1. Introduction
The recent years have seen a rapid increase in the incidence of diabetes around the world, due to many factors such as the continuous improvement in people’s living standards, changes in dietary structure, an increasingly rapid pace of life, and a sedentary lifestyle. Diabetes has become the third major chronic disease that seriously threatens human health, following cancer and cardiovascular disease [
1,
2]. According to statistics from the International Diabetes Federation (IDF), there were approximately 425 million patients with diabetes across the world in 2017. One in every 11 adults has diabetes, and one in every two patients is undiagnosed [
3]. As of 2016, diabetes directly caused 1.6 million deaths [
4], and it is estimated that by 2045, nearly 700 million people worldwide will suffer from diabetes, which will pose an increasing economic burden on health systems in most of countries. It is forecast that by 2030, at least
$490 billion will be spent on diabetes globally [
5].
Currently, the diagnosis rate of diabetes is low, as patients exhibit no obvious symptoms in the early onset of the disease, and many people do not realize they have the disease [
6]; thus, early detection and diagnosis are particularly needed. It is relatively simple to measure a person’s blood glucose under the existing medical conditions, but it takes a lot of human and material resources to detect the blood glucose of a large number of people in medical examination. Therefore, the prediction of the blood glucose of a large number of people in medical examination by machine learning can save a lot of unnecessary expenses (for example [
7]).
With the application of machine learning in medical fields, more and more people are applying emerging prediction methods to many different medical fields to help greatly reduce the workload of the related medical staff and improve the diagnosis efficiency of doctors. For example, Yu Daping and Liu applied the XGBoost model for the early diagnosis of lung cancer [
8]. Tjeng Wawan Cenggoro used the XGBoost model to predict and analyze colorectal cancer in Indonesia [
9]. Chang Wenbing forecasted the prognosis of hypertension using the XGBoost model [
10]. Ogunleye Adeola Azeez applied the XGBoost model for the diagnosis of chronic kidney disease [
11]. Wenbing Chang used the XGBoost model and clustering algorithm to analyze the probability of hypertension-related symptoms [
12]. Wang Bin predicted severe hand, foot, and mouth disease using the CatBoost model [
13].
As the XGBoost model generates a decision tree using the level-wise method [
14], it the splits leaves simultaneously on the same layer, but the splitting gain of most leaf nodes is low. In many cases where further splitting is unnecessary, the XGBoost model will continue to split, causing non-essential expenses. Compared with the LightGBM model, the XGBoost model occupies more memory and consumes more time when the dataset is large.
The traditional genetic algorithm and random search algorithm have different defects—the genetic algorithm is a natural adaptive optimization method that simulates the problem to be solved as a process of biological evolution, and gradually eliminates the solutions with low fitness function values through generating next-generation solutions by operations such as replication, crossover, and mutation [
15]. The genetic algorithm uses the search information of multiple search points at the same time, and adopts probabilistic search technology to obtain the optimal or sub-optimal solution of the optimization problem. It boasts of good search flexibility, global search capability, and is easily implemented [
16]. However, it has a poor local search ability, complicated process caused by many control variables, and there are no definite termination rules. The random searching algorithm (RandomizedSearchCV) performs a random search in a set parameter search space. It samples a fixed number of parameters from a specified distribution instead of trying all of the parameter values [
17]. However, the random searching algorithm exhibits a poor performance when the dataset is small. Therefore, this paper proposes an improved LightGBM model for blood glucose prediction.
The main contributions of this study are as follows: by preprocessing the data on various medical examination indicators of people receiving medical examination, this paper proposes a LightGBM model optimized by the Bayesian hyper-parameter optimization algorithm to predict the blood glucose level of people receiving medical examination. The LightGBM model optimized by the Bayesian hyper-parameter optimization algorithm achieves a higher accuracy than the XGBoost model, Catboost model, the LightGBM model optimized by genetic algorithm, and the LightGBM model optimized by random searching algorithm, which can help doctors give early warning to those potentially suffering from diabetes, so as to reduce the incidence of diabetes, increase the diagnosis rate, and provide new ideas for the in-depth study on diabetes. The proposed method was verified by the diabetes data from a grade-three first-class hospital in China from September to October 2017.
2. Materials and Methods
2.1. LightGBM Model
LightGBM is an improvement framework based on decision tree algorithm released by Microsoft in 2017. LightGBM and XGBoost both support parallel arithmetic, but LightGBM is more powerful than the previous XGBoost model, with a fast training speed and less memory occupation, which can reduce the communication cost of parallel learning. LightGBM is mainly featured by the decision tree algorithm based on gradient-based one-side sampling (GOSS), exclusive feature bundling (EFB), and a histogram and leaf-wise growth strategy with a depth limit.
The basic idea of GOSS (gradient-based one-side sampling) is to keep all of the large gradient samples and to perform random sampling on the small gradient samples according to proportion. The basic idea of the EFB (exclusive feature bundling) algorithm is to divide the features into a smaller number of mutually exclusive bundles, that is, it is impossible to find an accurate solution in polynomial time. Therefore, what it uses is an approximate solution, that is, a small number of sample points that are not mutually exclusive are allowed between features (for example, some corresponding sample points are not non-zero at the same time). Allowing a small part of conflict can obtain a smaller number of feature bundles, which further improves the computational effectiveness [
18].
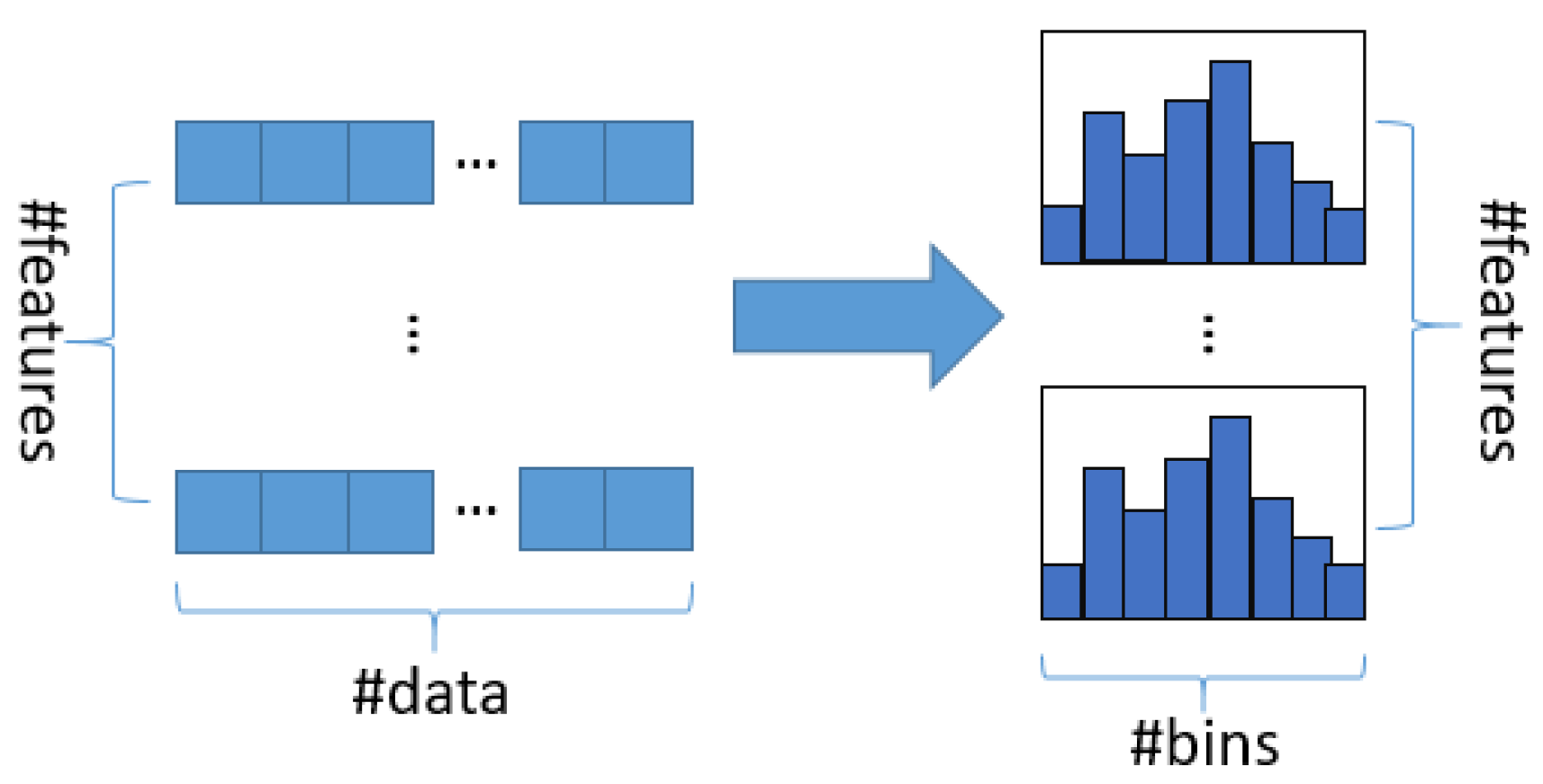
The basic idea of the histogram algorithm is to discretize continuous floating point features into k integers, and to construct a histogram with a width of k at the same time. When the data are traversed, statistics is accumulated in the histogram with the discretized value as index. After the data are traversed once, the histogram accumulates the required volume of statistics, and then the optimal segmentation point can be found through traverse according to the discrete value in the histogram, as shown in
Figure 1:
Most of the learning algorithm tree of decision tree is generated by the level-wise growth method, such as XGBoost, as shown in
Figure 2:
LightGBM uses a leaf-wise growth strategy with a depth limit to find a leaf node with the largest split gain in all of the current leaf nodes, then splits, and so on, as shown in
Figure 3:
Compared with the level-wise growth strategy, leaf-wise tree growth can reduce large errors and achieve a higher accuracy, thereby providing solutions to many problems. For example, Xile Gao used Stacked Denoising Auto Encoder (SDAE) and LightGBM models to recognize human activity [
19]; Sunghyeon Choi employed a random forest, XGBoost model, and LightGBM model to predict solar energy output [
20]; João Rala Cordeiro forecasted children’s height using a XGBoost model and LightGBM model [
21]; Ma Xiaojun et al. adopted a LightGBM model and XGBoost model to predict the default of P2P network loans [
22]; Chen Cheng et al. used LightGBM and a multi-information fusion model to predict the interaction between proteins [
23]; and Vikrant A. Dev applied a LightGBM model to stratum lithology classification [
24].
The following is the introduction to the theory of the LightGBM model’s objective function: yi is the objective value, i is the predicted value, T represents the number of leaf nodes, q denotes the structure function of the tree, and w is the leaf weight.
The objective function is as follows:
Use the Taylor expansion to define the objective function:
At this time, the objective function is the following:
Use the accumulation of n samples to traverse all of the leaf nodes:
where
Ij is the sample set in leaf node j, namely:
The partial derivative of the output
Wj of the
jth leaf node is obtained, and the minimum value is obtained as follows:
When the structure of the q (x) tree is determined, the function
Lt(q) is obtained, as follows:
The calculated gain is the following:
2.2. Bayesian Hyper-Parameter Optimization Algorithm
The basic idea of the Bayesian hyper-parameter optimization algorithm is to establish a substitute function based on the evaluation result of the past objective to find the minimum value of the objective function. The substitute function established in this process is easier to optimize than the original objective function, and the input value to be evaluated is selected by applying a certain standard to the proxy function [
25]. Although the genetic algorithm is currently widely used in the parameter optimization of machine learning [
26,
27,
28,
29], its disadvantages are also obvious, such as a poor ability of local search, many control variables, a complicated process, and no determined termination rules. The random searching algorithm is a traditional parameter optimization algorithm in machine learning [
30]. It uses random sampling within the search range for parameter optimization, but the effect is poor when the dataset is small. In contrast, Bayesian hyper-parameter optimization takes the result of the previous evaluation into account when trying another set of hyper-parameters, and has a simpler process of optimizing model parameters than the genetic algorithm, which can save a lot of time.
The HY_LightGBM model proposed in this paper uses one of the Bayesian optimization libraries in Python, Hyperopt, which uses Tree Parzen Estimation (TPE) as the optimization algorithm.
The optimization process of the Bayesian optimization algorithm is shown in
Figure 4.
By defining the objective function, domain space, and constructing the substitution function, the optimal parameters were finally obtained, with mean square error (MSE) as the evaluation indicator. The obtained optimal parameters were input into the LightGBM model to further improve the prediction ability of the model.
2.3. Improved LightGBM Model Based on Bayesian Hyper-Parameter Optimization Algorithm
The experiment was conducted on a computer with Intel I5 8400 2.8 GHz six-core six-thread central processing unit (CPU), 16G random-access memory (RAM), and Windows 10 operating system. The simulation platform is Pycharm, and Python was used for programming, with sklearn, pandas, and numpy libraries adopted.
Because LightGBM has many parameters, the manual adjustment of parameters will be complicated, and considering its parameters have a great impact on experimental results, it is particularly necessary to use Bayesian hyper-parameter optimization algorithm for parameter optimization. The Hyperopt used in this paper is one of the Bayesian optimization libraries in Python. It is also a class library used in distributed asynchronous algorithm configuration in Python, with a faster speed and better effect in finding the optimal parameters of the model than the traditional parameter optimization algorithms.
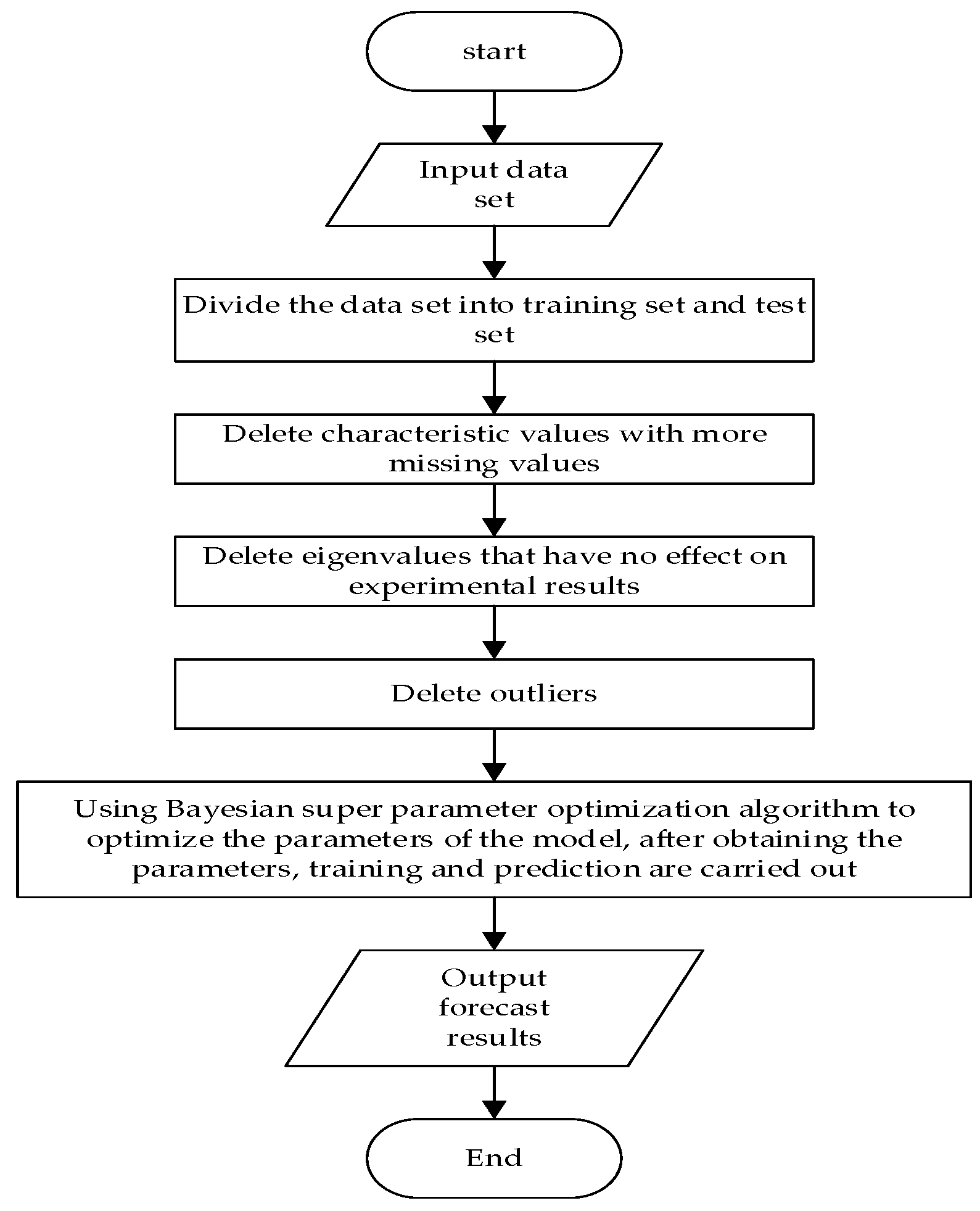
The specific steps are as follows:
- (1)
Divide the dataset into training set and test set, process the missing values, analyze the weight of the influence of the eigenvalues on the results, delete useless eigenvalues, and delete outliers;
- (2)
Use the Bayesian hyper-parameter optimization algorithm for the parameter optimization of the LightGBM model, and the HY_LightGBM model is constructed and trained;
- (3)
Use the HY_LightGBM model for prediction and output the prediction results.
- (4)
The specific experimental process is shown in
Figure 5.
2.4. Data Preprocessing
The dataset in this paper is the diabetes data from September to October 2017 in a grade-three first-class hospital, provided by the Tianchi competition platform as the data source, with a total of 7642 pieces of data and 42 eigenvalues, As shown in
Table 1. The eigenvalues include the following:
In this paper, the dataset was divided, with 6642 pieces of data as the training set, and the remaining 1000 pieces of data as the test set.
Firstly, the missing values in the original dataset were analyzed to obtain the proportion of the missing data. It can be seen from
Figure 6 that the missing proportion of five basic features—hepatitis B surface antigen, hepatitis B surface antibody, hepatitis B core antibody, hepatitis B e antigen, and hepatitis B e antibody—is over 70%, significantly exceeding the missing proportion of other basic features. Therefore, these features with large missing values were deleted, and those with smaller missing values were filled with medians.
Secondly, the weight of the influence of the data features was analyzed. The weight of the eigenvalues can be obtained through related functions. According to the eigenvalue weight of each item, the eigenvalues that have no effect on the results can be found. By processing the invalid eigenvalues, the accuracy of the experiment can be further improved. It can be seen from
Figure 7 that the date of medical examination and gender have no practical influence on the prediction results of the model. It can be known through common sense that ID also has no impact on the experimental results, so these features were deleted.
Again, to analyze the correlation coefficient of each eigenvalue, the darker the color, the stronger the correlation. From
Figure 8, we can see the correlation between each eigenvalue.
Finally, the basic features and the distribution of the blood glucose levels were learned, which can provide a more intuitive reference for feature engineering. The blood glucose values were used as the
Y-axis coordinates and the other basic features as the
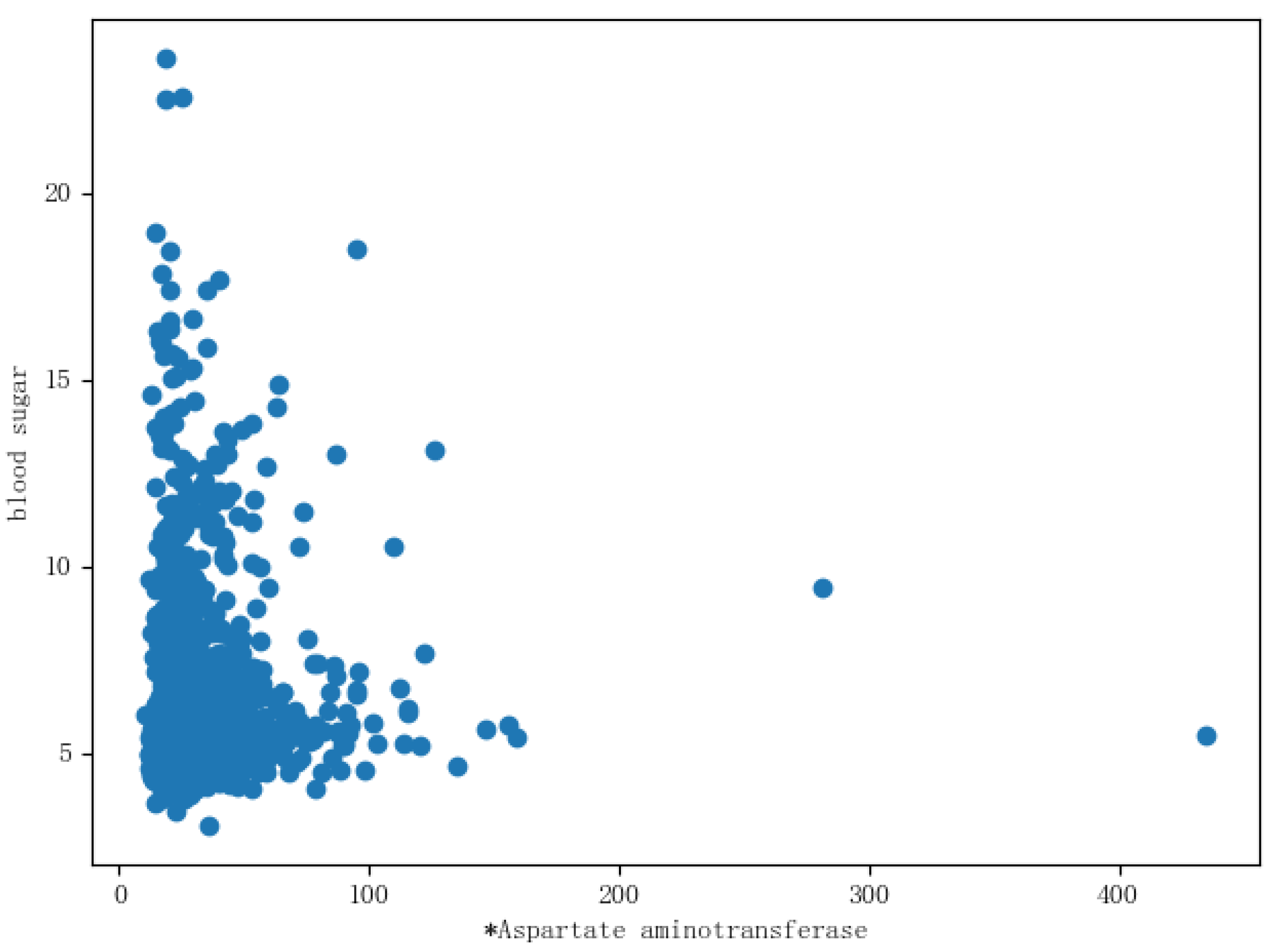
X-axis coordinates. Each basic feature and the distribution of the blood glucose value were listed for analysis, as shown in
Figure 9, where aspartate aminotransferase and the distribution of blood glucose are displayed. It can be seen from the figure that the distribution of the blood glucose value in aspartate aminotransferase included outliers, so the outliers were deleted. The outliers of the remaining features were also deleted.
2.5. Parameter Optimization Based on Bayesian Hyper-Parameter Optimization Algorithm
The optimal parameters of the LightGBM model were found by the Bayesian hyper-parameter optimization algorithm. Firstly, Hyperopt’s own function was used to define the parameter space, then the model and score acquirer were created, and finally, MSE was used as the evaluation indicator to obtain the optimal parameters of the LightGBM model. The optimal parameters of the LightGBM model obtained through the Bayesian hyper-parameter optimization algorithm are shown in
Table 2.
2.6. Blood Glucose Prediction by HY_LightGBM Model
Through the data preprocessing in
Section 2.4 and
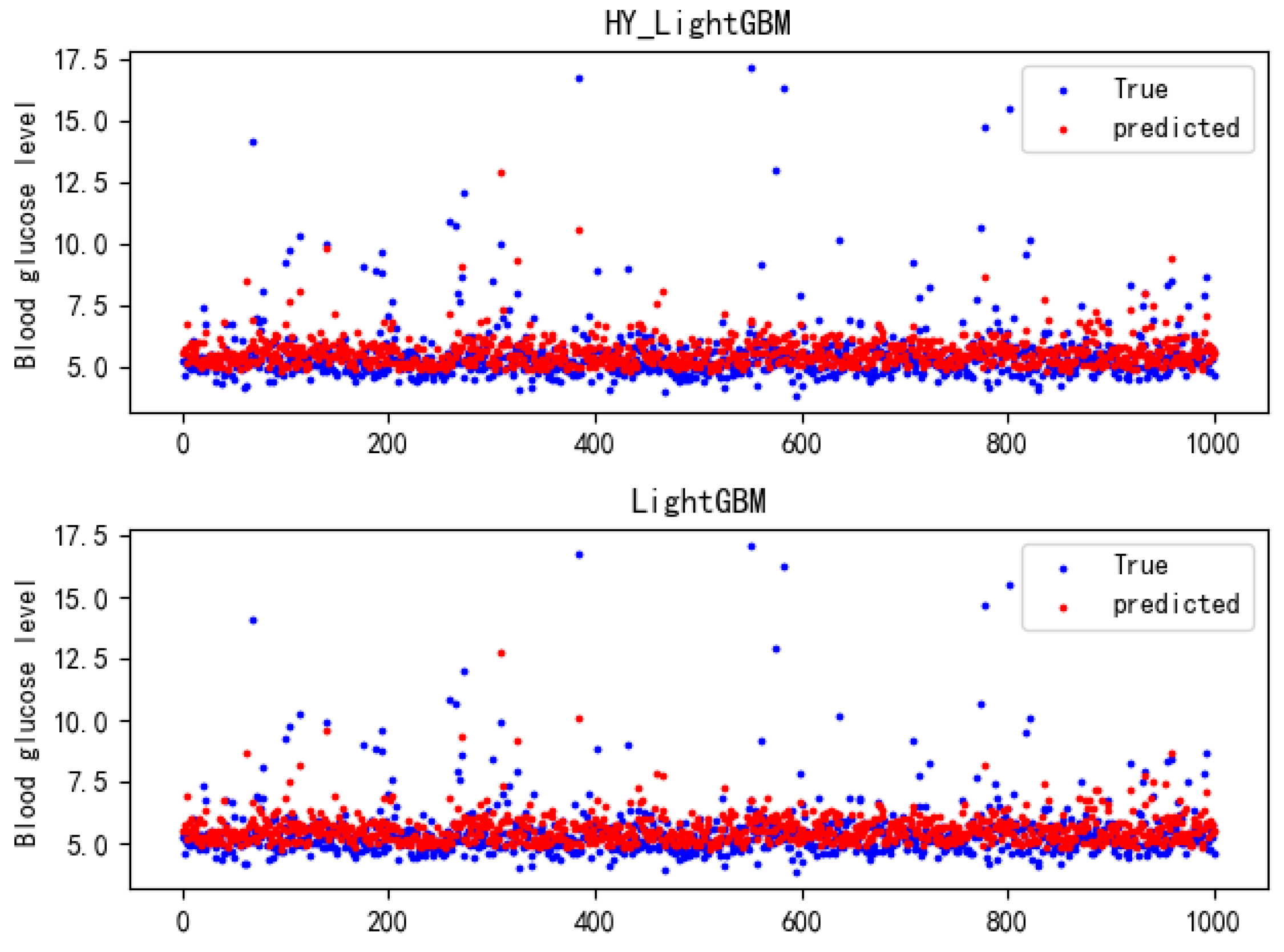
Section 2.5 and the Bayesian hyper-parameter optimization algorithm, the optimal parameters of the LightGBM model were determined and input into the LightGBM model for training and prediction. The blood glucose values output by it and the blood glucose values in the test set were evaluated by three evaluation indicators, namely: mean square error (MSE), root mean square error (RMSE), and determination coefficient R2 (R-Square).
2.7. Evaluation Indicators
The performance of the blood glucose prediction model was evaluated by the following three indicators: mean square error (MSE), root mean square error (RMSE), and determination coefficient R2 (R-Square). These are commonly used in regression tasks, and the smaller the mean square error (MSE) and the root mean square error (RMSE) value, the more accurate the prediction results.
4. Conclusions
This paper proposes an improved LightGBM prediction model based on the Bayesian hyper-parameter optimization algorithm, namely the HY_LightGBM model, where Bayesian hyper-parameter optimization was employed to find the optimal parameter combination for the model, which improved the prediction accuracy of the LightGBM model. The experiments proved that the method proposed in this paper achieves a higher prediction accuracy than the XGBoost model and CatBoost model, and has higher efficiency than the genetic algorithm and random searching algorithm.
The previous measurement of the blood glucose value needs to measure each person’s blood glucose value one by one, which requires a lot of manpower and material resources. After training, the model will no longer need the characteristic value of the blood glucose value. It can directly predict the blood glucose value of the physical examination personnel through the HY_LightGBM model and other physical examination indicators of the physical examination personnel.
The HY_LightGBM model, with a strong generalization ability, can also be applied to other types of auxiliary diagnosis and treatment, but the overall performance can be further improved. The next work will focus on the further optimization of the model using the idea of model fusion to improve the prediction accuracy of the model.