1. Introduction
As early as 1977, electrocardiogram (ECG) was identified for its potential for biometric authentication [
1]. ECG is relatively easy to acquire, for example, using finger sensors [
2]. Many methods have been implemented of ECG identification using feature extraction based on time, amplitude, and frequency [
3,
4,
5,
6], as well as using machine learning [
7,
8,
9]. Nevertheless, ECG biometrics still cannot be fully implemented in real-time security systems due to their high computational time and low accuracy.
ECG identification using multivariate analysis feature extraction has been reported in [
3]. Using principal components analysis (PCA) and soft independent modeling of class analogy (SIMCA), the ECG identification results vary from 90% to 100%. The accuracy of this method depends on two main factors, including the number of features and the type of lead used to obtain an ECG signal. In [
3], they used 12 leads and 30 types of time-based ECG features. Around 50 samples were evaluated from 20 ECG records. Although ECG identification can be performed with just one lead, this research needs to use at least three leads to improve identification accuracy [
3].
In [
5], the number of leads and the number of ECG features were reduced; specifically, the number of leads has been reduced one, and the number of features has been reduced to seven. After preprocessing and feature extraction, two different methods were normally used for identification purposes, i.e., template matching (TM) and the decision-based neural network (DBNN). Evaluation of 400 ECG beats from 20 people revealed that the accuracy of TM and DBNN was 95% and 80%, respectively. Moreover, combining both methods produced 100% identification accuracy.
In 2005, the ECG-ID database became publicly accessible [
10]. The database contains ECG recording files of 90 people using a single ECG lead. Recorded ECG signals from the same person are stored in one folder. Each folder contains two or more ECG recording files, except the 74th folder, containing only one record. After preprocessing and time-based feature extraction, ECG identification is conducted using PCA, linear discriminant analysis (LDA), and majority vote classifier (MVC). Moreover, the accuracy of this method is around 96%.
Apart from using the database of normal ECG heartbeats, ECG biometrics have also been tested using arrhythmia ECG signals [
11]. Using test data from the ECG records of 47 patients in the MIT-BIH database, 91.1% accuracy was reported. They applied five stages of blind signal processing (BSP), including preprocessing, wavelet transform, autocorrelation, component selection, and parallel one-dimension convolutional neural networks (1-D CNN). A similar identification method can be applied to other databases, including ECG records from the ECG-ID and MIT-BIH databases [
8,
9]. Bidirectional gated recurrent units (BGRUs) have been used to identify 47 patients from the MIT-BIH database and 90 ECG healthy people from the ECG-ID database with an accuracy of 98.40% and 98.60%, respectively [
8]. When, instead, a PCA neural network (PCANet) was used, accuracies of 100% and 97.75% were achieved when tested on ECG-ID and MIT-BIH databases, respectively [
9].
Recently, depthwise separable convolution (DSC) has been applied to a CNN model to reduce computation by reducing the number of model parameters and arithmetic calculations. DSC has been utilized to improve CNN model quality in image classification [
12] and neural machine translation [
13]. For cardiac arrhythmia classification, DSC has been tested on 16 ECG classes of the MIT-BIH database, outperforming other methods [
14]. Besides DSC, the CNN model can be further optimized using the residual technique. The residual technique has been used in CNN to simplify the training process and increase accuracy, as is evident in image classification [
15] and ECG classification [
16].
Residual DSC has been implemented in Xception [
12] and PydMobileNet [
17]. Similar algorithms with slight modifications will be used for ECG authentication in this paper. The first modification was on the filter dimension, i.e., 2D-CNN to 1D-CNN. The second modification was on the structure of CNN layers. In Xception and PydMobileNet, DSC was applied to single layers while, in our algorithm, DSC was implemented for the group of three layers, as has been explained for a cardiac arrhythmia classifier [
14]. A further modification was conducted for ECG authentication, in which a shortcut residual is applied to the three-layer DSC. Furthermore, for ECG authentication, none of the convolutional layers is similar to the ones in Xception and PydMobileNet. Each convolutional layer is an ordinary 1D convolution, not a DSC.
As discussed earlier, the common problems in ECG authentication include time-consuming signal processing and feature extraction. Another problem is accuracy; it is difficult to obtain 100% accuracy, which is required for authentication purposes. Therefore, in this paper, a fast and efficient algorithm will be proposed to overcome these problems using a residual depthwise separable convolution neural network. The training process required high computational resources, which can be alleviated using GPU. However, the testing process required low computational resources; a CPU alone is sufficient to be used during the authentication process.
2. Proposed ECG Authentication Algorithm Using Residual Depthwise Separable CNN
To simplify the ECG authentication process, the proposed method in this paper requires only two stages, including the segmentation stage and the classification stage. The segmentation stage carried out ECG beat detection and segmentation, while the classification stage performed feature extraction and classification. ECG beat detection is performed using Hamilton’s methods [
18,
19]. The segmentation is based on the relative position of the
, and occurs
before
and
after
depends on the sampling frequency and is set to 256 samples for both ECG-ID and MIT-BIH databases. Note that preprocessing has been performed on both databases to eliminate baseline drift and to remove powerline and high-frequency noises [
10]. Therefore, it can be assumed that the ECG signals from both databases were ready to be used for further processing.
The second stage, or classification stage, is realized by implementing the Residual Depthwise Convolutional Neural Network. The CNN used in this second stage is 1D-CNN that contains 28 layers. The 21 layers consist of seven layers of residual depthwise separable convolution neural network (RDSCNN) repeated three times, as illustrated in
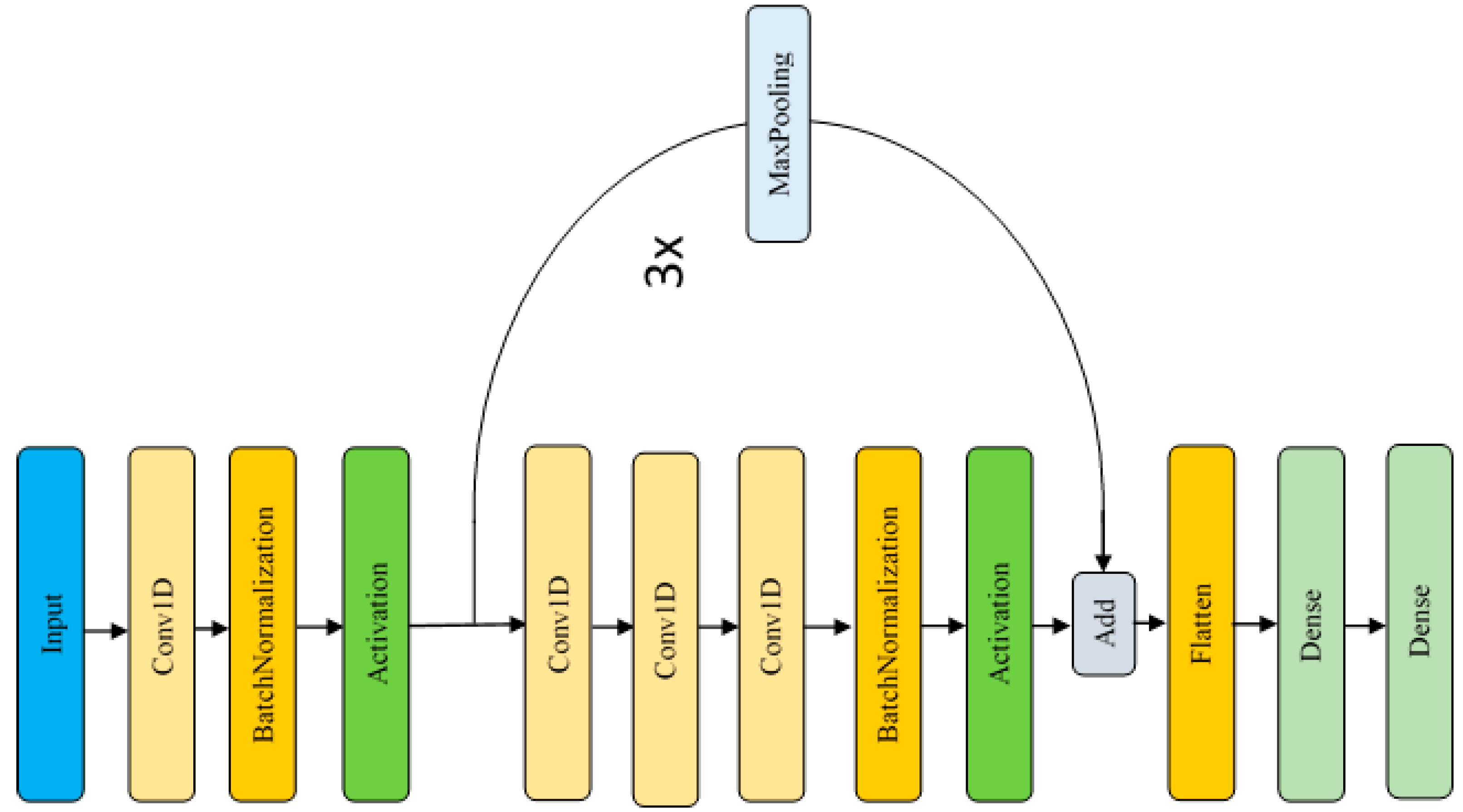
Figure 1.
As shown in
Figure 1, there are seven layers repeated three times, including a max pooling layer, three layers of 1D-CNN (Conv1D), a batch normalization layer, an activation layer, and an add layer. This structure is called as Residual Depthwise Separable Convolution implementation. The residual implementation uses a shortcut Max Pooling layer, while depthwise separable convolution implementation uses three Conv1D layers with the filter size [5 1 5], respectively. The summary of the proposed CNN model will be discussed in more detail in
Section 3.3.
There are 10 Conv1D layers in the proposed algorithm, which can be calculated using Equation (1) to (4), as follows [
20]:
where
is the layer number,
is the filter number at the particular layer,
is the index coefficient
of the filter,
is the input and
is the kernel (weight) from the
-th neuron at layer
to the
-th neuron at layer
.
is the output of the
-th neuron at layer
and
is the bias, which can be assumed to be zero.
performs full convolution in 1-D with
zero padding.
is the input difference after filtering and
flips the array.
is the delta error according to the loss function.
denotes error change with respect to the filter coefficients, while
denotes error change with respect to the bias.
To maximize the accuracy, the CNN model in this paper is optimized using two methods, i.e., Depthwise Separable [
12,
13] and Residual [
15]. The Depthwise Separable can be explained using Equation (5) to (8), as follows:
where
is the element index in layer
.
and
are the filters calculated from Equations (6) and (7). The calculation in Equation (5) can be replaced with Equation (8) through calculations of Equations (6) and (7). Although the output layer from Equation (5) is similar to Equations (6) to (8), the total number of parameters and the training time were decreased, as can be seen in
Table 1. Note that the depthwise separable technique will reduce the training time and reduce the accuracy as well. To compensate for the slight decrease in performance, the residual technique can be applied. Therefore, combining these two techniques, i.e., Depthwise Separable and Residual, not only reduces the number of parameters for training but also improves the accuracy.
Residual is a technique in CNN, in which one of the branches is a shortcut bypassing one or several other branch layers. Initially, this technique was intended to overcome the saturation of the increasing number of layers. Too many layers will lead to longer training iteration, lower speed, and lower accuracy. With the residual technique, the training iteration can be shorter, and the accuracy will increase. The general formulas for the identity function related to the residual shortcut are as follows [
15]:
where
is the input,
is the output layer and
is a filter or residual mapping which can be optimized.
is a collection of overlapped layers.
is a linear projection to adjust the
and
dimensions when a shortcut is performed. There is almost no change in the number of parameters and arithmetic operations, except for the addition operation which has a little computational load. Therefore, the application of this residual technique can minimize the training iteration while improving accuracy.
Generally, this residual technique is applied to CNN models with millions of training data, hundreds of layers, and tens of thousands of iteration epochs. However, in this paper, the residual technique can be applied to a smaller number of training data (between 900 and 4800), fewer layers (less than 30), and less iteration (100 epochs). As the number of biometric data to be authenticated increases, the number of layers, the number and size of filters, and the number of iterations can be increased as well.
3. Experimental Setup
This section describes the experimental setup to evaluate our proposed algorithm, including the ECG databases used, various experimental scenarios, and the summary of the CNN model.
3.1. ECG Databases and Computing Platform
Two popular Physionet databases were used to evaluate the performance of our proposed algorithm, i.e., ECG-ID [
10] and MIT-BIH [
21]. These two databases have been used by several researchers to test the performance of the ECG authentication algorithm. In this paper, each database is used to test five different conditions, with variations in the application of beat detection, as well as DSC and residual configurations.
The ECG-ID database contains 90 folders from 90 different people. Each folder contains several ECG records with a 500 Hz sampling frequency. Each record has been provided with 10 annotations related to the position of in the first 10 beats. The recording duration ranges from 20 s to 5 min. For testing purposes, only 179 files were used, i.e., two files from each folder, except the 74th folder which has only one file. For training purposes, ten beats of the first file in each folder were obtained, so that around 900 beats were used. For validation purposes, ten beats of the second file were acquired. As the 74th folder contains only one file, 10 beats from this file were used for both training and validation. The ECG beat extraction from 179 files from 90 folders produced 900 beats for training and 900 beats for validation. These validation beats were used to validate the CNN weights obtained in the training process.
Besides using the ECG database of healthy people, our proposed algorithm was also tested with ECG records with several cardiac arrhythmia beats taken from the MIT-BIH database. The database contains 48 files, in which each file contains around 30 min of ECG recordings of cardiac arrhythmia patients. Each data file is complemented with the annotation file containing the position in the associated ECG recording. There are two channels provided for each of these ECG records, but in this paper only channel 0 (first channel) was used. For training and testing purposes, each ECG recording is divided into two, in which each half contains 325,000 sample points. The first half is used for training, while the second half is used for testing. For training, only 100 beats of the first half were used, producing a total of 4800 ECG beats. For validation, only 10 beats of the second half were used, producing a total of 480 ECG beats. These validation beats were used to validate the CNN weights obtained in the training process.
The proposed algorithm was implemented in Python with Tensorflow with GPU [
22] and Keras libraries [
23], as well as other standard libraries, such as Numpy, Matplotlib, and Scipy. The experiments are performed on a computer with Intel Core i7-7700 CPU with a total of eight logical processors, memory of 8 GBytes, and an Nvidia GeForce GTX 1060 6GB DDR5 graphics card, using the Microsoft Windows 10 64-bit operating system.
3.2. Experimental Profiles
Using the two ECG databases, five experimental profiles with various CNN models were carried out as shown in
Table 1. In the first profile, DSC was not implemented and only residual was implemented. In the second profile, DSC was implemented and residual was not implemented. The third, fourth, and fifth profiles used both DSC and residual (our main proposed algorithm) with different methods in beat detection and segmentation. Manual beat detection used the annotated file marked by medical experts. Hamilton’s automatic beat detection and segmentation used Hamilton’s method [
18,
19]. The hybrid beat detection used manual beat detection using annotated files for the training and Hamilton’s automatic beat detection for the testing.
3.3. Residual Depthwise Separable Convolution Neural Network Model (RDSCNN)
The proposed residual depthwise separable convolution neural network (RDSCNN) algorithm is shown in
Figure 1 and summarized in
Table 2. There are two almost identical models applied to the two ECG databases. The difference is only in the number of neurons in the last layer configured according to the number of classes, i.e., 90 and 48 classes for ECG-ID and MIT-BIH database, respectively. For experimental scenario one (without DSC), the number of filters, size, and stride in layers 6, 13, and 20 should be modified (see column three in
Table 2) from [16 1 1] to [32 5 1]. For experimental scenario two (without residual), layers 10, 11, 17, 18, 24, and 25 should be removed. The rest of the experimental scenarios used the complete RDSCNN algorithm.
4. Results and Discussion
This section will elaborate on ECG beat segmentation and detection, training and validation, ECG biometric authentication experiments, and comparison with other methods.
4.1. Experiment on ECG Beat Detection and Segmentation
Using ECG-ID and MIT-BIH databases, the ECG record and its
positions were obtained, as shown in
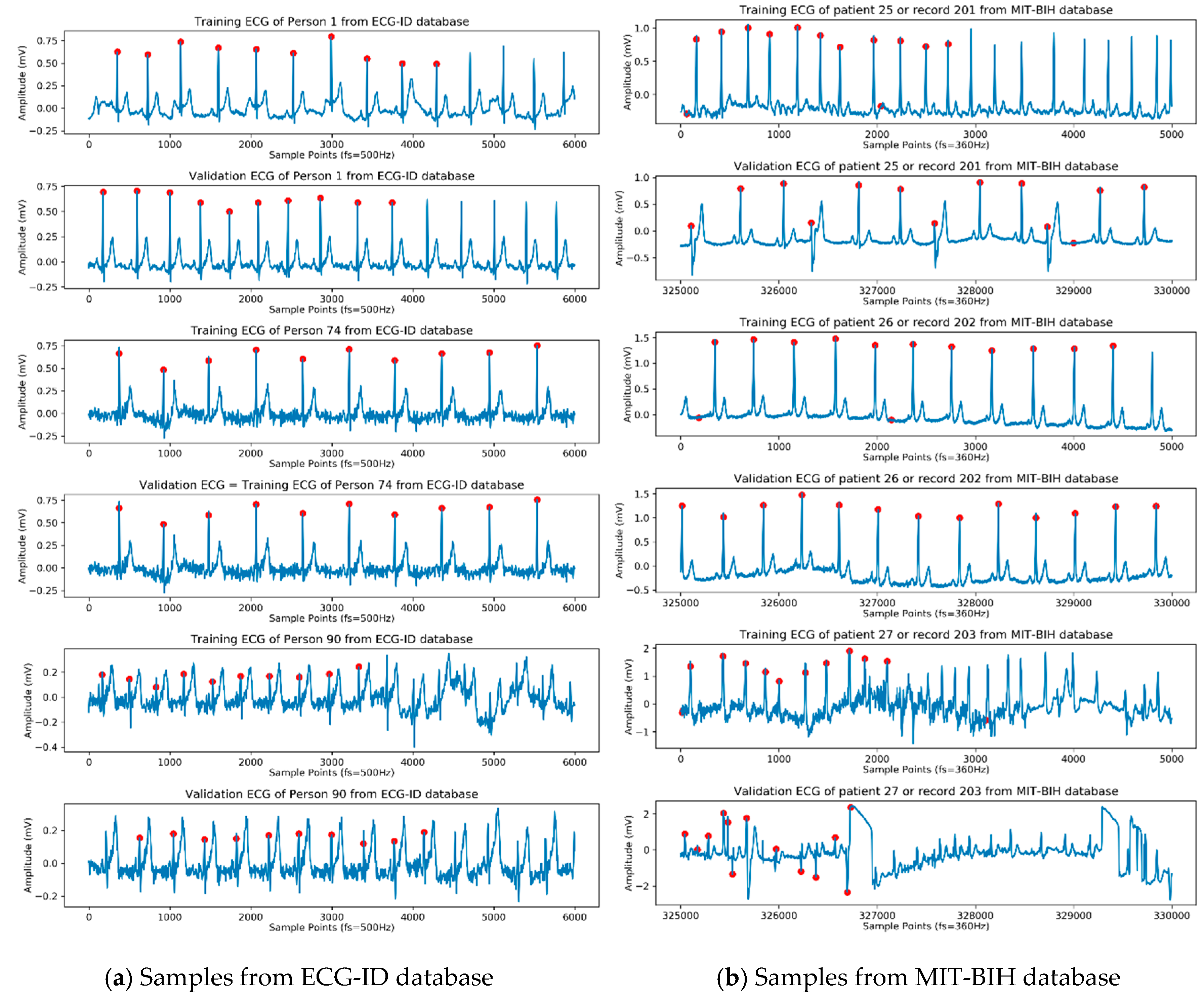
Figure 2. For the ECG-ID database, three samples were taken from 90 people. The first record was used for training, while the second record was used for validation. There were ten annotated
as shown in
Figure 2 for each person. In the MIT-BIH database, records 201 and 202 belonged to the same patient with different ECG beats, so they were classified as two classes. The red dots in
Figure 2 are an example of the
position detected manually by cardiologists. Using this
position, the ECG beat was detected and extracted. An example of ECG beat detection and segmentation for both databases is shown in
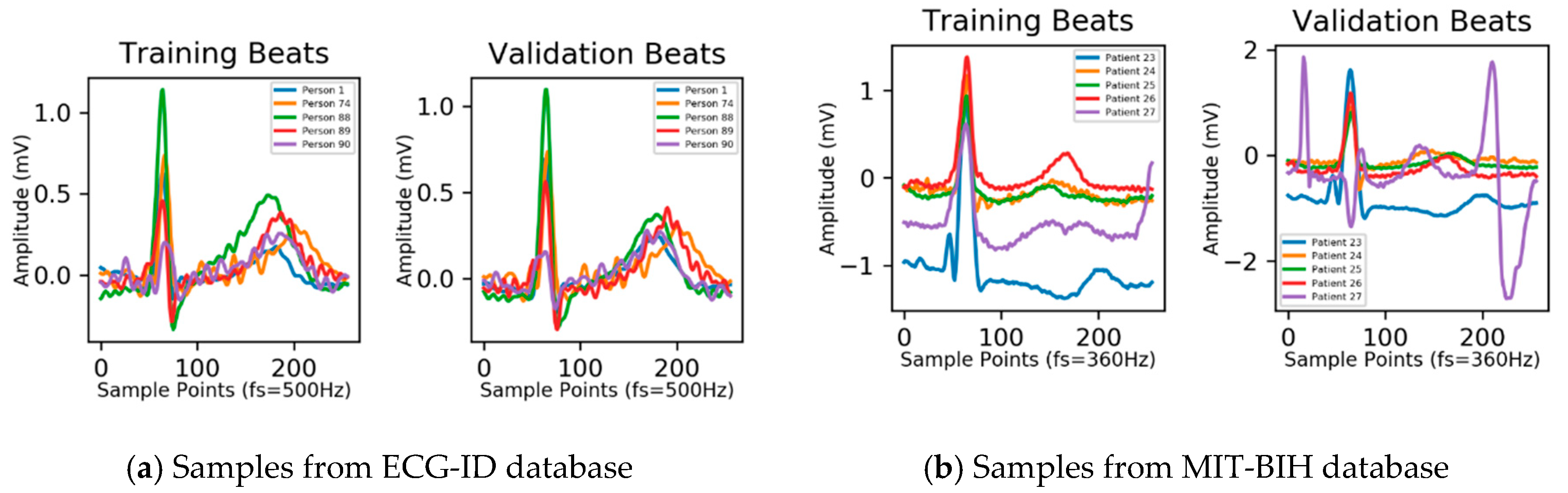
Figure 3.
As shown in
Figure 3a, ECG records from the ECG-ID database have been preprocessed to remove three types of noises, i.e., baseline drift, powerline noise, and high-frequency noise. Meanwhile,
Figure 3b shows that the effect of baseline drift and powerline noise is still visible in the ECG signal.
4.2. Experiment on the Training and Validation
As explained in
Section 3.1, the detected and segmented ECG beat was divided into two parts, i.e., training and validation. ECG beats for validation were conducted only after the training process was completed, by producing a weight file with the optimum value of all model parameters. There were a total of 33,226 and 30,496 parameters to be trained and optimized for the ECG-ID and MIT-BIH database, respectively. An example of ECG features extracted using the trained model can be seen in
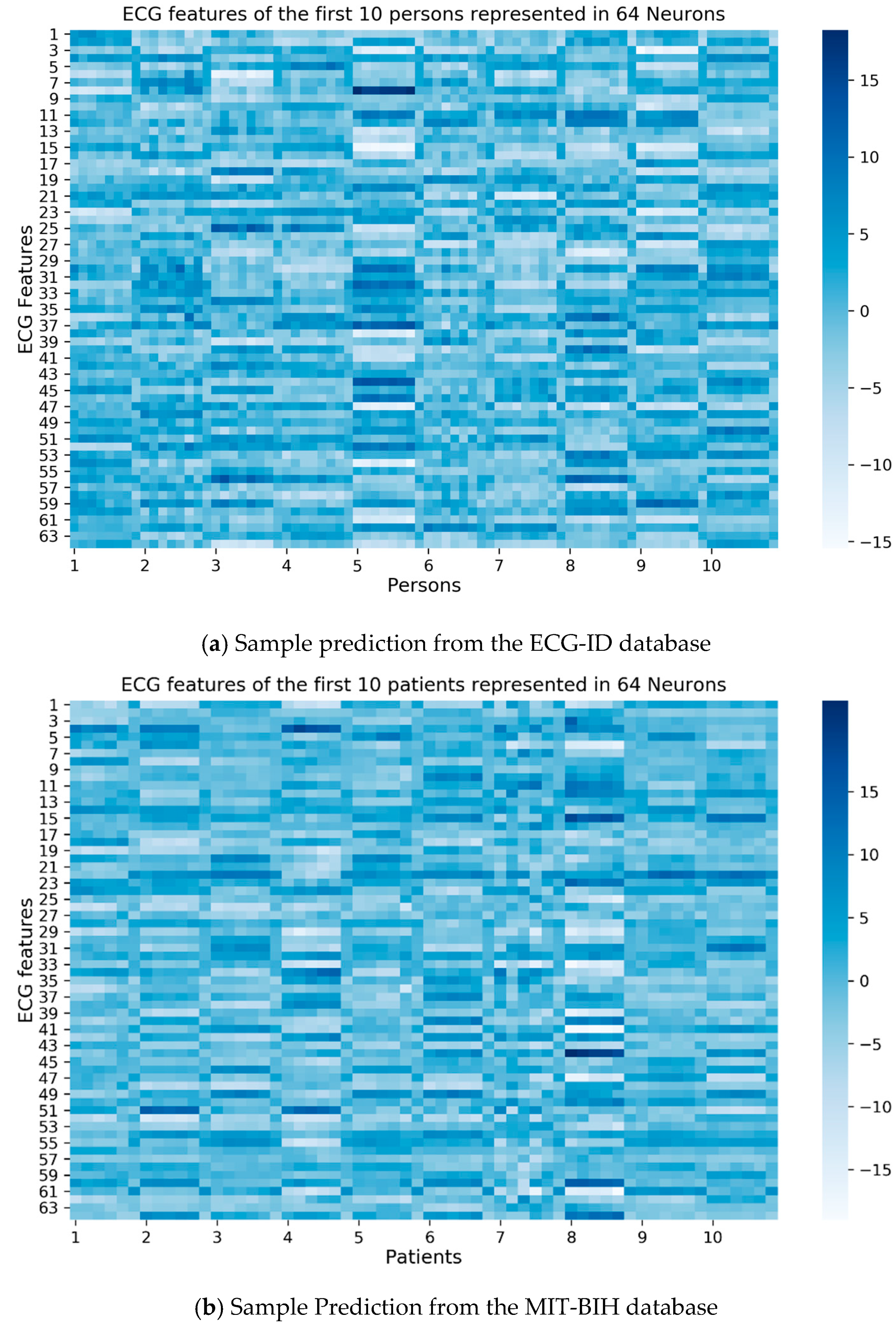
Figure 4. Referring to
Table 2, layer 27 (dense_1 layer) produces 64 values which can be used to represent the ECG beat feature of each person.
Figure 4a shows that there were eight columns for each person, while
Figure 4b shows that there were six columns for each person. The horizontal lines indicate the regularity of each person’s ECG beat feature.
Figure 4a shows that ECG beats from persons five, seven, and eight have visible horizontal lines, which indicate they are easier to identify.
Figure 4b shows that the regular ECG beat feature was seen for the first until the sixth patient. Meanwhile, the seventh patient does not show clear horizontal lines, indicating that the seventh patient’s ECG beats are rather difficult to identify.
Table 3 shows the authentication prediction in more detail for eight and six ECG beats of the ECG-ID and MIT-BIH databases, respectively. For ECG-ID, the sixth person could be authenticated as the 6th or 62nd person. Meanwhile, for MIT-BIH, the seventh patient could be authenticated as the 7th or 30th patient.
Based on the previous discussion, the ECG beat features in the 27th layer could be utilized for selection of the training and validation beats. This layer could be further investigated to produce a biometric key. Each biometric key of these 64 features can be quantized into 6 bits only, because the number keys are integer numbers between −20 to 20. Moreover, the number of bits and the number of features can be adjusted according to the number of persons to be authenticated.
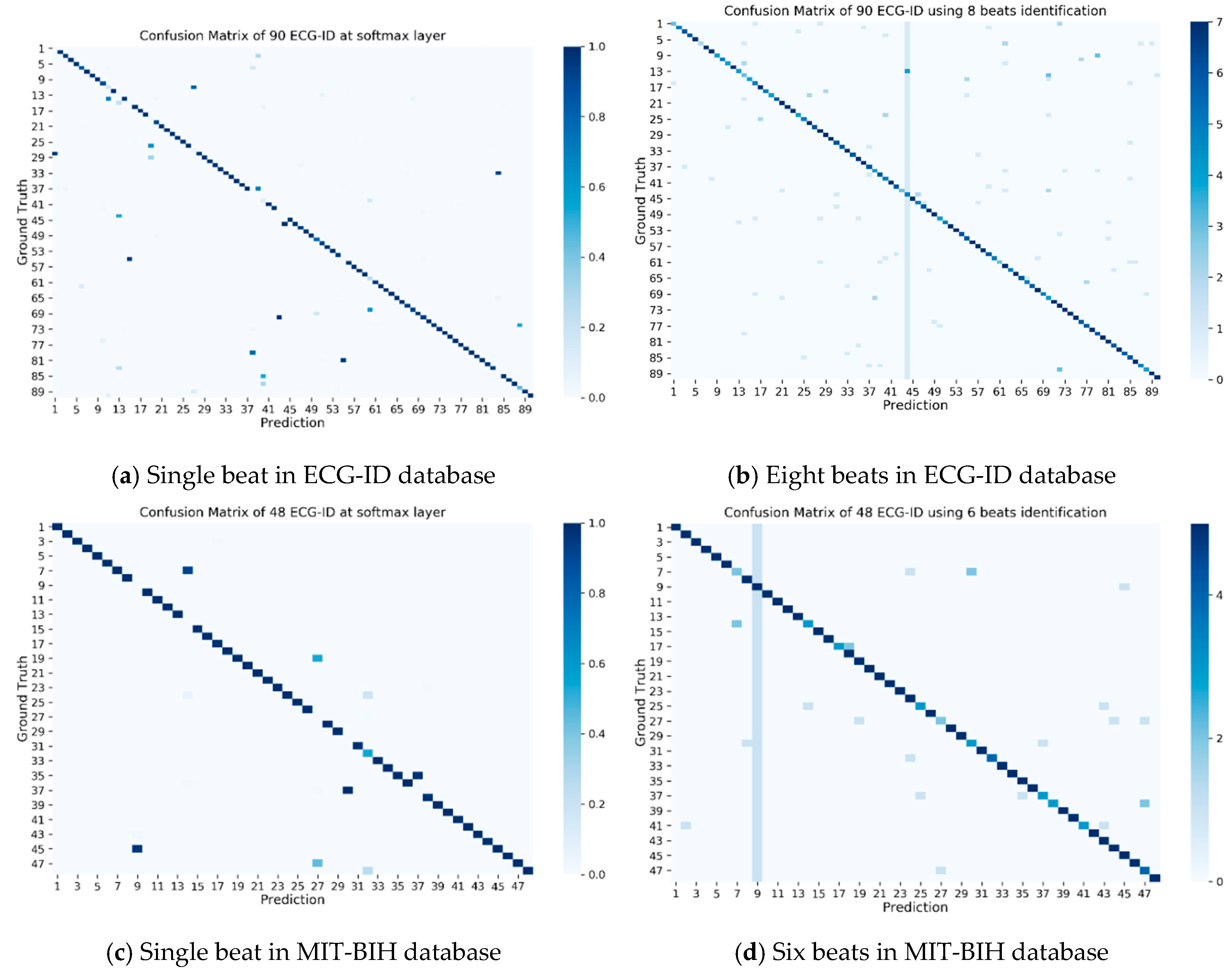
The summary of the authentication results of all validation beats can be seen in the confusion matrix shown in
Figure 5.
Figure 5a,b shows the confusion matrix of the authentication results for the 90 validation beats. These 90 beats were extracted from ECG records in the ECG-ID database. Each person is represented by eight beats or a total of 729 validation beats.
Figure 5c,d is the confusion matrix of the authentication results for the 48 validation beats. These 48 patient beats were extracted from ECG records in the MIT-BIH database. Each patient is represented by six beats or a total of 288 validation beats.
Figure 5a,c shows the confusion matrix for a single validation beat, i.e., the first beat of eight or six consecutive beats.
Figure 5b,d shows the confusion matrix for multiple validation beats. From
Figure 5, it can be shown that 100% accuracy can be obtained for the multiple validation beats, with six beats and eight beats for the ECG-ID and MIT-BIH databases, respectively. The optimum number of multiple beats will be evaluated in the next experiment. Nevertheless,
Figure 5 shows that multiple ECG beats produced better accuracy compared to a single ECG beat.
4.3. Experiment on the Effect of the Number of Consecutive ECG Beats
In this section, the effect of ECG beat detection methods, i.e., manual (using annotated files) vs. automatic Hamilton’s method, and the effect of the number of ECG beats on the accuracy, will be elaborated.
Table 4 shows the results for various numbers of ECG beats for two ECG beat detections and two ECG databases.
With manual ECG beat detection, 100% authentication accuracy can be obtained, starting from eight beats and above for the ECG-ID database and from six beats and above for the MIT-BIH database. This could be because the number of the persons to be identified from the ECG-ID database (90 persons) is almost twice the number of the patients to be identified from the MIT-BIH database (48 patients). In contrast, with Hamilton’s automatic detection, the authentication accuracy for both ECG-ID and MIT-BIH cannot reach 100%. This can happen because not all the ECG beats detected by Hamilton’s method are good beats, so they might require further preprocessing. Nevertheless, manual ECG beat detection (using annotated files) is only feasible in the lab or offline authentication, as the beat detection need to be verified manually by cardiologists. Therefore, the optimum numbers of ECG beat using Hamilton’s beat detection is seven and five for ECG-ID and MIT-BIH databases, respectively (the accuracy is highlighted in bold in
Table 4).
4.4. Experiment on the Various CNN Models and ECG Beat Detection
As described in
Section 3.2 (see
Table 1), five profiles were carried out for ECG-ID and MIT-BIH database, i.e., a total of 10 experiments.
Table 5 shows the experimental results for various CNN models (various experimental profiles). Of the particular interest is the third profile, which has the following configuration: using manual ECG beat detection (annotated files), using both depthwise separable CNN (DSC) and residual CNN. The third experimental profile produced good results for both the ECG-ID and MIT-BIH database, i.e., 100% accuracy, for eight and six ECG beats, respectively.
Although the third profile produced good authentication results, manual beat detection is only feasible in the lab or in offline authentication, as the beat detection needs to be conducted manually by cardiologists. Note that the first profile produced a good result as well, but it still uses manual beat detection, and the required consecutive beats per patient for the MIT-BIH database was higher (8) compared to the third scenario (6). Comparing the fourth and fifth profiles, it was found that Hamilton’s automatic beat detection produced better accuracy compared to hybrid beat detection (training using manual beat detection, while testing using Hamilton’s automatic beat detection). Therefore, the fourth experimental profile was found to be the optimal CNN configuration.
From measurement results using the computing platform described in
Section 3.1, automatic ECG beat detection in the ECG-ID database using Hamilton’s method required around 141,318
s to detect around 2412 ECG beats or 59
s per beat on average. In contrast, in the MIT-BIH database, beat detection required 10,827,746
s to detect around 108,709 beats, or 100
s per beat on average. Therefore, the maximum detection time required to segment 10 beats is around 1750
s. This is much faster than acquiring 10 ECG beats from someone’s finger.
4.5. Comparison with Other Algorithms
Table 6 shows the benchmarking results with the other 11 algorithms. Of the various algorithms, we highlighted the top three highest accuracies tested with both databases, i.e., Salloum & Kuo [
24], Bassiouoni et al. [
25], and Wang et al. [
9]. Note that, for a fair comparison, we used accuracy results from manual ECG beat detection and segmentation, as used by the other top three algorithms. Nevertheless, for real-time implementation, we could use Hamilton’s method for automatic beat detection and segmentation, as it only provides a slight decrease in accuracy performance.
In Salloum & Kuo [
24], ECG beat detection and segmentation was conducted using Tompkins’s algorithm [
29], in which they manually selected the best-quality eighteen or nine consecutive ECG beats to be fed to the Long Short Term Memory (LSTM) classifier. From each patient of MIT-BIH, they selected 18 ECG beats for training and testing. Each beat has 250 sample points, which is 125 points before and after
. For each subject of ECG-ID, they selected nine ECG beats for training and testing. Each beat has 300 sample points, which is 150 before and after
. Our proposed algorithm used a lower number of consecutive ECG beats, i.e., eight for ECG-ID and six for MIT-BIH, and no selection of the quality of ECG signal. Moreover, LSTM is rather more complicated compared to our proposed RDSCNN algorithm, so we can predict that our algorithm will be faster in the authentication process.
In Bassiouoni et al. [
25], the features were extracted from fiducial, non-fiducial, and its fusion. There were two different classifiers used, i.e., an artificial neural network (ANN) for MIT-BIH and a support vector machine (SVM) for ECG-ID. Only 30 out of 48 subjects in the MIT-BIH database were tested, while all 90 subjects in the ECG-ID database were tested. In summary, the algorithm in [
25] has three stages, including beat segmentation, feature extraction, and different classifiers for different databases. In contrast, our proposed algorithm has only two (faster) stages, including beat segmentation and an RDSCNN classifier, which can be applied for both databases.
Wang et al. [
9] used Tompkin’s method for ECG beat detection and segmentation starting from 0.24 s and 0.4 s before and after
, respectively. The QRST feature was extracted after the beats were decomposed using the discrete wavelet transform (DWT). They used a CNN model that functions like PCA, called PCA-Net, to extract ECG features from the results of DWT decomposition. For classification and authentication, SVM was used. The algorithm is rather complicated (higher computational time) and the accuracy for the ECG-ID database is only 97.75%.
As shown in
Table 6, our proposed algorithm using residual depthwise separable convolution neural network outperforms other algorithms, in which it can achieve 100% accuracy for both the ECG-ID and MIT-BIH databases. Furthermore, using Hamilton’s method for automatic beat segmentation, it requires around 100
s (see
Section 4.4). The classification process for eight consecutive ECG beats required around 560
s. Therefore, our proposed algorithms required around 660
s for the authentication process, i.e., automatic beat segmentation and classification. For a further comprehensive evaluation, other ECG databases could be used as highlighted in [
30], including six public domain databases and twelve private databases.
5. Conclusions
In this paper, we have presented a fast and accurate algorithm for ECG authentication using residual depthwise separable convolutional neural networks (RDSCNN). Two prominent databases were used for performance evaluation, i.e., 90 subjects from the ECG-ID database and 48 patients from the the MIT-BIH database. Two ECG beat detection and segmentation methods were evaluated, including manual, using annotated files, and automatic, using Hamilton’s method. It was found that the optimum number of consecutive ECG beats for automatic beat detection was seven and five for the ECG-ID and MIT-BIH databases, achieving authentication accuracy of 98.89% and 97.92%, respectively. Furthermore, using manual beat detection with eight (ECG-ID) and six (MIT-BIH) consecutive ECG beats, 100% authentication accuracy can be achieved. Our proposed algorithm is also rather fast, as it requires around 660 s to conduct authentication, i.e., automatic segmentation and classification. Further research will include optimizing current automatic ECG beat detection and segmentation to be as close as possible to the manual segmentation by cardiologists. Automatic selection of high-quality ECG beats for training and testing could also be conducted, for example by using the integer numbers of 64 ECG features and fractional numbers in the softmax layer. The number of consecutive ECG beats for authentication purposes could be further optimized, down to a single ECG beat.