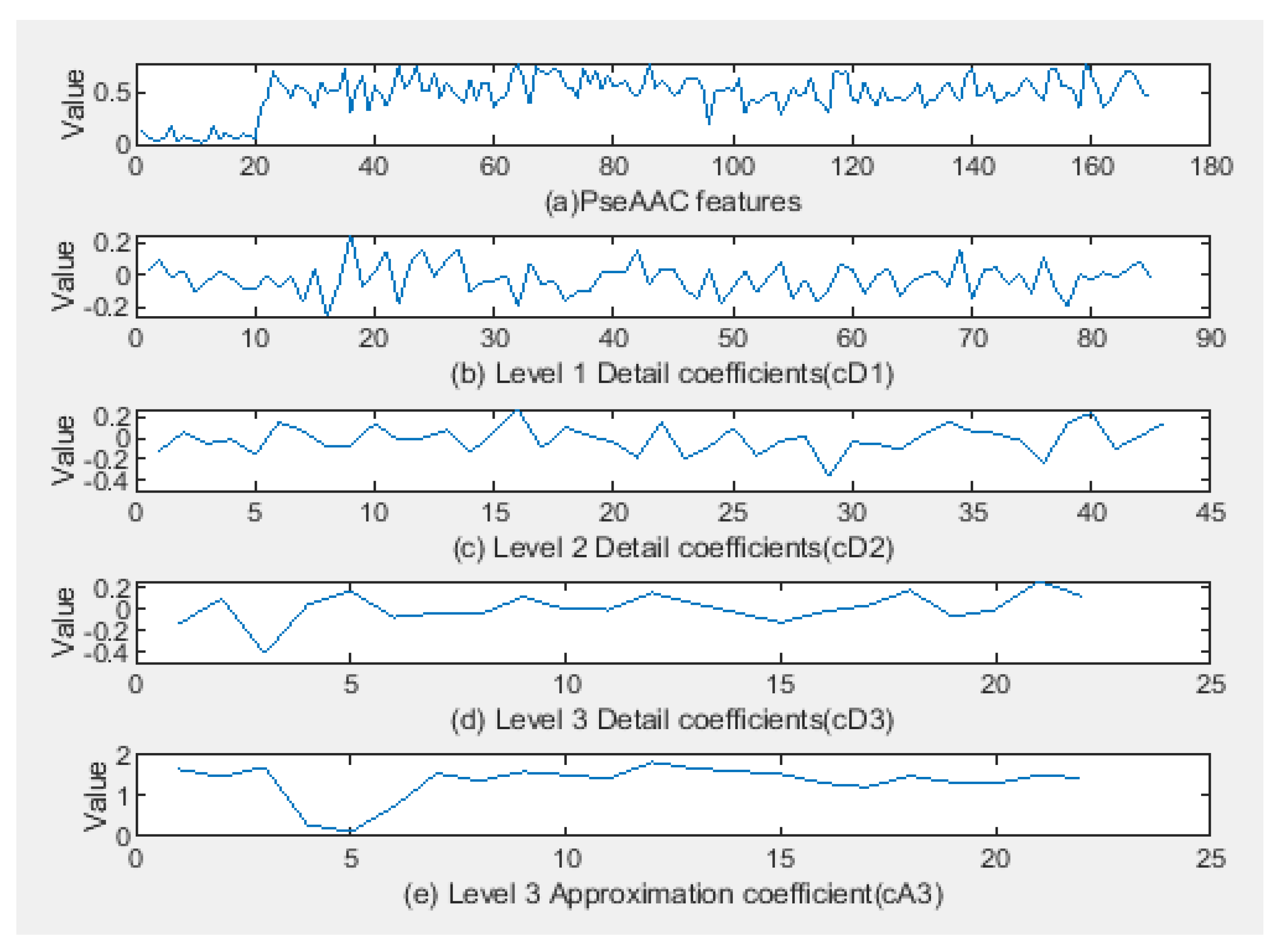
3.2. Feature Representation
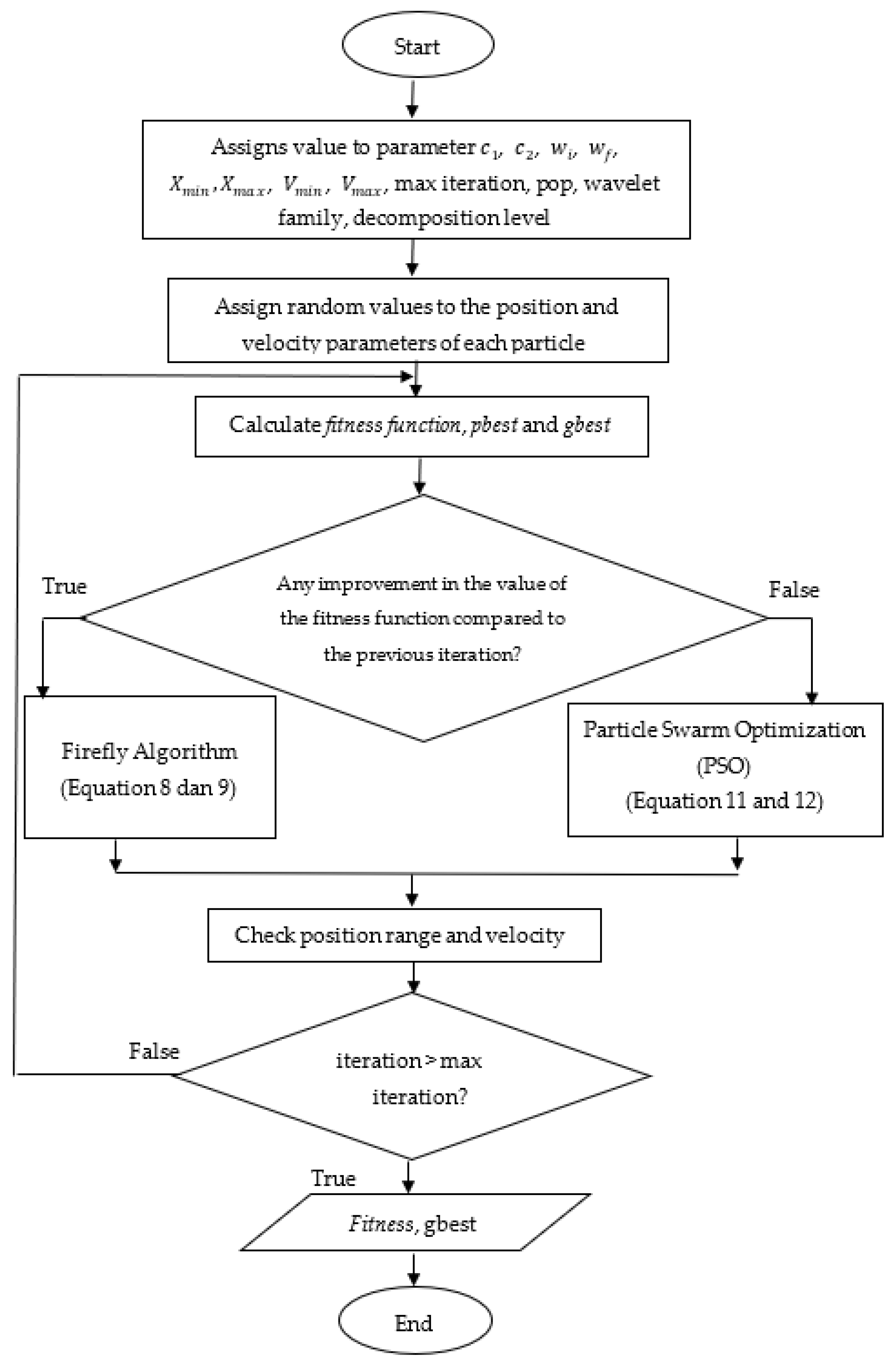
3.3. Hybrid Particle Swarm Optimization Algorithm and Firefly Algorithm (FAPSO)
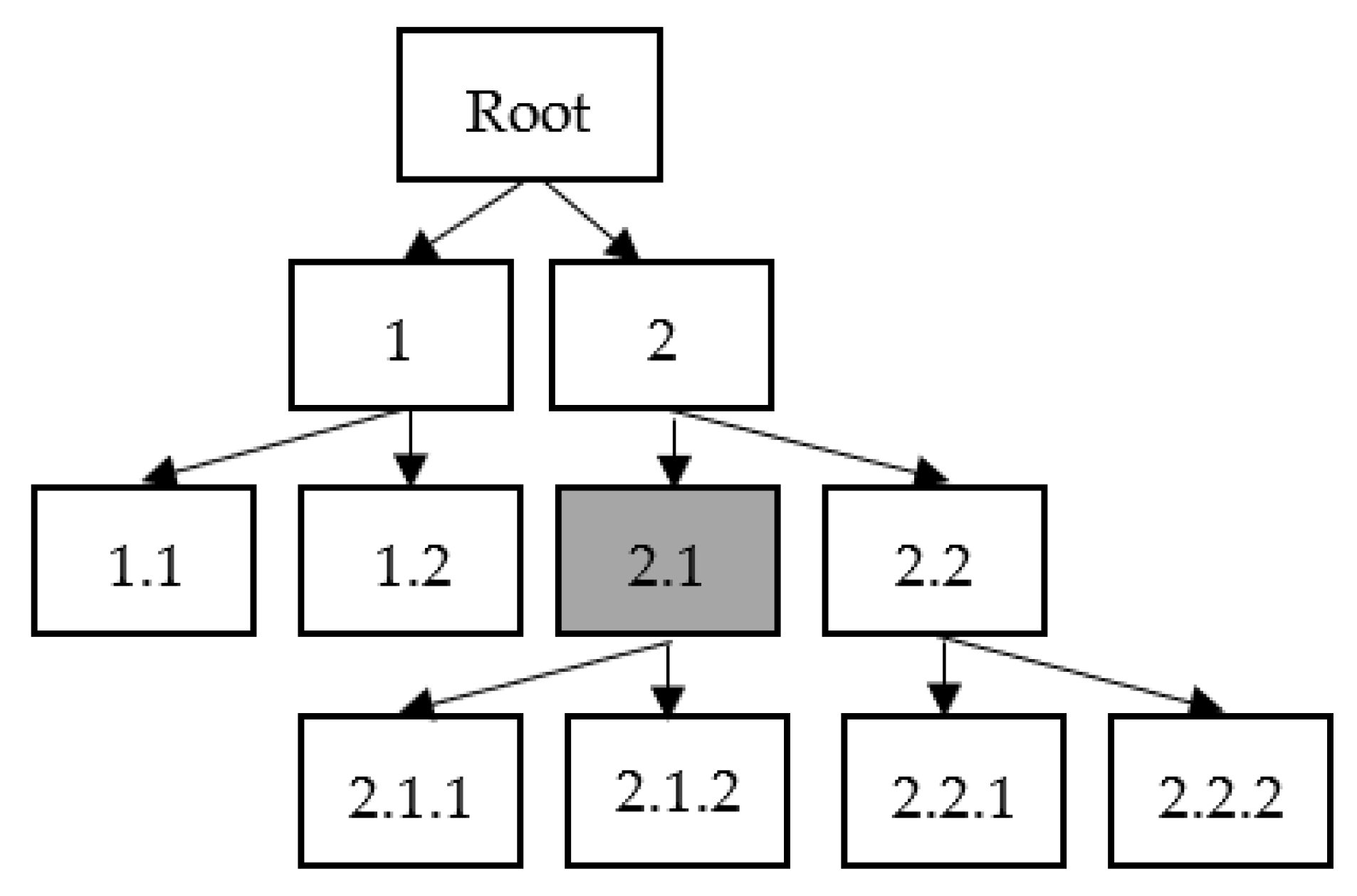
3.4. Hierarchical Classification
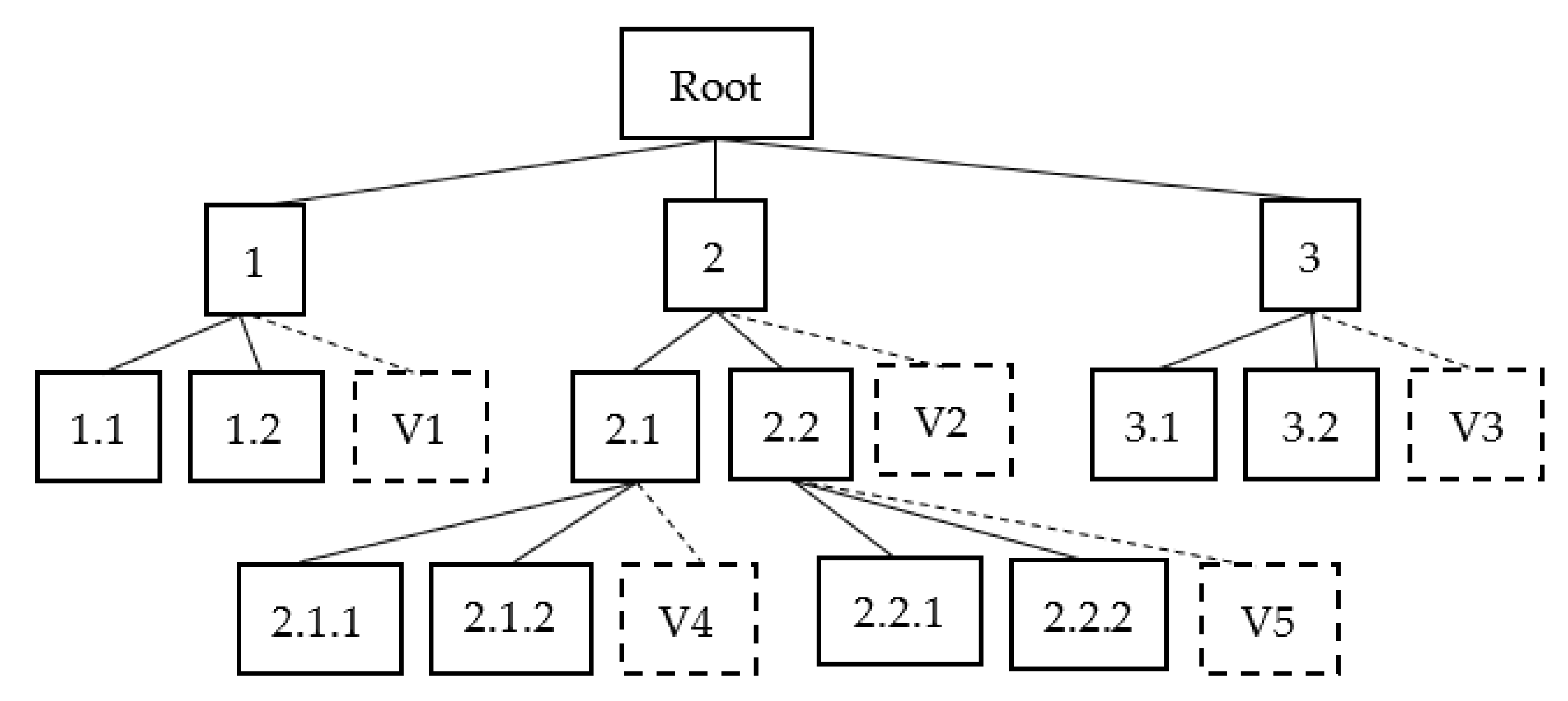
3.5. Hierarchical Classification with Virtual Class
3.6. Performance Evaluation
4. Results
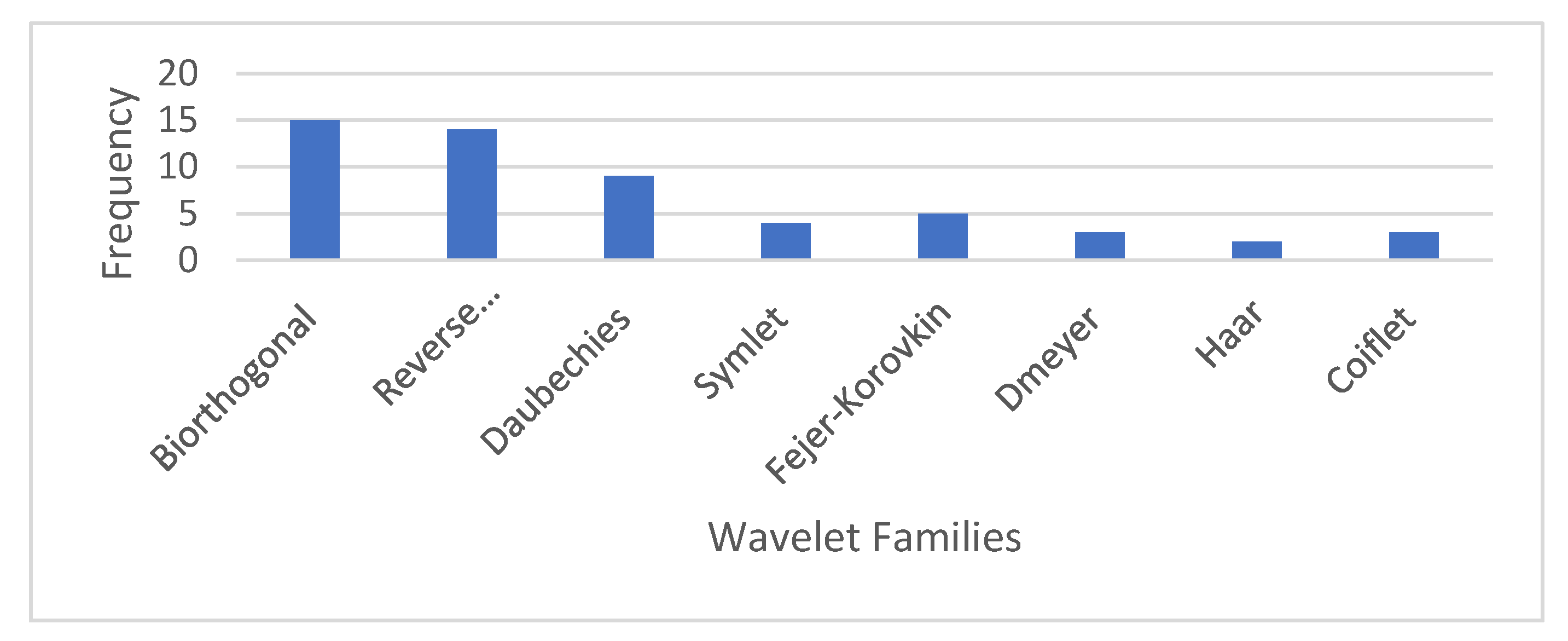
4.1. Selection of Wavelet Family and Decomposition Level
4.2. GPCR Classification Performance without Virtual Class
4.3. GPCR Classification Performance with Virtual Class
5. Discussion
6. Conclusions and Future Works
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| WT | Wavelet Transform |
| DWT | Discrete Wavelet Transform |
| SVM | Support Vector Machine |
| PSO | Particle Swarm Optimization |
| FA | Firefly Algorithm |
| FAPSO | Firefly Algorithm-Particle Swarm Optimization |
| VC | Virtual Class |
| DB | Daubechies |
| Coif | Coiflet |
| Sym | Symlet |
| Dmey | Discrete Meyer |
| FK | Fejer–Korovkin |
| Bior | Biorthogonal |
| RBior | Reverse Biorthogonal |
| GPCR | G-Protein Coupled Receptor |
| PseAAC | Pseudo Amino Acid Composition |
| LCPN | Local Classification at the Parent Node |
| NMLNP | Non-Mandatory Leaf Node Prediction |
References
- Naik, A. Hierarchical Classsification with Rare Categories and Inconsistencies. Ph.D. Thesis, George Mason University, Fairfax, VA, USA, 2017. [Google Scholar]
- Secker, A.; Davies, M.N.; Freitas, A.A.; Clark, E.; Timmis, J.; Flower, D.R. Hierarchical Classification of G-Protein-Coupled Receptors with Data-Driven Selection of Attributes and Classifiers. Int. J. Data Min. Bioinform. 2010, 4, 191–210. [Google Scholar] [CrossRef] [PubMed]
- Bekhouche, S.; Ben Ali, Y.M. Optimizing the Identification of GPCR Function. In Proceedings of the New Challenges in Data Sciences, Kenitra, Morocco, 28–29 March 2019; pp. 1–5. [Google Scholar] [CrossRef]
- Wang, T.; Li, L.; Huang, Y.A.; Zhang, H.; Ma, Y.; Zhou, X. Prediction of Protein-Protein Interactions from Amino Acid Sequences Based on Continuous and Discrete Wavelet Transform Features. Molecules 2018, 23, 823. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.-C. Pseudo Amino Acid Composition and Its Applications in Bioinformatics, Proteomics and System Biology. Curr. Proteom. 2009, 6, 262–274. [Google Scholar] [CrossRef]
- Ru, X.; Wang, L.; Li, L.; Ding, H.; Ye, X.; Zou, Q. Exploration of the Correlation between GPCRs and Drugs Based on a Learning to Rank Algorithm. Comput. Biol. Med. 2020, 119, 103660. [Google Scholar] [CrossRef]
- Ao, C.; Gao, L.; Yu, L. Identifying G-Protein Coupled Receptors Using Mixed-Feature Extraction Methods and Machine Learning Methods. IEEE Access 2020. early access. [Google Scholar] [CrossRef]
- Zhao, Z.Y.; Huang, W.Z.; Zhan, X.K.; Pan, J.; Huang, Y.A.; Zhang, S.W.; Yu, C.Q. An Ensemble Learning-Based Method for Inferring Drug-Target Interactions Combining Protein Sequences and Drug Fingerprints. Biomed Res. Int. 2021, 2021, 9933873. [Google Scholar] [CrossRef]
- Li, Y.; Huang, Y.A.; You, Z.H.; Li, L.P.; Wang, Z. Drug-Target Interaction Prediction Based on Drug Fingerprint Information and Protein Sequence. Molecules 2019, 24, 2999. [Google Scholar] [CrossRef]
- Davies, M.N.; Secker, A.; Freitas, A.A.; Mendao, M.; Timmis, J.; Flower, D.R. On the Hierarchical Classification of G Protein-Coupled Receptors. Bioinformatics 2007, 23, 3113–3118. [Google Scholar] [CrossRef]
- Yu, B.; Li, S.; Qiu, W.Y.; Chen, C.; Chen, R.X.; Wang, L.; Wang, M.H.; Zhang, Y. Accurate Prediction of Subcellular Location of Apoptosis Proteins Combining Chou’s PseAAC and PsePSSM Based on Wavelet Denoising. Oncotarget 2017, 8, 107640–107665. [Google Scholar] [CrossRef]
- Najeeb, S.; Raj, N. Wavelet Analysis in Current Cancer Genome to Identify Driver Mutation. Int. J. Eng. Res. Technol. 2017, 5, 1–7. [Google Scholar]
- Meng, T.; Soliman, A.T.; Shyu, M.L.; Yang, Y.; Chen, S.C.; Iyengar, S.S.; Yordy, J.S.; Iyengar, P. Wavelet Analysis in Current Cancer Genome Research: A Survey. IEEE/ACM Trans. Comput. Biol. Bioinforma. 2013, 10, 1442–1459. [Google Scholar] [CrossRef]
- Kulkarni, O.C.; Vigneshwar, R.; Jayaraman, V.K.; Kulkarni, B.D. Identification of Coding and Non-Coding Sequences Using Local Hölder Exponent Formalism. Bioinformatics 2005, 21, 3818–3823. [Google Scholar] [CrossRef]
- Chen, B.; Li, Y.; Zeng, N. Centralized Wavelet Multiresolution for Exact Translation Invariant Processing of ECG Signals. IEEE Access 2019, 7, 42322–42330. [Google Scholar] [CrossRef]
- Saini, S.; Dewan, L. Performance Comparison of First Generation and Second Generation Wavelets in the Perspective of Genomic Sequence Analysis. Int. J. Pure Appl. Math. 2018, 118, 417–442. [Google Scholar]
- Gayathri, T.T.; Christe, S.A. Wavelet Analysis in Prediction and Identification of Cancerous Genes. Int. J. Sci. Eng. Res. 2017, 8, 720–727. [Google Scholar]
- Hou, W.; Pan, Q.; Peng, Q.; He, M. A New Method to Analyze Protein Sequence Similarity Using Dynamic Time Warping. Genom. J. 2017, 109, 123–130. [Google Scholar] [CrossRef]
- Qiu, J.-D.; Sun, X.-Y.; Huang, J.-H.; Liang, R.-P. Prediction of the Types of Membrane Proteins Based on Discrete Wavelet Transform and Support Vector Machines. Protein J. 2010, 29, 114–119. [Google Scholar] [CrossRef]
- Elbir, A.; Ilhan, H.O.; Serbes, G.; Aydin, N. Short Time Fourier Transform Based Music Genre Classification. In Proceedings of the 2018 Electric Electronics, Computer Science, Biomedical Engineerings' Meeting (EBBT), Istanbul, Turkey, 8–19 April 2018; pp. 1–4. [Google Scholar] [CrossRef]
- Aggarwal, C.C. On Effective Classification of Strings with Wavelets. In Proceedings of the Eighth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Edmonton, AB, Canada, 23–26 July 2002; pp. 163–172. [Google Scholar] [CrossRef]
- Mai, T.D.; Ngo, T.D.; Le, D.D.; Duong, D.A.; Hoang, K.; Satoh, S. Using Node Relationships for Hierarchical Classification. In Proceedings of the 2016 IEEE International Conference on Image Processing (ICIP), Phoenix, AZ, USA, 25–28 September 2016; Volume 2016, pp. 514–518. [Google Scholar] [CrossRef]
- De Trad, C.; Fang, Q.; Cosic, I. An Overview of Protein Sequence Comparisons Using Wavelets. In Proceedings of the IEEE Engineering in Medicine and Biology Society, Istanbul, Turkey, 25–28 October 2001; pp. 115–119. [Google Scholar]
- Liò, P. Wavelets in Bioinformatics and Computational Biology: State of Art and Perspectives. Bioinformatics 2003, 19, 2–9. [Google Scholar] [CrossRef]
- Haimovich, A.D.; Byrne, B.; Ramaswamy, R.; Welsh, W.J. Wavelet Analysis of DNA Walks. J. Comput. Biol. 2006, 13, 1289–1298. [Google Scholar] [CrossRef]
- Germán-Salló, Z.; Strnad, G. Signal Processing Methods in Fault Detection in Manufacturing Systems. Procedia Manuf. 2018, 22, 613–620. [Google Scholar] [CrossRef]
- Alyasseri, Z.A.A.; Khader, A.T.; Al-Betar, M.A.; Abasi, A.K.; Makhadmeh, S.N. EEG Signals Denoising Using Optimal Wavelet Transform Hybridized with Efficient Metaheuristic Methods. IEEE Access 2020, 8, 10584–10605. [Google Scholar] [CrossRef]
- Aprillia, H.; Yang, H.T.; Huang, C.M. Optimal Decomposition and Reconstruction of Discrete Wavelet Transformation for Short-Term Load Forecasting. Energies 2019, 12, 4654. [Google Scholar] [CrossRef]
- Semnani, A.; Wang, L.; Ostadhassan, M.; Nabi-Bidhendi, M.; Araabi, B.N. Time-Frequency Decomposition of Seismic Signals via Quantum Swarm Evolutionary Matching Pursuit. Geophys. Prospect. 2019, 67, 1701–1719. [Google Scholar] [CrossRef]
- Jang, Y.I.; Sim, J.Y.; Yang, J.R.; Kwon, N.K. The Optimal Selection of Mother Wavelet Function and Decomposition Level for Denoising of Dcg Signal. Sensors 2021, 21, 1851. [Google Scholar] [CrossRef]
- He, H.; Tan, Y.; Wang, Y. Optimal Base Wavelet Selection for ECG Noise Reduction Using a Comprehensive Entropy Criterion. Entropy 2015, 17, 6093–6109. [Google Scholar] [CrossRef]
- Ngui, W.K.; Leong, M.S.; Hee, L.M.; Abdelrhman, A.M. Wavelet Analysis: Mother Wavelet Selection Methods. Appl. Mech. Mater. 2013, 393, 953–958. [Google Scholar] [CrossRef]
- Rhif, M.; Abbes, A.B.; Farah, I.R.; Martínez, B.; Sang, Y. Wavelet Transform Application for/in Non-Stationary Time-Series Analysis: A Review. Appl. Sci. 2019, 9, 1345. [Google Scholar] [CrossRef]
- Guarnizo, C.; Orozco, A.A.; Alvarez, M. Optimal Sampling Frequency in Wavelet-Based Signal Feature Extraction Using Particle Swarm Optimization. In Proceedings of the 35th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Osaka, Japan, 3–7 July 2013; Volume 2013, pp. 993–996. [Google Scholar] [CrossRef]
- Caramia, C.; De Marchis, C.; Schmid, M. Optimizing the Scale of a Wavelet-Based Method for the Detection of Gait Events from a Waist-Mounted Accelerometer under Different Walking Speeds. Sensors 2019, 19, 1869. [Google Scholar] [CrossRef]
- Zhang, Z.; Telesford, Q.K.; Giusti, C.; Lim, K.O.; Bassett, D.S. Choosing Wavelet Methods, Filters, and Lengths for Functional Brain Network Construction. PLoS ONE 2016, 11, e0157243. [Google Scholar] [CrossRef]
- Chen, D.; Wan, S.; Xiang, J.; Bao, F.S. A High-Performance Seizure Detection Algorithm Based on Discrete Wavelet Transform (DWT) and EEG. PLoS ONE 2017, 12, e0173138. [Google Scholar] [CrossRef]
- Oltean, G.; Ivanciu, L.N. Computational Intelligence and Wavelet Transform Based Metamodel for Efficient Generation of Not-yet Simulated Waveforms. PLoS ONE 2016, 11, e0146602. [Google Scholar] [CrossRef] [PubMed]
- Tao, H.; Zain, J.M.; Ahmed, M.M.; Abdalla, A.N.; Jing, W. A Wavelet-Based Particle Swarm Optimization Algorithm for Digital Image Watermarking. Integr. Comput. Aided. Eng. 2012, 19, 81–91. [Google Scholar] [CrossRef]
- Abdullah, N.A.; Rahim, N.A.; Gan, C.K.; Adzman, N.N. Forecasting Solar Power Using Hybrid Firefly and Particle Swarm Optimization (HFPSO) for Optimizing the Parameters in a Wavelet Transform-Adaptive Neuro Fuzzy Inference System (WT-ANFIS). Appl. Sci. 2019, 9, 3214. [Google Scholar] [CrossRef]
- Ngo, T.T.; Sadollah, A.; Kim, J.H. A Cooperative Particle Swarm Optimizer with Stochastic Movements for Computationally Expensive Numerical Optimization Problems. J. Comput. Sci. 2016, 13, 68–82. [Google Scholar] [CrossRef]
- Kora, P.; Rama Krishna, K.S. Hybrid Firefly and Particle Swarm Optimization Algorithm for the Detection of Bundle Branch Block. Int. J. Cardiovasc. Acad. 2016, 2, 44–48. [Google Scholar] [CrossRef]
- Aydilek, İ.B. A Hybrid Firefly and Particle Swarm Optimization Algorithm for Computationally Expensive Numerical Problems. Appl. Soft Comput. J. 2018, 66, 232–249. [Google Scholar] [CrossRef]
- Zhang, L.; Shah, S.K.; Kakadiaris, I.A. Hierarchical Multi-Label Classification Using Fully Associative Ensemble Learning. Pattern Recognit. 2017, 70, 89–103. [Google Scholar] [CrossRef]
- Zhu, S.; Wei, X.Y.; Ngo, C.W. Collaborative Error Reduction for Hierarchical Classification. Comput. Vis. Image Underst. 2014, 124, 79–90. [Google Scholar] [CrossRef]
- Nakano, F.K.; Pinto, W.J.; Pappa, G.L.; Cerri, R. Top-down Strategies for Hierarchical Classification of Transposable Elements with Neural Networks. In Proceedings of the 2017 International Joint Conference on Neural Networks, Anchorage, AK, USA, 14–19 May 2017; Volume 2017, pp. 2539–2546. [Google Scholar] [CrossRef]
- Ramírez-Corona, M.; Sucar, L.E.; Morales, E.F. Hierarchical Multilabel Classification Based on Path Evaluation. Int. J. Approx. Reason. 2016, 68, 179–193. [Google Scholar] [CrossRef]
- Ying, C.; Run Ying, D. Novel Top-down Methods for Hierarchical Text Classification. Procedia Eng. 2011, 24, 329–334. [Google Scholar] [CrossRef][Green Version]
- Stein, R.A.; Jaques, P.A.; Valiati, J.F. An Analysis of Hierarchical Text Classification Using Word Embeddings. Inf. Sci. 2019, 471, 216–232. [Google Scholar] [CrossRef]
- Alhosaini, K.; Azhar, A.; Alonazi, A.; Al-Zoghaibi, F. GPCRs: The Most Promiscuous Druggable Receptor of the Mankind. Saudi Pharm. J. 2021, 29, 539–551. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ling, C.; Gao, J. An Efficient CNN-Based Classification on G-Protein Coupled Receptors Using TF-IDF and N-Gram. In Proceedings of the 2017 IEEE Symposium on Computers and Communications (ISCC), Heraklion, Greece, 3–6 July 2017; pp. 924–931. [Google Scholar]
- Davies, M.; Secker, A.; Freitas, A. Optimizing Amino Acid Groupings for GPCR Classification. Bioinformatics 2008, 24, 1980–1986. [Google Scholar] [CrossRef] [PubMed]
- Karchin, R.; Karplus, K.; Haussler, D. Classifying G-Protein Coupled Receptors with Support Vector Machines. Bioinformatics 2002, 18, 147–159. [Google Scholar] [CrossRef]
- Cruz-Barbosa, R.; Ramos-Pérez, E.G.; Giraldo, J. Representation Learning for Class C G Protein-Coupled Receptors Classification. Molecules 2018, 23, 690. [Google Scholar] [CrossRef]
- Li, M.; Ling, C.; Xu, Q.; Gao, J. Classification of G-Protein Coupled Receptors Based on a Rich Generation of Convolutional Neural Network, N-Gram Transformation and Multiple Sequence Alignments. Amino Acids 2018, 50, 255–266. [Google Scholar] [CrossRef]
- Paki, R.; Nourani, E.; Farajzadeh, D. Classification of G Protein-Coupled Receptors Using Attention Mechanism. Gene Rep. 2020, 21, 100882. [Google Scholar] [CrossRef]
- Seo, S.; Oh, M.; Park, Y.; Kim, S. DeepFam: Deep Learning Based Alignment-Free Method for Protein Family Modeling and Prediction. Bioinformatics 2018, 34, i254–i262. [Google Scholar] [CrossRef]
- Qiu, J.-D.; Huang, J.-H.; Liang, R.-P.; Lu, X.-Q. Prediction of G-Protein-Coupled Receptor Classes Based on the Concept of Chou’s Pseudo Amino Acid Composition: An Approach from Discrete Wavelet Transform. Anal. Biochem. 2009, 390, 68–73. [Google Scholar] [CrossRef]
- Guo, Y.Z.; Li, M.; Lu, M.; Wen, Z.; Wang, K.; Li, G.; Wu, J. Classifying G Protein-Coupled Receptors and Nuclear Receptors on the Basis of Protein Power Spectrum from Fast Fourier Transform. Amino Acids 2006, 30, 397–402. [Google Scholar] [CrossRef]
- Tiwari, A.K. Prediction of G-Protein Coupled Receptors and Their Subfamilies by Incorporating Various Sequence Features into Chou’s General PseAAC. Comput. Methods Programs Biomed. 2016, 134, 197–213. [Google Scholar] [CrossRef] [PubMed]
- Naveed, M.; Khan, A.U. GPCR-MPredictor: Multi-Level Prediction of G Protein-Coupled Receptors Using Genetic Ensemble. Amino Acids 2012, 42, 1809–1823. [Google Scholar] [CrossRef] [PubMed]
- Zia-ur-Rehman, B.S.P.; Khan, A. Identifying GPCRs and Their Types with Chou’s Pseudo Amino Acid Composition: An Approach from Multi-Scale Energy Representation and Position Specific Scoring Matrix. Protein Pept. Lett. 2012, 19, 890–903. [Google Scholar] [CrossRef] [PubMed]
- Zekri, M.; Alem, K.; Souici-Meslati, L. Immunological Computation for Protein Function Prediction. Fundam. Inform. 2015, 139, 91–114. [Google Scholar] [CrossRef]
- Rehman, Z.-U.; Mirza, M.T.; Khan, A.; Xhaard, H. Predicting G-Protein-Coupled Receptors Families Using Different Physiochemical Properties and Pseudo Amino Acid Composition. Methods Enzymol. 2013, 522, 61–79. [Google Scholar] [CrossRef]
- Secker, A.; Davies, M.N.; Freitas, A.A.; Timmis, J.; Clark, E.; Flower, D.R. An Artificial Immune System for Clustering Amino Acids in the Context of Protein Function Classification. J. Math. Model. Algorithms 2009, 8, 103–123. [Google Scholar] [CrossRef]
- Gao, Q.B.; Ye, X.F.; He, J. Classifying G-Protein-Coupled Receptors to the Finest Subtype Level. Biochem. Biophys. Res. Commun. 2013, 439, 303–308. [Google Scholar] [CrossRef]
- Shen, H.B.; Chou, K.C. PseAAC: A Flexible Web Server for Generating Various Kinds of Protein Pseudo Amino Acid Composition. Anal. Biochem. 2008, 373, 386–388. [Google Scholar] [CrossRef]
- Dao, F.Y.; Yang, H.; Su, Z.D.; Yang, W.; Wu, Y.; Ding, H.; Chen, W.; Tang, H.; Lin, H. Recent Advances in Conotoxin Classification by Using Machine Learning Methods. Molecules 2017, 22, 1057. [Google Scholar] [CrossRef]
- Shaker, A. Comparison Between Orthogonal and Bi-Orthogonal Wavelets. J. Southwest Jiatong Univ. 2020, 55, 2. [Google Scholar] [CrossRef]
- Ahuja, N.; Lertrattanapanich, L.; Bose, N.K. Properties Determining Choice of Mother Wavelet. IEE Proc. Vis. Image Signal Process. 2005, 152, 205–212. [Google Scholar] [CrossRef]
- Dogra, A.; Goyal, B.; Agrawal, S. Performance Comparison of Different Wavelet Families Based on Bone Vessel Fusion. Asian J. Pharm. 2016, 2016, 9–12. [Google Scholar]
- Yu, B.; Lou, L.; Li, S.; Zhang, Y.; Qiu, W.; Wu, X.; Wang, M.; Tian, B. Prediction of Protein Structural Class for Low-Similarity Sequences Using Chou’s Pseudo Amino Acid Composition and Wavelet Denoising. J. Mol. Graph. Model. 2017, 76, 260–273. [Google Scholar] [CrossRef] [PubMed]
- Silla, C.N., Jr.; Freitas, A. A Survey of Hierarchical Classification across Different Application Domains. Data Min. Knowl. Discov. 2010, 22, 31–72. [Google Scholar] [CrossRef]
- Shen, W.; Wei, Z.; Li, Q.; Zhang, H.; Duoqian, M. Three-Way Decisions Based Blocking Reduction Models in Hierarchical Classification. Inf. Sci. 2020, 523, 63–76. [Google Scholar] [CrossRef]
- Liu, Y.; Dou, Y.; Jin, R.; Li, R.; Qiao, P. Hierarchical Learning with Backtracking Algorithm Based on the Visual Confusion Label Tree for Large-Scale Image Classification. Vis. Comput. 2021, 98, 897–917. [Google Scholar] [CrossRef]
- Yu, B.; Li, S.; Chen, C.; Xu, J.; Qiu, W.; Wu, X.; Chen, R. Prediction Subcellular Localization of Gram-Negative Bacterial Proteins by Support Vector Machine Using Wavelet Denoising and Chou’s Pseudo Amino Acid Composition. Chemom. Intell. Lab. Syst. 2017, 167, 102–112. [Google Scholar] [CrossRef]
- Gu, Q.; Ding, Y.-S.; Zhang, T.-L. Prediction of G-Protein-Coupled Receptor Classes in Low Homology Using Chous Pseudo Amino Acid Composition with Approximate Entropy and Hydrophobicity Patterns. Protein Pept. Lett. 2010, 17, 559–567. [Google Scholar] [CrossRef]
- Juba, B.; Le, H.S. Precision-Recall versus Accuracy and the Role of Large Data Sets. In Proceedings of the AAAI Conference on Artificial Intelligence, Honolulu, Hawaii, USA, 27 January–1 February 2019; Volume 33, pp. 4039–4048. [Google Scholar] [CrossRef]
- Secker, A.; Davies, M.N.; Freitas, A.A.; Timmis, J.; Mendao, M.; Flower, D.R. An Experimental Comparison of Classification Algorithms for the Hierarchical Prediction of Protein Function Classification of GPCRs. Mag. Br. Comput. Soc. Spec. Group AI 2007, 9, 17–22. [Google Scholar]
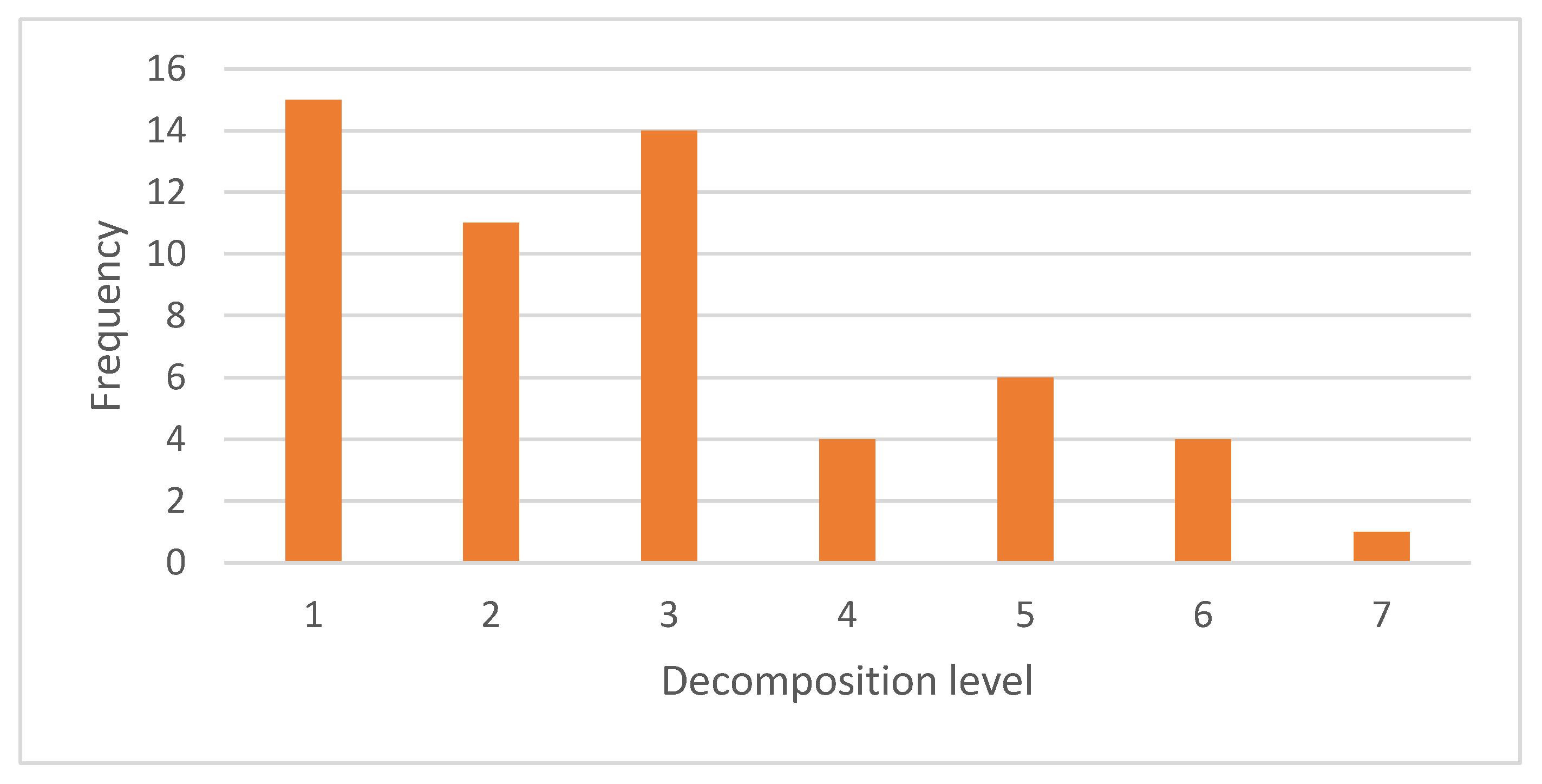
| Parameter | Value |
|---|---|
| 0.9 | |
| 0.5 | |
| 0.2 | |
| 2 | |
| 1 | |
| , | 1.49445 |
| Class Node | Wavelet Family and Decomposition Level |
|---|---|
| Root | (RBior 3.7,1), (Bior4.4,3), (Bio5.5,3), (Coif5,1), (Bior3.7,1) |
| Family A | (Coif4,2), (FK18,3), (Rbio2.2,2), (Db9,1), (Bior5.5,1) |
| Family B | (Db7,1), (Db2,3), (FK6,1), (Db1,1), (Rbio4.4,3) |
| Family C | (Dmey, 6), (Db1,1), (Rbio2.6,1), (Coif3,2), (Dmey,6) |
| Subfamily A Amine | (Bior3.9,1), (Sym10,2), (Rbio1.1,1), (Bior3.3,5), (Sym5,1) |
| Subfamily A Hormone | (Rbior2.2), (Sym9,2), (Bior 3.7,2), (Bior2.2,7), (Bior1.3,5) |
| Subfamily A Nucleotide | (Db3,5), (FK18,6), (Rbio5.5,5), (FK22,3), (Rbior3.8,3) |
| Subfamily A Peptide | (Db7,1), (Bior2.4,1), (Rbio1.1,1), (Db3,1), (FK4,1) |
| Subfamily A Thyro | (Db1,5), (Db3,3), (Db6,4), (Bior1.3, 3), (Rbio1.3,3) |
| Subfamily A Prostanoid | (Bior5.5,3), (RBio3.7,1), (RBio1.1,2), (Rbio1.1,2), (Rbio4.4,5) |
| Subfamily C CalcSense | (Dmey,3), (Rbio3.8,3), (Bior3.3,4), (Bior1.5,6), (Bior33,4) |
| Feature Extraction Method | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| PseAAC | 97.7 | 97.7 | 97.7 | 0.977 |
| FAPSO | 97.9 | 97.9 | 97.9 | 0.979 |
| Feature Extraction Method | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| PseAAC | 85.3 | 87.7 | 87.7 | 0.877 |
| FAPSO | 82.8 | 88.9 | 82.8 | 0.858 |
| Feature Extraction Method | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| PseAAC | 76.9 | 86.2 | 77.4 | 0.816 |
| FAPSO | 71.6 | 87.5 | 81.2 | 0.842 |
| Method | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| PseAAC+VC | 85.4 | 88.3 | 88.3 | 0.883 |
| FAPSO+VC | 86.9 | 94.1 | 90.1 | 0.921 |
| Method | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| PseAAC+VC | 78.8 | 89.6 | 82.6 | 0.86 |
| FAPSO+VC | 81.3 | 92.1 | 85.1 | 0.884 |
| Authors | Super Family | Family | Subfamily | Sub-Subfamily |
|---|---|---|---|---|
| [79] | - | 90.59% | 73.77% | 58.08% |
| [65] | - | 96.97% | 82.72% | 70.46% |
| [61] | 99.75% | 97.38% | 81.91% | 73.34% |
| [64] | - | 97.41% | 84.97% | 75.60% |
| [63] | 98% | 97.5% | 89.2% | 90.95% |
| [56] | - | 97.40% | 87.78% | 81.13% |
| [57] | - | 97.17% | 86.82% | 81.17% |
| This study | - | 97.9% | 86.9% | 81.3% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kamal, N.A.M.; Bakar, A.A.; Zainudin, S. Optimization of Discrete Wavelet Transform Feature Representation and Hierarchical Classification of G-Protein Coupled Receptor Using Firefly Algorithm and Particle Swarm Optimization. Appl. Sci. 2022, 12, 12011. https://doi.org/10.3390/app122312011
Kamal NAM, Bakar AA, Zainudin S. Optimization of Discrete Wavelet Transform Feature Representation and Hierarchical Classification of G-Protein Coupled Receptor Using Firefly Algorithm and Particle Swarm Optimization. Applied Sciences. 2022; 12(23):12011. https://doi.org/10.3390/app122312011
Chicago/Turabian StyleKamal, Nor Ashikin Mohamad, Azuraliza Abu Bakar, and Suhaila Zainudin. 2022. "Optimization of Discrete Wavelet Transform Feature Representation and Hierarchical Classification of G-Protein Coupled Receptor Using Firefly Algorithm and Particle Swarm Optimization" Applied Sciences 12, no. 23: 12011. https://doi.org/10.3390/app122312011
APA StyleKamal, N. A. M., Bakar, A. A., & Zainudin, S. (2022). Optimization of Discrete Wavelet Transform Feature Representation and Hierarchical Classification of G-Protein Coupled Receptor Using Firefly Algorithm and Particle Swarm Optimization. Applied Sciences, 12(23), 12011. https://doi.org/10.3390/app122312011





