Molecular Effects of Inorganic and Methyl Mercury in Aquatic Primary Producers: Comparing Impact to A Macrophyte and A Green Microalga in Controlled Conditions
Abstract
1. Introduction
2. Bioaccumulation of Hg in Aquatic Primary Producers
3. Effects of Hg Exposure on Different Levels of Biological Organization in Aquatic Primary Producers
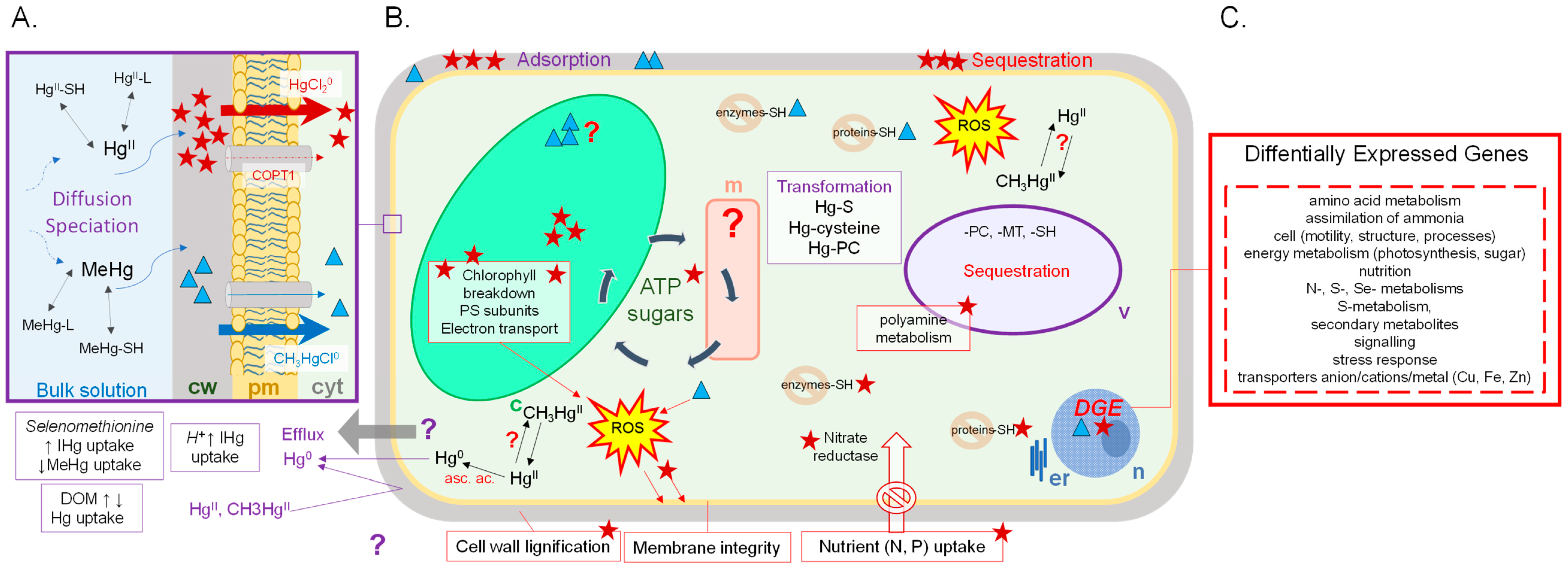
3.1. Physiological Responses
3.2. Response at the Protein and the Gene Level
4. A Case Study: Comparison of Responses to IHg and MeHg Exposure in C. reinhardtii and E. nuttallii
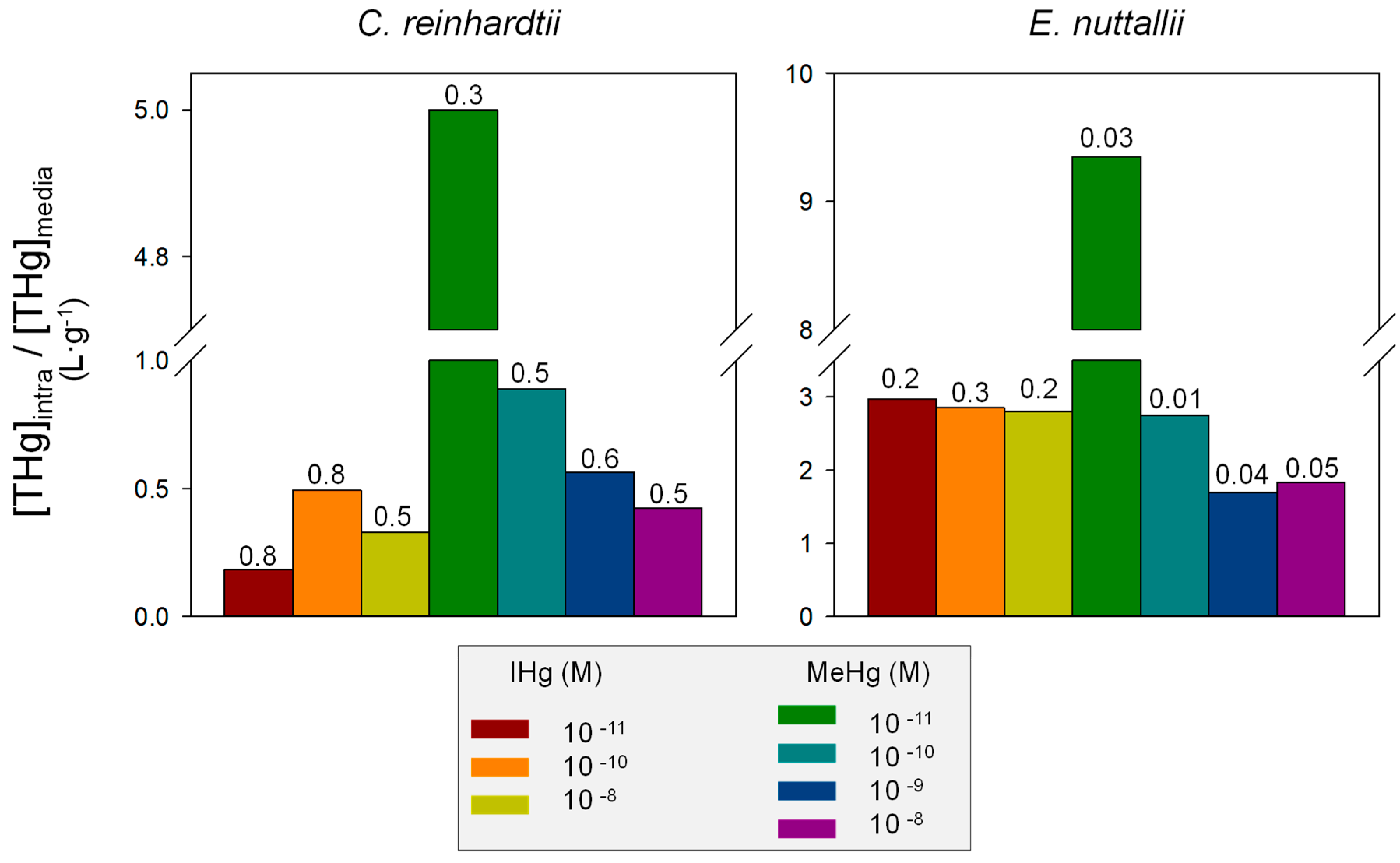
4.1. Comparison of IHg and MeHg Bioaccumulation
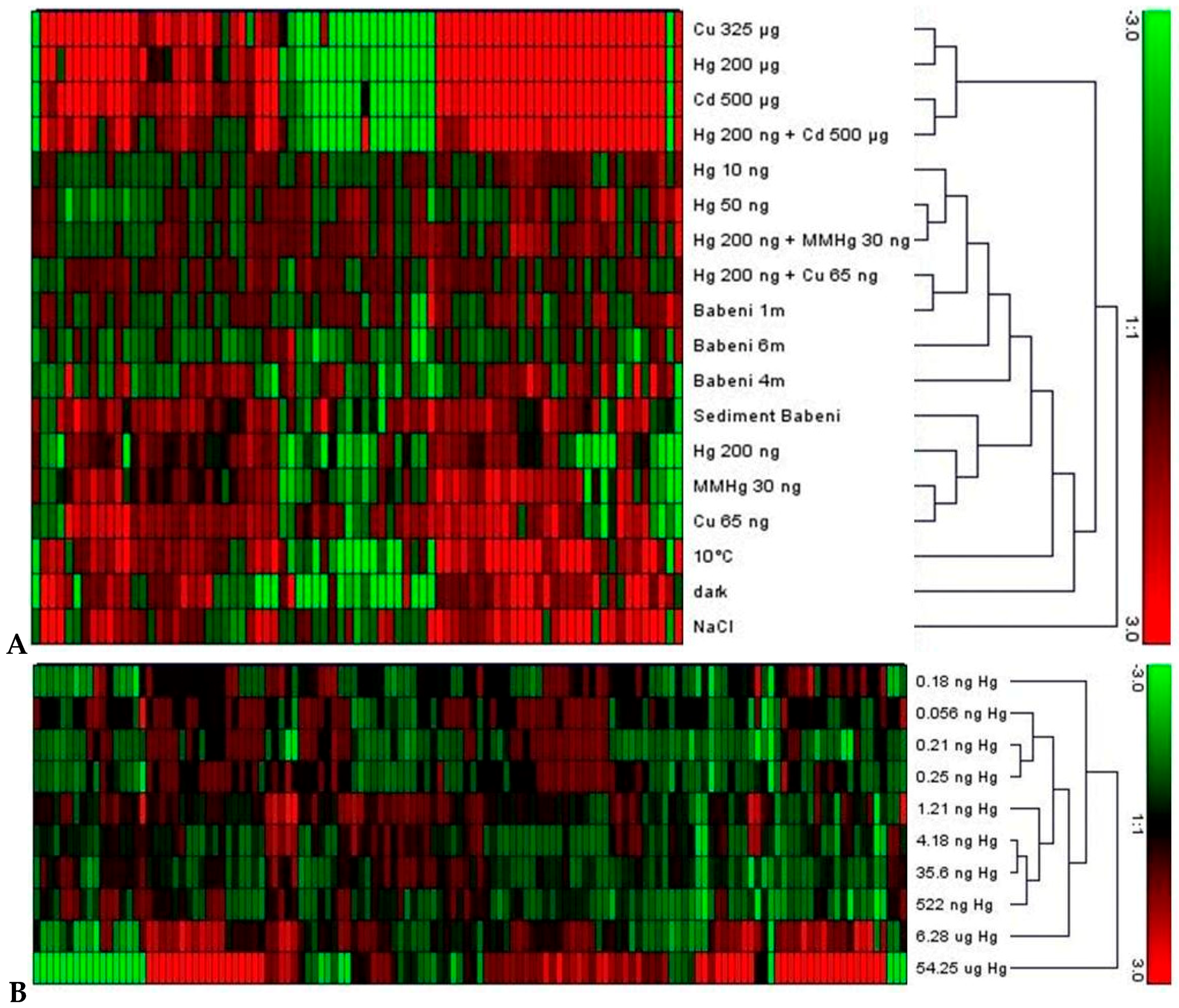
4.2. Comparison of Whole Transcriptome Analyses
4.3. Comparison of Physiological Data
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Dranguet, P.; Fluck, R.; Regier, N.; Cosio, C.; Le Faucheur, S.; Slaveykova, V.I. Towards mechanistic understanding of mercury availability and toxicity to aquatic primary producers. Chimia 2014, 68, 799–805. [Google Scholar] [CrossRef] [PubMed]
- Kocman, D.; Wilson, S.; Amos, H.; Telmer, K.; Steenhuisen, F.; Sunderland, E.; Mason, R.; Outridge, P.; Horvat, M. Toward an assessment of the global inventory of present-day mercury releases to freshwater environments. Int. J. Environ. Res. Public Health 2017, 14, 138. [Google Scholar] [CrossRef] [PubMed]
- Cossa, D.; Averty, B.; Pirrone, N. The origin of methylmercury in open Mediterranean waters. Limnol. Oceanogr. 2009, 54, 837–844. [Google Scholar] [CrossRef]
- Bravo, A.G.; Cosio, C.; Amouroux, D.; Zopfi, J.; Cheualley, P.A.; Spangenberg, J.E.; Ungureanu, V.G.; Dominik, J. Extremely elevated methyl mercury levels in water, sediment and organisms in a Romanian reservoir affected by release of mercury from a chlor-alkali plant. Water Res. 2014, 49, 391–405. [Google Scholar] [CrossRef] [PubMed]
- Bank, M.S. Mercury in the Environment: Pattern and Process, 1st ed.; University of California Press: Berkeley, CA, USA, 2012; 358p. [Google Scholar]
- Dranguet, P.; Cosio, C.; Le Faucheur, S.; Beauvais-Fluck, R.; Freiburghaus, A.; Worms, I.A.M.; Petit, B.; Civic, N.; Docquier, M.; Slaveykova, V.I. Transcriptomic approach for assessment of the impact on microalga and macrophyte of in-situ exposure in river sites contaminated by chlor-alkali plant effluents. Water Res. 2017, 121, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Beauvais-Fluck, R.; Slaveykova, V.I.; Skyllberg, U.; Cosio, C. Molecular effects, speciation and competition of inorganic and methyl mercury in the aquatic plant Elodea nuttallii. Environ. Sci. Technol. 2018, 52, 8876–8884. [Google Scholar] [CrossRef] [PubMed]
- Powell Kipton, J.; Brown Paul, L.; Byrne Robert, H.; Gajda, T.; Hefter, G.; Sjöberg, S.; Wanner, H. Chemical speciation of environmentally significant heavy metals with inorganic ligands. Part 2005, 1, The Hg2+–Cl–, OH–, CO32–, SO42–, and PO43– aqueous systems (IUPAC Technical Report). Pure Appl. Chem. 2009. [Google Scholar] [CrossRef]
- Le Faucheur, S.; Tremblay, Y.; Fortin, C.; Campbell, P.G.C. Acidification increases mercury uptake by a freshwater alga, Chlamydomonas reinhardtii. Environ. Chem. 2011, 8, 612–622. [Google Scholar] [CrossRef]
- Bravo, A.G.; Loizeau, J.L.; Bouchet, S.; Richard, A.; Rubin, J.F.; Ungureanu, V.G.; Amouroux, D.; Dominik, J. Mercury human exposure through fish consumption in a reservoir contaminated by a chlor-alkali plant: Babeni reservoir (Romania). Environ. Sci. Pollut. Res. 2010, 17, 1422–1432. [Google Scholar] [CrossRef] [PubMed]
- Marshall, B.G.; Forsberg, B.R.; Thome-Souza, M.; Peleja, R.; Moreira, M.Z.; Freitas, C.E. Evidence of mercury biomagnification in the food chain of the cardinal tetra Paracheirodon axelrodi (Osteichthyes: Characidae) in the Rio Negro, central Amazon, Brazil. J. Fish Biol. 2016, 89, 220–240. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Cai, Y.; O’Driscoll, N.J. Environmental Chemistry and Toxicology of Mercury; Wiley: Hoboken, NJ, USA, 2012; p. 600. [Google Scholar]
- Crane, M.; Babut, M. Environmental quality standards for water framework directive priority substances: Challenges and opportunities. Integr. Environ. Assess. Manag. 2007, 3, 290–296. [Google Scholar] [CrossRef] [PubMed]
- Snape, J.R.; Maund, S.J.; Pickford, D.B.; Hutchinson, T.H. Ecotoxicogenomics: The challenge of integrating genomics into aquatic and terrestrial ecotoxicology. Aquat. Toxicol. 2004, 67, 143–154. [Google Scholar] [CrossRef] [PubMed]
- Field, C.B.; Behrenfeld, M.J.; Randerson, J.T.; Falkowski, P. Primary production of the biosphere: Integrating terrestrial and oceanic components. Science 1998, 281, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Noges, T.; Luup, H.; Feldmann, T. Primary production of aquatic macrophytes and their epiphytes in two shallow lakes (Peipsi and Vrtsjarv) in Estonia. Aquat. Ecol. 2010, 44, 83–92. [Google Scholar] [CrossRef]
- Bravo, A.G.; Loizeau, J.-L.; Dranguet, P.; Makri, S.; Björn, E.; Ungureanu, V.G.; Slaveykova, V.I.; Cosio, C. Persistent Hg contamination and occurrence of Hg methylating gene (hgcA) downstream a chlor-alkali plant in the Olt River (Romania). Environ. Sci. Pollut. Res. 2016, 23, 10529–10541. [Google Scholar] [CrossRef] [PubMed]
- Gregoire, D.S.; Poulain, A.J. A little bit of light goes a long way: The role of phototrophs on mercury cycling. Metallomics 2014, 6, 396–407. [Google Scholar] [CrossRef] [PubMed]
- Le Faucheur, S.; Campbell, P.G.C.; Fortin, C.; Slaveykova, V.I. Interactions between mercury and phytoplankton: Speciation, bioavailability, and internal handling. Environ. Toxicol. Chem. 2014, 33, 1211–1224. [Google Scholar] [CrossRef] [PubMed]
- Cosio, C.; Flück, R.; Regier, N.; Slaveykova, V.I. Effects of macrophytes on the fate of mercury in aquatic systems. Environ. Toxicol. Chem. 2014, 33, 1225–1237. [Google Scholar] [CrossRef] [PubMed]
- Benbassat, D.; Mayer, A.M. Volatilization of mercury by algae. Physiol. Plant. 1975, 33, 128–131. [Google Scholar] [CrossRef]
- Regier, N.; Larras, F.; Bravo, A.G.; Ungureanu, V.-G.; Amouroux, D.; Cosio, C. Mercury bioaccumulation in the aquatic plant Elodea nuttallii in the field and in microcosm: Accumulation in shoots from the water might involve copper transporters. Chemosphere 2013, 90, 595–602. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Li, G.; Bishopp, A.; Heenatigala, P.P.M.; Hu, S.; Chen, Y.; Wu, Z.; Kumar, S.; Duan, P.; Yao, L.; et al. A comparison of growth on mercuric chloride for three Lemnaceae species reveals differences in growth dynamics that effect their suitability for use in either monitoring or remediating Ecosystems Contaminated with Mercury. Front. Chem. 2018, 6, 112. [Google Scholar] [CrossRef] [PubMed]
- Lominchar, M.A.; Sierra, M.J.; Jimenez-Moreno, M.; Guirado, M.; Martin-Doimeadios, R.C.R.; Millan, R. Mercury species accumulation and distribution in Typha domingensis under real field conditions (Almaden, Spain). Environ. Sci. Pollut. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Wang, Q.; Zhang, Z.; Zhou, X. Mercury distribution along the food chain of a wetland ecosystem at Sanjiang Plain, Northeast China. Bull. Environ. Contam. Toxicol. 2017, 98, 162–166. [Google Scholar]
- Slaveykova, V.I.; Wilkinson, K.J. Physicochemical aspects of lead bioaccumulation by Chlorella vulgaris. Environ. Sci. Technol. 2002, 36, 969–975. [Google Scholar] [CrossRef] [PubMed]
- Boudou, A.; Delnomdedieu, M.; Georgescauld, D.; Ribeyre, F.; Saouter, E. Fundamental roles of biological barriers in mercury accumulation and transfer in fresh-water ecosystems—(Analysis at organism, organ, cell and molecular-levels). Water Air Soil Pollut. 1991, 56, 807–822. [Google Scholar] [CrossRef]
- Morel, F.M.M.; Kraepiel, A.M.L.; Amyot, M. The chemical cycle and bioaccumulation of mercury. Annu. Rev. Ecol. Syst. 1998, 29, 543–566. [Google Scholar] [CrossRef]
- Mason, R.P.; Reinfelder, J.R.; Morel, F.M.M. Uptake, toxicity, and trophic transfer of mercury in a coastal diatom. Environ. Sci. Technol. 1996, 30, 1835–1845. [Google Scholar] [CrossRef]
- Moye, H.A.; Miles, C.J.; Phlips, E.J.; Sargent, B.; Merritt, K.K. Kinetics and uptake mechanisms for monomethylmercury between freshwater algae and water. Environ. Sci. Technol. 2002, 36, 3550–3555. [Google Scholar] [CrossRef] [PubMed]
- Hassett, R.; Kosman, D.J. Evidence for Cu(II) reduction as a component of copper uptake by Saccharomyces cerevisae. J. Biol. Chem. 1995, 270, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Page, M.D.; Kropat, J.; Hamel, P.P.; Merchant, S.S. Two Chlamydomonas CTR copper transporters with a novel Cys-Met motif are localized to the plasma membrane and function in copper assimilation. Plant Cell 2009, 21, 928–943. [Google Scholar] [CrossRef] [PubMed]
- Larras, F.; Regier, N.; Planchon, S.; Poté, J.; Renaut, J.; Cosio, C. Physiological and proteomic changes suggest an important role of cell walls in the high tolerance to metals of Elodea nuttallii. J. Hazard. Mater. 2013, 263 Pt 2, 575–583. [Google Scholar] [CrossRef]
- Castro, R.; Pereira, S.; Lima, A.; Corticeiro, S.; Valega, M.; Pereira, E.; Duarte, A.; Figueira, E. Accumulation, distribution and cellular partitioning of mercury in several halophytes of a contaminated salt marsh. Chemosphere 2009, 76, 1348–1355. [Google Scholar] [CrossRef] [PubMed]
- Carrasco-Gil, S.; Alvarez-Fernandez, A.; Sobrino-Plata, J.; Millan, R.; Carpena-Ruiz, R.O.; Leduc, D.L.; Andrews, J.C.; Abadia, J.; Hernandez, L.E. Complexation of Hg with phytochelatins is important for plant Hg tolerance. Plant Cell Environ. 2011, 34, 778–791. [Google Scholar] [CrossRef] [PubMed]
- Beauvais-Fluck, R.; Slaveykova, V.I.; Cosio, C. Cellular toxicity pathways of inorganic and methyl mercury in the green microalga Chlamydomonas reinhardtii. Sci. Rep. 2017, 7, 8034. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.B.; Vajpayee, P.; Tripathi, R.D.; Rai, U.N.; Kumar, A.; Singh, N.; Behl, H.M.; Singh, S.P. Mercury bioaccumulation induces oxidative stress and toxicity to submerged macrophyte Potamogeton crispus L. Bull. Environ. Contam. Toxicol. 2000, 65, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Ferrat, L.; Romeo, M.; Gnassia-Barelli, M.; Pergent-Martini, C. Effects of mercury on antioxidant mechanisms in the marine phanerogam Posidonia oceanica. Dis. Aquat. Org. 2002, 50, 157–160. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-A.; Chi, W.-C.; Huang, T.-L.; Lin, C.-Y.; Quynh Nguyeh, T.T.; Hsiung, Y.-C.; Chia, L.-C.; Huang, H.-J. Mercury-induced biochemical and proteomic changes in rice roots. Plant Physiol. Biochem. 2012, 55, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Regier, N.; Beauvais-Flück, R.; Slaveykova, V.I.; Cosio, C. Elodea nuttallii exposure to mercury exposure under enhanced ultraviolet radiation: Effects on bioaccumulation, transcriptome, pigment content and oxidative stress. Aquat. Toxicol. 2016, 180, 218–226. [Google Scholar] [CrossRef] [PubMed]
- Siebman, C.; Velev, O.; Slaveykova, V. Two-dimensional algal collection and assembly by combining AC-Dielectrophoresis with fluorescence detection for contaminant-induced oxidative stress sensing. Biosensors 2015, 5, 319–336. [Google Scholar] [CrossRef] [PubMed]
- Patra, M.; Sharma, A. Mercury toxicity in plants. Bot. Rev. 2000, 66, 379–422. [Google Scholar] [CrossRef]
- Beauvais-Fluck, R.; Slaveykova, V.I.; Cosio, C. Effects of two-hour exposure to environmental and high concentrations of methylmercury on the transcriptome of the macrophyte Elodea nuttallii. Aquat. Toxicol. 2018, 194, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Samson, G.; Morissette, J.C.; Popovic, R. Determination of 4 apparent mercury interaction sites in photosystem-II by using a new modification of the Stern-Volmer analysis. Biochem. Biophys. Res. Commun. 1990, 166, 873–878. [Google Scholar] [CrossRef]
- Samson, G.; Popovic, R. Inhibitory effects of mercury on Photosystem-II photochemistry in Dunaliella tertiolecta under in vivo conditions. J. Photochem. Photobiol. B-Biol. 1990, 5, 303–310. [Google Scholar] [CrossRef]
- Sersen, F.; Kralova, K.; Bumbalova, A. Action of mercury on the photosynthetic apparatus of spinach chloroplasts. Photosynthetica 1998, 35, 551–559. [Google Scholar] [CrossRef]
- Singh, R.; Dubey, G.; Singh, V.; Srivastava, P.; Kumar, S.; Prasad, S. High light intensity augments mercury toxicity in cyanobacterium Nostoc muscorum. Biol. Trace Elem. Res. 2012, 149, 262–272. [Google Scholar] [CrossRef] [PubMed]
- Juneau, P.; Dewez, D.; Matsui, S.; Kim, S.-G.; Popovic, R. Evaluation of different algal species sensitivity to mercury and metolachlor by PAM-fluorometry. Chemosphere 2001, 45, 589–598. [Google Scholar] [CrossRef]
- Poschenrieder, C.; Cabot, C.; Martos, S.; Gallego, B.; Barcelo, J. Do toxic ions induce hormesis in plants? Plant Sci. 2013, 212, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Beauvais-Fluck, R.; Slaveykova, V.I.; Cosio, C. Transcriptomic and physiological responses of the green microalga Chlamydomonas reinhardtii during short-term exposure to subnanomolar methylmercury concentrations. Environ. Sci. Technol. 2016, 50, 7126–7134. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zeng, Y.; Qu, J.Y.; Wang, W.-X. Mercury effects on Thalassiosira weissflogii: Applications of two-photon excitation chlorophyll fluorescence lifetime imaging and flow cytometry. Aquat. Toxicol. 2012, 110–111, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Mortimer, D.C.; Czuba, M. Structural damage to leaf chloroplasts of Elodea densa caused by methylmercury accumulated from water. Ecotoxicol. Environ. Saf. 1982, 6, 193–195. [Google Scholar] [CrossRef]
- Küpper, H.; Küpper, F.; Spiller, M. Environmental relevance of heavy metal-substituted chlorophylls using the example of water plants. J. Exp. Bot. 1996, 47, 259–266. [Google Scholar] [CrossRef]
- Rai, P.K.; Tripathi, B.D. Comparative assessment of Azolla pinnata and Vallisneria spiralis in Hg removal from GB Pant Sagar of Singrauli Industrial region, India. Environ. Monit. Assess. 2009, 148, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Chandra, P. Bioaccumulation and toxicity of mercury in rooted-submerged macrophyte Vallisneria spiralis. Environ. Pollut. 1998, 103, 327–332. [Google Scholar] [CrossRef]
- Patra, M.; Bhowmik, N.; Bandopadhyay, B.; Sharma, A. Comparison of mercury, lead and arsenic with respect to genotoxic effects on plant systems and the development of genetic tolerance. Environ. Exp. Bot. 2004, 52, 199–223. [Google Scholar] [CrossRef]
- Schaumlöffel, D. The Position of Metallomics within Other Omics Fields. In Metallomics Analytical Techniques and Speciation Methods; John Wiley & Sons: Hoboken, NJ, USA, 2016; pp. 3–16. [Google Scholar]
- Schirmer, K.; Fischer, B.B.; Madureira, D.J.; Pillai, S. Transcriptomics in ecotoxicology. Anal. Bioanal. Chem. 2010, 397, 917–923. [Google Scholar] [CrossRef] [PubMed]
- Heidenreich, B.; Mayer, K.; Sandermann, H.; Ernst, D. Mercury-induced genes in Arabidopsis thaliana: Identification of induced genes upon long-term mercuric ion exposure. Plant Cell Environ. 2001, 24, 1227–1234. [Google Scholar] [CrossRef]
- Venkatachalam, P.; Srivastava, A.K.; Raghothama, K.G.; Sahi, S.V. Genes induced in response to mercury-ion-exposure in heavy metal hyperaccumulator Sesbania drummondii. Environ. Sci. Technol. 2009, 43, 843–850. [Google Scholar] [CrossRef] [PubMed]
- Montero-Palmero, M.B.; Martín-Barranco, A.; Escobar, C.; Hernández, L.E. Early transcriptional responses to mercury: A role for ethylene in mercury-induced stress. New Phytol. 2014, 201, 116–130. [Google Scholar] [CrossRef] [PubMed]
- Regier, N.; Baerlocher, L.; Munsterkotter, M.; Farinelli, L.; Cosio, C. Analysis of the Elodea nuttallii transcriptome in response to mercury and cadmium pollution: Development of sensitive tools for rapid ecotoxicological testing. Environ. Sci. Technol. 2013, 47, 8825–8834. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, Z.M. Mercury toxicity, molecular response and tolerance in higher plants. Biometals 2012, 25, 847–857. [Google Scholar] [CrossRef] [PubMed]
- Gouveia, D.; Chaumot, A.; Charnot, A.; Almunia, C.; Francois, A.; Navarro, L.; Armengaud, J.; Salvador, A.; Geffard, O. Ecotoxico-proteomics for aquatic environmental monitoring: First in situ application of a new proteomics-based multibiomarker assay using caged amphipods. Environ. Sci. Technol. 2017, 51, 13417–13426. [Google Scholar] [CrossRef] [PubMed]
- Lindemann, C.; Fiksen, O.; Andersen, K.H.; Aksnes, D.L. Scaling laws in phytoplankton nutrient uptake affinity. Front. Mar. Sci. 2016, 3, 26. [Google Scholar] [CrossRef]
- Hossain, Z.; Makino, T.; Komatsu, S. Proteomic study of beta-aminobutyric acid-mediated cadmium stress alleviation in soybean. J. Proteom. 2012, 75, 4151–4164. [Google Scholar] [CrossRef] [PubMed]
- Hossain, Z.; Komatsu, S. Contribution of proteomic studies towards understanding plant heavy metal stress response. Front. Plant Sci. 2013, 3, 310. [Google Scholar] [CrossRef] [PubMed]
- Feiz, L.; Irshad, M.; Pont-Lezica, R.F.; Canut, H.; Jamet, E. Evaluation of cell wall preparations for proteomics: A new procedure for purifying cell walls from Arabidopsis hypocotyls. Plant Methods 2006, 2, 10. [Google Scholar] [CrossRef] [PubMed]
- Witzel, K.; Shahzad, M.; Matros, A.; Mock, H.-P.; Mühling, K.H. Comparative evaluation of extraction methods for apoplastic proteins from maize leaves. Plant Methods 2011, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Clemens, S.; Ma, J.F. Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu. Rev. Plant Biol. 2016, 67, 489–512. [Google Scholar] [CrossRef] [PubMed]

| [Hg], Duration | Method | Main Results | Physiology | Reference |
|---|---|---|---|---|
| MACROPHYTES | ||||
| Oryza sativa | ||||
| 25 µM IHg, 3 h | P: 2-D differential gel electrophoresis | 14 up-: e.g., protein disulfide isomerase, peroxidase, ascorbate peroxidase, 11 down-regulated proteins: e.g., fructokinase, cysteine synthase, enolase | lipid peroxidation ↑ root growth ↓ | ([63]) |
| Elodea nuttallii | ||||
| 0.35 nM IHg, 24 h | P: 2D-differential gel electrophoresis | 4 up-, 18 down-regulated identified proteins: photosynthesis (e.g., light harvesting complex), defense/stress (e.g., POD), lignin synthesis (e.g., phenylcoumaran benzylic ether reductase), glycolysis, carbon fixation, cytoskeleton organization (e.g., actin) | lignification of cell walls ↑ | ([33]) |
| 0.1 nM MeHg, 24 h | no effect | no effect | ||
| 0.004, 0.4 and 4 µM IHg, 24 h | T: RNA-Seq | dose-dependent up-regulation: e.g., interaction with the environment, HSP70 dose-dependent down-regulation: N-assimilation, metal transport, S metabolism | none measured | ([62]) |
| 0.4 nM IHg, 24 h | T: RNA-Seq | 79 up-, 48 down-regulated contigs no significant enriched biological pathway | POD activity ↓ SOD activity ↑ | ([40]) |
| 0.4 µM IHg, 24 h | 4472 up-: gene expression, fatty acid oxidation, 2273 down-regulated contigs: chloroplast, photosynthesis, chlorophyll biosynthesis | |||
| 1 and 10 nM MeHg, 2 h | T: RNA-Seq | 4389 and 16853 regulated genes: sugar, amino acid and secondary metabolism (e.g., cinnamic acid, flavonoids) at both concentrations. Genes coding for photosynthesis, membrane integrity, metal homeostasis, water transport and anti-oxidative enzymes at 10−8 M. | POD activity ↑ anthocyanin ↑ | ([43]) |
| 10 pM to 10 nM IHg and MeHg, 2 h | T: RNA-Seq | IHg: Up to 1677, MeHg up to 18,557 regulated genes: energy metabolism, development, transport, secondary metabolism. No specific GO categories for MeHg or IHg. | IHg: chlorophyll ↓ MeHg: antioxidant ↑ | ([7]) |
| Field study 12 pM THg, 2 h | T: RNA-Seq | 8700 regulated genes: anti-oxidative response, gene regulation, energy metabolism, secondary metabolism, hormone metabolism, transport and stress, | no measurable bioaccumulation | ([6]) |
| ALGAE | ||||
| Chlamydomonas reinhardtii | ||||
| 0.036 nM MeHg 2 h | T: RNA-Seq | 2080 up-, 1868 down-regulated: cell motility, transport, fatty acid degradation, phospholipids biosynthesis, cell organization, energy metabolism, RedOx, secondary metabolism | Photosynthesis efficiency ↑ | ([50]) |
| 0.37 nM MeHg 2 h | 2415 up-, 2369 down-regulated: cell motility, transport, fatty acid degradation and synthesis, energy metabolism, RedOx, secondary metabolsim | |||
| 10 pM to 10 nM IHg and MeHg, 2 h | T: RNA-Seq | Stronger regulation by MeHg than IHg; 8461 regulated: gene expression (nucleotide to protein synthesis, signaling), cell processes (motility, division, development), energy metabolism (photosynthesis, sugar metabolism), lipid metabolism, amino acid metabolism, stress and transport. No specific GO category for IHg. | Photosynthesis efficiency ↑ MeHg: ROS ↑ | ([36]) |
| Field study 12 pM THg, 2 h | T: RNA-Seq | regulated genes energy metabolism, cell motility, transport, amino acids metabolism, and other metabolisms (lipids, hormones, vitamins, isoprenoids), gene regulation, stress and RedOx, cell structure, major CHO metabolism, lipid metabolism | no measurable bioaccumulation | ([6]) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Beauvais-Flück, R.; Slaveykova, V.I.; Cosio, C. Molecular Effects of Inorganic and Methyl Mercury in Aquatic Primary Producers: Comparing Impact to A Macrophyte and A Green Microalga in Controlled Conditions. Geosciences 2018, 8, 393. https://doi.org/10.3390/geosciences8110393
Beauvais-Flück R, Slaveykova VI, Cosio C. Molecular Effects of Inorganic and Methyl Mercury in Aquatic Primary Producers: Comparing Impact to A Macrophyte and A Green Microalga in Controlled Conditions. Geosciences. 2018; 8(11):393. https://doi.org/10.3390/geosciences8110393
Chicago/Turabian StyleBeauvais-Flück, Rebecca, Vera I. Slaveykova, and Claudia Cosio. 2018. "Molecular Effects of Inorganic and Methyl Mercury in Aquatic Primary Producers: Comparing Impact to A Macrophyte and A Green Microalga in Controlled Conditions" Geosciences 8, no. 11: 393. https://doi.org/10.3390/geosciences8110393
APA StyleBeauvais-Flück, R., Slaveykova, V. I., & Cosio, C. (2018). Molecular Effects of Inorganic and Methyl Mercury in Aquatic Primary Producers: Comparing Impact to A Macrophyte and A Green Microalga in Controlled Conditions. Geosciences, 8(11), 393. https://doi.org/10.3390/geosciences8110393






