Microbial Populations of Fresh and Cold Stored Donkey Milk by High-Throughput Sequencing Provide Indication for A Correct Management of This High-Value Product
Abstract
1. Introduction
2. Materials and Methods
2.1. Milk Sampling
2.2. Microbiological Analysis by Culture-Dependent Methods
2.3. DNA Extraction
2.4. Amplicon Library Preparation and Pyrosequencing
2.5. Bioinformatics and Statistical Analysis
3. Results and Discussion
3.1. Sequences Analysis
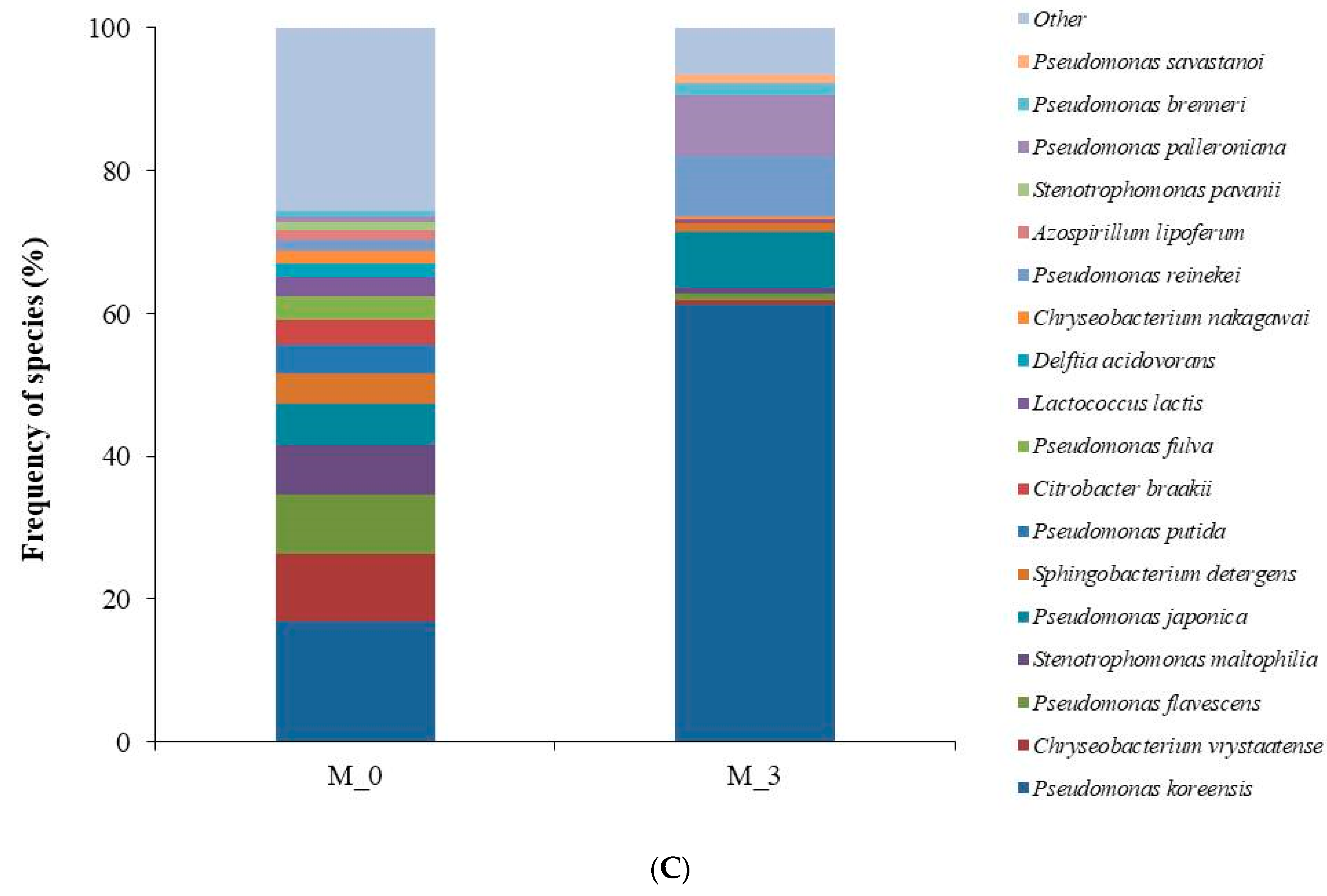
3.2. Analysis of Bacterial Diversity
Author Contributions
Funding
Conflicts of Interest
References
- Aspri, M.; Economou, N.; Papademas, P. Donkey milk: An overview on functionality, technology and future prospects. Food Rev. Int. 2016, 33, 316–333. [Google Scholar] [CrossRef]
- Camillo, F.; Rota, A.; Biagini, L.; Tesi, M.; Fanelli, D.; Panzani, D. The current situation and trend of donkey industry in Europe. J. Equine Vet. Sci. 2018, 65, 44–49. [Google Scholar] [CrossRef]
- Miraglia, N.; Salimei, E.; Fantuz, F. Equine milk production and valorization of marginal areas—A review. Animals 2020, 10, 353. [Google Scholar] [CrossRef] [PubMed]
- Giacometti, F.; Bardasi, L.; Merialdi, G.; Morbarigazzi, M.; Federici, S.; Piva, S.; Serraino, A. Shelf life of donkey milk subjected to different treatment and storage conditions. J. Dairy Sci. 2016, 99, 4291–4299. [Google Scholar] [CrossRef] [PubMed]
- Turchi, B.; Pedonese, F.; Torracca, B.; Fratini, F.; Mancini, S.; Galiero, A.; Montalbano, B.; Cerri, D.; Nuvoloni, R. Lactobacillus plantarum and Streptococcus thermophilus as starter cultures for a donkey milk fermented beverage. Int. J. Food Microbiol. 2017, 256, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Faccia, M.; Gambacorta, G.; Martemucci, G.; Natrella, G.; D’Alessandro, A.G. Technological attempts at producing cheese from donkey milk. J. Dairy Res. 2018, 85, 327–330. [Google Scholar] [CrossRef] [PubMed]
- Altomonte, I.; Salaria, F.; Licitra, R.; Martini, M. Donkey and human milk: Insights into their compositional similarities. Int. Dairy J. 2019, 89, 111–118. [Google Scholar] [CrossRef]
- Souroullas, K.; Aspri, M.; Papademas, P. Donkey milk as a supplement in infant formula: Benefits and technological challenges. Food Res. Int. 2018, 109, 416–425. [Google Scholar] [CrossRef] [PubMed]
- Conte, F.; Panebianco, A. Potential hazards associated with raw donkey milk consumption: A review. Int. J. Food Sci. 2019, 2019, 5782974. [Google Scholar] [CrossRef]
- Quigley, L.; O’Sullivan, O.; Stanton, C.; Beresford, T.P.; Ross, R.P.; Fitzgerald, G.F.; Cotter, P.D. The complex microbiota of raw milk. FEMS Microbiol. Rev. 2013, 37, 664–698. [Google Scholar] [CrossRef]
- Quigley, L.; O’Sullivan, O.; Beresford, T.P.; Ross, R.P.; Fitzgerald, G.F.; Cotter, P.D. Molecular approaches to analysing the microbial composition of raw milk and raw milk cheese. Int. J. Food Microbiol. 2011, 150, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, Z.; Lei, F.; Wang, B.; Jiang, S.; Peng, Q.; Zhang, J.; Shao, Y. Bacterial diversity in goat milk from the Guanzhong area of China. J. Dairy Sci. 2017, 100, 7812–7824. [Google Scholar] [CrossRef] [PubMed]
- Alexandraki, V.; Kazou, M.; Angelopoulou, A.; Arena, M.P.; Capozzi, V.; Russo, P.; Fiocco, D.; Spano, G.; Papadimitriou, K.; Tsakalidou, E. The Microbiota of Non-cow Milk and Products. In Non-Bovine Milk and Milk Products; Tsakalidou, E., Papadimitriou, K., Eds.; Academic Press: London, UK, 2016; pp. 117–159. [Google Scholar]
- Cavallarin, L.; Giribaldi, M.; Soto-Del Rio, M.; de los Valle, D.E.; Barbarino, G.; Gennero, M.S.; Civera, T. A survey on the milk chemical and microbiological quality in dairy donkey farms located in NorthWestern Italy. Food Control 2015, 50, 230–235. [Google Scholar] [CrossRef]
- Malissiova, E.; Arsenos, G.; Papademas, P.; Fletouris, D.; Manouras, A.; Aspri, M.; Nikolopoulou, A.; Giannopoulou, A.; Arvanitoyannis, J.S. Assessment of donkey milk chemical, microbiological and sensory attributes in Greece and Cyprus. Int. J. Dairy Technol. 2016, 69, 143–146. [Google Scholar] [CrossRef]
- Soto del Rio, M.D.L.D.; Dalmasso, A.; Civera, T.; Bottero, M.T. Characterization of bacterial communities of donkey milk by high-throughput sequencing. Int. J. Food Microbiol. 2017, 251, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Machado, S.G.; Baglinière, F.; Marchand, S.; Van Coillie, E.; Vanetti, M.C.; De Block, J.; Heyndrickx, M. The Biodiversity of the Microbiota Producing Heat-Resistant Enzymes Responsible for Spoilage in Processed Bovine Milk and Dairy Products. Front. Microbiol. 2017, 8, 302. [Google Scholar] [CrossRef]
- Yuan, L.; Sadiq, F.A.; Burmølle, M.; Wang, N.I.; He, G. Insights into psychrotrophic bacteria in raw milk: A review. J. Food Prot. 2019, 82, 1148–1159. [Google Scholar] [CrossRef]
- Raats, D.; Offek, M.; Minz, D.; Halpern, M. Molecular analysis of bacterial communities in raw cow milk and the impact of refrigeration on its structure and dynamics. Food Microbiol. 2011, 28, 465–471. [Google Scholar] [CrossRef]
- Porcellato, D.; Aspholm, M.; Skeie, S.B.; Monshaugen, M.; Brendehaug, J.; Mellegård, H. Microbial diversity of consumption milk during processing and storage. Int. J. Food Microbiol. 2018, 266, 21–30. [Google Scholar] [CrossRef]
- Fernández de Palencia, P.; López, P.; Corbí, A.L.; Peláez, C.; Requena, T. Probiotic strains: Survival under simulated gastrointestinal conditions, in vitro adhesion to Caco-2 cells and effect on cytokine secretion. Eur. Food Res. Technol. 2008, 227, 1475–1484. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of a general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2012, 41, e1. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Marcel, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Huang, Y.; Niu, B.; Gao, Y.; Fu, L.; Li, W. CD-HIT Suite: A web server for clustering and comparing biological sequences. Bioinformatics 2010, 26, 680–682. [Google Scholar] [CrossRef]
- Hantsis-Zacharov, E.; Shakéd, T.; Senderovich, Y.; Halpern, M. Chryseobacterium oranimense sp. nov., a psychrotolerant, proteolytic and lipolytic bacterium isolated from raw cow’s milk. Int. J. Syst. Evol. Microbiol. 2008, 58, 2635–2639. [Google Scholar] [CrossRef]
- Yuan, L.; Sadiq, F.A.; Liu, T.J.; Li, Y.; Gu, J.S.; Yang, H.Y.; He, G.Q. Spoilage potential of psychrotrophic bacteria isolated from raw milk and the thermo-stability of their enzymes. J. Zhejiang Univ. Sci. B 2018, 19, 630–642. [Google Scholar] [CrossRef]
- Masiello, S.N.; Martin, N.H.; Trmčić, A.; Wiedmann, M.; Boor, K.J. Identification and characterization of psychrotolerant coliform bacteria isolated from pasteurized fluid milk. J. Dairy Sci. 2016, 99, 130–140. [Google Scholar] [CrossRef]
- Schmidt, V.S.; Kaufmann, V.; Kulozik, U.; Scherer, S.; Wenning, M. Microbial biodiversity, quality and shelf life of microfiltered and pasteurized extended shelf life (ESL) milk from Germany, Austria and Switzerland. Int. J. Food Microbiol. 2012, 154, 1–9. [Google Scholar] [CrossRef]
- Odeyemi, O.A.; Alegbeleye, O.O.; Strateva, M.; Stratev, D. Understanding spoilage microbial community and spoilage mechanisms in foods of animal origin. Compr. Rev. Food Sci. Food Saf. 2019. [Google Scholar] [CrossRef]
- Zhang, R.; Huo, W.; Zhu, W.; Mao, S. Characterization of bacterial community of raw milk from dairy cows during subacute ruminal acidosis challenge by high-throughput sequencing. J. Sci. Food Agric. 2015, 95, 1072–1079. [Google Scholar] [CrossRef] [PubMed]
- Schukken, Y.; Chuff, M.; Moroni, P.; Gurjar, A.; Santisteban, C.; Welcome, F.; Zadoks, R. The “other” Gram-negative bacteria in mastitis Klebsiella, Serriata, and more. Vet. Clin. N. Am. Food A 2012, 28, 239–256. [Google Scholar] [CrossRef] [PubMed]
- Fernández, M.; Porcel, M.; de la Torre, J.; Molina-Henares, M.A.; Daddaoua, A.; Llamas, M.A.; Roca, A.; Carriel, V.; Garzón, I.; Ramos, J.L.; et al. Analysis of the pathogenic potential of nosocomial Pseudomonas putida strains. Front. Microbiol. 2015, 6, 871. [Google Scholar] [CrossRef] [PubMed]
- Brooke, J.S. Stenotrophomonas maltophilia: An emerging global opportunistic pathogen. Clin. Microbiol. Rev. 2012, 25, 2–41. [Google Scholar] [CrossRef] [PubMed]
- Hirai, J.; Uechi, K.; Hagihara, M.; Sakanashi, D.; Kinjo, T.; Haranaga, S.; Fujita, J. Bacteremia due to Citrobacter braakii: A case report and literature review. J. Infect. Chemother. 2016, 22, 819–821. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, V.S.; Wenning, M.; Scherer, S. Sphingobacterium lactis sp. nov. and Sphingobacterium alimentarium sp. nov., isolated from raw milk and a dairy environment. Int. J. Syst. Evol. Microbiol. 2012, 62, 1506–1511. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Sadiq, F.A.; Liu, T.; Flint, S.; Chen, J.; Yang, H.; Gu, J.; Zhang, G.; He, G. Psychrotrophic bacterial populations in Chinese raw dairy milk. LWT Food Sci. Technol. 2017, 84, 409–418. [Google Scholar] [CrossRef]
- Kaur, M.; Singh, H.; Sharma, S.; Mishra, S.; Tanuku, N.R.S.; Pinnaka, A.K. Sphingobacterium bovisgrunnientis sp. nov., isolated from yak milk. Int. J. Syst. Evol. Microbiol. 2018, 68, 636–642. [Google Scholar] [CrossRef]
- Liu, W.; Xi, X.; Sudu, Q.; Kwok, L.; Guo, Z.; Hou, Q.; Menhe, B.; Sun, T.; Zhang, H. High-throughput sequencing reveals microbial community diversity of Tibetan naturally fermented yak milk. Ann. Microbiol. 2015, 65, 1741–1751. [Google Scholar] [CrossRef]
- Anandham, R.; Heo, J.; Krishnamoorthy, R.; SenthilKumar, M.; Gopal, N.O.; Kim, S.J.; Kwon, S.W. Azospirillum ramasamyi sp. nov., a novel diazotrophic bacterium isolated from fermented bovine products. Int. J. Syst. Evol. Microbiol. 2019, 69, 1369–1375. [Google Scholar] [CrossRef]
- McInnis, E.A.; Kalanetra, K.M.; Mills, D.A.; Maga, E.A. Analysis of raw goat milk microbiota: Impact of stage of lactation and lysozyme on microbial diversity. Food Microbiol. 2015, 46, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Quigley, L.; McCarthy, R.; O’Sullivan, O.; Beresford, T.P.; Fitzgerald, G.F.; Ross, R.P.; Stanton, C.; Cotter, P.D. The microbial content of raw and pasteurized cow milk as determined by molecular approaches. J. Dairy Sci. 2013, 96, 4928–4937. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Y.; Zhao, L.; Jiang, L.; Dong, M.L.; Ren, F.Z. The antimicrobial activity of donkey milk and its microflora changes during storage. Food Control 2008, 19, 1191–1195. [Google Scholar] [CrossRef]
- von Neubeck, M.; Baur, C.; Krewinkel, M.; Stoeckel, M.; Kranz, B.; Stressler, T.; Fischer, L.; Hinrichs, J.; Scherer, S.; Wenning, M. Biodiversity of refrigerated raw milk microbiota and their enzymatic spoilage potential. Int. J. Food Microbiol. 2015, 211, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Xin, L.; Meng, Z.; Zhang, L.; Cui, Y.; Han, X.; Yi, H. The diversity and proteolytic properties of psychrotrophic bacteria in raw cows’ milk from North China. Int. Dairy J. 2017, 66, 34–41. [Google Scholar] [CrossRef]
- Zhang, D.; Palmer, J.; Teh, K.H.; Biggs, P.; Flint, S. 16S rDNA high-throughput sequencing and MALDI-TOF MS are complementary when studying psychrotrophic bacterial diversity of raw cows’ milk. Int. Dairy J. 2019, 97, 86–91. [Google Scholar] [CrossRef]
- Rasolofo, E.A.; St-Gelais, D.; LaPointe, G.; Roy, D. Molecular analysis of bacterial population structure and dynamics during cold storage of untreated and treated milk. Int. J. Food Microbiol. 2010, 138, 108–118. [Google Scholar] [CrossRef]
- Alothman, M.; Lusk, K.A.; Silcock, P.; Bremer, P.J. Comparing PTR-MS profile of milk inoculated with pure or mixed cultures of spoilage bacteria. Food Microbiol. 2017, 64, 155–163. [Google Scholar] [CrossRef]
- Uniacke-Lowe, T.; Fox, P.F. Equid milk. In Encyclopedia of Dairy Sciences, 2nd ed.; Fuquay, J.W., Fox, P.F., McSweeney, P.L.H., Eds.; Academic Press: San Diego, CA, USA, 2011; pp. 518–529. [Google Scholar]
- Iannella, G. Donkey cheese made through pure camel chymosin. Afr. J. Food Sci. 2015, 9, 421–425. [Google Scholar]
- Saric, L.C.; Saric, B.M.; Mandic, A.I.; Hadnadev, M.S.; Gubic, J.M.; Milovanovic, I.L.; Tomic, J.M. Characterization of extra-hard cheese produced from donkeys’ and caprine milk mixture. Dairy Sci. Technol. 2016, 96, 227–241. [Google Scholar] [CrossRef][Green Version]
- EFSA. Panel on Biological Hazards. Scientific Opinion on the public health risks related to the consumption of raw drinking milk. EFSA J. 2015, 13, 3940. [Google Scholar] [CrossRef]
- Giribaldi, M.; Antoniazzi, S.; Gariglio, G.M.; Coscia, A.; Bertino, E.; Cavallarin, L. A preliminary assessment of HTST processing on donkey milk. Vet. Sci. 2017, 4, 50. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, A.G.; Martemucci, G.; Loizzo, P.; Faccia, M. Production of cheese from donkey milk as influenced by addition of transglutaminase. J. Dairy Sci. 2019, 102, 10867–10876. [Google Scholar] [CrossRef] [PubMed]
- Capozzi, V.; Fragasso, M.; Romaniello, R.; Berbegal, C.; Russo, P.; Spano, G. Spontaneous food fermentations and potential risks for human health. Fermentation 2017, 3, 49. [Google Scholar] [CrossRef]
- Capozzi, V.; Fragasso, M.; Russo, P. Microbiological safety and the management of microbial resources in artisanal foods and beverages: The need for a transdisciplinary assessment to conciliate actual trends and risks avoidance. Microorganisms 2020, 88, 306. [Google Scholar] [CrossRef] [PubMed]
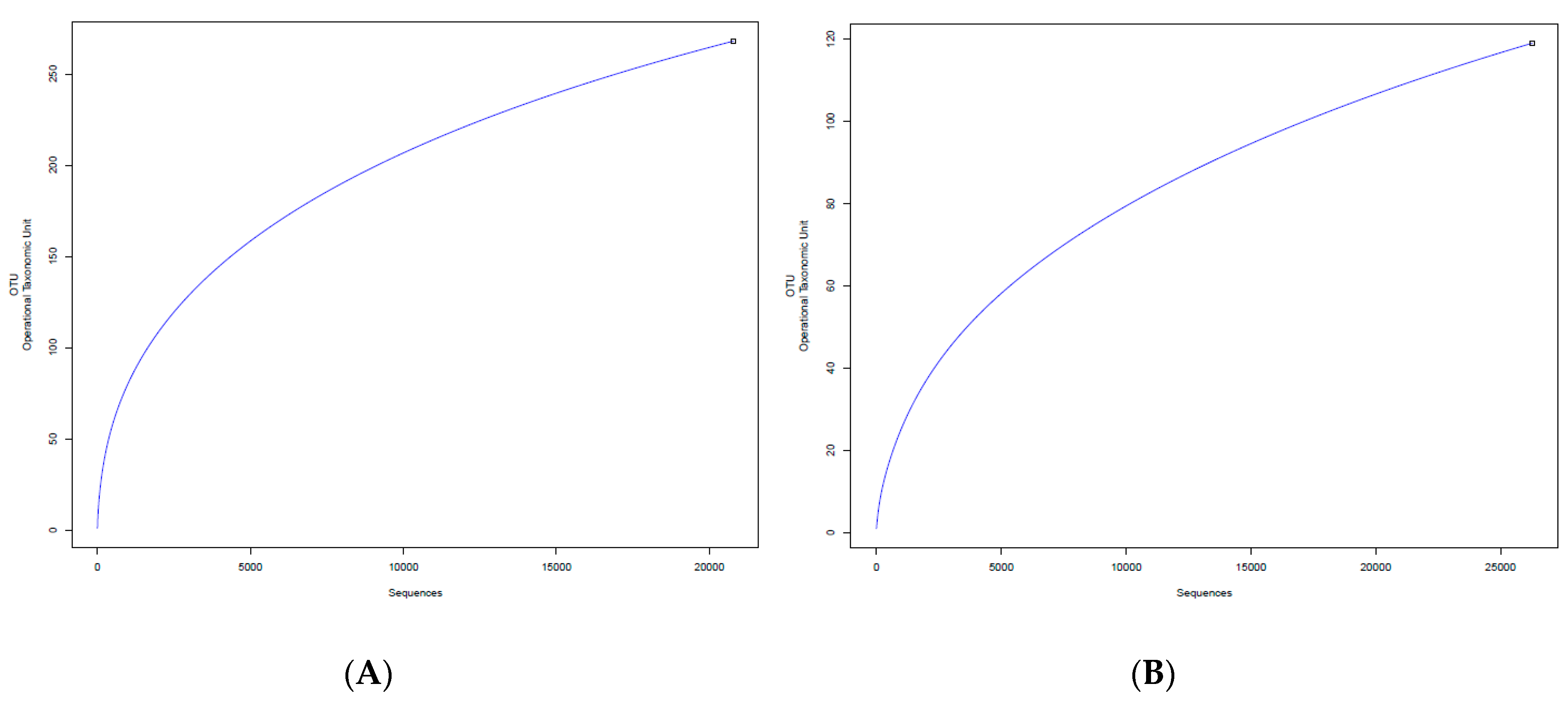
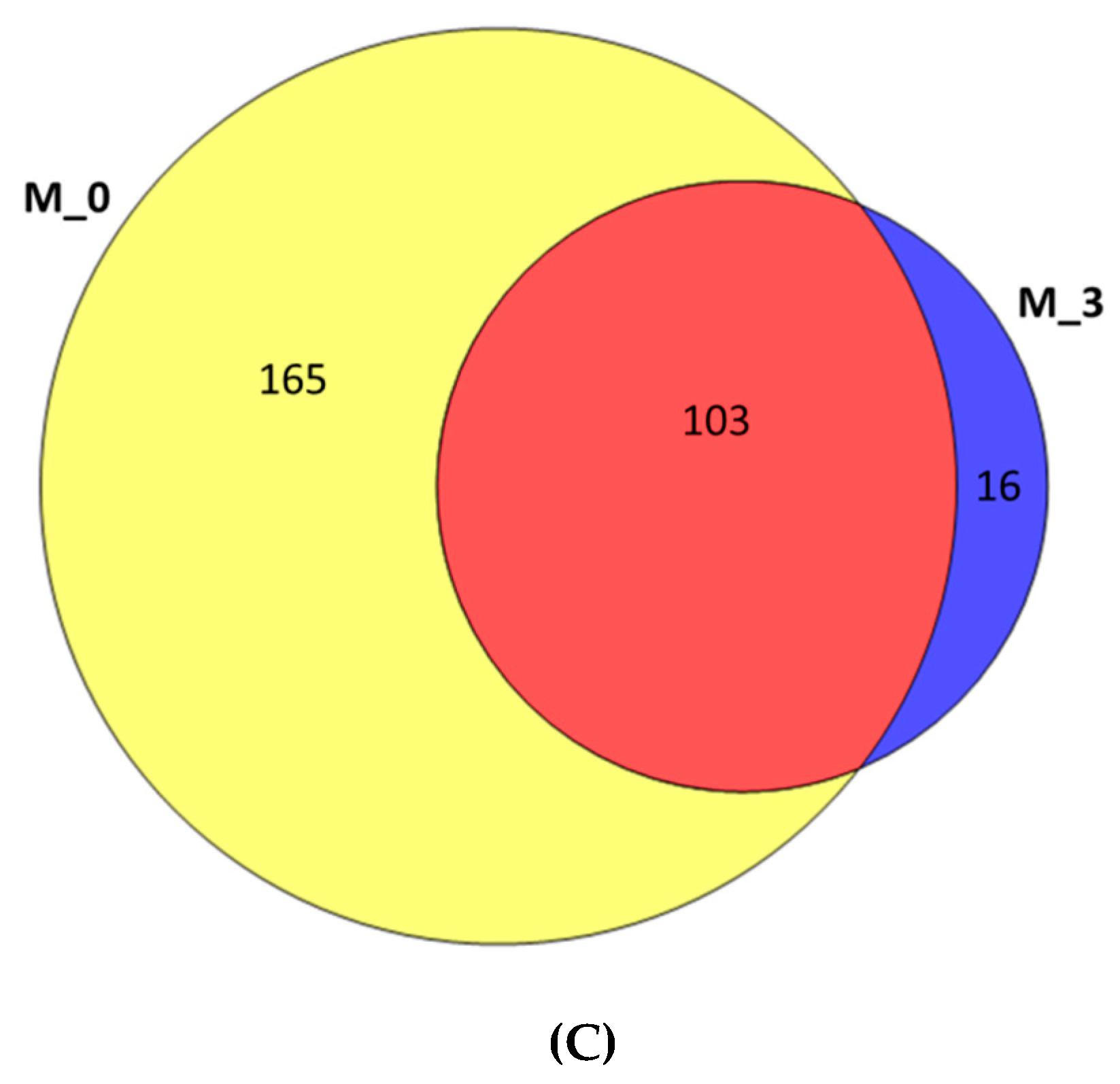
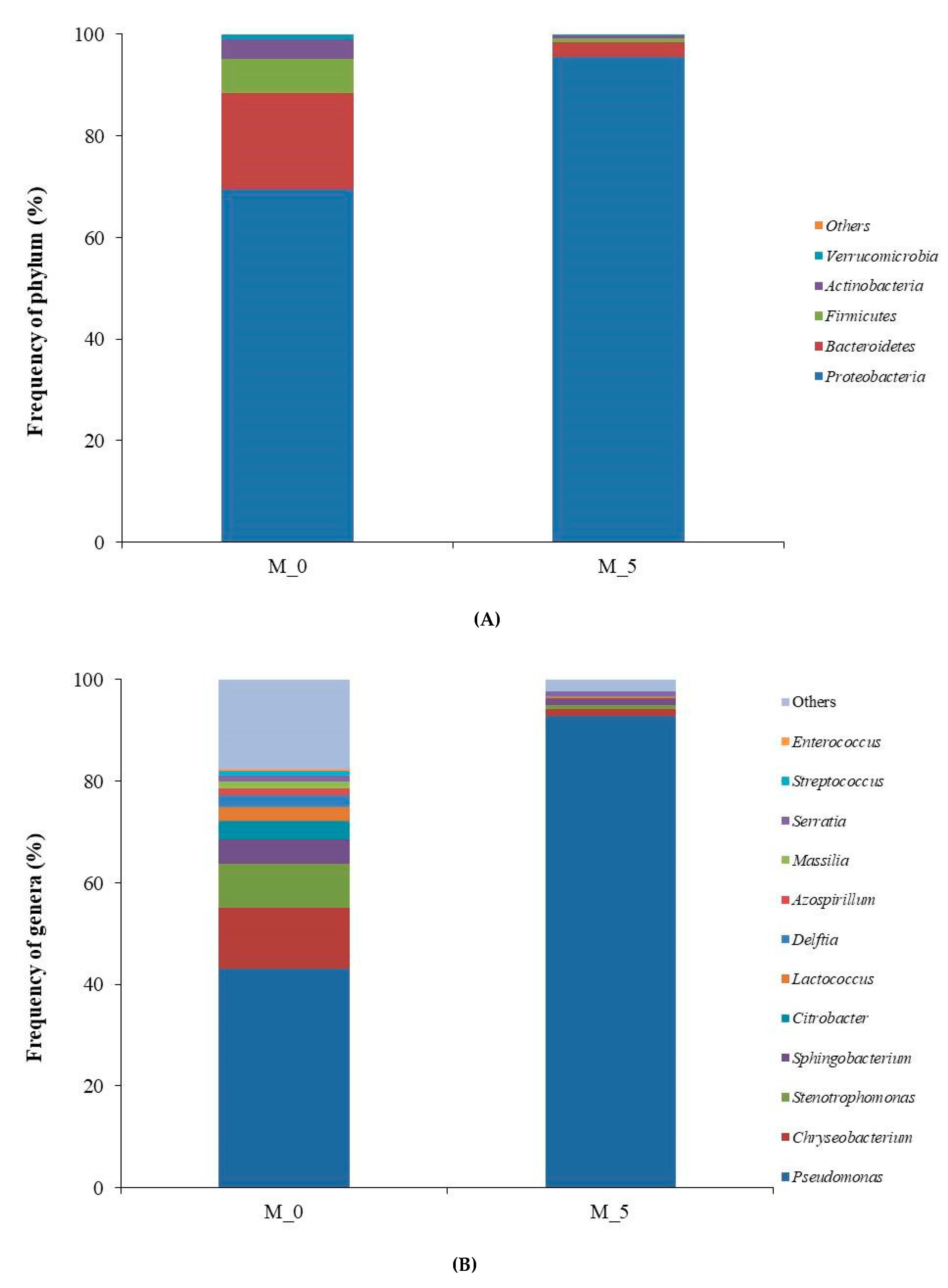
| Sample | Sequences | Av. Length | Total Mb | Av. Quality | OTUs | Shannon Value |
|---|---|---|---|---|---|---|
| Mt0–16S | 22,483 | 443.12 | 9.96 | 35.69 | 268 | 2.580 |
| Mt3–16S | 27,141 | 446.46 | 12.12 | 36.07 | 119 | 0.478 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Russo, P.; Fiocco, D.; Albenzio, M.; Spano, G.; Capozzi, V. Microbial Populations of Fresh and Cold Stored Donkey Milk by High-Throughput Sequencing Provide Indication for A Correct Management of This High-Value Product. Appl. Sci. 2020, 10, 2314. https://doi.org/10.3390/app10072314
Russo P, Fiocco D, Albenzio M, Spano G, Capozzi V. Microbial Populations of Fresh and Cold Stored Donkey Milk by High-Throughput Sequencing Provide Indication for A Correct Management of This High-Value Product. Applied Sciences. 2020; 10(7):2314. https://doi.org/10.3390/app10072314
Chicago/Turabian StyleRusso, Pasquale, Daniela Fiocco, Marzia Albenzio, Giuseppe Spano, and Vittorio Capozzi. 2020. "Microbial Populations of Fresh and Cold Stored Donkey Milk by High-Throughput Sequencing Provide Indication for A Correct Management of This High-Value Product" Applied Sciences 10, no. 7: 2314. https://doi.org/10.3390/app10072314
APA StyleRusso, P., Fiocco, D., Albenzio, M., Spano, G., & Capozzi, V. (2020). Microbial Populations of Fresh and Cold Stored Donkey Milk by High-Throughput Sequencing Provide Indication for A Correct Management of This High-Value Product. Applied Sciences, 10(7), 2314. https://doi.org/10.3390/app10072314