Portable through Bottle SORS for the Authentication of Extra Virgin Olive Oil
Abstract
1. Introduction
2. Experimental Methods and Materials
2.1. Oils
2.1.1. Sample Preparation for Vial-Mode Analysis
2.1.2. Sample Preparation for Through-Barrier Analysis
2.2. Raman Spectroscopy
- CBEx handheld Raman spectrometer from Snowy Range (Laramie, WY, USA), equipped with 785 nm excitation laser wavelength, with laser power of 70 mW on the sample, was used to collected spectra over the 400–2300 cm−1 range with 12–14 cm−1 spectral resolution. The acquisition time was 2 s for vials-mode analysis;
- CBEx handheld Raman spectrometer (Snowy Range) operating at 1064 nm, with laser power 300 mW on the sample, was employed to collect data in the 400 to 2300 cm−1 range with 12–14 cm−1 spectral resolution. The acquisition time was 15 s for vial-mode analysis; and
- Resolve (Cobalt Light System, Oxfordshire, UK now part of Agilent) handheld SORS system, equipped with an 830 nm excitation laser wavelength, with laser power of 450 mW on the sample, was employed to collect spectra in the 350–2000 cm−1 range with 3 cm−1 spectral resolution. The acquisition time was less than 2 min for vial-mode, and 2 min for through-barrier analysis.
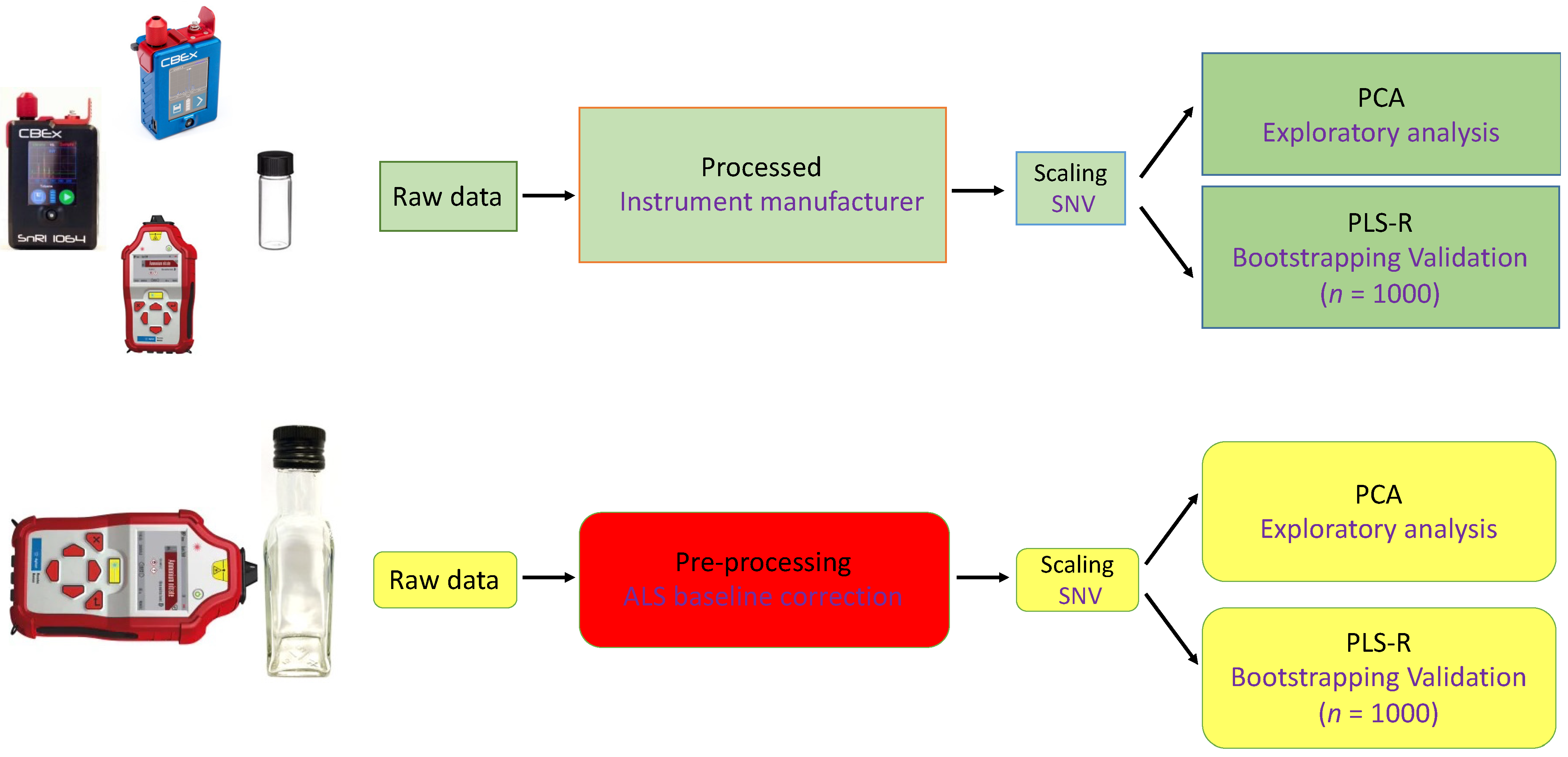
2.3. Data Analysis
3. Results and Discussion
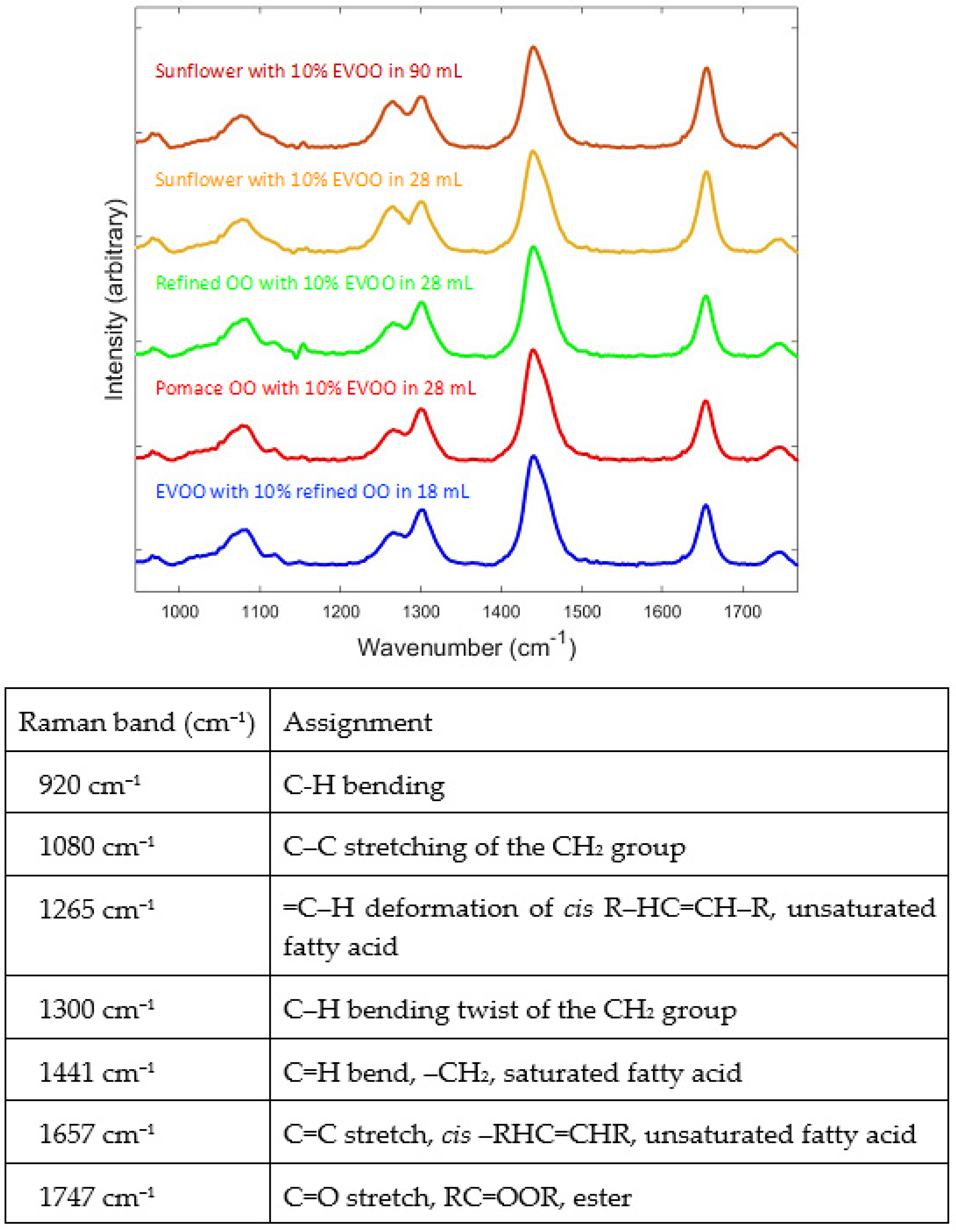
3.1. Visible Inspection of the Raman and SORS Spectra
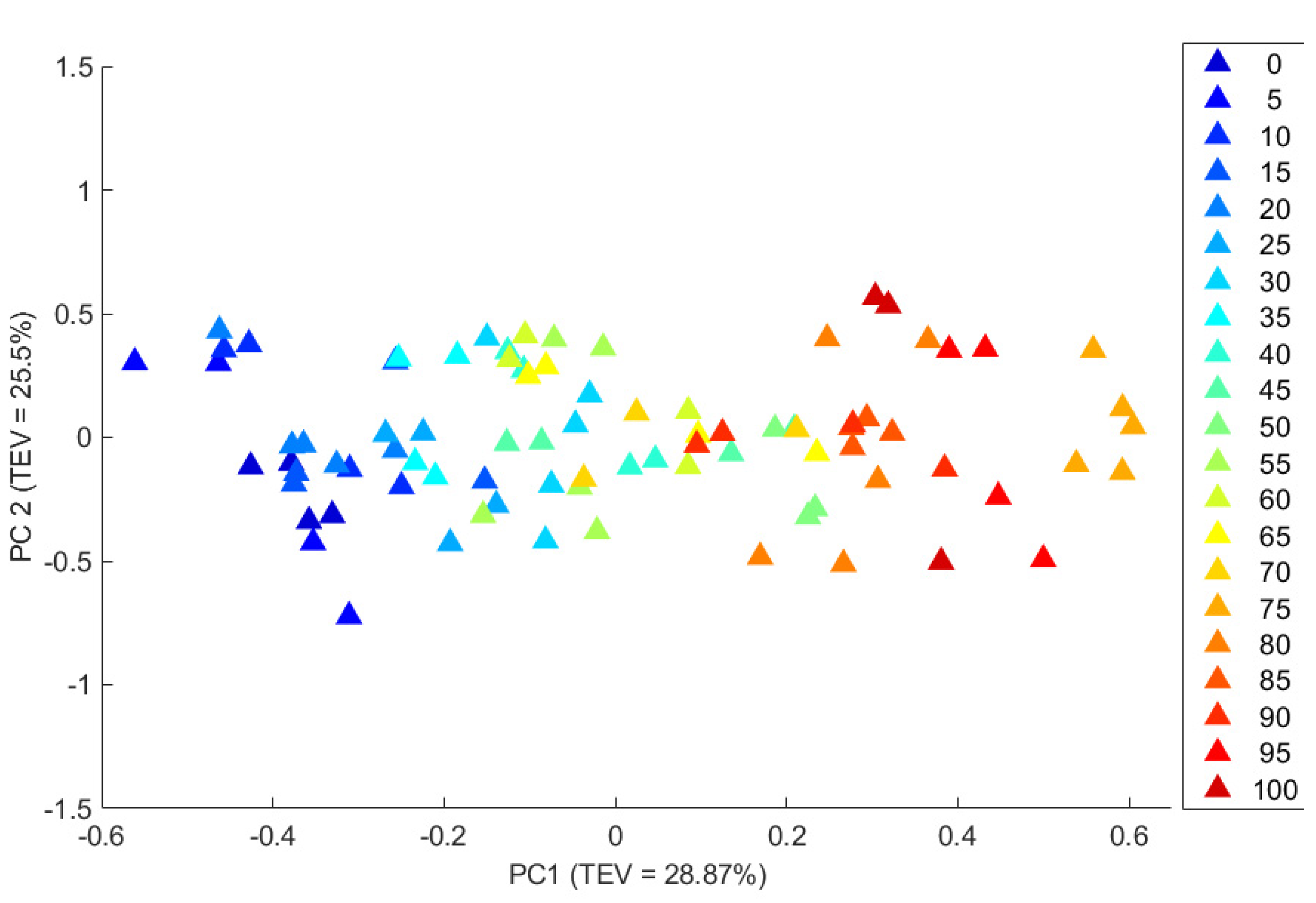
3.2. Principal Components Analysis (PCA) of the Raman and SORS Spectra
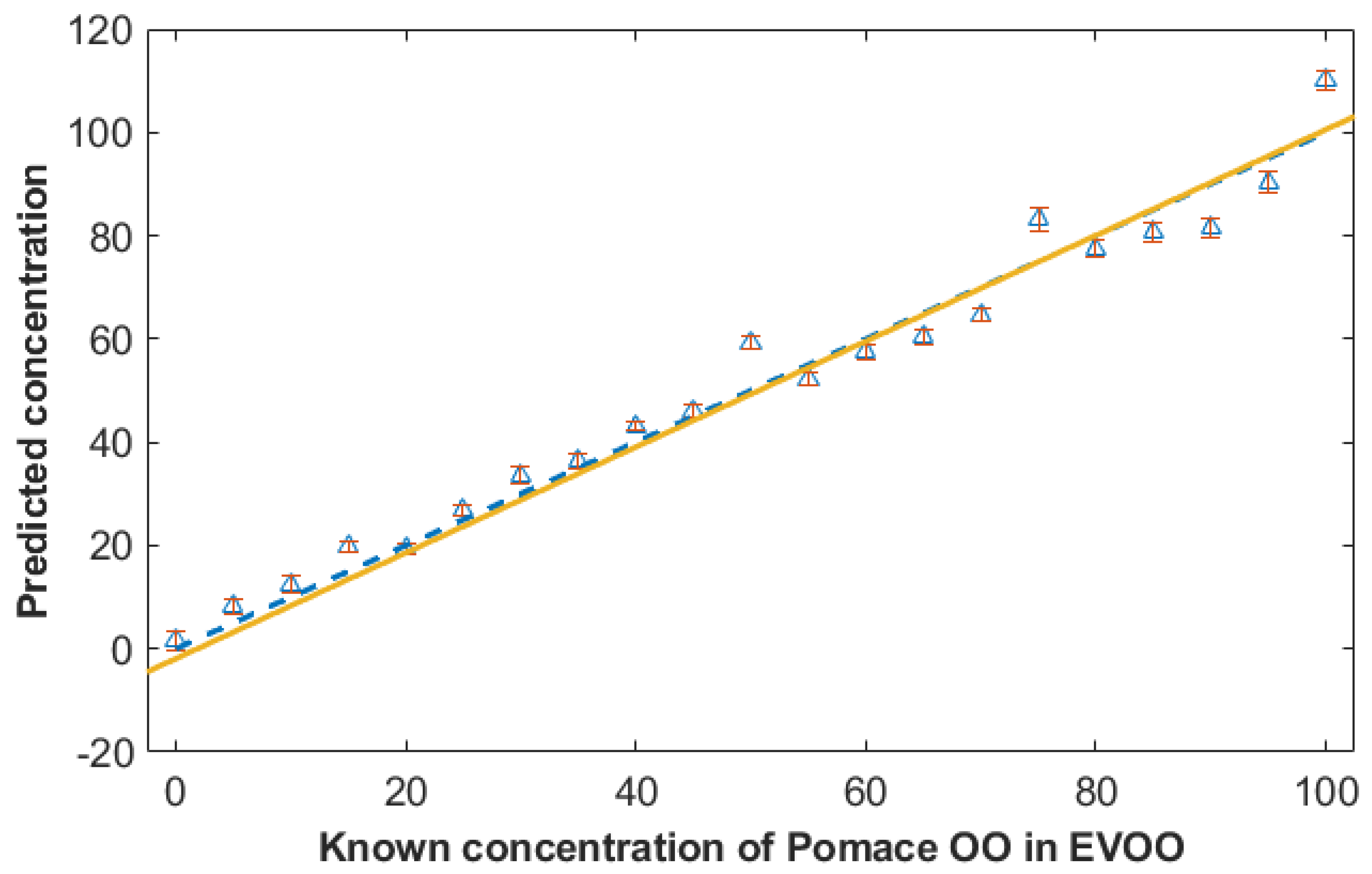
3.3. Quantification of the Level of the Adulterant Using Partial Least Squares Regression (PLSR)
3.4. Quantification of Adulterants Using SORS and PLSR
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ellis, D.I.; Brewster, V.L.; Dunn, W.B.; Allwood, J.W.; Golovanov, A.P.; Goodacre, R. Fingerprinting food: Current technologies for the detection of food adulteration and contamination. Chem. Soc. Rev. 2012, 41, 5706–5727. [Google Scholar] [CrossRef]
- Olive Oil Times. Five Years Later, UC Davis Report Still Sends Shockwaves. Available online: https://www.oliveoiltimes.com/olive-oil-making-and-milling/five-years-later-uc-davis-report-still (accessed on 20 July 2021).
- de la Paz, M.P.; Philen, R.M.; Borda, A.I. Toxic oil syndrome: The perspective after 20 years. Epidemiol. Rev. 2001, 23, 231–247. [Google Scholar] [CrossRef]
- Vanstone, N.; Moore, A.; Martos, P.; Neethirajan, S. Detection of the adulteration of extra virgin olive oil by near-infrared spectroscopy and chemometric techniques. Food Qual. Saf. 2018, 2, 189–198. [Google Scholar] [CrossRef]
- Aparicio, R.; Aparicio-Ruíz, R. Authentication of vegetable oils by chromatographic techniques. J. Chromatogr. A 2000, 881, 93–104. [Google Scholar] [CrossRef]
- Kiritsakis, A.K. Flavor components of olive oil—A review. J. Am. Oil Chem. Soc. 1998, 75, 673–681. [Google Scholar] [CrossRef]
- Zou, M.-Q.; Zhang, X.-F.; Qi, X.-H.; Ma, H.-L.; Dong, Y.; Liu, C.-W.; Guo, X.; Wang, H. Rapid authentication of olive oil adulteration by Raman spectrometry. J. Agric. Food Chem. 2009, 57, 6001–6006. [Google Scholar] [CrossRef]
- Gallina Toschi, T.; Bendini, A.; Lercker, G. Evaluation of 3,5-stigmastadiene content of edible oils: Comparison between the traditional capillary gas chromatographic method and the on-line high performance liquid chromatography-capillary gas chromatographic analysis. Chromatographia 1996, 43, 195–199. [Google Scholar] [CrossRef]
- Ellis, D.I.; Muhamadali, H.; Haughey, S.A.; Elliott, C.T.; Goodacre, R. Point-and-shoot: Rapid quantitative detection methods for on-site food fraud analysis—Moving out of the laboratory and into the food supply chain. Anal. Methods 2015, 7, 9401–9414. [Google Scholar] [CrossRef]
- Gamazo-Vázquez, J.; García-Falcón, M.S.; Simal-Gándara, J. Control of contamination of olive oil by sunflower seed oil in bottling plants by GC-MS of fatty acid methyl esters. Food Control 2003, 14, 463–467. [Google Scholar] [CrossRef]
- Saba, A.; Mazzini, F.; Raffaelli, A.; Mattei, A.; Salvadori, P. Identification of 9(E),11(E)-18:2 fatty acid methyl ester at trace level in thermal stressed olive oils by GC coupled to acetonitrile CI-MS and CI-MS/MS, a possible marker for adulteration by addition of deodorized olive oil. J. Agric. Food Chem. 2005, 53, 4867–4872. [Google Scholar] [CrossRef] [PubMed]
- Jabeur, H.; Zribi, A.; Makni, J.; Rebai, A.; Abdelhedi, R.; Bouaziz, M. Detection of chemlali extra-virgin olive oil adulteration mixed with soybean oil, corn oil, and sunflower oil by using GC and HPLC. J. Agric. Food Chem. 2014, 62, 4893–4904. [Google Scholar] [CrossRef] [PubMed]
- Fragaki, G.; Spyros, A.; Siragakis, G.; Salivaras, E.; Dais, P. Detection of extra virgin olive oil adulteration with lampante olive oil and refined olive oil using nuclear magnetic resonance spectroscopy and multivariate statistical analysis. J. Agric. Food Chem. 2005, 53, 2810–2816. [Google Scholar] [CrossRef]
- Andrikopoulos, N.K.; Giannakis, I.G.; Tzamtzis, V. Analysis of olive oil and seed oil triglycerides by capillary gas chromatography as a tool for the detection of the adulteration of olive oil. J. Chromatogr. Sci. 2001, 39, 137–145. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Duraipandian, S.; Petersen, J.; Lassen, M. Authenticity and concentration analysis of extra virgin olive oil using spontaneous Raman spectroscopy and multivariate data analysis. Appl. Sci. 2019, 9, 2433. [Google Scholar] [CrossRef]
- López-Díez, E.C.; Bianchi, G.; Goodacre, R. Rapid quantitative assessment of the adulteration of virgin olive oils with hazelnut oils using Raman spectroscopy and chemometrics. J. Agric. Food Chem. 2003, 51, 6145–6150. [Google Scholar] [CrossRef]
- Mosca, S.; Dey, P.; Salimi, M.; Gardner, B.; Palombo, F.; Stone, N.; Matousek, P. Spatially offset Raman spectroscopy—How deep? Anal. Chem. 2021, 93, 6755–6762. [Google Scholar] [CrossRef]
- Matousek, P.; Clark, I.P.; Draper, E.R.C.; Morris, M.D.; Goodship, A.E.; Everall, N.; Towrie, M.; Finney, W.F.; Parker, A.W. Subsurface probing in diffusely scattering media using spatially offset Raman spectroscopy. Appl. Spectrosc. 2005, 59, 393–400. [Google Scholar] [CrossRef]
- Eliasson, C.; Matousek, P. Noninvasive authentication of pharmaceutical products through packaging using spatially offset Raman spectroscopy. Anal. Chem. 2007, 79, 1696–1701. [Google Scholar] [CrossRef]
- Stokes, R.J.; Bailey, M.; Bonthron, S.; Stone, T.; Maskall, G.; Presly, O.; Roy, E.; Tombling, C.; Loeffen, P.W. New capability for hazardous materials ID within sealed containers using a portable spatially offset Raman spectroscopy (SORS) device. In Optics and Photonics for Counterterrorism, Crime Fighting, and Defence XII, Proceedings of the SPIE, Edinburgh, UK, 26–27 September 2016; Burgess, D., Owen, G., Bouma, H., Carlysle-Davies, F., Stokes, R.J., Yitzhaky, Y., Eds.; SPIE: Bellingham, WA, USA, 2016; Volume 9995. [Google Scholar]
- Bloomfield, M.; Andrews, D.; Loeffen, P.; Tombling, C.; York, T.; Matousek, P. Non-invasive identification of incoming raw pharmaceutical materials using Spatially Offset Raman Spectroscopy. J. Pharm. Biomed. Anal. 2013, 76, 65–69. [Google Scholar] [CrossRef]
- Izake, E.L. Forensic and homeland security applications of modern portable Raman spectroscopy. Forensic Sci. Int. 2010, 202, 1–8. [Google Scholar] [CrossRef]
- Ellis, D.I.; Eccles, R.; Xu, Y.; Griffen, J.; Muhamadali, H.; Matousek, P.; Goodall, I.; Goodacre, R. Through-container, extremely low concentration detection of multiple chemical markers of counterfeit alcohol using a handheld SORS device. Sci. Rep. 2017, 7, 12082. [Google Scholar] [CrossRef] [PubMed]
- Lohumi, S.; Lee, H.; Kim, M.S.; Qin, J.; Cho, B.K. Through-packaging analysis of butter adulteration using line-scan spatially offset Raman spectroscopy. Anal. Bioanal. Chem. 2018, 410, 5663–5673. [Google Scholar] [CrossRef]
- Liu, Z.; Huang, M.; Zhu, Q.; Qin, J.; Kim, M.S. Detection of adulterated sugar with plastic packaging based on spatially offset Raman imaging. J. Sci. Food Agric. 2021. [Google Scholar] [CrossRef]
- Afseth, N.K.; Bloomfield, M.; Wold, J.P.; Matousek, P. A novel approach for subsurface through-skin analysis of salmon using spatially offset Raman spectroscopy (SORS). Appl. Spectrosc. 2014, 68, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Huang, M.; Zhu, Q.; Qin, J.; Kim, M.S. Nondestructive freshness evaluation of intact prawns (Fenneropenaeus chinensis) using line-scan spatially offset Raman spectroscopy. Food Control 2021, 126, 108054. [Google Scholar] [CrossRef]
- Barnes, R.J.; Dhanoa, M.S.; Lister, S.J. Standard normal variate transformation and de-trending of near-infrared diffuse reflectance spectra. Appl. Spectrosc. 1989, 43, 772–777. [Google Scholar] [CrossRef]
- Martens, H.; Stark, E. Extended multiplicative signal correction and spectral interference subtraction: New preprocessing methods for near infrared spectroscopy. J. Pharm. Biomed. Anal. 1991, 9, 625–635. [Google Scholar] [CrossRef]
- Richardson, P.I.C.; Muhamadali, H.; Ellis, D.I.; Goodacre, R. Rapid quantification of the adulteration of fresh coconut water by dilution and sugars using Raman spectroscopy and chemometrics. Food Chem. 2019, 272, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Vera, D.N.; Jiménez-Carvelo, A.M.; Cuadros-Rodríguez, L.; Ruisánchez, I.; Callao, M.P. Authentication of the geographical origin of extra-virgin olive oil of the Arbequina cultivar by chromatographic fingerprinting and chemometrics. Talanta 2019, 203, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Hotelling, H. Analysis of a complex of statistical variables into principal components. J. Educ. Psychol. 1933, 24, 417–441. [Google Scholar] [CrossRef]
- Esbensen, K.H.; Geladi, P. Principal component analysis: Concept, geometrical interpretation, mathematical background, algorithms, history, practice. In Comprehensive Chemometrics; Brown, S.D., Tauler, R., Walczak, B., Eds.; Elsevier: Oxford, UK, 2009; pp. 211–226. [Google Scholar]
- Martens, H.; Naes, T. Multivariate Calibration; John Wiley & Sons: Chichester, UK, 1989; ISBN 0471-90979-3. [Google Scholar]
- Geladi, P.; Kowalski, B.R. Partial least-squares regression: A tutorial. Anal. Chim. Acta 1986, 185, 1–17. [Google Scholar] [CrossRef]
- Gromski, P.S.; Muhamadali, H.; Ellis, D.I.; Xu, Y.; Correa, E.; Turner, M.L.; Goodacre, R. A tutorial review: Metabolomics and partial least squares-discriminant analysis—A marriage of convenience or a shotgun wedding. Anal. Chim. Acta 2015, 879, 10–23. [Google Scholar] [CrossRef]
- Ellis, D.I.; Ellis, J.; Muhamadali, H.; Xu, Y.; Horn, A.B.; Goodacre, R. Rapid, high-throughput, and quantitative determination of orange juice adulteration by Fourier-transform infrared spectroscopy. Anal. Methods 2016, 8, 5581–5586. [Google Scholar] [CrossRef]
- Clayton, C.A.; Hines, J.W.; Elkins, P.D. Detection limits with specified assurance probabilities. Anal. Chem. 1987, 59, 2506–2514. [Google Scholar] [CrossRef]
- Ortiz, M.C.; Sarabia, L.A.; Herrero, A.; Sánchez, M.S.; Sanz, M.B.; Rueda, M.E.; Giménez, D.; Meléndez, M.E. Capability of detection of an analytical method evaluating false positive and false negative (ISO 11843) with partial least squares. Chemom. Intell. Lab. Syst. 2003, 69, 21–33. [Google Scholar] [CrossRef]
- Ellis, D.I.; Muhamadali, H.; Allen, D.P.; Elliott, C.T.; Goodacre, R. A flavour of omics approaches for the detection of food fraud. Curr. Opin. Food Sci. 2016, 10, 7–15. [Google Scholar] [CrossRef]
- Philippidis, A.; Poulakis, E.; Papadaki, A.; Velegrakis, M. Comparative study using Raman and visible spectroscopy of cretan extra virgin olive oil adulteration with sunflower oil. Anal. Lett. 2017, 50, 1182–1195. [Google Scholar] [CrossRef]
- Muik, B.; Lendl, B.; Molina, D.A.; Ayora-Cañada, M.J. Direct, reagent-free determination of free fatty acid content in olive oil and olives by Fourier transform Raman spectrometry. Anal. Chim. Acta 2003, 487, 211–220. [Google Scholar] [CrossRef]
- Blanch, G.P.; Caja, M.D.M.; del Castillo, M.L.R.; Herraiz, M. Comparison of different methods for the evaluation of the authenticity of olive oil and hazelnut oil. J. Agric. Food Chem. 1998, 46, 3153–3157. [Google Scholar] [CrossRef]
- Cox, A.; Wohlschlegel, A.; Jack, L.; Smart, E. The Cost of Food Crime. Available online: https://www.food.gov.uk/research/research-projects/the-cost-of-food-crime (accessed on 20 July 2021).
- Casadei, E.; Valli, E.; Panni, F.; Donarski, J.; Farrús Gubern, J.; Lucci, P.; Conte, L.; Lacoste, F.; Maquet, A.; Brereton, P.; et al. Emerging trends in olive oil fraud and possible countermeasures. Food Control 2021, 124, 107902. [Google Scholar] [CrossRef]
- Eliasson, C.; Macleod, N.A.; Matousek, P. Noninvasive detection of concealed liquid explosives using Raman spectroscopy. Anal. Chem. 2007, 79, 8185–8189. [Google Scholar] [CrossRef] [PubMed]
- Loeffen, P.; Maskall, G.; Bonthron, S.; Bloomfield, M.; Tombling, C.; Matousek, P. The performance of spatially offset Raman spectroscopy for liquid explosive detection. In Optics and Photonics for Counterterrorism, Crime Fighting, and Defence XII, Proceedings of the SPIE, Edinburgh, UK, 26–27 September 2016; SPIE: Bellingham, WA, USA, 2016; Volume 9995, p. 99950D. [Google Scholar]
- Loeffen, P.; Maskall, G.; Bonthron, S.; Bloomfield, M.; Tombling, C.; Matousek, P. Chemical and explosives point detection through opaque containers using spatially offset Raman spectroscopy (SORS). In Chemical, Biological, Radiological, Nuclear, and Explosives (CBRNE) Sensing XII, Proceedings of the SPIE Defense, Security, and Sensing, Orlando, FL, USA, 25–29 April 2011; SPIE: Bellingham, WA, USA, 2011; Volume 8018, p. 80181E. [Google Scholar]
- Bloomfield, M.; Loeffen, P.W.; Matousek, P. Detection of concealed substances in sealed opaque plastic and coloured glass containers using SORS. In Optics and Photonics for Counterterrorism and Crime Fighting VI and Optical Materials in Defence Systems Technology VII, Proceedings of the SPIE, Toulouse, France, 20–23 September 2010; SPIE: Bellingham, WA, USA, 2010; Volume 7838, p. 783808. [Google Scholar]
- Commission Implementing Regulation (EU) 2021/808 of 22 March 2021 on the Performance of Analytical Methods for Residues of Pharmacologically Active Substances Used in Food-Producing Animals and on the Interpretation of Results as Well as on the Methods to Be Used for Sampling and Repealing Decisions 2002/657/EC and 98/179/EC (Text with EEA Relevance). Available online: https://op.europa.eu/s/pmU5 (accessed on 20 July 2021).
| 785 nm | # PLS Factors | R2 | Q2 | RMSEC | RMSEP | LOD (%) |
|---|---|---|---|---|---|---|
| EVOO & Sunflower O | 6 | 0.99 | 0.99 | 3.64 | 4.23 | 2.92 |
| EVOO & Pomace OO | 3 | 0.95 | 0.89 | 7.21 | 10.20 | 19.94 |
| EVOO & Refined OO | 4 | 0.90 | 0.93 | 9.67 | 8.00 | 12.02 |
| 1064 nm | ||||||
| EVOO & Sunflower O | 2 | 0.98 | 0.99 | 4.07 | 3.71 | 6.19 |
| EVOO & Pomace OO | 4 | 0.97 | 0.95 | 5.32 | 6.50 | 11.50 |
| EVOO & Refined OO | 4 | 0.97 | 0.92 | 5.27 | 8.13 | 11.29 |
| 830 nm | ||||||
| EVOO & Sunflower O | 5 | 1.00 | 1.00 | 1.93 | 1.98 | 3.97 |
| EVOO & Pomace OO * | 3 | 0.98 | 0.92 | 4.30 | 5.34 | 10.39 |
| EVOO & Refined OO * | 6 | 0.93 | 0.96 | 4.84 | 6.35 | 11.39 |
| Mixture [Volume in Bottle] | #PLS Factors | R2 | Q2 | RMSEC | RMSEP | LOD (%) |
|---|---|---|---|---|---|---|
| Sunflower O in EVOO: 0–90% EVOO [90 mL in bottle] | 6 | 0.99 | 0.98 | 2.51 | 3.27 | 8.59 |
| Sunflower O in EVOO: 10–90% EVOO [28 mL in bottle] | 6 | 1.00 | 1.00 | 1.01 | 1.37 | 2.24 |
| Pomace OO in EVOO: 0–90% EVOO [28 mL in bottle] | 4 | 0.97 | 0.97 | 4.68 | 4.91 | 10.64 |
| Refined OO in EVOO: 0–90% EVOO [28 mL in bottle] | 6 | 0.92 | 0.84 | 7.37 | 10.15 | 13.60 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Varnasseri, M.; Muhamadali, H.; Xu, Y.; Richardson, P.I.C.; Byrd, N.; Ellis, D.I.; Matousek, P.; Goodacre, R. Portable through Bottle SORS for the Authentication of Extra Virgin Olive Oil. Appl. Sci. 2021, 11, 8347. https://doi.org/10.3390/app11188347
Varnasseri M, Muhamadali H, Xu Y, Richardson PIC, Byrd N, Ellis DI, Matousek P, Goodacre R. Portable through Bottle SORS for the Authentication of Extra Virgin Olive Oil. Applied Sciences. 2021; 11(18):8347. https://doi.org/10.3390/app11188347
Chicago/Turabian StyleVarnasseri, Mehrvash, Howbeer Muhamadali, Yun Xu, Paul I. C. Richardson, Nick Byrd, David I. Ellis, Pavel Matousek, and Royston Goodacre. 2021. "Portable through Bottle SORS for the Authentication of Extra Virgin Olive Oil" Applied Sciences 11, no. 18: 8347. https://doi.org/10.3390/app11188347
APA StyleVarnasseri, M., Muhamadali, H., Xu, Y., Richardson, P. I. C., Byrd, N., Ellis, D. I., Matousek, P., & Goodacre, R. (2021). Portable through Bottle SORS for the Authentication of Extra Virgin Olive Oil. Applied Sciences, 11(18), 8347. https://doi.org/10.3390/app11188347








