A Multiplex PCR-Based Next Generation Sequencing-Panel to Identify Mutations for Targeted Therapy in Breast Cancer Circulating Tumor Cells
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Enrichment and Enumeration of CTCs
2.3. CTC Isolation
2.4. Whole-Genome Amplification
2.5. Multiplex PCR-Based Next Generation Sequencing
2.6. Validation Experiments with Spiked Cells
2.7. Statistical Analysis
3. Results
3.1. Development of the Assay
3.2. Validation of the Assay
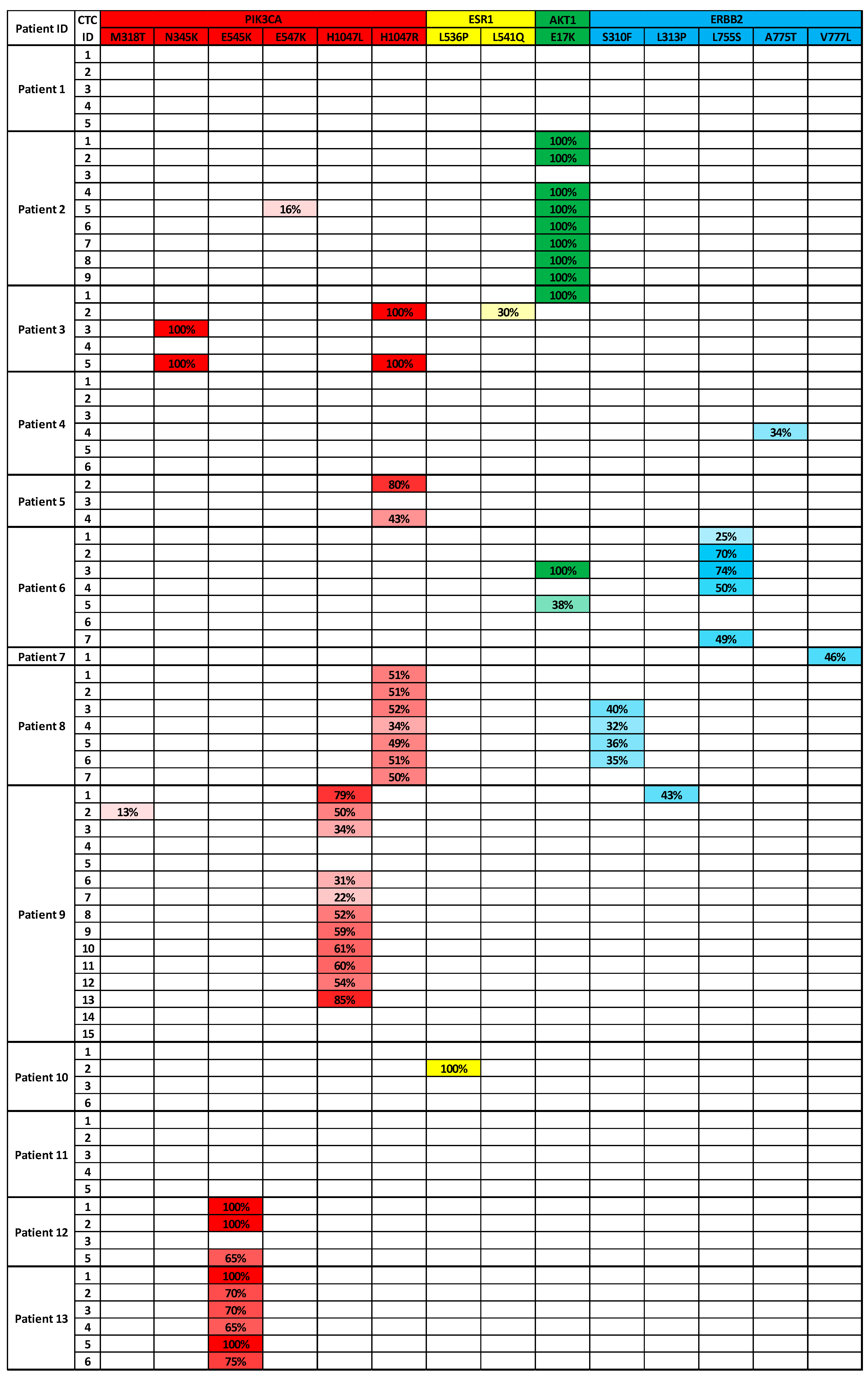
3.3. Analysis of CTCs from Metastatic Breast Cancer Patients to Identify Targeted Therapies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Perakis, S.; Speicher, M.R. Emerging concepts in liquid biopsies. BMC Med. 2017, 15, 75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turajlic, S.; Sottoriva, A.; Graham, T.; Swanton, C. Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 2019, 20, 404–416. [Google Scholar] [CrossRef] [PubMed]
- Palmirotta, R.; Lovero, D.; Cafforio, P.; Felici, C.; Mannavola, F.; Pellè, E.; Quaresmini, D.; Tucci, M. Liquid biopsy of cancer: A multimodal diagnostic tool in clinical oncology. Ther. Adv. Med. Oncol. 2018, 10, 1758835918794630. [Google Scholar] [CrossRef] [PubMed]
- Pantel, K.; Alix-Panabières, C. Circulating tumour cells in cancer patients: Challenges and perspectives. Trends Mol. Med. 2010, 16, 398–406. [Google Scholar] [CrossRef]
- Barradas, A.M.C.; Terstappen, L.W.M.M. Towards the biological understanding of CTC: Capture technologies, definitions and potential to create metastasis. Cancers 2013, 5, 1619–1642. [Google Scholar] [CrossRef]
- Yu, M.; Stott, S.; Toner, M.; Maheswaran, S.; Haber, D.A. Circulating tumor cells: Approaches to isolation and characterization. J. Cell Biol. 2011, 192, 373–382. [Google Scholar] [CrossRef]
- Allard, W.J.; Matera, J.; Miller, M.C.; Repollet, M.; Connelly, M.C.; Rao, C.; Tibbe, A.G.J.; Uhr, J.W.; Terstappen, L.W.M.M. Tumor Cells Circulate in the Peripheral Blood of All Major Carcinomas but not in Healthy Subjects or Patients with Nonmalignant Diseases. Clin. Cancer Res. 2005, 10, 6897–6904. [Google Scholar] [CrossRef] [Green Version]
- Wan, J.C.M.; Massie, C.; Garcia-Corbacho, J.; Mouliere, F.; Brenton, J.D.; Caldas, C.; Pacey, S.; Baird, R.; Rosenfeld, N. Liquid biopsies come of age: Towards implementation of circulating tumour DNA. Nat. Rev. Cancer 2017, 17, 223–238. [Google Scholar] [CrossRef]
- FDA Oncology Update. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6684051/ (accessed on 1 March 2020).
- Janku, F.; Yap, T.A.; Meric-Bernstam, F. Targeting the PI3K pathway in cancer: Are we making headway? Nat. Rev. Clin. Oncol. 2018, 15, 273–291. [Google Scholar] [CrossRef]
- Zhao, L.; Vogt, P.K. Helical domain and kinase domain mutations in p110α of phosphatidylinositol 3-kinase induce gain of function by different mechanisms. Proc. Natl. Acad. Sci. USA 2008, 105, 2652–2657. [Google Scholar] [CrossRef] [Green Version]
- Arpino, G.; Wiechmann, L.; Osborne, C.K.; Schiff, R. Crosstalk between the Estrogen Receptor and the HER Tyrosine Kinase Receptor Family: Molecular Mechanism and Clinical Implications for Endocrine Therapy Resistance. Endocr. Rev. 2008, 29, 217–233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toy, W.; Shen, Y.; Won, H.; Green, B.; Sakr, R.a.; Will, M.; Gala, K.; Fanning, S.; King, T.a.; Hudis, C.; et al. ESR1 ligand binding domain mutations in hormone-resistant breast cancer. Nat. Genet. 2013, 45, 1439–1445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franken, A.; Honisch, E.; Reinhardt, F.; Meier-Stiegen, F.; Yang, L.; Jaschinski, S.; Esposito, I.; Alberter, B.; Polzer, B.; Huebner, H.; et al. Detection of ESR1 Mutations in Single Circulating Tumor Cells on Estrogen Deprivation Therapy but Not in Primary Tumors from Metastatic Luminal Breast Cancer Patients. J. Mol. Diagn. 2019, 22, 111–121. [Google Scholar] [CrossRef]
- Carpten, J.D.; Faber, A.L.; Horn, C.; Donoho, G.P.; Briggs, S.L.; Robbins, C.M.; Hostetter, G.; Boguslawski, S.; Moses, T.Y.; Savage, S.; et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 2007, 448, 439–444. [Google Scholar] [CrossRef] [PubMed]
- Bose, R.; Kavuri, S.M.; Searleman, A.C.; Shen, W.; Shen, D.; Koboldt, D.C.; Monsey, J.; Goel, N.; Aronson, A.B.; Li, S.; et al. Activating HER2 Mutations in HER2 Gene Amplification Negative Breast Cancer. Cancer Discov. 2013, 3, 224–237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neumann, M.H.D.; Schneck, H.; Decker, Y.; Schömer, S.; Franken, A.; Endris, V.; Pfarr, N.; Weichert, W.; Niederacher, D.; Fehm, T.; et al. Isolation and characterization of circulating tumor cells using a novel workflow combining the CellSearch® system and the CellCelectorTM. Biotechnol. Prog. 2017, 33, 125–132. [Google Scholar] [CrossRef]
- Afgan, E.; Baker, D.; van den Beek, M.; Blankenberg, D.; Bouvier, D.; Čech, M.; Chilton, J.; Clements, D.; Coraor, N.; Eberhard, C.; et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 2016, 44, W3–W10. [Google Scholar] [CrossRef] [Green Version]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef] [Green Version]
- Franken, A.; Driemel, C.; Behrens, B.; Meier-stiegen, F.; Endris, V.; Stenzinger, A.; Niederacher, D.; Fischer, J.C.; Stoecklein, N.H.; Ruckhaeberle, E.; et al. Label-Free Enrichment and Molecular Characterization of Viable Circulating Tumor Cells from Diagnostic Leukapheresis Products. Clin. Chem. 2019, 65, 549–558. [Google Scholar] [CrossRef] [Green Version]
- Lampignano, R.; Yang, L.; Neumann, M.H.D.; Franken, A.; Fehm, T.; Niederacher, D.; Neubauer, H. A novel workflow to enrich and isolate patient-matched EpCAMhighand EpCAMlow/negativeCTCs enables the comparative characterization of the PIK3CA status in metastatic breast cancer. Int. J. Mol. Sci. 2017, 18, 1885. [Google Scholar] [CrossRef] [Green Version]
- Paolillo, C.; Mu, Z.; Rossi, G.; Schiewer, M.J.; Nguyen, T.; Austin, L.; Capoluongo, E.; Knudsen, K.; Cristofanilli, M.; Fortina, P. Detection of activating estrogen receptor gene (ESR1) mutations in single circulating tumor cells. Clin. Cancer Res. 2017, 23, 6086–6093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shaw, J.A.; Guttery, D.S.; Hills, A.; Fernandez-Garcia, D.; Page, K.; Rosales, B.M.; Goddard, K.S.; Hastings, R.K.; Luo, J.; Ogle, O.; et al. Mutation analysis of cell-free DNA and single circulating tumor cells in metastatic breast cancer patients with high circulating tumor cell counts. Clin. Cancer Res. 2017, 23, 88–96. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Onidani, K.; Shoji, H.; Kakizaki, T.; Yoshimoto, S.; Okaya, S.; Miura, N.; Sekikawa, S.; Furuta, K.; Lim, C.T.; Shibahara, T.; et al. Monitoring of cancer patients via next-generation sequencing of patient-derived circulating tumor cells and tumor DNA. Cancer Sci. 2019, 110, 2590–2599. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- André, F.; Ciruelos, E.; Rubovszky, G.; Campone, M.; Loibl, S.; Rugo, H.S.; Iwata, H.; Conte, P.; Mayer, I.A.; Kaufman, B.; et al. Alpelisib for PIK3CA-Mutated, Hormone Receptor–Positive Advanced Breast Cancer. N. Engl. J. Med. 2019, 380, 1929–1940. [Google Scholar] [CrossRef]
- Chandarlapaty, S.; Sakr, R.A.; Giri, D.; Patil, S.; Heguy, A.; Morrow, M.; Modi, S.; Norton, L.; Rosen, N.; Hudis, C.; et al. Frequent Mutational Activation of the PI3K-AKT Pathway in Trastuzumab-Resistant Breast Cancer. Clin. Cancer Res. 2012, 18, 6784–6791. [Google Scholar] [CrossRef] [Green Version]
- Fribbens, C.; O’Leary, B.; Kilburn, L.; Hrebien, S.; Garcia-Murillas, I.; Beaney, M.; Cristofanilli, M.; Andre, F.; Loi, S.; Loibl, S.; et al. Plasma ESR1 Mutations and the Treatment of Estrogen Receptor-Positive Advanced Breast Cancer. J. Clin. Oncol. 2016, 34, 2961–2968. [Google Scholar] [CrossRef]
- Hyman, D.M.; Smyth, L.M.; Donoghue, M.T.A.; Westin, S.N.; Bedard, P.L.; Emma, J.; Bando, H.; El-khoueiry, A.B.; Mita, A.; Schellens, J.H.M.; et al. AKT Inhibition in Solid Tumors with AKT1 Mutations. J. Clin. Oncol. 2019, 35, 2251–2262. [Google Scholar] [CrossRef]
- Zardavas, D.; Fumagalli, D.; Loi, S. Phosphatidylinositol 3-kinase/AKT/mammalian target of rapamycin pathway inhibition: A breakthrough in the management of luminal (ER+/HER2−) breast cancers? Curr. Opin. Oncol. 2012, 24, 623–634. [Google Scholar] [CrossRef]
- Troxell, M.L. PIK3CA/AKT1 Mutations in Breast Carcinoma: A Comprehensive Review of Experimental and Clinical Studies. J. Clin. Exp. Pathol. 2012. [Google Scholar] [CrossRef]
- Mukohara, T. Pi3k mutations in breast cancer: Prognostic and therapeutic implications. Breast Cancer Targets Ther. 2015, 7, 111–123. [Google Scholar] [CrossRef] [Green Version]
- Samuels, Y.; Waldman, T. Oncogenic mutations of PIK3CA in human cancers. Curr. Top. Microbiol. Immunol. 2010, 346, 21–41. [Google Scholar]
- Tzanikou, E.; Markou, A.; Politaki, E.; Koutsopoulos, A.; Psyrri, A.; Mavroudis, D.; Georgoulias, V.; Lianidou, E. PIK3CA hotspot mutations in circulating tumor cells and paired circulating tumor DNA in breast cancer: A direct comparison study. Mol. Oncol. 2019, 13, 2515–2530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adzhubei, I.; Jordan, D.M.; Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 2013, 76, 7.20.1–7.20.41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tavtigian, S.V.; Deffenbaugh, A.M.; Yin, L.; Judkins, T.; Scholl, T.; Samollow, P.B.; De Silva, D.; Zharkikh, A.; Thomas, A. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J. Med. Genet. 2006, 43, 295–305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, Y.; Chan, A.P. PROVEAN web server: A tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015, 31, 2745–2747. [Google Scholar] [CrossRef] [Green Version]

| Characteristics | Total | in % | |
|---|---|---|---|
| Patients | 13 | 100 | |
| Age at blood draw | |||
| Mean | 61.2 | ||
| Median | 61 | ||
| Range | 46–78 | ||
| Tumor size | |||
| pT1 | 4 | 30.8 | |
| pT2–4 | 8 | 61.5 | |
| na | 1 | 7.7 | |
| Nodal status at time of diagnosis | |||
| 0 | 4 | 30.8 | |
| 1–3 | 7 | 53.8 | |
| na | 2 | 15.4 | |
| Metastasis status at time of diagnosis | |||
| 0 | 10 | 77.0 | |
| 1 | 2 | 15.4 | |
| na | 1 | 7.7 | |
| Histology | |||
| NST | 8 | 61.5 | |
| Invasive-lobular | 5 | 38.5 | |
| Grading | |||
| 1–2 | 9 | 69.2 | |
| 3 | 2 | 15.4 | |
| na | 2 | 23.1 | |
| Subtype | |||
| Luminal | 11 | 84.6 | |
| Triple negative | 2 | 15.4 | |
| CTC count (per 7.5 mL blood) | |||
| Median | 94 | ||
| Range | 6–15,340 | ||
| Forward Primer | Reverse Primer | |
|---|---|---|
| PIK3CA Exon 5 | 5′ AAGACTCGGCAGCATCTCCAGCATTTCCACAGCTACACCA 3′ | 5′ GCGATCGTCACTGTTCTCCAGATGTTCTCCTAACCATCTGA 3′ |
| PIK3CA Exon 10 | 5′ AAGACTCGGCAGCATCTCCAGGGAAAATGACAAAGAACAG 3′ | 5′ GCGATCGTCACTGTTCTCCAATTTTAGCACTTACCTGTGAC 3′ |
| PIK3CA Exon 21 | 5′ AAGACTCGGCAGCATCTCCATTGATGACATTGCATACATTCG 3′ | 5′ GCGATCGTCACTGTTCTCCAGTGGAAGATCCAATCCATTT 3′ |
| ESR1 Exon 5 | 5′ AAGACTCGGCAGCATCTCCATTGACCCTCCATGATCAGGT 3′ | 5′ GCGATCGTCACTGTTCTCCAGCTACTCCTAAGCTACAGCC 3′ |
| ESR1 Exon 7 | 5′ AAGACTCGGCAGCATCTCCATCTCTCACTCTCTCTCTGCG 3′ | 5′ GCGATCGTCACTGTTCTCCAGATGTGGGAGAGGATGAGGA 3′ |
| ESR1 Exon 8 | 5′ AAGACTCGGCAGCATCTCCAAGTAGTCCTTTCTGTGTCTTC 3′ | 5′ GCGATCGTCACTGTTCTCCAAATGCGATGAAGTAGAGCCC 3′ |
| AKT1 Exon 3 | 5′ AAGACTCGGCAGCATCTCCAGTAGAGTGTGCGTGGCTC 3′ | 5′ GCGATCGTCACTGTTCTCCACCCCAAATCTGAATCCCGAG 3′ |
| ERBB2 Exon 8 | 5′ AAGACTCGGCAGCATCTCCAGGCTACATGTTCCTGATCTCC 3′ | 5′ GCGATCGTCACTGTTCTCCAGGGTCTGAGGAAGGATAGGA 3′ |
| ERBB2 Exon 18 | 5′ AAGACTCGGCAGCATCTCCAAAGTACACGATGCGGAGACT 3′ | 5′ GCGATCGTCACTGTTCTCCAACCTTCACCTTCCTCAGCTC 3′ |
| ERBB2 Exon 19 | 5′ AAGACTCGGCAGCATCTCCAATCCTCCTCTTTCTGCCCAG 3′ | 5′ GCGATCGTCACTGTTCTCCAAGTCTAGGTTTGCGGGAGTC 3′ |
| ERBB2 Exon 20 | 5′ AAGACTCGGCAGCATCTCCATGGTTTGTGATGGTTGGGAG 3′ | 5′ GCGATCGTCACTGTTCTCCAGACATGGTCTAAGAGGCAGC 3′ |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Franken, A.; Rivandi, M.; Yang, L.; Jäger, B.; Krawczyk, N.; Honisch, E.; Niederacher, D.; Fehm, T.; Neubauer, H. A Multiplex PCR-Based Next Generation Sequencing-Panel to Identify Mutations for Targeted Therapy in Breast Cancer Circulating Tumor Cells. Appl. Sci. 2020, 10, 3364. https://doi.org/10.3390/app10103364
Franken A, Rivandi M, Yang L, Jäger B, Krawczyk N, Honisch E, Niederacher D, Fehm T, Neubauer H. A Multiplex PCR-Based Next Generation Sequencing-Panel to Identify Mutations for Targeted Therapy in Breast Cancer Circulating Tumor Cells. Applied Sciences. 2020; 10(10):3364. https://doi.org/10.3390/app10103364
Chicago/Turabian StyleFranken, André, Mahdi Rivandi, Liwen Yang, Bernadette Jäger, Natalia Krawczyk, Ellen Honisch, Dieter Niederacher, Tanja Fehm, and Hans Neubauer. 2020. "A Multiplex PCR-Based Next Generation Sequencing-Panel to Identify Mutations for Targeted Therapy in Breast Cancer Circulating Tumor Cells" Applied Sciences 10, no. 10: 3364. https://doi.org/10.3390/app10103364
APA StyleFranken, A., Rivandi, M., Yang, L., Jäger, B., Krawczyk, N., Honisch, E., Niederacher, D., Fehm, T., & Neubauer, H. (2020). A Multiplex PCR-Based Next Generation Sequencing-Panel to Identify Mutations for Targeted Therapy in Breast Cancer Circulating Tumor Cells. Applied Sciences, 10(10), 3364. https://doi.org/10.3390/app10103364





