A Method to Obtain Parameters of One-Column Jansen–Rit Model Using Genetic Algorithm and Spectral Characteristics
Abstract
1. Introduction
2. Materials and Methods
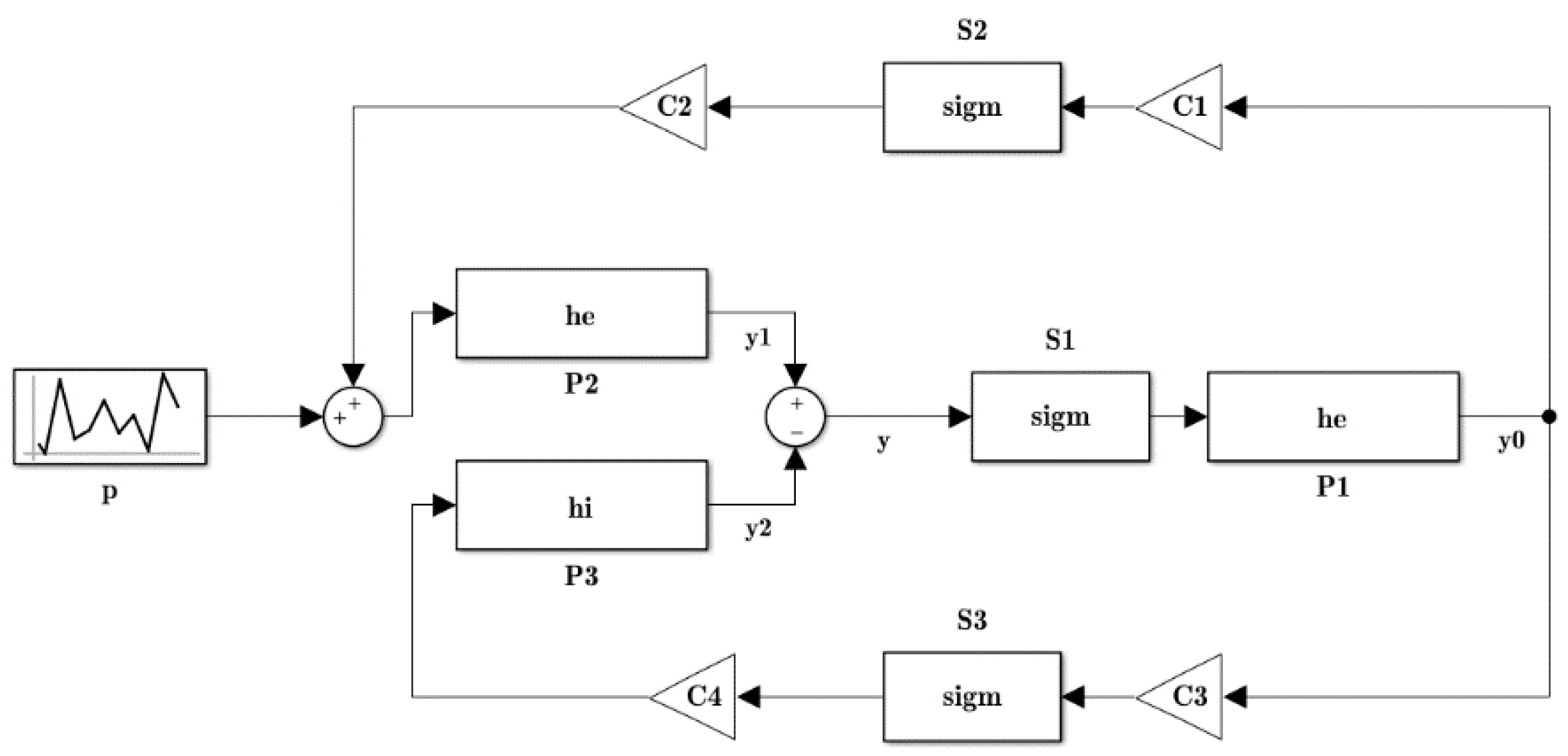
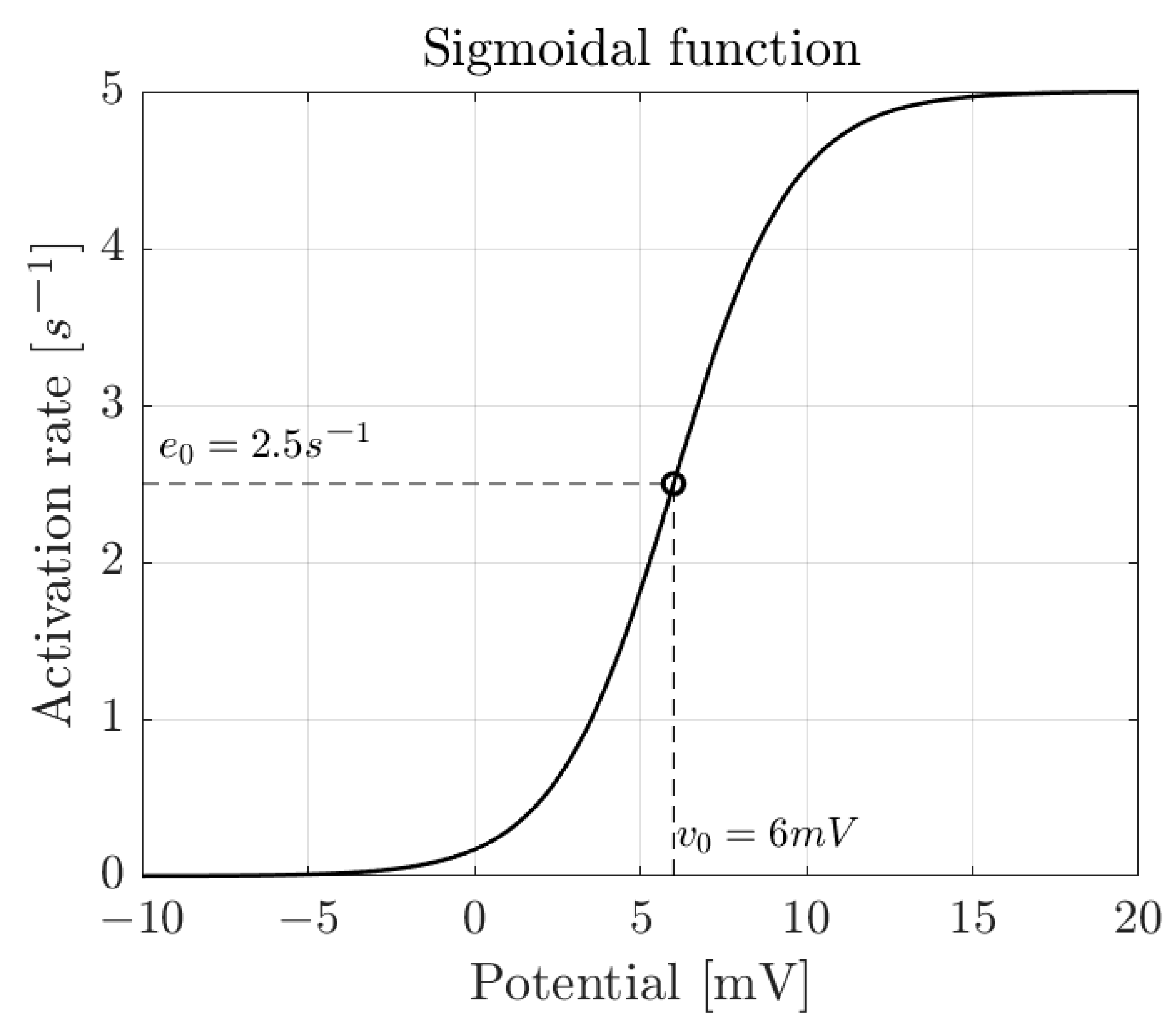
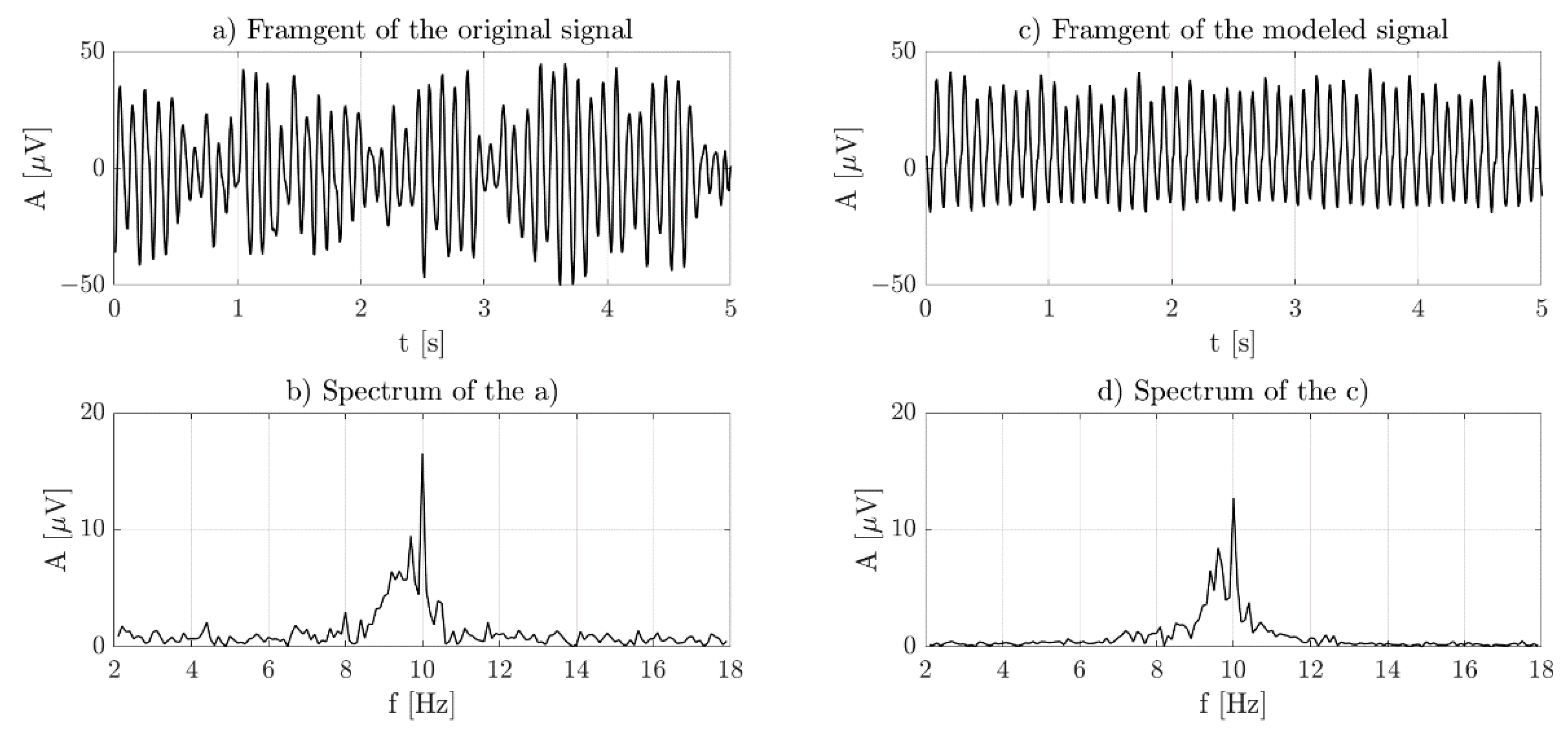
2.1. Modeled Signal
2.2. Model Parameter Obtaining Process
2.2.1. Function Minimized in This Study
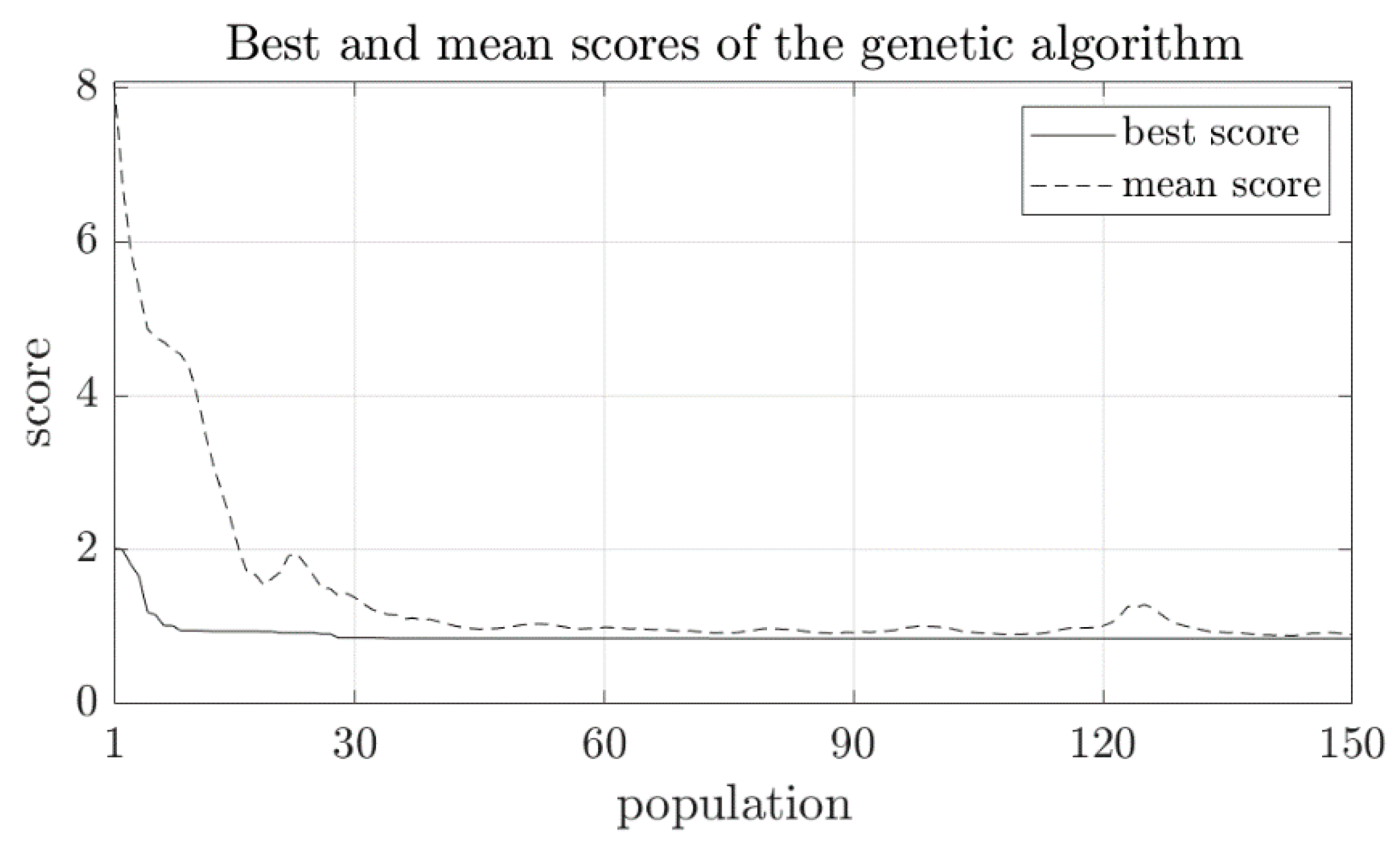
2.2.2. The Genetic Algorithm
2.2.3. Computation Complexity
2.3. Repeatability of the Method
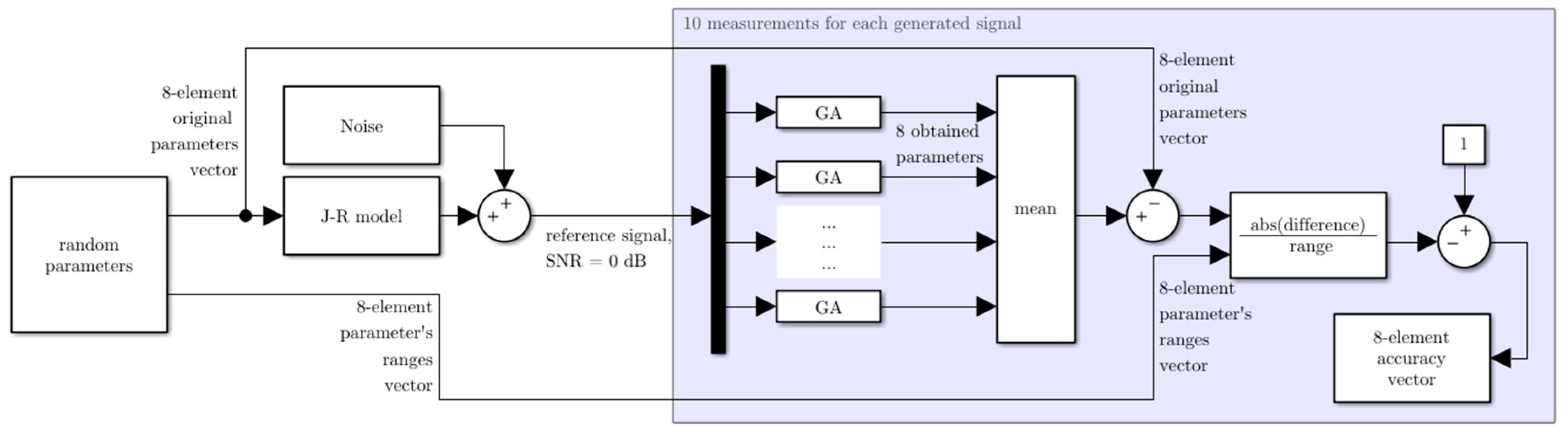
2.4. Accuracy of the Method
2.5. Reference Signals Generation Process
2.6. Real, Measured Signal
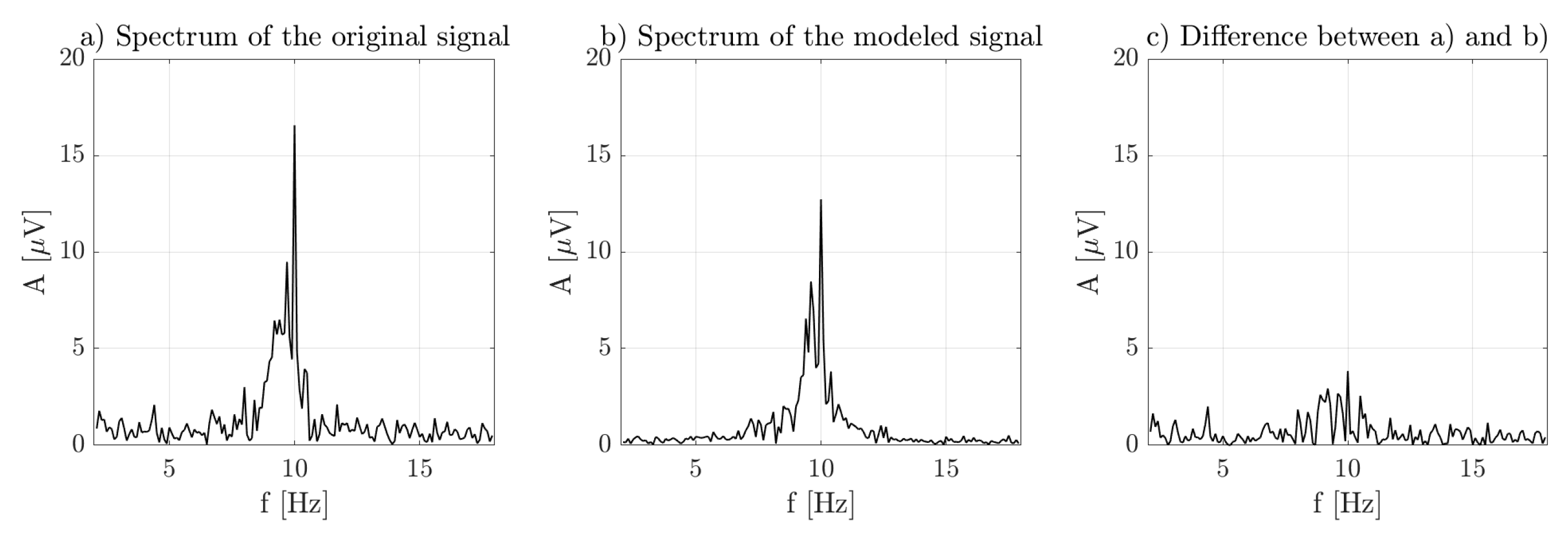
3. Results
4. Discussion and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Louis, E.K.S.; Frey, L.C.; Britton, J.W.; Hopp, J.L.; Korb, P.; Koubeissi, M.Z.; Lievens, W.E.; Pestana-Knight, E.M.; Foundation, C.C.C. Electroencephalography (EEG): An Introductory Text and Atlas of Normal and Abnormal Findings in Adults, Children, and Infants; American Epilepsy Society: Chicago, IL, USA, 2016. [Google Scholar]
- Paszkiel, S. Using the Raspberry PI2 Module and the Brain-Computer Technology for Controlling a Mobile Vehicle. In Automation 2019; Szewczyk, R., Zieliński, C., Kaliczyńska, M., Eds.; Springer International Publishing: Cham, Switzerland, 2020; Volume 920, pp. 356–366. [Google Scholar] [CrossRef]
- Paszkiel, S. Characteristics of Question of Blind Source Separation Using Moore-Penrose Pseudoinversion for Reconstruction of EEG Signal. In Automation 2017; Szewczyk, R., Zieliński, C., Kaliczyńska, M., Eds.; Springer International Publishing: Cham, Switzerland, 2017; Volume 550, pp. 393–400. [Google Scholar]
- Paszkiel, S. Augmented Reality of Technological Environment in Correlation with Brain Computer Interfaces for Control Processes. In Recent Advances in Automation, Robotics and Measuring Techniques; Szewczyk, R., Zieliński, C., Kaliczyńska, M., Eds.; Springer International Publishing: Cham, Switzerland, 2014; Volume 267, pp. 197–203. [Google Scholar]
- Shayegh, F.; Fattahi, R.A.; Sadri, S.; Ansari-Asl, K. A brief survey of computational models of normal and epileptic EEG signals: A guideline to model-based seizure prediction. J. Med. Signals Sens. 2011, 1, 62–72. [Google Scholar] [CrossRef] [PubMed]
- Beurle, R.L. Properties of a mass of cells capable of regenerating pulses. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 1956, 240, 55–94. [Google Scholar] [CrossRef]
- Wilson, H.R.; Cowan, J.D. Excitatory and Inhibitory Interactions in Localized Populations of Model Neurons. Biophys. J. 1972, 12, 1–24. [Google Scholar] [CrossRef]
- Amari, S.-I. Homogeneous nets of neuron-like elements. Biol. Cybern. 1975, 17, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, F.H.L.; Hoeks, A.; Smits, H.; Zetterberg, L.H. Model of brain rhythmic activity: The alpha-rhythm of the thalamus. Biol. Cybern. 1974, 15, 27–37. [Google Scholar] [CrossRef]
- Grimbert, F.; Faugeras, O. Analysis of Jansen’s Model of a Single Cortical Column. INRIA, RR-5597. May 2006. Available online: https://hal.inria.fr/inria-00070410 (accessed on 18 June 2020).
- Wendling, F.; Bartolomei, F.; Bellanger, J.J.; Chauvel, P. Epileptic fast activity can be explained by a model of impaired GABAergic dendritic inhibition. Eur. J. Neurosci. 2002, 15, 1499–1508. [Google Scholar] [CrossRef] [PubMed]
- Svozilova, V.; Mezl, M. Modeling of the EEG signal. Elektrorevue 2016, 18, 43–51. [Google Scholar]
- Jansen, B.H.; Zouridakis, G.; Brandt, M.E. A neurophysiologically-based mathematical model of flash visual evoked potentials. Biol. Cybern. 1993, 68, 275–283. [Google Scholar] [CrossRef]
- Wendling, F.; Bellanger, J.J.; Bartolomei, F.; Chauvel, P. Relevance of nonlinear lumped-parameter models in the analysis of depth-EEG epileptic signals. Biol. Cybern. 2000, 83, 367–378. [Google Scholar] [CrossRef]
- Todo, R.S. Personalization of a Biophysically Realistic Computational Model of the Brain Activity Using EEG and MRI Data. 2018. Available online: https://repositori.upf.edu/handle/10230/44674 (accessed on 18 December 2020).
- Jafarian, A.; Freestone, D.R.; Nesic, D.; Grayden, D.B. Slow-Fast Duffing Neural Mass Model. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; Volume 2019, pp. 142–145. [Google Scholar]
- Al-Hossenat, A.; Wen, P.; Li, Y. Modelling and simulating different bands of EEG signals with the virtual brain. Int. J. Electr. Electron. Data Commun. 2019, 7, 5. [Google Scholar]
- Al-Hossenat, A.; Wen, P.; Li, Y. Simulation α of EEG Using Brain Network Model; Swinburne University of Technology: Melbourne, Australia, 2017; pp. 336–345. [Google Scholar]
- Jedynak, M.; Pons, A.J.; Garcia-Ojalvo, J.; Goodfellow, M. Temporally correlated fluctuations drive epileptiform dynamics. NeuroImage 2017, 146, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Zhang, D.; Meng, J.; Zhu, X. Interactions between two neural populations: A mechanism of chaos and oscillation in neural mass model. Neurocomputing 2011, 74, 1026–1034. [Google Scholar] [CrossRef]
- Basu, I.; Crocker, B.; Farnes, K.; Robertson, M.M.; Paulk, A.C.; Vallejo, D.I.; Dougherty, D.D.; Cash, S.S.; Eskandar, E.N.; Kramer, M.M.; et al. A neural mass model to predict electrical stimulation evoked responses in human and non-human primate brain. J. Neural Eng. 2018, 15, 066012. [Google Scholar] [CrossRef] [PubMed]
- Ye, W. Dynamics of a revised neural mass model in the stop-signal task. Chaos Solitons Fractals 2020, 139, 110004. [Google Scholar] [CrossRef]
- Postoyan, R.; Chong, M.; Nešić, D.; Kuhlmann, L. Parameter and state estimation for a class of neural mass models. In Proceedings of the 2012 IEEE 51st IEEE Conference on Decision and Control (CDC), Wailea, HI, USA, 10–13 December 2012; pp. 2322–2327. [Google Scholar]
- Freestone, D.; Kuhlmann, L.; Chong, M.; Nešić, D.; Grayden, D.; Aram, P.; Postoyan, R.; Cook, M.J.; Elger, C.E.; Tetzlaff, R.; et al. Patient-Specific Neural Mass Modeling—Stochastic And Deterministic Methods. In Recent Advances in Predicting and Preventing Epileptic Seizures; Technische Universität: Dresden, Germany, 2013; pp. 63–82. [Google Scholar] [CrossRef]
- Kuhlmann, L.; Freestone, D.R.; Manton, J.; Heyse, B.; Vereecke, H.E.M.; Lipping, T.; Struys, M.M.; Liley, D.T. Neural mass model-based tracking of anesthetic brain states. NeuroImage 2016, 133, 438–456. [Google Scholar] [CrossRef] [PubMed]
- Freeman, W.J. Mass Action in the Nervous System: Examination of the Neurophysiological Basis of Adaptive Behavior through the EEG; Academic Press: New York, NY, USA, 1975. [Google Scholar]
- Quaranta, G.; Lacarbonara, W.; Masri, S.F. A review on computational intelligence for identification of nonlinear dynamical systems. Nonlinear Dyn. 2020, 99, 1709–1761. [Google Scholar] [CrossRef]
- Genetic Algorithm—MATLAB & Simulink. Available online: https://www.mathworks.com/help/gads/genetic-algorithm.html (accessed on 2 December 2020).
- Wolak, M.E.; Fairbairn, D.J.; Paulsen, Y.R. Guidelines for estimating repeatability. Methods Ecol. Evol. 2012, 3, 129–137. [Google Scholar] [CrossRef]
- McGraw, K.O.; Wong, S.P. Forming Inferences about Some Intraclass Correlation Coefficients. Psychol. Methods 1996, 1, 30. [Google Scholar] [CrossRef]
- Koo, T.K.; Li, M.Y. A Guideline of Selecting and Reporting Intraclass Correlation Coefficients for Reliability Research. J. Chiropr. Med. 2016, 15, 155–163. [Google Scholar] [CrossRef]
- Zabcikova, M. Measurement of Visual and Auditory Stimuli Using EEG Headset Emotiv Epoc+. MATEC Web Conf. 2019, 292, 01023. [Google Scholar] [CrossRef]
- Marcuse, L.; Fields, M.; Yoo, J. Rowan’s Primer of EEG, 2nd ed.; Elsevier: Amsterdam, The Netherlands, 2015. [Google Scholar]
- Greenfield, L.J.; Geyer, J.D.; Carney, P.R. Reading EEGs. A Practical Approach; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2012. [Google Scholar]
- Foster, J.J.; Sutterer, D.W.; Serences, J.T.; Vogel, E.K.; Awh, E. Alpha-Band Oscillations Enable Spatially and Temporally Resolved Tracking of Covert Spatial Attention. Psychol. Sci. 2017, 28, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Paszkiel, S.; Hunek, W.; Shylenko, A. Project and Simulation of a Portable Device for Measuring Bioelectrical Signals from the Brain for States Consciousness Verification with Visualization on LEDs. In Challenges in Automation, Robotics and Measurement Techniques; Szewczyk, R., Zieliński, C., Kaliczyńska, M., Eds.; Springer International Publishing: Cham, Switzerland, 2016; Volume 440, pp. 25–35. [Google Scholar] [CrossRef]
- Paszkiel, S.; Dobrakowski, P.; Łysiak, A. The Impact of Different Sounds on Stress Level in the Context of EEG, Cardiac Measures and Subjective Stress Level: A Pilot Study. Brain Sci. 2020, 10, 728. [Google Scholar] [CrossRef] [PubMed]

| Algorithm | Time | Found Optimum |
|---|---|---|
| Genetic algorithm | 1.000 | 0.161 |
| Simulated annealing | 0.440 | 1.000 |
| Particle swarm | 0.978 | 0.189 |
| Surrogate optimization | 0.236 | 0.275 |
| Parameter | Lower Search Range | Upper Search Range |
|---|---|---|
| A (mV) | 2.25 | 4.25 |
| B (mV) | 12 | 32 |
| C | 70 | 675 |
| v0 (mV) | 5 | 7 |
| e0 (s−1) | 2 | 3 |
| r | 0.5 | 0.6 |
| Lower noise limit (pps) | 50 | 300 |
| Noise range (pps) | 200 | 1000 |
| Parameter | Accuracy (Mean) | Accuracy (Std) | ICC |
|---|---|---|---|
| A (mV) | 0.794 | 0.272 | 0.897 |
| B (mV) | 0.724 | 0.343 | 0.910 |
| C | 0.853 | 0.189 | 0.978 |
| v0 (mV) | 0.756 | 0.306 | 0.865 |
| e0 (s−1) | 0.662 | 0.403 | 0.676 |
| r | 0.700 | 0.384 | 0.733 |
| Lower noise limit (pps) | 0.761 | 0.320 | 0.877 |
| Noise range (pps) | 0.863 | 0.191 | 0.966 |
| Parameter | Lower Range | Upper Range | Default Value | Obtained Value |
|---|---|---|---|---|
| A (mV) | 2.25 | 4.25 | 3.25 | 4.21 |
| B (mV) | 12 | 32 | 22 | 12.06 |
| C | 70 | 675 | 135 | 406.50 |
| v0 (mV) | 5 | 7 | 6 | 6.60 |
| e0 (s−1) | 2 | 3 | 2.5 | 2.92 |
| r | 0.5 | 0.6 | 0.56 | 0.6 |
| Lower noise limit (pps) | 50 | 300 | 120 | 278.77 |
| Noise range (pps) | 200 | 1000 | 200 | 897.34 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Łysiak, A.; Paszkiel, S. A Method to Obtain Parameters of One-Column Jansen–Rit Model Using Genetic Algorithm and Spectral Characteristics. Appl. Sci. 2021, 11, 677. https://doi.org/10.3390/app11020677
Łysiak A, Paszkiel S. A Method to Obtain Parameters of One-Column Jansen–Rit Model Using Genetic Algorithm and Spectral Characteristics. Applied Sciences. 2021; 11(2):677. https://doi.org/10.3390/app11020677
Chicago/Turabian StyleŁysiak, Adam, and Szczepan Paszkiel. 2021. "A Method to Obtain Parameters of One-Column Jansen–Rit Model Using Genetic Algorithm and Spectral Characteristics" Applied Sciences 11, no. 2: 677. https://doi.org/10.3390/app11020677
APA StyleŁysiak, A., & Paszkiel, S. (2021). A Method to Obtain Parameters of One-Column Jansen–Rit Model Using Genetic Algorithm and Spectral Characteristics. Applied Sciences, 11(2), 677. https://doi.org/10.3390/app11020677






