1. Introduction
The double-stranded helix of deoxyribonucleic acid (dsDNA) can exist in a great variety of different geometries (conformations), depending on the base stacking interactions, hydrogen bonding, and long-range intra- and inter-backbone forces renormalized by water-mediated interactions with the entities situated in the DNA hydration shells [
1,
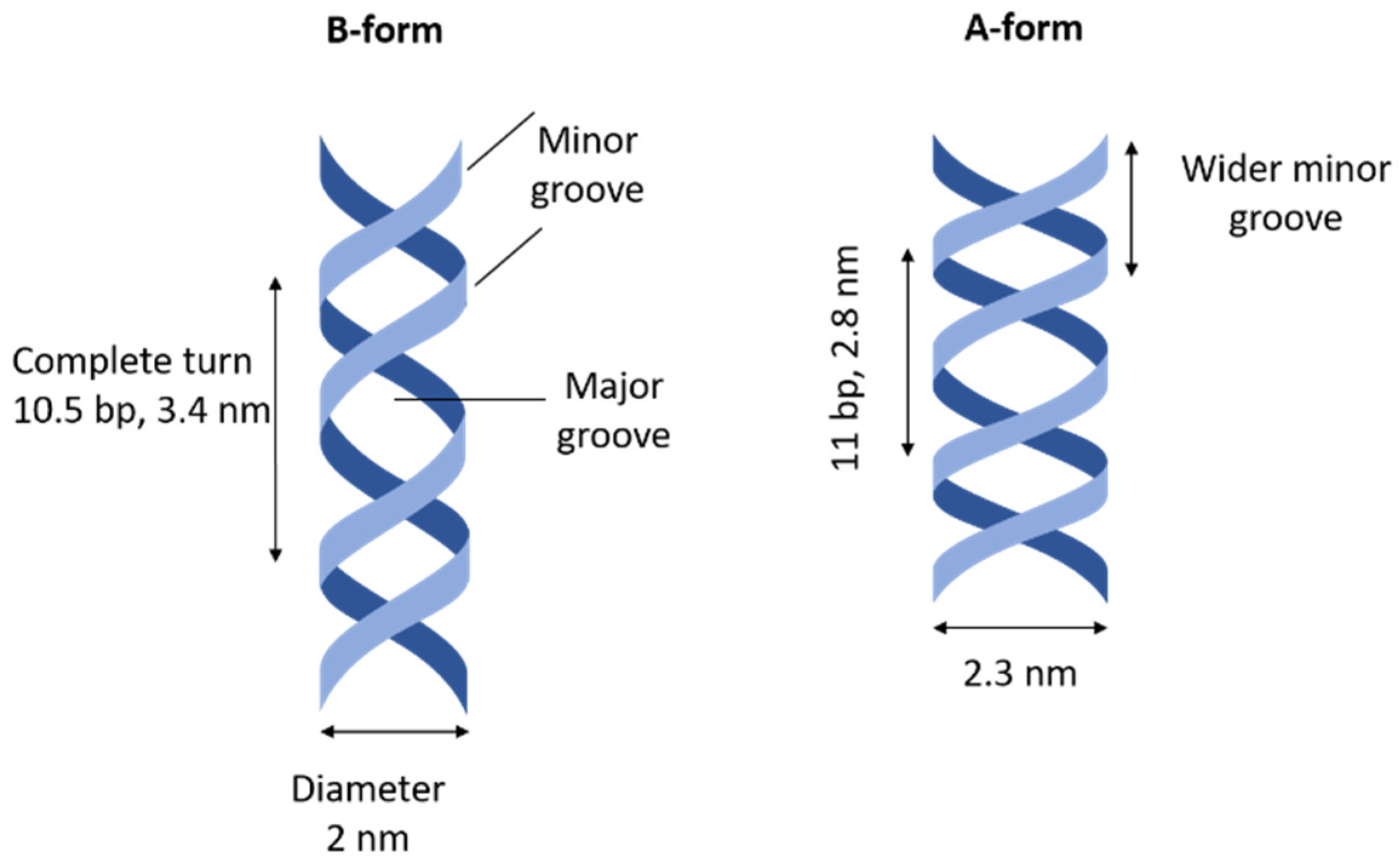
2]. Particularly important and biologically relevant are A-form and B-form helices because they can be found in prokaryotic and eukaryotic cells. In both A- and B-form, the helix is right-handed and the sizes of the major and minor grooves are distinctly defined; A-form dsDNA is broader, shorter, and has a wider minor grove than B-form dsDNA [
1,
3]. In A-form, the sugar-based block adopts the C
3’ endo/anti conformation, and in the B-form, the C
2′ endo/anti conformation. A-form is usually associated with the decreased number of water molecules per phosphate (lower hydration), whereas, in a fully hydrated state, dsDNA adopts B-form [
4]. All the subtleties in the structure between the two forms determine the accessibility of other molecules to DNA binding sites and reveal distinct surfaces of DNA susceptible to different environments. Both forms may be detected simultaneously because segments of DNA can adopt different forms; in this case, %A- and %B-form can be estimated [
2,
5]. Certain conditions may trigger structural A → B and B → A transitions in living cells and organisms and in vitro. These conformational transitions may be a vital step in some processes such as transcription and binding of specific proteins or drugs or can provide protection from desiccation in simple organisms such as bacteria [
2]. Consequently, conformational transitions are often fundamental in the studies of drug- and anti-cancer drug-DNA interaction, resistance to DNA damage induced by radiation, heat, and desiccation, and are extremely important tools in the detection of cancer markers and other abnormalities.
Over the years, structural techniques such as X-ray crystallography and nuclear magnetic resonance spectroscopy (NMR) provided valuable information on conformational heterogeneity in the structures of dsDNA [
6]. While X-ray improved understanding of different crystal structures of DNA, high-resolution NMR experiments provided detailed information on the dsDNA conformation in solution. Both techniques established phosphate–ester conformation as the most variable part of nucleic acid structure [
7]. Thus, a great part of understanding the structure and dynamics of DNA, in addition to protein–DNA recognition and drug–DNA interactions, lies in untangling important roles of phosphate–sugar moieties in DNA. Even though powerful, X-ray crystallography requires the macromolecule to be ordered in a crystal lattice; thus, the conformation of DNA in the crystal structure may not be identical to the one observed in solution or DNA confined in reduced spatial dimensions, such as thin films [
6]. Additionally, NMR studies are usually restricted to short DNA sequences as there is increased difficulty in interpretation of the spectra of biomolecules with large molecular weights [
7,
8]. Thus, different experimental methods such as infrared spectroscopy (IR), UV spectrophotometry, dielectric spectroscopy, etc. have been utilized in order to elucidate DNA structural features in diverse environments.
Infrared spectroscopy methods such as Fourier transform infrared spectroscopy (FTIR) and Raman spectroscopy have long been utilized as non-destructive and sensitive techniques for probing DNA structure and function in different concentrations, ionic strengths, and without limitations to DNA length. In particular, “textbook” A- and B-form geometries were associated with different spectral characteristics; markers of A- and B-form dsDNA were established that are now common in the literature [
2]. Even though there is consensus as to what constitutes spectral profiles of the two forms, inconsistencies can be found because spectra can be obtained from different types of DNA samples, i.e., short oligonucleotides, polynucleotides, genomic or synthetic DNA, DNA isolated from various cells, and tissues. Moreover, spectra can be obtained from various physical states of DNA such as solutions, gels, thin films, and crystals; all of this contributes to the final spectral profile of the studied DNA sample with each state providing different advantages.
In DNA thin films, hydration conditions of the samples can be distinctly controlled by various simple techniques such as air-drying or drying under a stream of inert gasses such as nitrogen, or more advanced techniques that imply the use of home-built humidity chambers or cells where DNA thin films are prepared over the course of several hours (usually after 24 h) [
9]. Thus, a great variety of hydration conditions from fully hydrated to fully dehydrated state can be obtained and dehydration and rehydration experiments can be performed. Additionally, impeding absorptions from the bulk water can be avoided, and with the right substrate, the spectral range can be greatly increased. Another important advantage of DNA thin film is the ability to use these samples in (imaging) techniques such as FTIR microspectroscopy [
10,
11] and atomic force microscopy (AFM) [
12]. These fast-growing techniques require the deposition of DNA on a substrate (thus forming DNA thin films) and have great capabilities of resolving different structural properties of DNA.
Recently, there is a rise in research utilizing different spectroscopic techniques in the field of biomedicine with intention of expanding and improving diagnostic methods making them faster, less expensive, and less invasive [
13]. Even though fluorescence spectroscopy has long been used in clinical settings, due to the development of techniques such as immunofluorescence and fluorescence-activated cell sorting (FACS), IR spectroscopy techniques are not yet common in clinical applications. (FT)IR-based diagnostics is a promising field due to the technique being inexpensive, non-destructive, fast, and easy to operate [
14]. However, in order to establish FTIR spectroscopy techniques in clinical applications several setbacks, such as precise acquisition and unambiguous interpretation of the spectra, statistically acceptable number, and sample size, must be overcome.
FTIR spectroscopy has proven useful in detecting conformational transitions in DNA thin films and related biological samples with the intent of detecting cancer markers [
2,
15,
16,
17] and other abnormalities caused by various types of DNA damage in cells and tissues [
3,
18], before and after drug administration [
19], and tracking anti-cancer drug efficiency [
20,
21]. These studies are dependent on the ability to identify and preserve DNA in certain conformation and rely on the correct interpretation of conformation-related markers [
2]. Thus, in order to improve the ability of IR spectroscopy techniques to be more accurately used in cancer diagnostics, damage detection, and drug efficiency, there is a need to further improve the interpretation of the spectra of DNA in different hydration and conformation states and to distinguish between the two.
In this study, utilizing FTIR spectroscopy, we provide detailed analysis of dynamics of A- to B-form transitions in DNA thin films obtained from sodium salt lyophilized DNA in NaCl solution under different hydration conditions based on a statistical analysis of a substantial number of spectra and extensive band shape analysis (peak fitting) in both the phosphate (1150–1000 cm−1) and sugar–phosphate region (900–750 cm−1). Systematically controlled hydration of DNA thin films provides information on dynamics and reorganization of hydration shells; the conformational transition from A-form to more ordered B-form is observed when sufficient time for conformation transition to take place is allowed. Additionally, a comparison of the spectral signatures and information obtained by peak fitting reveals and distinguishes between hydration and conformation sensitive bands, providing valuable information for various diagnostics purposes.
2. Materials and Methods
Dry salmon testes lyophilized DNA threads obtained from Sigma-Aldrich Corp., St. Louis, MO, USA (DNA sodium salt) were dissolved for 48 h in 10 mM NaCl solution at 4 °C. Routinely performed sample characterization such as protein content, pH, and conductivity was performed and obtained as presented in our previous research [
22]. The final DNA concentration of all solutions was 5 g/L (15 mM DNA phosphate and 7.5 mM DNA base pairs).
In order to obtain thin films, 30 μL of the respective DNA stock solutions were deposited on optical grade silicon transmission windows and placed inside a desiccator chamber under an active vacuum (rotary vacuum pump) from 3 min to 40 min after which thin films were recorded (a) immediately after their removal from the desiccator chamber and (b) 5 min after their removal from the desiccator chamber, thus allowing the samples to come to ambient conditions (atmospheric pressure, room temperature and humidity), i.e., to stabilize.
FTIR spectra of thin films were recorded on a PerkinElmer Waltham, MA, USA Spectrum GX spectrometer equipped with a nitrogen-cooled Mercury-Cadmium-Telluride (MCT) detector and a KBr beam splitter. Spectra were measured at 25 °C and recorded in transmission mode with 4 cm
−1 resolution and 64 co-added scans and later processed in Kinetics add-on for Matlab 2010, MathWorks, Natick, MA, USA. Raw data were baseline-corrected and normalized using the band at 966 cm
−1 (deoxyribose C–C stretching mode) as an internal reference because it showed negligible spectral changes in position or intensity. All spectra obtained under the same conditions were averaged using an eFTIR software, EssentialFTIR, Monona, WI, USA and standard deviations were calculated using OriginPro 2019, OriginLab Corp., Northampton, MA, USA. For easier identification, average spectra obtained under different active vacuum times are abbreviated as 3-min film, 5-min film, etc. and, if not stated otherwise, are assumed to be obtained (b) “after stabilization.” Additional information on thin film preparation and statistical analysis (number of spectra etc.) can be found in
Appendix A.
Band shape analysis (peak fitting) of the phosphate (1150–1000 cm
−1) and sugar–phosphate region (900–750 cm
−1) was performed using OriginPro 2019 and eFTIR software (for details see
Appendix B).
4. Discussion
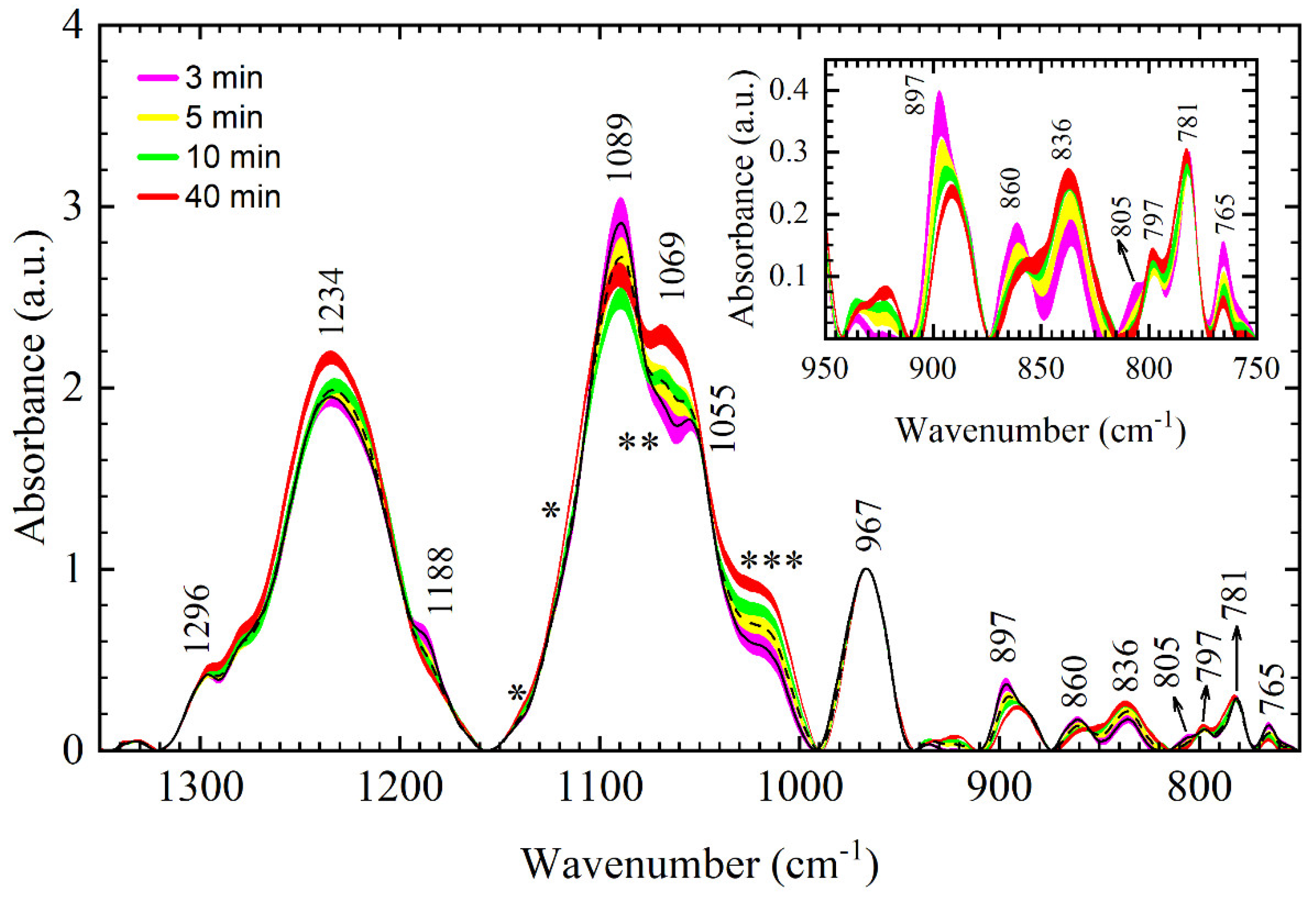
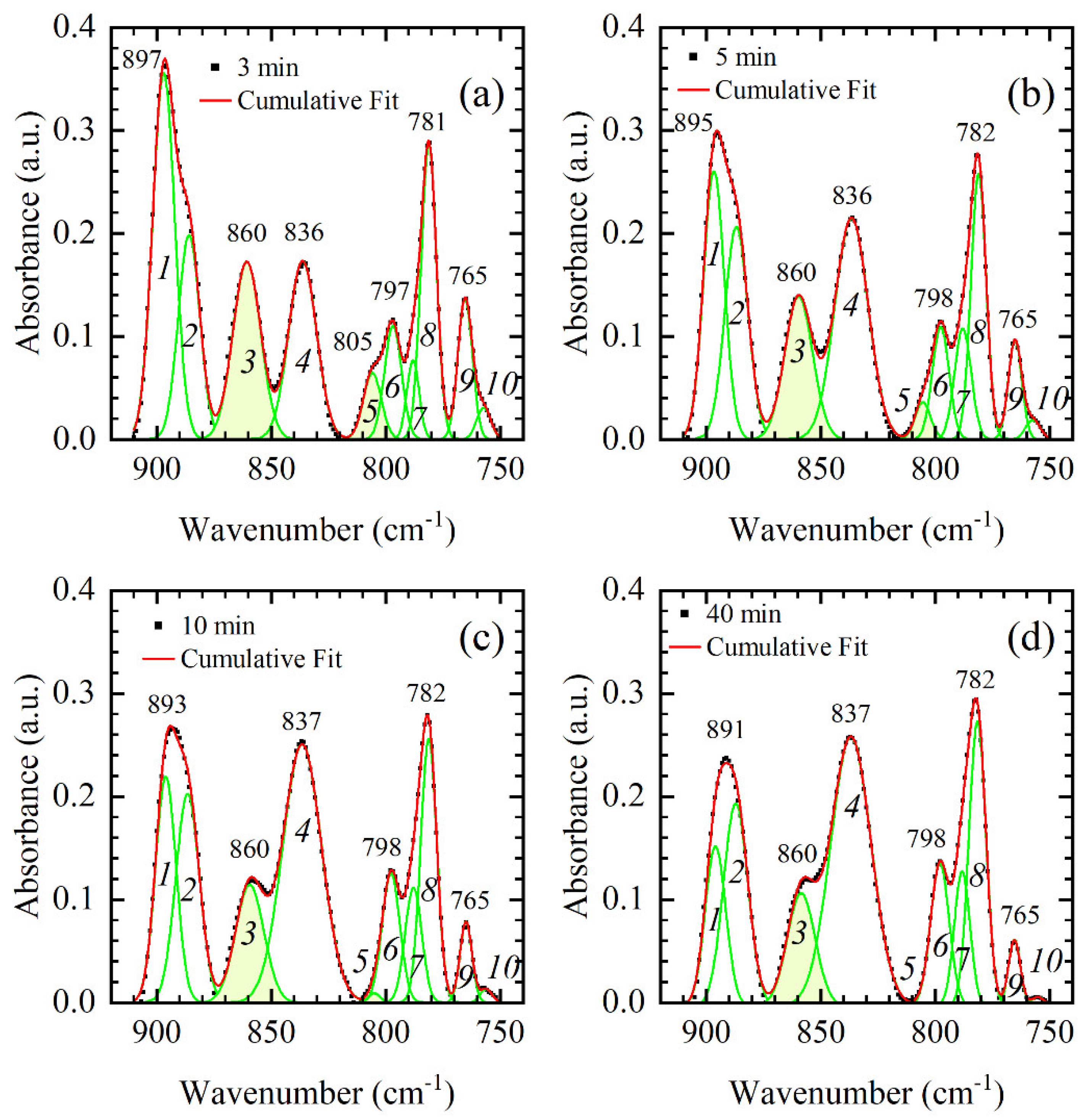
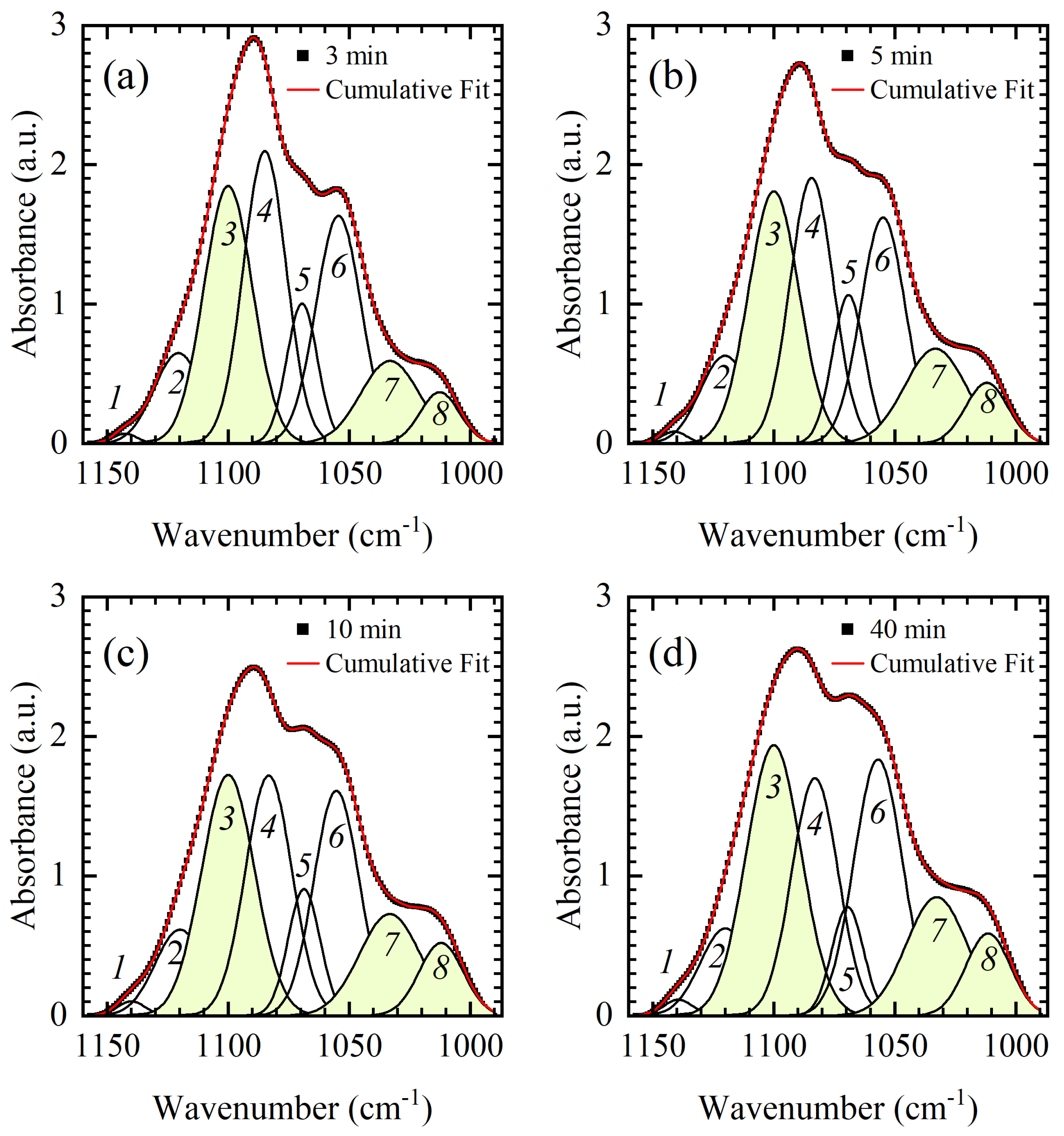
In the previous section, the main differences between DNA thin films were detected and ascribed to either conformational changes or changes in hydration. All thin films studied, 3, 5, 10, 15, 20, 25, and 40 min, can be roughly classified into two groups with respect to their vibrational signature in the backbone region, i.e., 3-min and 5-min films in the first, and 10–40 min films in the second. In the first group, 3-min and 5-min film showed similar spectral features reflecting A-like dsDNA conformation (
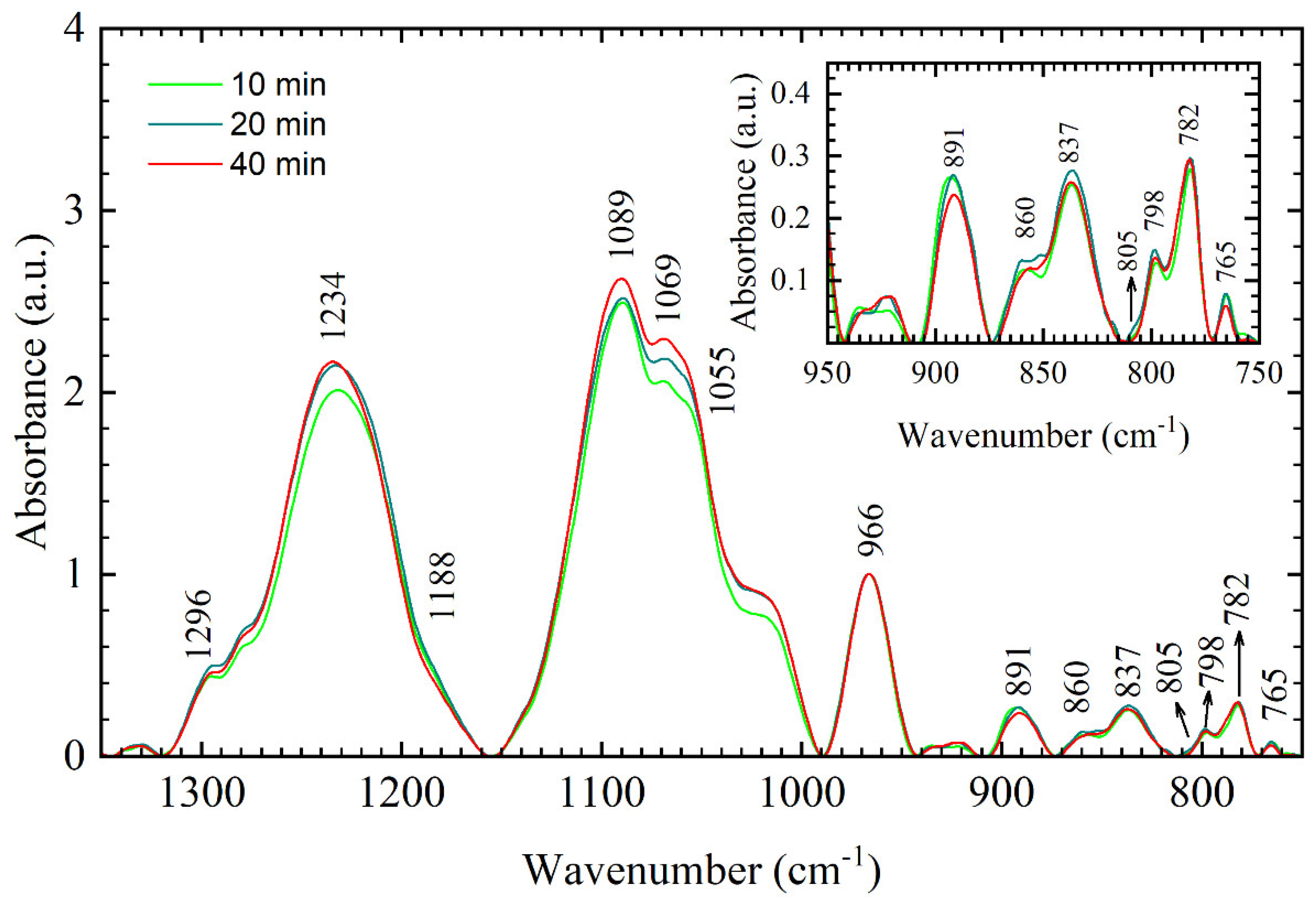
Figure 4). On the other hand, 10–40 min films showed no significant variations in B-like conformation and only a small variation in the phosphodiester bands with respect to the time spent in the vacuum chamber (
Figure 3). Importantly, 10–40 min films have a different spectral signature in the 900–750 cm
−1 region than 3-min and 5-min films (i.e., compare panels of
Figure 5). Band shape analysis presented in
Figure 4,
Figure 5 and
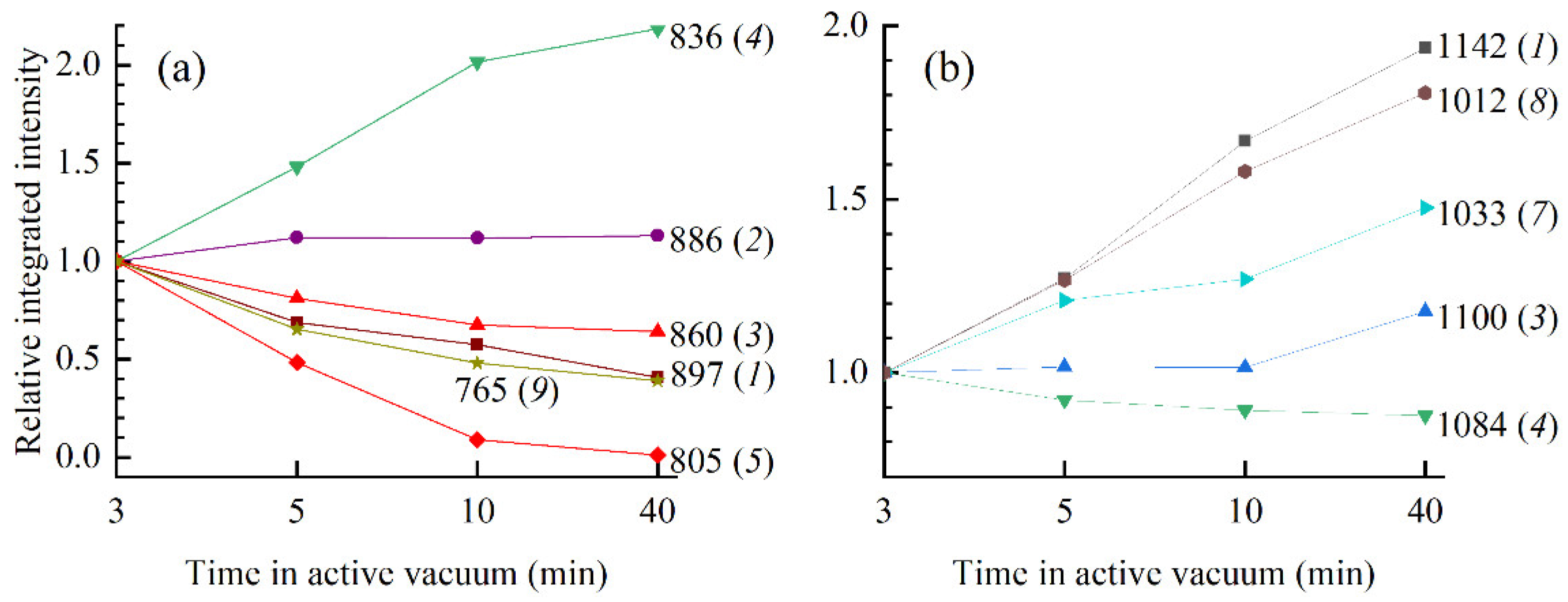
Figure 6 has proven very illuminating in resolving bands and constituent modes sensitive to changes in conformation and hydration. The band at 897 cm
−1 (3 -min film) was found very sensitive to desiccation time; dynamics of modes
1 and
2 resulted in spectral reshaping that can easily be tracked in the spectra and thus might serve as a new spectral signature of conformational transitions. On the other hand, a shoulder near 1030 cm
−1 and a band at 765 cm
−1 showed great sensitivity to changes in hydration conditions without saturation, making them good candidates for tracking changes on the backbone that might have taken place as a result of changes in DNA hydration shells.
In this section, we focus our discussion on two important issues— why the 3-min film reflects more A-like conformation than the 10-min film, and why almost no conformational changes are observed in the spectra after 10 min. It was previously found that the conformational B → A and A → B transitions are related to the changes in hydration, but authors also highlight the importance of dynamics of conformational transitions [
2,
15,
27]. Due to the slow kinetics of the transition, attainment of equilibrium between B- and A-forms caused by rapid changes in hydration may be prevented [
27]. This is why most conformation studies that use specially designed chambers in which hydration is controlled by the use of special salts require slow and step-wise protocols in which samples are maintained from several up to 24 h [
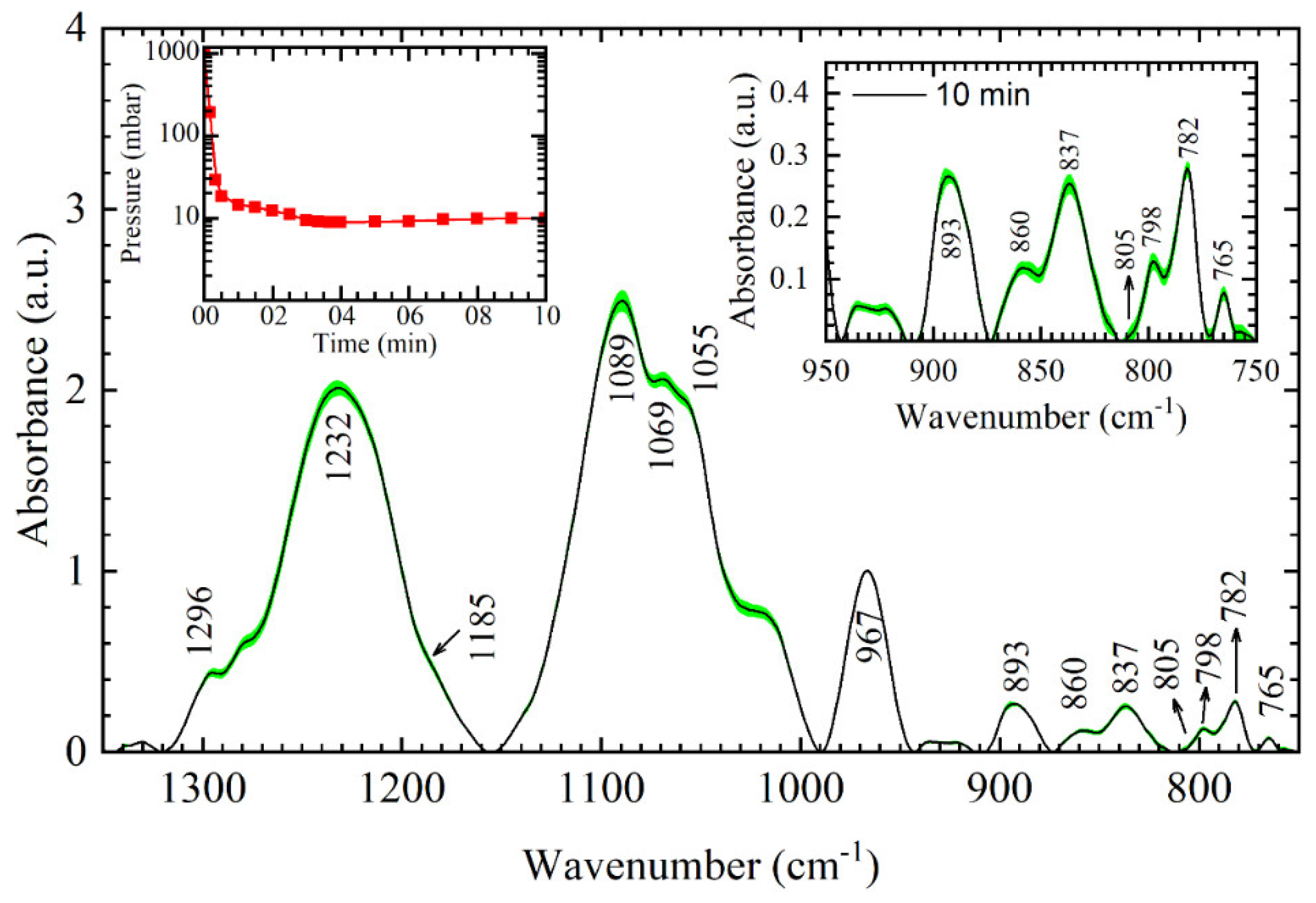
9]. In our study, the opposite is achieved in the desiccator chamber. DNA liquid samples are “shocked” within the first 2 min with the rapid changes in the atmosphere where pressure changes from 1000–10 mbar in the desiccator chamber (left inset of
Figure 1). Approaching 3 min, the formation of the thin film is visible in the chamber and constant pressure of 10 mbar is established. The change of DNA aggregate state from solution to the thin film is thus stopped and 3-min film spectra are taken; DNA spectra reflect more disordered A-like conformation. The level of disorder in the DNA structure has previously been calculated with respect to integrated intensity ratios of the deoxyribose bands at 1069 cm
−1 and 1055 cm
−1 [
20]. Because thin films are allowed more time in the desiccator chamber, the integrated intensity ratio 1069/1055 decreases from 0.42 in the 3-min film to 0.28 in the 40-min film, indicating transition into a more ordered state. If enough time is allowed, after approximately 10 min, structural organization of DNA is complete and 10–40 min thin films reflect similar, B-like conformation. Again, note that after 3–5 min, there are no more changes in the pressure in the desiccator chamber (left inset of
Figure 1); thin films merely stay up to 30 additional minutes in the same atmosphere, which does not have as strong an influence on the spectra as the first few minutes of the process. Consequently, 10–40 min films show similar spectral features in the conformation-related 900–750 cm
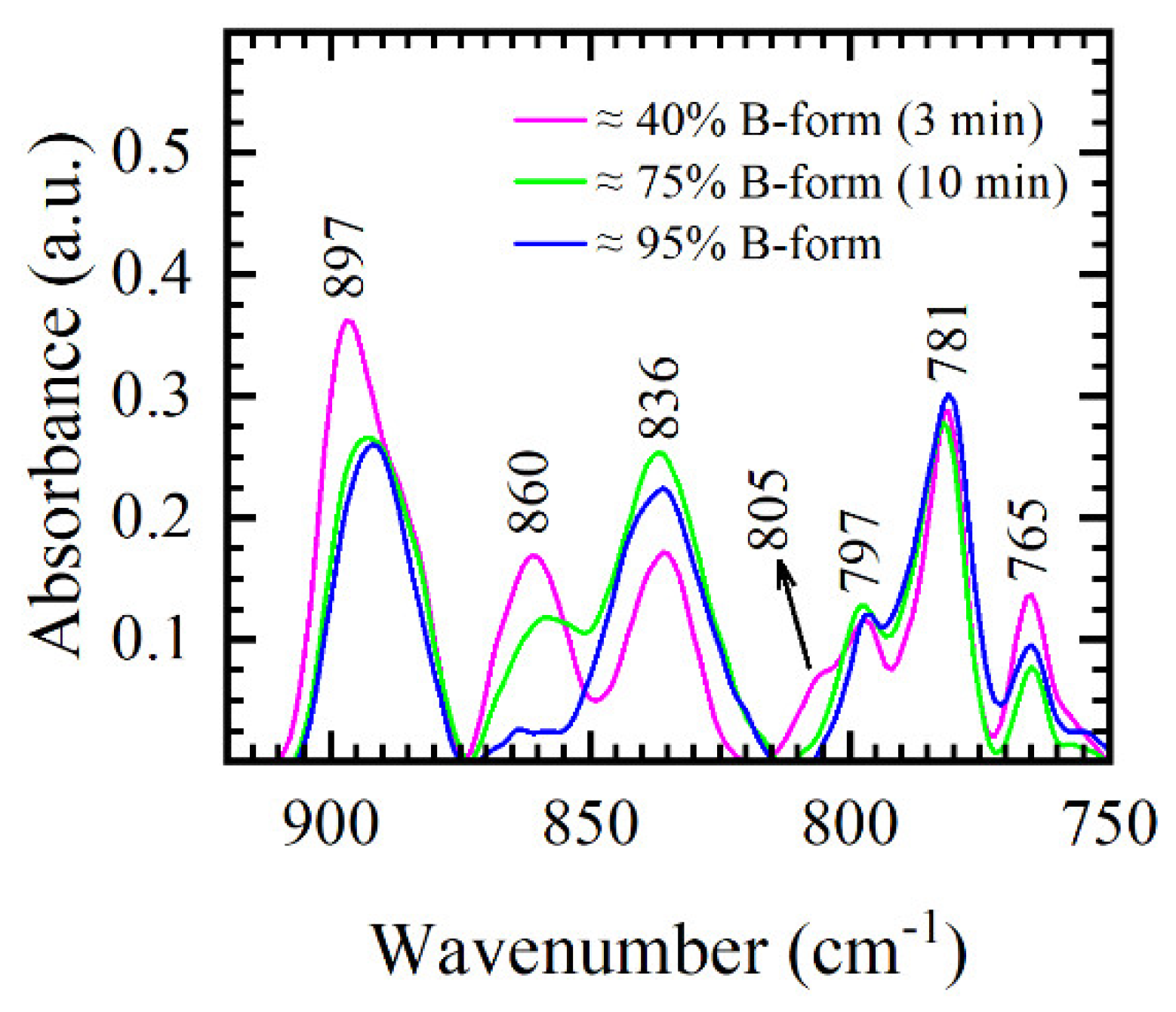
−1 region. Different stages of B-form formation in 3-min and 10-min films, as indicated by the FTIR spectra and calculated using integrated absorptions of 860 cm
−1, 805 cm
−1, and 836 cm
−1 bands, are presented in
Figure 8 and compared to DNA manganese salt thin films obtained in our previous work [
31]. Due to increased screening of Mn
2+ ions, in DNA manganese salt prepared by similar protocol (10 min desiccation time and same DNA type and concentration) complete formation of B-form is evident; 860 cm
−1 and 805 cm
−1 markers are missing in the spectra and ratio of integrated absorptions yield 94% B-form. However, in DNA sodium salt presented in this study, only 805 cm
−1 marker completely vanishes, while 860 cm
−1 marker experiences intensity and integrated intensity decrease, as evident in
Figure 8. Thus, even though 10-min film reflects mostly B-like conformation, some traces of A-form are identified in the spectra. The coexistence of A- and B-forms in DNA in which sections of phosphate groups are in different conformations was also reported by different NMR [
32] and FTIR studies [
2].
Finally, we address the issue of resolving hydration-, conformation-, and ambient-related signatures in the spectra. In a fully hydrated state, DNA is surrounded by several hydration shells. The primary hydration shell of DNA is different from the bulk water and at 80% relative humidity it is considered complete with about 20 water molecules per nucleotide [
1]. In this hydrated state, DNA assumes B-form and changes in hydration may trigger conformational transitions that occur stepwise and cooperatively [
1]. One of the first studies that studied the structural organization of water molecules upon rehydration by Falk et al. concluded that water is first absorbed by the PO
2− groups, then by C–O and C–C groups of the phosphodiester linkage, and finally by the DNA bases [
33]. Pevsner and Diem found their study complementing and observed spectral changes of DNA phase transitions in terms of two concurrent effects—conformational change and hydration of the phosphate groups [
9]. Their study suggested that when the radical conformational transition takes place (fully hydrated DNA sample in B-form opposed to dry DNA pellet in A-form) substantial shifts in the frequency of asymmetric stretching PO
2− vibration from 1225 cm
−1 to 1238 cm
−1 and symmetric stretching PO
2− vibration from 1088 cm
−1 to 1093 cm
−1 occur. On the other hand, hydration effects are recognized by infrared absorption intensity changes of the two phosphate vibrations due to an increase in the local dielectric of the surroundings of the phosphate groups [
9]. A nice correlation with our data can be found for their two hydrated pallets, i.e., 1090 cm
−1 and 1235 cm
−1 and 1088 cm
−1 and 1229 cm
−1, respectively, compared to our thin films 1088 cm
−1 and 1230–1234 cm
−1. However, radical shifts of asymmetric and symmetric vibrations reported in [
9] are not found in our research, supporting less drastic changes to the conformation and the coexistence of both forms in thin films. Unfortunately, the authors do not provide any information on the bands below 950 cm
−1; therefore, the region that contains important A- and B-form markers [
34] cannot be compared to their study.
Our results further support the interpretation of spectral changes with respect to two competing effects (hydration and conformation), although with several subtleties. Our study does not cover DNA transitions in extreme conditions but rather in-between and, more importantly, extends interpretation of the spectra below 950 cm−1 so that correlation with the sugar vibrations can be established. As substantial shifts of asymmetric and symmetric PO2− vibrations are missing in our spectra, phosphate bands alone cannot be used in monitoring conformational transitions; asymmetric PO2− vibration is found in the 1230–1234 cm−1 interval reflecting only “fine-tuning” of the conformation, while symmetric PO2− vibration remains at 1089 cm−1. On the other hand, conformation-related markers at 860 cm−1, 836 cm−1, and 805 cm−1 track conformational transition more effectively; a decrease in A-form from ≈60% to ≈25% in 3-min and 10-min films, respectively, is observed. Additionally, backbone vibration at 893 cm−1 is also found to be sensitive to conformational transitions because it experiences severe spectral reshaping due to dissimilar integrated intensity changes of constituent modes. The band at 893 cm−1 is thus a valuable conformation-related marker, especially if the information on the lower frequency modes is missing in the spectra. Finally, note that changes in 893, 860, 836, and 805 cm−1 eventually saturate, indicating the end of conformational transitions after approximately 10 min in the desiccation chamber. Hydration-related effects, on the other hand, do not seem to saturate. Asymmetric and symmetric PO2− vibration at ≈1234 cm−1 and 1089 cm−1, respectively, and backbone vibrations near 1030 cm−1 and 765 cm−1 exhibit continuous changes without saturation and consequently show great sensitivity to hydration conditions near PO2− groups rather than conformational transitions.
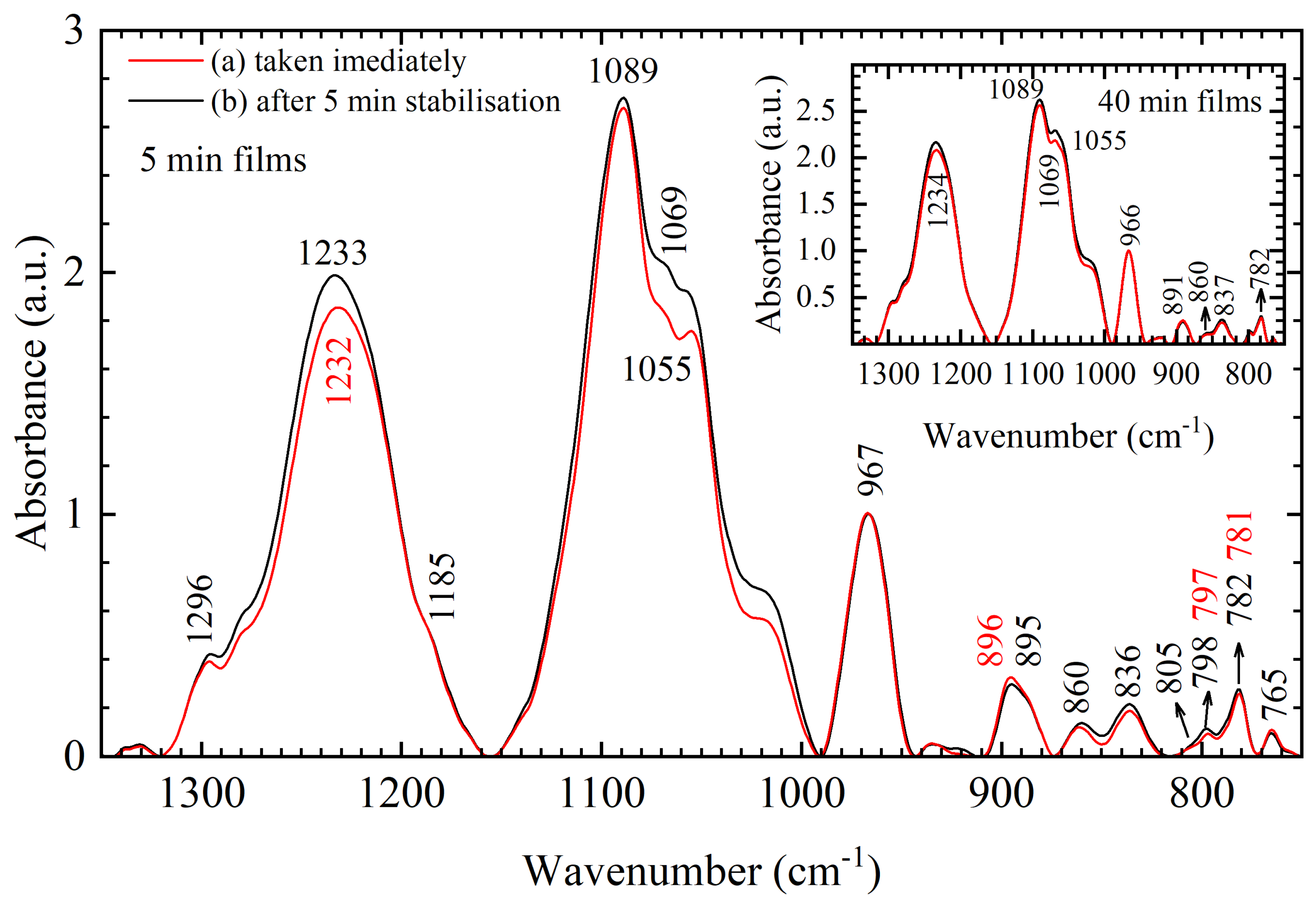
Vibrations of the phosphate groups were also found to be sensitive to ambient conditions, which is best evident from the analysis of 180 samples of 5-min DNA thin films. The 5-min films were obtained in two ways—(a) immediately after their removal from the desiccator chamber and (b) 5 min after their removal from the desiccator chamber, i.e., “before” and “after” stabilization (
Figure 9 and
Figure A1). Average spectra of 5-min films, obtained by either (a) or (b), are in the same conformation as evident by the lack of significant frequency shifts in the entire spectral region from 1350–750 cm
−1 and lack of changes in the intensities of conformation related markers in the 900–750 cm
−1 region (
Figure 9). Only differences in absorption intensities of asymmetric and symmetric PO
2− vibration and backbone vibrations near 1030 cm
−1 are evident in the spectra and are most likely caused by the atmospheric changes that happen as films transition from a vacuum atmosphere to an ambient atmosphere. Furthermore, note that spectral differences between films obtained by (a) and (b) are greatly reduced in 40-min films (inset of
Figure 9), i.e., the influence of ambient conditions can be reduced by increasing the desiccation time. This spectral behavior supports the standpoint in which hydration-, conformation- and ambient-related effects are distinguished during the structural organization of DNA and highlights the importance of conformation-related markers in the, often overlooked, 900–750 cm
−1 region. More on ambient-related spectral features can be found in the second paragraph of
Appendix A and
Figure A1.
Finally, we comment on the importance of reproducibility of the method, i.e., controlling structural transitions utilizing chambers under active vacuum with respect to the sensitivity of the phosphodiester bands. In the light of this study, and with the distinct intention of utilizing FTIR techniques in cancer diagnostics and diagnostics of DNA damage and other abnormalities, special care must be taken in the acquisition and interpretation of the spectra as both drying techniques and small ambient changes may induce a number of spectral changes. Some authors, such as Wood et al., also stressed and recognized the importance of sample-handling in the case of cells fixed by fixatives as ethanol/methanol and samples obtained by air-drying because great variations in the spectra might occur [
2]. This is best evident in
Figure 9 and
Figure A1, in which an increase in the time of thin films in the desiccator chamber almost completely removed the artifacts arising due to ambient conditions (40-min film) versus a similar technique that produced more variability (5-min film). This might prove to be of great importance in situations when one deals with a large number of spectra in terms of differentiating experimental and control groups and in the case of different types of classification and modeling such as k nearest neighbors (KNN), support vector machine algorithms (SVMDA), principal component analysis (PCA), partial least squares (PLS), etc. that are extensively used in biomedical research. Because one of the main obstacles of establishing FTIR spectroscopy in routine applications in clinical environments lies in the unambiguous interpretation of the spectra, the appearance of spectral variations and scattering artifacts must be elucidated and minimized.