1. Introduction
The effective reproduction number (
) is a key epidemiological metric used to assess the current rate of disease transmission within a population. By tracking changes in transmission, it enables public health officials to assess the impact of interventions such as vaccination and masking, guiding timely adjustments to strategies aimed at controlling diseases. It represents the average number of secondary infections that a single infectious individual generates at a given point in time [
1,
2]. Traditionally,
estimates rely on clinical data such as reported cases, hospitalizations, or deaths. However, these estimates can be subject to temporal biases due to variations in testing availability, reporting delays, test-seeking behavior, and changes in healthcare practices [
3,
4]. Although hospitalizations and deaths are less influenced by testing availability, they can be highly sensitive to external factors such as hospital capacity and resource constraints [
3].
The U.S. coronavirus disease 2019 (COVID-19) public health emergency declaration expired on 11 May 2023 [
5]. As of 18 December 2023, the California Department of Public Health (CDPH) ceased updates of COVID-19 cases, shifting its focus toward monitoring hospitalizations, emergency department visits, and wastewater data [
6].
Wastewater-based epidemiology (WBE) continues to be a valuable and cost-effective approach for monitoring the spread of infectious diseases [
7,
8,
9]. WBE methodology involves the collection of sewage samples from wastewater treatment plants (WWTPs) or sewer lines, followed by molecular analysis to quantify pathogen concentrations. This approach provides a powerful means to track community health trends by analyzing biological and chemical markers present in wastewater [
10]. During the COVID-19 pandemic, many studies showed that SARS-CoV-2 RNA concentrations in wastewater correlated with the number of confirmed cases, making wastewater monitoring a useful way to track the spread of the virus [
11,
12,
13,
14]. However, research showed that the relationship between wastewater data and COVID-19 clinical cases varied over time [
14,
15,
16,
17,
18]. This relationship depended on several factors, including testing availability and methods, public health policies, people’s behavior, vaccination levels, immunity from past infections, and new variants that affect virus shedding in feces [
15,
19,
20,
21]. In particular, inadequate or inconsistent testing was identified as a major driver of instability in the ratio between wastewater signals and reported case counts [
14].
To improve the prediction of COVID-19 cases from wastewater data, various machine learning approaches have recently been explored [
22,
23]. For instance, Rezaeitavabe et al. [
23] found that k-nearest neighbors and random forest models achieved the highest predictive accuracy across demographic and socioeconomic groups, while simpler linear models underperformed. However, these approaches require fitting separate models for each site and do not account for testing volume or incorporate interpretable epidemiological structure. Consequently, their ability to generalize across regions or to support mechanistic estimation of transmission metrics such as
remains limited.
To overcome these challenges, we introduce a unified hierarchical framework that jointly accounts for spatial heterogeneity, temporal autocorrelation, and variability in testing access. While previous studies have addressed some of these components [
20,
24,
25], our work is the first to integrate all of them within a single model.
Our framework integrates daily testing volume, test positivity rates, and SARS-CoV-2 wastewater concentrations using a hierarchical Beta-Binomial formulation that shares information across twenty California counties while retaining county-specific effects. Importantly, using a Beta-Binomial response model naturally accommodates extra-binomial variation arising from variability in viral concentrations in wastewater, spatial heterogeneity, and correlated measurement error driven by county-level differences in the probability of reporting positive cases—sources of dependence that are appropriately captured by the Beta-Binomial formulation but would violate the constant-
p and independence assumptions of a standard binomial likelihood. These are sources of variability that a binomial model cannot capture without ad hoc dispersion adjustments [
26,
27]. This model also incorporates a temporal random effects to capture short-term temporal dependence in transmission and explicitly accounts for overdispersion through a covariate-dependent dispersion term linked to wastewater levels and testing volume. By directly modeling testing behavior, our framework adjusts case estimates for changes in testing access and behavior, moving beyond the assumption of stable testing conditions. The resulting testing-adjusted case counts are then used to compute
following the established methodology of Cori et al. [
1]. We apply this model to data from twenty California counties participating in WastewaterSCAN, a statewide wastewater-based surveillance program initiated in 2020 [
28].
2. Materials and Methods
We use reported COVID-19 cases and SARS-CoV-2 concentrations in wastewater from 20 counties in California. This data spans from 1 March 2023 to 17 December 2023, after which public reporting of COVID-19 cases was discontinued. The start date was chosen to coincide with the earliest available wastewater data. This timeframe was selected to align both data sources and to restrict the analysis to a period with reliable and comprehensive clinical reporting.
Figure 1 provides a schematic overview of the analytical workflow used in this study.
2.1. Data Sources
Clinical Data. We used publicly available data for daily county-level COVID-19 cases from the Official California State Government website, which sourced COVID-19 data from the California Health & Human Services Agency (CalHHS) [
6]. We considered both “Total Tests” and “Positive Tests”, based on the specimen collection date, to capture overall testing volume and confirmed cases. The TPR is calculated by dividing the number of positive tests by the total number of tests administered, or (Positive Tests)/(Total Tests). To estimate population-level case rates for each county, we divided the number of new cases by the county population and scaled the result to reflect cases per 10,000 individuals. From this point forward, we refer to the standardized case counts as the “case rates”.
SARS-CoV-2 RNA concentrations. We analyzed SARS-CoV-2 RNA concentration in wastewater data from twenty-eight WWTPs across twenty California counties, each conducting surveillance three to seven times per week, between 1 March 2023 and 6 August 2024. Wastewater data were obtained from CPDH [
29] and WastewaterSCAN, a national WDS program that began in California in 2020 [
28]. We included only WWTPs that conducted active SARS-CoV-2 monitoring and had at least five consecutive months of overlapping clinical case data prior to 17 December 2023. A total of twenty-eight plants across twenty counties met these inclusion criteria. We selected this period because it corresponds with the availability of published protocols for sampling, analysis, and data processing.
To account for variability in fecal content and nucleic acid recovery efficiency, SARS-CoV-2 RNA concentrations in each wastewater sample were normalized to the concentration of pepper mild mottle virus (PMMoV) RNA, a fecal biomarker consistently present in human waste. This normalization (N/PMMoV) enhances comparability across samples by adjusting for differences in sample quality and processing. PMMoV is a highly abundant and stable fecal indicator in wastewater and is routinely measured alongside SARS-CoV-2 to standardize viral concentrations. Its inclusion allows for normalization across samples, helping to control for differences in population size, wastewater flow, and analytical variability [
28].
Normalized SARS-CoV-2 RNA concentrations in wastewater were subsequently smoothed using the np R package version 0.60.18. We applied kernel regression via the npreg function to obtain a non-parametric estimate of the trend. First, we used the npregbw function to compute least-squares cross-validated bandwidths for the local constant estimator. These bandwidths were then used to fit the model with npreg. Finally, we applied the predict function to generate fitted values for all dates in the original dataset, ensuring a smooth representation of wastewater concentration trends.
2.2. Time-Lag Cross-Correlation
To determine the temporal relationship between the smoothed wastewater signal and clinical indicators (cases and test positivity rate), we conducted cross-correlation analyses across all counties for the study period between 1 March 2023 and 17 December 2023. This allowed us to identify leading or lagging patterns between wastewater trends and clinical outcomes. We calculated Spearman’s correlation coefficients for time lags ranging from
to
days over a moving 12-week window. This lag range reflects the biologically plausible interval between infection, viral shedding, and case reporting and aligns with prior wastewater surveillance studies [
14,
30]. We selected the lag with the highest mean correlation. Negative lags indicate that increments in the wastewater signal precede rises in reported clinical cases, suggesting the potential of wastewater surveillance as an early warning indicator. In contrast, positive lags suggest that the wastewater signal follows clinical trends, with viral concentrations in wastewater increasing after a rise in reported cases. The identified lag values were subsequently used to temporally align clinical case data with wastewater trends, ensuring accurate synchronization for downstream analyses and enhancing the validity of comparative and predictive assessments.
2.3. Statistical Model
This study uses a hierarchical regression framework to model COVID-19 cases across California counties, integrating wastewater viral concentrations as a key covariate for early detection of transmission trends. Let
denote the number of reported COVID-19 cases in county
on day
. We model
using a Beta-Binomial distribution. The Beta-Binomial model was popularized by [
26] and is a compound distribution that combines the binomial and beta distributions, making it useful for modeling overdispersed binomial data (i.e., data where the variance exceeds the mean):
where
is the number of tests conducted in county
i on day
j,
is the average probability of success that represents proportion of all COVID-19 tests performed that are positive, and
is an extra dispersion parameter to account for variability beyond what is expected under a standard binomial distribution,
where larger values of
indicate lower overdispersion, and as
the model converge to binomial model. The mean is given by
, with
where
represents the wastewater viral concentration,
and
are county-specific coefficients, and
is a temporal random effect, modeled as a first-order autoregressive (AR(1)) process, which captures deviations due to dynamic changes in transmission over time. County-specific intercepts and slopes are included as random effects to account for spatial heterogeneity in baseline transmission and the relationship with wastewater concentrations.
where
captures the baseline infection level for county
i, with
representing the average baseline across all counties and
denoting the variance of the county-specific deviations from
; and
quantifies the association between the smoothed wastewater signal and case counts, where
is the population-average slope and
the variance in county-specific deviations from
.
This hierarchical formulation enables spatial and temporal flexibility in estimating the relationship between wastewater signals and reported COVID-19 cases. County-specific effects account for heterogeneity in vaccination coverage, demographics, testing availability, and reporting practices. At the same time, temporal variation captures the impact of emerging variants, public health interventions, and changes in population immunity.
The Beta-Binomial regression model is implemented using the glmmTMB package in R [
31], which provides flexible specification of both the mean and dispersion components of the model. We specified fixed and random effects for the mean structure, and used the dispformula argument to model the dispersion parameter
as a function of covariates. The logarithm of the dispersion parameter is modeled linearly as
where
represents county-specific fixed effects,
is the coefficient associated with wastewater viral concentration
, and
is the coefficient associated with the total number of tests performed per 10,000 population,
. We expected overdispersion to vary by county, with higher testing reducing variability and higher wastewater viral levels increasing it due to greater transmission and higher case counts.
2.4. Estimation of from Predicted Cases
Gostic et al. [
3] offers a comprehensive review of the key challenges and recommended strategies for accurate and timely estimation of the effective reproduction number,
. For near real-time estimation, they endorse the method proposed by Cori et al. [
1]. This method estimates
as follows:
where
is the number of new infections (cases) at time
t, and
is the generation interval distribution, representing the probability that an individual infected
s days ago infects a new individual (secondary case) today. In practice,
is a proxy for the true number of infections
, as it typically includes only detected cases and excludes undiagnosed or asymptomatic infections. A common assumption is
, with
p as an unknown constant. As a dimensionless ratio,
is invariant to this scaling [
4,
32], and estimates remain consistent regardless of
p [
18,
20].
To estimate using testing-adjusted incidence, we first compute the daily test positivity rate (TPR), which serves as a proxy for infection prevalence. Daily incidence is then approximated as , where T is the average number of tests conducted during the study period. This reconstructed time series is used to estimate . As testing data are reported by specimen collection date—which typically precedes the date of case reporting—we do not model the reporting delay distribution.
Although
depends on incidence, it is unaffected by the scaling factor
T, allowing us to compute
directly from the TPR as follows:
It is worth noting that our objective is not to estimate true incidence but to accurately capture infection trends to support the reliable estimation of . captures changes in transmission dynamics (increasing or decreasing); however, it does not provide information on the absolute number of cases or overall disease burden.
Huisman et al. [
11] introduced a method to estimate
from wastewater data through a deconvolution of the shedding load distribution (SLD). They showed that the optimal SLD can be determined by analyzing the relationship between wastewater-based estimates (
) and case-based estimates (
). Once the SLD is estimated from historical wastewater and case data, it enhances the accuracy of
compared to predefined SLDs. Building on this work, Champredon et al. [
33] developed the ern R package version 2.1.2 which estimates
directly from wastewater data. While the package relies on the EpiEstim [
1] framework for the core
calculation, it is specifically designed to incorporate assumptions about the SLD, following the idea of Huisman et al. [
11] with the addition of a locally estimated scatterplot smoothing (LOESS) smooth. Both studies underscore that
estimates can vary substantially depending on the assumed SLD.
We estimated the
using the method of Cori et al. [
1], implemented through the EpiEstim package version 2.2.5. Estimates were computed from both observed cases and predicted cases derived from wastewater data, using the Beta-Binomial regression model. Our analysis used a daily time step with a 7-day sliding window, assuming a gamma-distributed generation interval with a mean of 6.5 days and a standard deviation of 4.0, consistent with prior COVID-19 studies [
4].
For comparison, we generated
estimates using the ern package with its default COVID-19 parameters. The generation interval was modeled as a gamma distribution (mean = 6.84, SD = 0.7486, shape = 2.39, shape SD = 0.3573), and the fecal shedding distribution also followed a gamma distribution (mean = 12.90215, SD = 1.136829, shape = 1.759937, shape SD = 0.2665988). The LOESS smoothing span was set based on data length (span = 21/N, with N the number of observations) as in [
34]. We then assessed the consistency of these estimates with those derived from our hierarchical model.
2.5. Model Evaluation
Three versions of the Beta-Binomial model were implemented to predict the number of COVID-19 cases in 20 counties across California. These versions differ in the structure of the logit link specified in Equation (
2). In Model 1 (M1), the logit of the probability is defined as
where
is a fixed effect shared across all counties (i.e.,
in Equation (
2)). Model 2 (M2) builds upon M1 by adding a temporal random effect to capture time-dependent variations,
, modeled as a first-order autoregressive process (AR(1)):
. Model 3 (M3) builds upon M2 by allowing the slope
to vary by county, treating it as a random effect:
.
Note that the mean of the Beta-Binomial model is given by . With an estimate of the TPR, we approximate the expected number of cases as . To assess the impact of testing volume on case predictions, we compared model outputs under fixed testing levels, which allows for smoother trend estimation by reducing variability introduced by fluctuations in daily testing. For each county, we used the 5th, 50th, and 95th percentiles of daily test counts during the study period to represent low, average, and high testing scenarios, respectively. We also generated predictions using the real number of tests conducted during each day. While the magnitude of predicted cases varies with the assumed testing level, the estimation of remains largely stable due to its scale-invariant property, as discussed earlier.
To assess model performance, we partitioned the data into training and testing sets. The models were trained on data from 1 March 2023 through 12 November 2023, and their predictive accuracy was assessed over the following 35-day period, from 13 November 2023 to 17 December 2023, using unseen data. This approach allowed us to assess each model’s capacity to generalize beyond the training data and assess its long-term forecasting accuracy across all counties.
For comparison with the proposed Beta-Binomial hierarchical model, we additionally implemented random forest (RF) and k-nearest neighbors (k-NN). This selection follows evidence from Rezaeitavabe et al. [
23], who reported that RF and k-NN often exhibit strong predictive performance among eight evaluated machine learning models. Consistent with their approach, we trained separate models for each of the 20 counties and then averaged the predictive metrics across counties to obtain an overall measure of performance. Before model training, we conducted 5-fold cross-validation combined with grid search to perform hyperparameter tuning and model selection.
We used the mean Absolute percentage error (MAPE) and Spearman’s rank correlation () to compare observed and predicted values. Spearman’s measures the strength and direction of the monotonic relationship, indicating how well the rankings of observed and predicted values align. MAPE measures the average prediction error as a percentage of the observed values; lower values indicate better predictive accuracy. In addition, for models M1–M3, model fit was evaluated using the Akaike information criterion (AIC) and the Bayesian information criterion (BIC), where lower values indicate a better trade-off between model complexity and goodness of fit. We also calculated the percent differences in MAPE between models to highlight the influence of model components.
We did not report AIC or BIC for the random forest and k-nearest neighbors models because these algorithms are non-parametric and do not have a likelihood function or a well-defined number of parameters. AIC and BIC can only be computed for parametric models estimated via maximum likelihood, which applies to our Beta-Binomial models but not to RF or k-NN. As noted by Hastie et al. [
35], information criteria such as AIC are meaningful only when a likelihood-based loss is available. Similarly, James et al. [
36] emphasize that cross-validation is preferred when the assumptions behind AIC/BIC do not hold or when a model cannot be specified through a likelihood.
For this reason, while AIC/BIC were used for model comparison within the Beta-Binomial family, the RF and k-NN models were, instead, tuned and evaluated using cross-validated predictive error.
3. Results
3.1. COVID-19 Cases and SARS-CoV-2 Wastewater Concentration
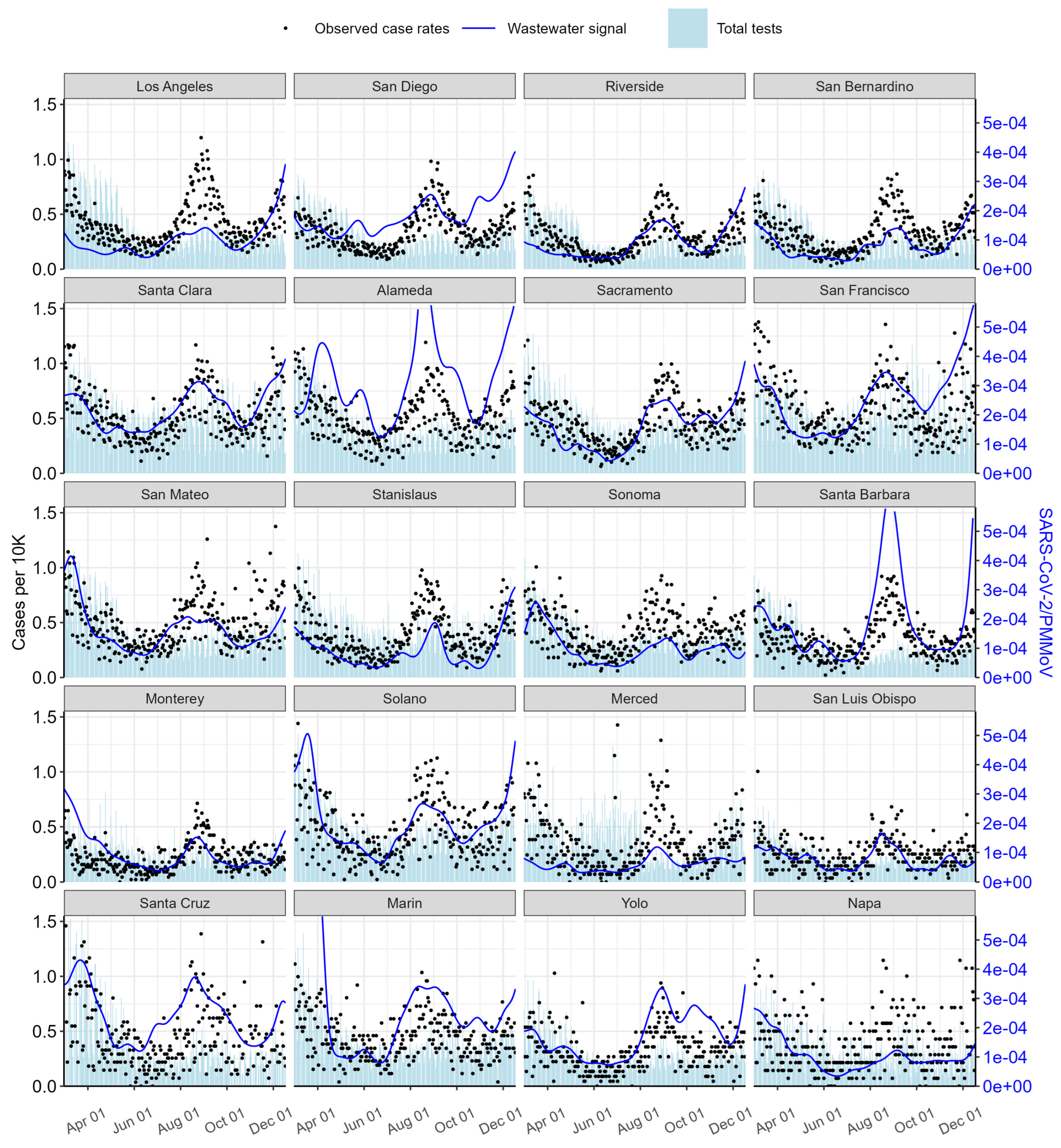
Figure 2 presents time series of COVID-19 metrics for the counties included in this study across California. In each panel, red dots represent reported case rates, blue lines depict the smoothed wastewater signal, and light blue bars indicate the total number of clinical tests performed. The figure illustrates how wastewater trends align closely with reported cases across different counties; sorted by population size. It also highlights how shifts in testing practices, such as reduced testing frequency and the rise of unreported at-home tests, can contribute to substantial underestimation of true infection rates.
Figure A2, in
Appendix A presents the reported TPR.
3.2. Model Prediction Performance
Table 1 summarizes the predictive performance of the proposed hierarchical models (M1–M3) and the two machine learning benchmarks, reporting both the mean absolute percentage error (MAPE) and Spearman’s rank correlation (
) as key evaluation metrics. To highlight the contribution of each model component, we also report the percent change in MAPE between models.
Overall, the hierarchical models clearly outperform both random forest and k-NN in terms of prediction accuracy and rank correlation. Incorporating temporal dependence through an AR(1) structure (Model M2) improves performance relative to the baseline model (M1), resulting in a 10.05% reduction in MAPE and a slight increase in Spearman’s correlation. This indicates that short-term autocorrelation plays an important role in capturing the dynamics of test positivity rates.
Figure A5, in
Appendix A, illustrates the temporal evolution of the AR(1) intercept parameter (
) across counties, capturing shifts in the latent autoregressive structure of the model over time. These trends offer valuable insights into local transmission patterns and variability in reporting.
Although models M2 and M3 achieve nearly identical predictive performance, we selected M2 as the final model based on model parsimony and information criteria. Both models produce the same MAPE (32.99) and Spearman correlation (0.721), indicating that the wastewater–county interaction term in M3 does not improve accuracy. Consistent with this, AIC and BIC values are slightly lower for M2, reflecting a better balance between goodness-of-fit and model complexity. Because the interaction effect () did not yield measurable improvement and increased model complexity without added benefit, we retained the simpler and more interpretable M2 for all subsequent analyses.
In contrast, the machine learning models exhibit substantially poorer predictive performance. Random forest and k-NN produce markedly higher MAPE values—41.29% and 33.15% higher than Model M2, respectively—and and lower rank correlations compared with the hierarchical models. Their inferior performance likely reflects the absence of explicit temporal structure, their reliance on county-specific models without partial pooling, and the lack of mechanistic links between wastewater concentrations, testing behavior, and infection trends.
Figure 3 displays the predicted case rates for each county based on Model M2, under varying testing levels. These levels correspond to the 5th (red), 50th (orange), and 95th (green) percentiles of daily tests conducted during the study period. Predictions using the actual number of tests performed are shown with blue and red bars for the training and testing period, respectively. The predicted case trends align closely with observed data, indicating that the forecasting model performs well. Predictions for the TPR are presented in
Figure A3 in
Appendix A.
The corresponding estimated
, calculated from predicted cases as the product of the predicted TPR, based on Model M2, and the daily total number of tests per county, is presented in
Figure 4. Assuming higher testing levels predicted more cases, as anticipated, yet the overall transmission trends remained consistent across different testing scenarios (
Figure 3). Consequently, the average
estimates remained consistent across varying testing levels, as shown in
Figure A4 in
Appendix A. Although the average
values remain comparable across low, average, and high testing scenarios, the confidence intervals broaden with decreasing testing levels, indicating increased uncertainty when testing is limited.
When infection prevalence is low, the epidemic “signal”, appearing as sparse clinical case counts and/or low viral concentrations in wastewater, is frequently masked by background noise. As a result, estimates of
often drift toward 1 and are accompanied by wide credible intervals, indicating greater uncertainty about the actual epidemic trend during periods of low prevalence. This challenge is especially evident in counties such as Napa, Santa Cruz, San Luis Obispo, and Yolo. Wastewater monitoring is generally more reliable in urban areas, where larger populations, closer proximity to laboratories, and better-equipped treatment facilities enhance sampling coverage and data quality.
Figure A6 in
Appendix C shows the MAPE and Spearman’s correlation between predicted and observed case rates, plotted against county population size. Both metrics indicate improved model performance in more populous counties: correlations are higher and MAPE values are lower, suggesting more reliable case estimation in densely populated areas.
3.3. Overdispersion Modeling
In
Appendix B, we present parameter estimates for the dispersion model.
Table A1 supports our hypothesis: the coefficient for wastewater viral concentration (
) is consistently negative, indicating that higher viral loads increase variability in case detection across counties. In contrast, the coefficient for testing (
) is positive in the more comprehensive models (M2 and M3), suggesting that increased testing reduces overdispersion. The median county-specific effect is 8.21 (IQR = 7.92), reflecting variation and confirming that overdispersion varies by county.
4. Discussion
This study highlights the value of wastewater-based disease surveillance (WDS) as a powerful tool for tracking infectious disease trends, particularly as clinical testing becomes less widespread and reliable. We developed and validated a series of hierarchical regression models that combine SARS-CoV-2 RNA concentrations in wastewater with reported COVID-19 testing data to predict case trends and estimate the effective reproduction number () at the county level throughout California.
The results indicate that the hierarchical Beta-Binomial model with AR(1) structure (M2) provides the best balance of predictive accuracy, interpretability, and statistical fit. Borrowing information across counties and explicitly modeling temporal dependence yield clear improvements over both simpler hierarchical models and flexible machine learning approaches.
Importantly, our results reinforce that estimation is scale-invariant and can be reliably derived from test positivity rates (TPR), even in the absence of absolute case counts. This is particularly relevant given the discontinuation of county-level COVID-19 case reporting in California as of December 2023. By leveraging TPR as a proxy for incidence and adjusting for testing volume, we preserved the structural integrity of estimates. Our comparison with estimates from the ern package confirmed the reliability of our approach and its applicability to real-time public health decision making. We also found that model performance varied by county population size, with higher prediction accuracy in counties with larger populations. This is likely due to stronger and more stable wastewater signals, better-equipped treatment infrastructure, and more consistent testing practices in those areas. These findings underscore the need to carefully interpret in rural or sparsely populated regions where data may be noisy or incomplete.
This study establishes wastewater surveillance as a vital, reliable method for tracking transmission dynamics, especially in the current context where COVID-19 is endemic and routine clinical testing has become much less frequent. In this context, wastewater monitoring provides a non-invasive, cost-effective, and timely means to capture community-level infection trends. This approach helps fill important gaps left by reduced testing and enables sustained vigilance against potential future outbreaks. However, several limitations require careful consideration.
First, some wastewater treatment plants temporarily paused sampling during the study period, creating data gaps that our modeling approach could not fully address. These interruptions may have introduced uncertainties in the wastewater signal, potentially affecting the precision of transmission estimates for affected counties. Developing methods to impute or model around these interruptions—such as using spatial interpolation or leveraging auxiliary data—remains an important direction for future work. Second, although our hierarchical framework effectively captures spatial and temporal variations in transmission, it does not explicitly incorporate critical external factors such as population mobility, behavioral changes, or the emergence of new viral variants. These factors can significantly influence transmission dynamics by altering contact patterns, susceptibility, and viral spread. Ignoring them may limit the model’s ability to fully explain fluctuations in infection rates or predict sudden changes in transmission. Incorporating such contextual information in future models would enhance their accuracy and provide a more comprehensive understanding of the drivers behind epidemic trends. Lastly, future research should explore the integration of Bayesian hierarchical or advanced machine learning models that explicitly incorporate relevant covariates such as mobility patterns, behavioral changes, vaccination rates, and variant prevalence. Including these factors would allow models to capture complex, real-world influences on transmission dynamics, leading to potentially more accurate and timely forecasts.
While this study was conducted using data from California, the hierarchical modeling framework is designed for broader applicability. The use of county-level random effects allows the model to learn local patterns from available data, making it adaptable to other geographic units such as cities, neighborhoods, or sewershed catchments. This structure is particularly valuable for stabilizing estimates in areas with sparse or noisy surveillance. Although adaptation to regions with limited clinical testing infrastructure would require at least minimal clinical data for calibration, alternative proxies, such as hospitalization counts or syndromic indicators, could be incorporated when testing data are scarce. The hierarchical structure also supports application in locations with different sewer system configurations, as the model can borrow strength across sites when wastewater data are limited. However, its performance in settings with decentralized, fragmented, or non-sewered sanitation systems depends on the representativeness of the wastewater signal. Successful implementation in such contexts would require appropriate normalization approaches and sampling strategies tailored to local infrastructure to ensure that measured viral concentrations reflect community-level transmission.
Overall, our framework adds robust evidence to the expanding consensus that wastewater surveillance is a cost-effective, scalable, and timely approach for tracking infectious disease trends. As traditional clinical surveillance systems reduce testing and reporting, WDS is an essential complementary tool, capable of providing early warnings and continuous situational awareness at the community level.
5. Conclusions
This study presents a hierarchical Beta-Binomial model that integrates SARS-CoV-2 RNA concentrations in wastewater with reported COVID-19 testing data to predict case trends and estimate the effective reproduction number () across California counties. Our framework addresses key challenges in wastewater-based epidemiology by explicitly accounting for spatio-temporal variation through county-specific random effects, autoregressive temporal structure, and covariate-dependent dispersion terms that link overdispersion to wastewater levels and testing volume.
Incorporating an autoregressive temporal component substantially improved predictive performance, while more complex spatial interactions provided negligible benefits, underscoring the dominant role of temporal dynamics in shaping epidemic trends. The selected model (M2) demonstrates that incorporating temporal autocorrelation significantly improves predictive performance, while county-specific slopes for wastewater effects provide negligible gains. This parsimonious framework outperforms machine learning alternatives (random forest and k-NN) in both prediction accuracy and correlation with observed data, highlighting the value of hierarchical structures that share information across counties while accommodating local heterogeneity.
Crucially, our results confirm that estimation remains reliable when derived from test positivity rates, even as absolute case reporting declines. This scale-invariance property ensures robust transmission monitoring in the post-reporting era, as evidenced by consistent estimates across varying testing scenarios and alignment with wastewater-only estimates from established methods. However, performance varies with population density, with urban counties showing more reliable predictions due to stronger wastewater signals and consistent testing practices. This underscores the need for careful interpretation in rural areas where data sparsity introduces greater uncertainty. As COVID-19 transitions to endemic monitoring and clinical reporting diminishes, our framework establishes wastewater surveillance as an essential component of sustainable public health infrastructure. The hierarchical modeling approach provides a flexible foundation for tracking transmission dynamics, offering early warning capabilities and continuous community-level insights that complement traditional surveillance systems.
Author Contributions
Conceptualization, J.C.M.-L., M.L.D.-T., A.M.-L., J.C., H.N.B. and M.N.; methodology, J.C.M.-L., M.L.D.-T., A.M.-L., H.N.B. and M.N.; software, J.C.M.-L. and M.L.D.-T.; validation, J.C.M.-L., M.L.D.-T., A.M.-L., J.C., H.N.B. and M.N.; formal analysis, J.C.M.-L., M.L.D.-T. and A.M.-L.; investigation, J.C.M.-L., M.L.D.-T., A.M.-L., J.C., H.N.B. and M.N.; resources, H.N.B. and M.N.; data curation, J.C.M.-L. and M.L.D.-T.; writing—original draft preparation, J.C.M.-L. and M.L.D.-T.; writing—review and editing, J.C.M.-L., M.L.D.-T., A.M.-L., J.C., H.N.B. and M.N.; visualization, J.C.M.-L. and M.L.D.-T.; supervision, J.C.M.-L., M.L.D.-T. and M.N.; project administration, J.C.M.-L. and M.L.D.-T.; funding acquisition, M.N. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
Acknowledgments
This research was supported by a grant from the University of California Alianza MX (UC Alianza MX).
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| COVID-19 | Coronavirus disease 2019 |
| RNA | Ribonucleic acid |
| SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 |
| PMMoV | Pepper mild mottle virus |
| TPR | Test positivity rate |
| WBE | Wastewater-based epidemiology |
| WDS | Wastewater-based disease surveillance |
| WWTPs | Wastewater treatment plants |
| LOESS | Locally estimated scatterplot smoothing |
| AR(1) | First-order autoregressive process |
| SD | Standard deviation |
| SLD | Shedding load distribution |
| AIC | Akaike information criterion |
| BIC | Bayesian information criterion |
| MAPE | Mean absolute percentage error |
| CDPH | California Department of Public Health |
Appendix A. Figures
Appendix A.1. Wastewater Treatment Plants (WWTPs) Locations
Figure A1.
Locations of twenty-eight WWTPs monitoring SARS-CoV-2 in twenty California counties from 1 March 2023 to 17 December 2023. The map displays county-level cumulative COVID-19 cases per 10,000 population in the study period, together with the population served by each facility. Counties excluded from the study are outlined in gray.
Figure A1.
Locations of twenty-eight WWTPs monitoring SARS-CoV-2 in twenty California counties from 1 March 2023 to 17 December 2023. The map displays county-level cumulative COVID-19 cases per 10,000 population in the study period, together with the population served by each facility. Counties excluded from the study are outlined in gray.
Appendix A.2. COVID-19 Test Positivity Rate (TPR) and Wastewater Signal
Figure A2.
COVID-19 test positivity rate (TPR) (black dots) and wastewater signal (blue lines). Blue bars indicate total tests conducted per county. Counties are sorted by population size.
Figure A2.
COVID-19 test positivity rate (TPR) (black dots) and wastewater signal (blue lines). Blue bars indicate total tests conducted per county. Counties are sorted by population size.
Appendix A.3. Predicted COVID-19 Test Positivity Rate (TPR)
Figure A3.
Predicted COVID-19 TPR. The red lines represent the mean predictions of TPR and black dots indicate the observed TPR. Counties are sorted by population size.
Figure A3.
Predicted COVID-19 TPR. The red lines represent the mean predictions of TPR and black dots indicate the observed TPR. Counties are sorted by population size.
Appendix A.4. Predicted Rt by Testing Level
Figure A4.
estimates based on predicted cases using the 5 (red), 50 (orange), and 95 (green) percentiles of tests conducted per county, sorted by population size. Shaded bands represent 95% confidence intervals. While the mean estimates are similar across testing levels, the confidence intervals widen under lower testing scenarios, reflecting increased uncertainty with limited data.
Figure A4.
estimates based on predicted cases using the 5 (red), 50 (orange), and 95 (green) percentiles of tests conducted per county, sorted by population size. Shaded bands represent 95% confidence intervals. While the mean estimates are similar across testing levels, the confidence intervals widen under lower testing scenarios, reflecting increased uncertainty with limited data.
Figure A5.
Temporal evolution of the AR(1) intercept parameter () across counties. Each line corresponds to a different county and reflects changes in the latent autoregressive structure of the model over time. The parameter captures persistent deviations in case counts not explained by covariates, providing insight into underlying county-specific temporal dependencies.
Figure A5.
Temporal evolution of the AR(1) intercept parameter () across counties. Each line corresponds to a different county and reflects changes in the latent autoregressive structure of the model over time. The parameter captures persistent deviations in case counts not explained by covariates, providing insight into underlying county-specific temporal dependencies.
Appendix B. Overdispersion Modeling
We hypothesized that overdispersion would vary across counties, with higher testing rates associated with reduced variability—indicating greater certainty in case detection—and higher viral concentrations in wastewater linked to increased variability, reflecting greater transmission potential and higher case counts. The parameter estimates in
Table A1 support this hypothesis: the coefficient for the viral loads (
) is consistently negative, suggesting that higher viral loads correspond to increased variability in case detection across counties. In contrast, the coefficient for the number of tests (
) is positive in the most comprehensive models (M2 and M3), indicating that more extensive testing is associated with lower overdispersion.
Table A1.
Parameter estimates from the log-dispersion model of the Beta-Binomial regression.
Table A1.
Parameter estimates from the log-dispersion model of the Beta-Binomial regression.
| | Mean Model | Overdispersion Model |
|---|
| Model | | | | |
| M1 = WW + County 1 | 3.18 | 1.46 | −3.13 | −2.81 |
| M2 = M1+ AR(1) | 0.80 | 0.89 | −0.85 | 3.84 |
| M3 = M2 + WW × County 1 | 0.80 | 0.89 | −0.85 | 3.84 |
Appendix C. Spearman’s Correlation and Mean Absolute Percentage Error
Figure A6.
Spearman’s correlation and mean absolute percentage error (MAPE) between predicted and observed COVID-19 cases across counties, shown with population size.
Figure A6.
Spearman’s correlation and mean absolute percentage error (MAPE) between predicted and observed COVID-19 cases across counties, shown with population size.
Appendix D. Literature Review
Table A2.
Summary of wastewater-based epidemiology predictive modeling studies.
Table A2.
Summary of wastewater-based epidemiology predictive modeling studies.
| Study Location | Predictive Model Type | Model Input | Spatial Structure | Temporal Model | Test/Testing Adjustment | Model Output | Ref |
|---|
| Chesapeake, USA | Copula-based Time Series Model (Gaussian Copula + ARMA) | SARS-CoV-2 N gene concentration; daily COVID-19 cases, hospitalizations, deaths | No (city-level aggregate) | Yes (ARMA) | No | Predicted COVID-19 case counts and trends with confidence intervals | [25] |
| England (45 STW sites) | Gradient Boosted Regression Tree; phenomenological prevalence model | SARS-CoV-2 RNA (N1 gene), biochemical & demographic covariates | Yes (45 sites mapped to sub-regions) | Yes (daily interpolation from 3-4 samples/week) | No (validated against independent prevalence survey) | Estimated SARS-CoV-2 prevalence across subregions | [24] |
| Davis, California, USA | Sequential Bayesian Beta Regression | Smoothed wastewater SARS-CoV-2 concentration (N/PMMoV); daily COVID-19 cases and tests | No (two separate sewershed analyses) | Yes (sequential/adaptive updating) | Yes (Explicitly via TPR) | Estimated Test Positivity Rate (TPR), WW transmission thresholds, Effective Reproductive Number (Re) | [20] |
| Ohio, USA (55 sites) | Multiple Machine Learning Models (k-NN, Random Forest, XGBoost best performers) | SARS-CoV-2 N2 concentration; demographic & socioeconomic indicators; daily COVID-19 cases | Yes (via site-level geographic and sociodemographic features) | No (lag analysis) | No | Predicted number of COVID-19 clinical cases | [23] |
| Athens, USA | Non-linear Machine Learning Models (Feed-Forward Neural Networks) | Wastewater SARS-CoV-2 concentration (various normalizations); daily COVID-19 cases | No (sewershed-level aggregate) | No (lag analysis) | No | Predicted number of COVID-19 clinical cases | [22] |
| Zurich, Switzerland & San Jose, USA | Bayesian Deconvolution (Expectation-Maximization) & EpiEstim | Smoothed SARS-CoV-2 RNA concentrations (N1, N2, S, ORF1a genes) from wastewater and sewage sludge | No (WWTP catchment aggregate) | Yes (Deconvolution with shedding load distribution) | No | Estimated Effective Reproductive Number, Incidence | [11] |
References
- Cori, A.; Ferguson, N.M.; Fraser, C.; Cauchemez, S. A New Framework and Software to Estimate Time-Varying Reproduction Numbers During Epidemics. Am. J. Epidemiol. 2013, 178, 1505–1512. [Google Scholar] [CrossRef] [PubMed]
- Wallinga, J.; Teunis, P. Different Epidemic Curves for Severe Acute Respiratory Syndrome Reveal Similar Impacts of Control Measures. Am. J. Epidemiol. 2004, 160, 509–516. [Google Scholar] [CrossRef] [PubMed]
- Gostic, K.M.; McGough, L.; Baskerville, E.B.; Abbott, S.; Joshi, K.; Tedijanto, C.; Kahn, R.; Niehus, R.; Hay, J.A.; De Salazar, P.M.; et al. Practical considerations for measuring the effective reproductive number, Rt. PLoS Comput. Biol. 2020, 16, e1008409. [Google Scholar] [CrossRef]
- Pitzer, V.E.; Chitwood, M.; Havumaki, J.; Menzies, N.A.; Perniciaro, S.; Warren, J.L.; Weinberger, D.M.; Cohen, T. The Impact of Changes in Diagnostic Testing Practices on Estimates of COVID-19 Transmission in the United States. Am. J. Epidemiol. 2021, 190, 1908–1917. [Google Scholar] [CrossRef]
- CDC. CDC Archive. 2023. Available online: https://archive.cdc.gov/www_cdc_gov/coronavirus/2019-ncov/your-health/end-of-phe.html (accessed on 23 May 2025).
- CDPH. COVID-19 Time-Series Metrics by County and State-Datasets-California Health and Human Services Open Data Portal. 2025. Available online: https://data.chhs.ca.gov/dataset/covid-19-time-series-metrics-by-county-and-state (accessed on 23 May 2025).
- Boehm, A.B.; Hughes, B.; Duong, D.; Chan-Herur, V.; Buchman, A.; Wolfe, M.K.; White, B.J. Wastewater concentrations of human influenza, metapneumovirus, parainfluenza, respiratory syncytial virus, rhinovirus, and seasonal coronavirus nucleic-acids during the COVID-19 pandemic: A surveillance study. Lancet Microbe 2023, 4, e340–e348. [Google Scholar] [CrossRef]
- Safford, H.R.; Shapiro, K.; Bischel, H.N. Wastewater analysis can be a powerful public health tool—if it’s done sensibly. Proc. Natl. Acad. Sci. USA 2022, 119, e2119600119. [Google Scholar] [CrossRef]
- Kumblathan, T.; Liu, Y.; Uppal, G.K.; Hrudey, S.E.; Li, X.F. Wastewater-Based Epidemiology for Community Monitoring of SARS-CoV-2: Progress and Challenges. ACS Environ. Au 2021, 1, 18–31. [Google Scholar] [CrossRef]
- Li, G.; Denise, H.; Diggle, P.; Grimsley, J.; Holmes, C.; James, D.; Jersakova, R.; Mole, C.; Nicholson, G.; Smith, C.R.; et al. A spatio-temporal framework for modelling wastewater concentration during the COVID-19 pandemic. Environ. Int. 2023, 172, 107765. [Google Scholar] [CrossRef]
- Huisman, J.S.; Scire, J.; Caduff, L.; Fernandez-Cassi, X.; Ganesanandamoorthy, P.; Kull, A.; Scheidegger, A.; Stachler, E.; Boehm, A.B.; Hughes, B.; et al. Wastewater-Based Estimation of the Effective Reproductive Number of SARS-CoV-2. Environ. Health Perspect. 2022, 130, 057011. [Google Scholar] [CrossRef]
- McMahan, C.S.; Self, S.; Rennert, L.; Kalbaugh, C.; Kriebel, D.; Graves, D.; Colby, C.; Deaver, J.A.; Popat, S.C.; Karanfil, T.; et al. COVID-19 wastewater epidemiology: A model to estimate infected populations. Lancet Planet. Health 2021, 5, e874–e881. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, M.K.; Topol, A.; Knudson, A.; Simpson, A.; White, B.; Vugia, D.J.; Yu, A.T.; Li, L.; Balliet, M.; Stoddard, P.; et al. High-Frequency, High-Throughput Quantification of SARS-CoV-2 RNA in Wastewater Settled Solids at Eight Publicly Owned Treatment Works in Northern California Shows Strong Association with COVID-19 Incidence. mSystems 2021, 6, e00829-21. [Google Scholar] [CrossRef]
- Xiao, A.; Wu, F.; Bushman, M.; Zhang, J.; Imakaev, M.; Chai, P.R.; Duvallet, C.; Endo, N.; Erickson, T.B.; Armas, F.; et al. Metrics to relate COVID-19 wastewater data to clinical testing dynamics. Water Res. 2022, 212, 118070. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Haak, L.; Carine, M.; Pagilla, K.R. Temporal assessment of SARS-CoV-2 detection in wastewater and its epidemiological implications in COVID-19 case dynamics. Heliyon 2024, 10, e29462. [Google Scholar] [CrossRef]
- Kadonsky, K.F.; Naughton, C.C.; Susa, M.; Olson, R.; Singh, G.L.; Daza-Torres, M.L.; Montesinos-López, J.C.; Garcia, Y.E.; Gafurova, M.; Gushgari, A.; et al. Expansion of wastewater-based disease surveillance to improve health equity in California’s Central Valley: Sequential shifts in case-to-wastewater and hospitalization-to-wastewater ratios. Front. Public Health 2023, 11, 1141097. [Google Scholar] [CrossRef]
- D’Aoust, P.M.; Tian, X.; Towhid, S.T.; Xiao, A.; Mercier, E.; Hegazy, N.; Jia, J.J.; Wan, S.; Kabir, M.P.; Fang, W.; et al. Wastewater to clinical case (WC) ratio of COVID-19 identifies insufficient clinical testing, onset of new variants of concern and population immunity in urban communities. Sci. Total Environ. 2022, 853, 158547. [Google Scholar] [CrossRef] [PubMed]
- Daza-Torres, M.L.; Montesinos-López, J.C.; Kim, M.; Olson, R.; Bess, C.W.; Rueda, L.; Susa, M.; Tucker, L.; García, Y.E.; Schmidt, A.J.; et al. Model training periods impact estimation of COVID-19 incidence from wastewater viral loads. Sci. Total Environ. 2023, 858, 159680. [Google Scholar] [CrossRef]
- Graham, M.S.; Sudre, C.H.; May, A.; Antonelli, M.; Murray, B.; Varsavsky, T.; Kläser, K.; Canas, L.S.; Molteni, E.; Modat, M.; et al. Changes in symptomatology, reinfection, and transmissibility associated with the SARS-CoV-2 variant B. 1.1. 7: An ecological study. Lancet Public Health 2021, 6, e335–e345. [Google Scholar] [CrossRef]
- Montesinos-López, J.C.; Daza-Torres, M.L.; García, Y.E.; Herrera, C.; Bess, C.W.; Bischel, H.N.; Nuño, M. Bayesian sequential approach to monitor COVID-19 variants through test positivity rate from wastewater. mSystems 2023, 8, e00018-23. [Google Scholar] [CrossRef] [PubMed]
- Colman, E.; Kao, R. The impact of signal variability on COVID-19 epidemic growth rate estimation from wastewater surveillance data. PLoS ONE 2025, 20, e0322057. [Google Scholar] [CrossRef]
- Rezaeitavabe, F.; Rezaie, M.; Modayil, M.; Pham, T.; Ice, G.; Riefler, G.; Coschigano, K.T. Beyond linear regression: Modeling COVID-19 clinical cases with wastewater surveillance of SARS-CoV-2 for the city of Athens and Ohio University campus. Sci. Total Environ. 2024, 912, 169028. [Google Scholar] [CrossRef]
- Rezaeitavabe, F.; Coschigano, K.T.; Riefler, G. Predicting COVID-19 in Ohio: Insights from wastewater, demographic and socioeconomic data. Sci. Total Environ. 2025, 969, 178938. [Google Scholar] [CrossRef]
- Morvan, M.; Jacomo, A.L.; Souque, C.; Wade, M.J.; Hoffmann, T.; Pouwels, K.; Lilley, C.; Singer, A.C.; Porter, J.; Evens, N.P.; et al. An analysis of 45 large-scale wastewater sites in England to estimate SARS-CoV-2 community prevalence. Nat. Commun. 2022, 13, 4313. [Google Scholar] [CrossRef]
- Jeng, H.A.; Singh, R.; Diawara, N.; Curtis, K.; Gonzalez, R.; Welch, N.; Jackson, C.; Jurgens, D.; Adikari, S. Application of wastewater-based surveillance and copula time-series model for COVID-19 forecasts. Sci. Total Environ. 2023, 885, 163655. [Google Scholar] [CrossRef] [PubMed]
- Skellam, J.G. A Probability Distribution Derived from the Binomial Distribution by Regarding the Probability of Success as Variable between the Sets of Trials. J. R. Stat. Soc. Ser. B (Methodol.) 2018, 10, 257–261. [Google Scholar] [CrossRef]
- Johnson, N.L.; Kemp, A.W.; Kotz, S. Univariate Discrete Distributions; John Wiley & Sons: Hoboken, NJ, USA, 2005. [Google Scholar]
- Boehm, A.B.; Wolfe, M.K.; Wigginton, K.R.; Bidwell, A.; White, B.J.; Hughes, B.; Duong, D.; Chan-Herur, V.; Bischel, H.N.; Naughton, C.C. Human viral nucleic acids concentrations in wastewater solids from Central and Coastal California USA. Sci. Data 2023, 10, 396. [Google Scholar] [CrossRef] [PubMed]
- CA.gov. California Open Data Portal: CDPH-Wastewater Surveillance Data, California. 2025. Available online: https://data.ca.gov/dataset/covid-19-wastewater-surveillance-data-california (accessed on 23 May 2025).
- Kumar, M.; Joshi, M.; Patel, A.K.; Joshi, C.G. Unravelling the early warning capability of wastewater surveillance for COVID-19: A temporal study on SARS-CoV-2 RNA detection and need for the escalation. Environ. Res. 2021, 196, 110946. [Google Scholar] [CrossRef] [PubMed]
- McGillycuddy, M.; Warton, D.I.; Popovic, G.; Bolker, B.M. Parsimoniously Fitting Large Multivariate Random Effects in glmmTMB. J. Stat. Softw. 2025, 112, 1–19. [Google Scholar] [CrossRef]
- Capistrán, M.A.; Capella, A.; Christen, J.A. Filtering and improved Uncertainty Quantification in the dynamic estimation of effective reproduction numbers. Epidemics 2022, 40, 100624. [Google Scholar] [CrossRef]
- Champredon, D.; Papst, I.; Yusuf, W. ern: An R package to estimate the effective reproduction number using clinical and wastewater surveillance data. PLoS ONE 2024, 19, e0305550. [Google Scholar] [CrossRef]
- Hill, D.T.; Zhu, Y.; Dunham, C.; Moran, E.J.; Zhou, Y.; Collins, M.B.; Kmush, B.L.; Larsen, D.A. Estimating the effective reproduction number from wastewater (Rt): A methods comparison. Epidemics 2025, 52, 100839. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, 2nd ed.; Springer: New York, NY, USA, 2009. [Google Scholar]
- James, G.; Witten, D.; Hastie, T.; Tibshirani, R. An Introduction to Statistical Learning: With Applications in R, 2nd ed.; Springer: New York, NY, USA, 2021. [Google Scholar]
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).