Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria
Abstract
1. Introduction
2. Materials and Methods
2.1. The Samples
2.2. Bacterial Strains
2.3. Bacterial Cultivation
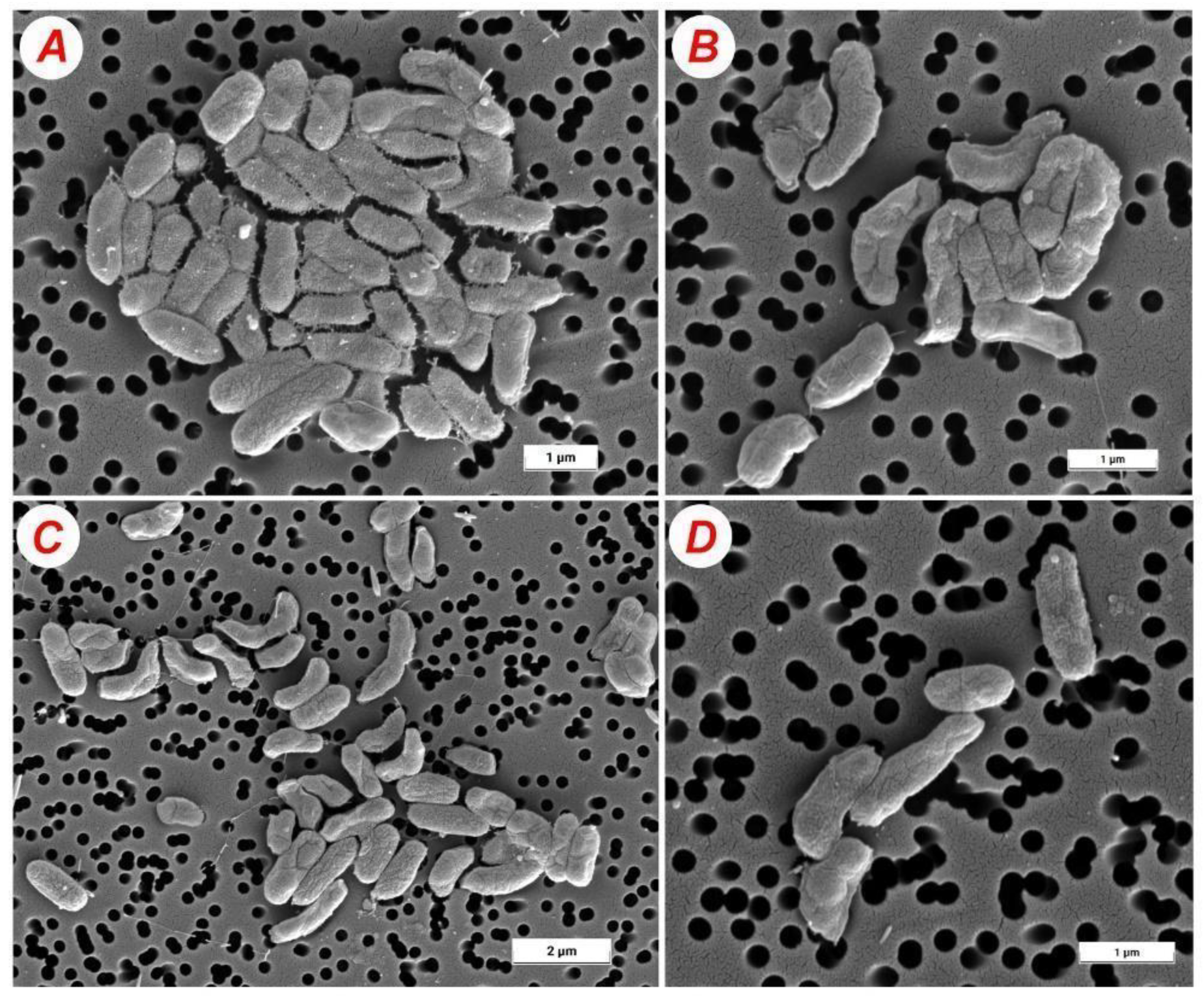
2.4. Morphology of Bacterial Cells
2.5. Measurement of Microelements in Bacterial Cells
2.6. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lennard-Jones, J.E. Classification of inflammatory bowel disease. Scand. J. Gastroenterol. 1989, 24, 2–6. [Google Scholar] [CrossRef]
- Xavier, R.J.; Podolsky, D.K. Unravelling the pathogenesis of inflammatory bowel disease. Nature 2007, 448, 427–434. [Google Scholar] [CrossRef]
- Katz, J.A.; Itoh, J.; Fiocchi, C. Pathogenesis of inflammatory bowel disease. Curr. Opin. Gastroenterol. 1999, 15, 291. [Google Scholar] [CrossRef] [PubMed]
- Danese, S.; Fiocchi, C. Etiopathogenesis of inflammatory bowel diseases. World J. Gastroenterol. 2006, 12, 4807. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Martínková, K.; Vítězová, M.; Rittmann, S.K.-M.R. Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies. J. Clin. Med. 2021, 10, 462. [Google Scholar] [CrossRef]
- Kushkevych, I.; Sangrador, J.C.; Dordević, D.; Rozehnalová, M.; Černý, M.; Fafula, R.; Vítězová, M.; Rittmann, S.K.-M.R. Evaluation of physiological parameters of intestinal sulfate-reducing bacteria isolated from patients suffering from IBD and healthy people. J. Clin. Med. 2020, 9, 1920. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Vítězová, M.; Kos, J.; Kollár, P.; Jampilek, J. Effect of selected 8-hydroxyquinoline-2-carboxanilides on viability and sulfate metabolism of Desulfovibrio piger. J. Appl. Biomed. 2018, 16, 241–246. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kollar, P.; Suchy, P.; Parak, T.; Pauk, K.; Imramovsky, A. Activity of selected salicylamides against intestinal sulfate-reducing bacteria. Neuro Endocrinol. Lett. 2015, 36, 106–113. [Google Scholar]
- Kushkevych, I.V. Kinetic Properties of Pyruvate Ferredoxin Oxidoreductase of Intestinal Sulfate-Reducing Bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Pol. J. Microbiol. 2015, 64, 107–114. [Google Scholar] [CrossRef]
- Kotrsová, V.; Kushkevych, I. Possible methods for evaluation of hydrogen sulfide toxicity against lactic acid bacteria. Biointerface Res. Appl. Chem. 2019, 9, 4066–4069. [Google Scholar]
- Kushkevych, I.; Vítězová, M.; Fedrová, P.; Vochyanová, Z.; Paráková, L.; Hošek, J. Kinetic properties of growth of intestinal sulphate-reducing bacteria isolated from healthy mice and mice with ulcerative colitis. Acta Vet. Brno 2017, 86, 405–411. [Google Scholar] [CrossRef]
- Gibson, G.R.; Cummings, J.H.; Macfarlane, G.T. Growth and activities of sulphate-reducing bacteria in gut contents of health subjects and patients with ulcerative colitis. FEMS Microbiol. Ecol. 1991, 86, 103–112. [Google Scholar] [CrossRef]
- Kováč, J.; Vítězová, M.; Kushkevych, I. Metabolic activity of sulfate-reducing bacteria from rodents with colitis. Open Med. 2018, 13, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Vítězová, M.; Kollár, P. Cross-correlation analysis of the Desulfovibrio growth parameters of intestinal species isolated from people with colitis. Biologia 2018, 73, 1137–1143. [Google Scholar] [CrossRef]
- Cummings, J.H.; Macfarlane, G.T.; Macfarlane, S. Intestinal Bacteria and Ulcerative Colitis. Curr. Issues Intest. Microbiol. 2003, 4, 9–20. [Google Scholar] [PubMed]
- Pitcher, M.C.; Cummings, J.H. Hydrogen sulphide: A bacterial toxin in ulcerative colitis? Gut 1996, 39, 1–4. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Toxicity of hydrogen sulfide toward sulfate-reducing bacteria Desulfovibrio piger Vib-7. Arch. Microbiol. 2019, 201, 389–397. [Google Scholar] [CrossRef]
- Coutinho, C.M.L.M.; Coutinho-Silva, R.; Zinkevich, V.; Pearce, C.B.; Ojcius, D.M.; Beech, I. Sulphate-reducing bacteria from ulcerative colitis patients induce apoptosis of gastrointestinal epithelial cells. Microb. Pathog. 2017, 112, 126–134. [Google Scholar] [CrossRef]
- Loubinoux, J.; Bronowicji, J.P.; Pereira, I.A. Sulphate-reducing bacteria in human feces and their association with inflammatory diseases. FEMS Microbiol. Ecol. 2002, 40, 107–112. [Google Scholar] [CrossRef]
- Rowan, F.E.; Docherty, N.G.; Coffey, J.C.; O’Connell, P.R. Sulphate-reducing bacteria and hydrogen sulphide in the aetiology of ulcerative colitis. Br. J. Surg. 2009, 96, 151–158. [Google Scholar] [CrossRef]
- Scaldaferri, F.; Fiocchi, C. Inflammatory bowel disease: Progress and current concepts of etiopathogenesis. J. Dig. Dis. 2007, 8, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Fiocchi, C. Inflammatory bowel disease: Etiology and pathogenesis. Gastroenterology 1998, 115, 182–205. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Kollar, P.; Vítězová, M.; Drago, L. Hydrogen sulfide as a toxic product in the small–large intestine axis and its role in IBD development. J. Clin. Med. 2019, 8, 1054. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Kotrsová, V.; Dordević, D.; Buňková, L.; Vítězová, M.; Amedei, A. Hydrogen sulfide effects on the survival of Lactobacilli with emphasis on the development of inflammatory bowel diseases. Biomolecules 2019, 9, 752. [Google Scholar] [CrossRef] [PubMed]
- Dordević, D.; Jančíková, S.; Vítězová, M.; Kushkevych, I. Hydrogen sulfide toxicity in the gut environment: Meta-analysis of sulfate-reducing and lactic acid bacteria in inflammatory processes. J. Adv. Res. 2020, 27, 55–69. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Possible synergy effect of hydrogen sulfide and acetate produced by sulfate-reducing bacteria on inflammatory bowel disease development. J. Adv. Res. 2020, 27, 71–78. [Google Scholar] [CrossRef]
- Beauchamp, R.O.; Bus, J.S.; Popp, J.A.; Boreiko, C.J.; Andjelkovich, D.A.; Leber, P. A critical review of the literature on hydrogen sulfide toxicity. CRC Crit. Rev. Toxicol. 1984, 13, 25–97. [Google Scholar] [CrossRef]
- Blachier, F.; Davila, A.M.; Mimoun, S. Luminal sulfide and large intestine mucosa: Friend or foe? Amino Acids 2010, 39, 335–347. [Google Scholar] [CrossRef]
- Attene-Ramos, M.S.; Wagner, E.D.; Plewa, M.J.; Gaskins, H.R. Evidence that hydrogen sulfide is a genotoxic agent. Mol. Cancer Res. 2006, 4, 9–14. [Google Scholar] [CrossRef]
- Kushkevych, I.; Leščanová, O.; Dordević, D.; Jančíková, S.; Hošek, J.; Vítězová, M.; Buňková, L.; Drago, L. The Sulfate-Reducing Microbial Communities and Meta-Analysis of Their Occurrence during Diseases of Small–Large Intestine Axis. J. Clin. Med. 2019, 8, 1656. [Google Scholar] [CrossRef]
- Kushkevych, I.; Fafula, R.; Parak, T.; Bartoš, M. Activity of Na+/K+-activated Mg2+-dependent ATP hydrolase in the cell-free extracts of the sulfate-reducing bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Acta Vet. Brno 2015, 84, 3–12. [Google Scholar] [CrossRef]
- Barondeau, D.P.; Kassmann, C.J.; Bruns, C.K.; Tainer, J.A.; Getzoff, E.D. Nickel superoxide dismutase structure and mechanism. Biochemistry 2004, 43, 8038–8047. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.J.; Kim, E.J.; Suh, B.; Choi, J.H.; Roe, J.H. Duplicate genes for Fe-containing superoxide dismutase in Streptomyces coelicolor A3 (2). Gene 1999, 231, 87–93. [Google Scholar] [CrossRef]
- Ge, B.; Scheller, F.W.; Lisdat, F. Electrochemistry of immobilized CuZn-SOD and Fe-SOD and their interaction with superoxide radicals. Biosens. Bioelectron. 2003, 18, 295–302. [Google Scholar] [CrossRef]
- Abdulina, D.; Kováč, J.; Iutynska, G.; Kushkevych, I. ATP sulfurylase activity of sulfate-reducing bacteria from various ecotopes. 3 Biotech 2020, 10, 55. [Google Scholar] [CrossRef] [PubMed]
- Parey, K.; Demmer, U.; Warkentin, E. Structural, biochemical and genetic characterization of dissimilatory ATP sulfurylase from Allochromatium vinosum. PLoS ONE 2013, 8, e74707. [Google Scholar] [CrossRef]
- Gadd, G.M. Metals and microorganisms: A problem of definition. FEMS Microbiol. Lett. 1992, 79, 197–203. [Google Scholar] [CrossRef]
- Cervantes, C.; Silver, S. Plasmid chromate resistance and chromate reduction. Plasmid 1992, 27, 65–71. [Google Scholar] [CrossRef]
- De Flora, S.; Bennicelli, C.; Bagnasco, M. Genotoxicity of mercury compounds. Mut. Res. 1994, 317, 57–79. [Google Scholar] [CrossRef]
- Kováč, J.; Kushkevych, I. New modification of cultivation medium for isolation and growth of intestinal sulfate-reducing bacteria. In Proceedings of the International PhD Students Conference Mendel Net, Brno, Czech Republic, 6–7 November 2016; pp. 702–707. [Google Scholar]
- Postgate, J.R. The Sulfate-Reducing Bacteria, 2nd ed.; Cambridge University Press: Cambridge, UK, 1984; p. 208. [Google Scholar]
- Mauerhofer, L.-M.; Pappenreiter, P.; Paulik, C.; Seifert, A.H.; Bernacchi, S.; Rittmann, S.K.-M.R. Methods for quantification of growth and productivity in anaerobic microbiology and biotechnology. Folia Microbiol. 2019, 64, 321–360. [Google Scholar] [CrossRef]
- Abdulina, D.R.; Purish, L.M.; Asaulenko, L.G.; Iutynska, G.O. Sulfidogenic Microbial Communities From Technogenically Transformed Soils. Microbiol. Biotechnol. 2016, 34, 16–29. [Google Scholar]
- Kozhevnikova, M.F.; Levenets, V.V.; Diorditsa, V.A.; Rolyk, I.L.; Schur, A.A. Chemometric approach to processing data obtained from using of nuclear-physical methods of substances analysis. Skladni Syst. I Protsesy 2008, 1, 48–54. [Google Scholar]
- Gorban, A.N.; Zinovyev, A.Y. Principal Graphs and Manifolds. In Handbook of Research on Machine Learning Applications and Trends: Algorithms, Methods, and Techniques; IGI Global: Hershey, PA, USA, 2009; pp. 28–59. [Google Scholar]
- Bailey, N.T.J. Statistical Methods in Biology; Cambridge University Press: Cambridge, UK, 1995; p. 255. [Google Scholar]
- Brenner, D.J.; Krieg, N.R.; Staley, J.T.; Garrity, G.M. Bergey’s Manual of Systematic Bacteriology. Volume Two: The Proteobacteria, Part C: The Alpha-, Beta-, Delta-, and Epsilonproteobacteria, 2nd ed.; Springer: Boston, MA, USA, 2005; p. 1388. [Google Scholar]
- Eiler, A.; Farnleitner, A.H.; Zechmeister, T.C.; Herzig, A.; Hurban, C.; Wesner, W.; Krachler, R.; Velimirov, B.; Kirschner, A.K.T. Factors controlling extremely productive heterotrophic bacterial communities in shallow soda pools. Microbiol. Ecol. 2003, 46, 43–54. [Google Scholar] [CrossRef] [PubMed]
- Hubert, M.; Engelen, S. Robust PCA and classification in biosciences. Bioinformatics 2004, 20, 1728–1736. [Google Scholar] [CrossRef] [PubMed]
- Florin, T.H.; Neale, G.; Goretski, S. Sulfate in food and beverages. J. Food Compos. Anal. 1993, 6, 140–151. [Google Scholar] [CrossRef]
- Kushkevych, I.V. Activity and kinetic properties of phosphotransacetylase from intestinal sulfate-reducing bacteria. Acta Biochem. Pol. 2015, 62, 1037–1108. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Analysis of pH dose-dependent growth of sulfate-reducing bacteria. Open Med. 2019, 14, 66–74. [Google Scholar] [CrossRef]
- Tomasova, L.; Konopelski, P.; Ufnal, M. Gut bacteria and hydrogen sulfide: The new old players in circulatory system homeostasis. Molecules 2016, 21, 1558. [Google Scholar] [CrossRef]
- Černý, M.; Vítězová, M.; Vítěz, T.; Bartoš, M.; Kushkevych, I. Variation in the distribution of hydrogen producers from the clostridiales order in biogas reactors depending on different input substrates. Energies 2018, 11, 3270. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kollar, P.; Ferreira, A.L.; Palma, D.; Duarte, A.; Lopes, M.M.; Bartos, M.; Pauk, K.; Imramovsky, A.; Jampilek, J. Antimicrobial effect of salicylamide derivatives against intestinal sulfate-reducing bacteria. J. Appl. Biomed. 2016, 14, 125–130. [Google Scholar] [CrossRef]
- Kushkevych, I.; Cejnar, J.; Treml, J.; Dordević, D.; Kollar, P.; Vítězová, M. Recent Advances in Metabolic Pathways of Sulfate Reduction in Intestinal Bacteria. Cells 2020, 9, 698. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Kollar, P. Analysis of physiological parameters of Desulfovibrio strains from individuals with colitis. Open Life Sci. 2018, 13, 481–488. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Bartoš, M. Production of biogas: Relationship between methanogenic and sulfate-reducing microorganisms. Open Life Sci. 2017, 12, 82–91. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Kováč, J.; Kaucká, P.; Jesionek, W.; Bartoš, M.; Barton, L. A new combination of substrates: Biogas production and diversity of the methanogenic microorganisms. Open Life Sci. 2018, 13, 119–128. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kos, J.; Kollar, P.; Kralova, K.; Jampilek, J. Activity of ring-substituted 8-hydroxyquinoline-2-carboxanilides against intestinal sulfate-reducing bacteria Desulfovibrio piger. Med. Chem. Res. 2018, 27, 278–284. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kováč, J.; Vítězová, M.; Vítěz, T.; Bartoš, M. The diversity of sulfate-reducing bacteria in the seven bioreactors. Arch. Microbiol. 2018, 200, 945–950. [Google Scholar] [CrossRef]
- Trevors, J.T.; Stratton, G.W.; Gadd, G.M. Cadmium transport, resistance, and toxicity in bacteria, algae, and fungi. Can. J. Microbiol. 1986, 32, 447–464. [Google Scholar] [CrossRef]
- Nowak, A.; Przebulewska, K.; Szopa, E.; Stacewicz, A. Influence of the heavy metals (Hg, Cd, Cu, Pb) on the growth and ferment activity of soil bacteria. Folia Univ. Agric. Ctetin. Agric. 2001, 88, 165–173. [Google Scholar]
- Inbar, O.; Ron, E.Z. Induction of cadmium tolerance in Escherichia coli K12. FEMS Microbiol. Lett. 1993, 113, 197–200. [Google Scholar] [CrossRef][Green Version]
- Oremland, R.S.; Steinberg, N.A.; Maest, A.S. Measurement of in situ rates of selenate removal by dissimilatory bacterial reduction in sediments. Environ. Sci. Technol. 1990, 24, 1157–1164. [Google Scholar] [CrossRef]
- Abdel Azim, A.; Rittmann, S.K.-M.R.; Fino, D.; Bochmann, G. The physiological effect of heavy metals and volatile fatty acids on Methanococcus maripaludis S2. Biotechnol. Biofuels 2018, 11, 301. [Google Scholar] [CrossRef] [PubMed]
- Tebo, B.M.; Obraztsova, A.Y. Sulfate-reducing bacterium grows with Cr (VI), U (VI), Mn (IV), and Fe (III) as electron acceptors. FEMS Microbiol. Lett. 1998, 162, 193–198. [Google Scholar] [CrossRef]
- Bharathi, P.A.L.; Sathe, V.; Chandramohan, D. Effect of lead, mercury and cadmium on a sulphate-reducing bacterium. Environ. Pollut. 1990, 67, 361–374. [Google Scholar] [CrossRef]
- Karnachuk, O.V. Influence of hexavalent chromium on hydrogen sulfide formation by sulfate-reducing bacteria. Microbiology 1995, 64, 262–265. [Google Scholar]
- Lloyd, J.R. Microbial reduction of metals and radionuclids. FEMS Microbiol. Rev. 2003, 27, 411–425. [Google Scholar] [CrossRef]
- Lovely, D.R. Dissimilatory metal reduction. Annu. Rev. Microbiol. 1993, 47, 263–290. [Google Scholar] [CrossRef]
- Zaitseva, S.V.; Abidueva, E.Y.; Buryukhaev, S.P.; Namsaraev, B.B. Factors controlling the activity of the microbial community of the alkaline lake Beloe (Trans-baikal region). Microbiology 2012, 81, 468–476. [Google Scholar] [CrossRef]
- Lovley, D.R.; Phillips, E.J.P. Reduction of uranium by Desulfovibrio desulfuricans. Appl. Environ. Microbiol. 1992, 58, 850–856. [Google Scholar] [CrossRef]
- Gangola, P.; Rosen, B.P. Maintenance of intracellular calcium in Escherichia coli. J. Biol. Chem. 1987, 262, 12570–12574. [Google Scholar] [CrossRef]
- Dominguez, D.C. Calcium signalling in bacteria. Mol. Mikrobiol. 2004, 54, 291–297. [Google Scholar] [CrossRef]
- Mullen, M.D.; Wolf, D.C.; Ferris, F.G.; Beveridge, T.J.; Flemming, C.A.; Bailey, G.W. Bacterial sorption of heavy metals. Appl. Environ. Microbiol. 1989, 55, 3143–3149. [Google Scholar] [CrossRef] [PubMed]
- Dubbs, J.M.; Mongkolsuk, S. Peroxiredoxins in bacterial antioxidant defense. In Peroxiredoxin Systems; Springer: Dordrecht, The Netherlands, 2007; pp. 143–193. [Google Scholar]
| Patient | Sex (M/F) | Diagnosis | Age (Years) | Weight (kg) | Year of Diagnosis | State of the Disease * | Isolated SRB Strains |
|---|---|---|---|---|---|---|---|
| Diseased persons | |||||||
| Patient 1 | M | CD | 24 | 73 | 2006 | Long-term remission | D. vulgaris M9.1 Desulfovibrio sp. M9.2 D. vulgaris M9.3 |
| Patient 2 | F | CD | 21 | 57 | 2011 | Short-term remission | SRB M1.1 D. vulgaris M1.2 |
| Patient 3 | M | CD | 34 | 104 | 2013 | Short-term remission | SRB M2.1 SRB M2.2 SRB M2.3 |
| Patient 4 | M | CD | 34 | 70 | 2018 | Disease flare | D. vulgaris M10.1 SRB M10.2 |
| Patient 5 | M | UC | 44 | 61 | 2014 | Long-term remission | D. vulgaris M7.1 D. vulgaris M7.2 D. vulgaris M7.3 |
| Patient 6 | M | UC | 35 | 92 | 2018 | Short-term remission | Desulfovibrio sp. M5.1 D. vulgaris M5.2 |
| Patient 7 | F | UC | 20 | 70 | 2003 | Short-term remission | D. vulgaris M8 |
| Patient 8 | F | UC | 66 | 78 | 2012 | Short-term remission | D. vulgaris M3.1 D. vulgaris M3.2 |
| Patient 9 | F | UC | 41 | 53 | 2005 | Disease flare | SRB M4.1 SRB M4.2 D. vulgaris M4.3 |
| Patient 10 | F | UC | 39 | 120 | 2013 | Disease flare | D. vulgaris M6.1 Desulfovibrio sp. M6.2 SRB M6.3 |
| Healthy persons | |||||||
| Control 1 | M | no IBD | 46 | 108 | – | Healthy | SRB Z8.1 D. vulgaris Z8.2 |
| Control 2 | M | no IBD | 53 | 72 | – | Healthy | SRB Z9.1 D. vulgaris Z9.2 D. vulgaris Z9.3 |
| Control 3 | M | no IBD | 42 | 95 | – | Healthy | D. vulgaris Z11.1 SRB Z11.2 SRB Z11.3 |
| Control 4 | M | no IBD | 26 | 78 | – | Healthy | D. vulgaris Z12.1 SRB Z12.2SRB Z12.3 |
| Groups | Parameters | Na+ | K+ | Ca2+ | Mg2+ | Zn2+ | Fe2+ | Cu+ | Mn2+ | Co2+ | Ni2+ | Mo2+ |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CD | X | 4.46 | 21.84 | 15.66 | 1.07 | 0.17 | 3.96 | 0.02 | 0.033 | 0.0005 | 0 | 0.0011 |
| Δx (t lim 2.26, n = 9) | 0.22 | 3.17 | 13.31 | 0.24 | 0.22 | 2.62 | 0.01 | 0.01 | 0.0004 | 0 | 0.0011 | |
| UC | X | 4.22 | 17.30 | 7.36 | 0.77 | 0.08 | 1.47 | 0.023 | 0.027 | 0.0005 | 0.0052 | 0.0001 |
| Δx (t lim 2.16, n = 14) | 0.28 | 0.89 | 2.29 | 0.06 | 0.06 | 1.02 | 0.01 | 0.002 | 0.0002 | 0.0061 | 0.0003 | |
| H | X | 4.16 | 18.37 | 11.41 | 0.94 | 0.25 | 8.41 | 0.04 | 0.03 | 0.001 | 0.01 | 0.0022 |
| Δx (t lim 2.23, n = 11) | 0.24 | 1.38 | 3.67 | 0.13 | 0.11 | 3.69 | 0.039 | 0.004 | 0.0003 | 0.0144 | 0.0010 |
| Importance | Factors | Power | Principal Components | ||
|---|---|---|---|---|---|
| PC1 | PC2 | PC3 | |||
| 1 | Mn | 0.87 | 0.874 | 0.333 | 0.068 |
| 2 | Mg | 0.86 | 0.836 | 0.388 | −0.138 |
| 3 | Co | 0.86 | 0.729 | −0.456 | 0.3411 |
| 4 | Ca | 0.84 | 0.718 | 0.337 | −0.468 |
| 5 | Mo | 0.81 | 0.806 | −0.323 | 0.233 |
| 6 | Fe | 0.78 | 0.765 | −0.317 | 0.297 |
| 7 | Na | 0.77 | 0.066 | 0.460 | 0.753 |
| 8 | Cu | 0.76 | 0.565 | −0.598 | −0.279 |
| 9 | K | 0.72 | 0.475 | 0.701 | −0.074 |
| 10 | Zn | 0.68 | 0.796 | 0.200 | −0.121 |
| 11 | Ni | 0.59 | 0.296 | −0.699 | −0.110 |
| Eigenvalues | 5.02 (S) | 2.38 (S) | 1.18 | ||
| Total variance (%) | 45.65 | 21.65 | 10.74 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kushkevych, I.; Abdulina, D.; Dordević, D.; Rozehnalová, M.; Vítězová, M.; Černý, M.; Svoboda, P.; Rittmann, S.K.-M.R. Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria. Appl. Sci. 2021, 11, 1152. https://doi.org/10.3390/app11031152
Kushkevych I, Abdulina D, Dordević D, Rozehnalová M, Vítězová M, Černý M, Svoboda P, Rittmann SK-MR. Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria. Applied Sciences. 2021; 11(3):1152. https://doi.org/10.3390/app11031152
Chicago/Turabian StyleKushkevych, Ivan, Daryna Abdulina, Dani Dordević, Monika Rozehnalová, Monika Vítězová, Martin Černý, Pavel Svoboda, and Simon K.-M. R. Rittmann. 2021. "Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria" Applied Sciences 11, no. 3: 1152. https://doi.org/10.3390/app11031152
APA StyleKushkevych, I., Abdulina, D., Dordević, D., Rozehnalová, M., Vítězová, M., Černý, M., Svoboda, P., & Rittmann, S. K.-M. R. (2021). Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria. Applied Sciences, 11(3), 1152. https://doi.org/10.3390/app11031152








