A Novel 13 bp Deletion within the NR6A1 Gene Is Significantly Associated with Growth Traits in Donkeys
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Animals and Data Collection
2.3. DNA Extraction and DNA Pool Construction
2.4. PCR Amplification and Sequenci
2.5. Genotyping
2.6. Statistical Analysis
3. Results
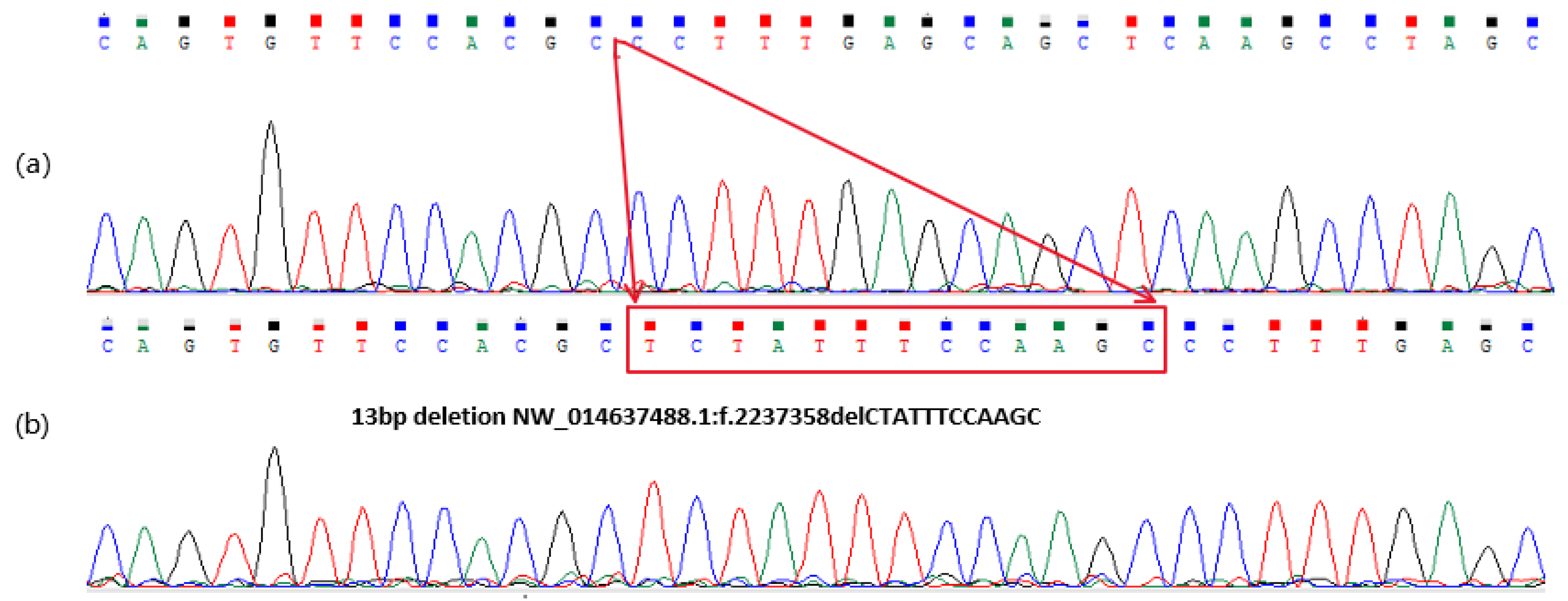
3.1. Sequence Variants Identified in the Donkey NR6A1 Gene
3.2. Polymorphisms and Genetic Diversity
3.3. Association Analysis of Polymorphisms with Growth Traits of the Donkey
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Gudbjartsson, D.F.; Walters, G.B.; Thorleifsson, G.; Stefansson, H.; Halldorsson, B.V.; Zusmanovich, P.; Sulem, P.; Thorlacius, S.; Gylfason, A.; Steinberg, S.; et al. Many sequence variants affecting diversity of adult human height. Nat. Genet. 2008, 40, 609–615. [Google Scholar] [CrossRef] [PubMed]
- Lango, A.H.; Estrada, K.; Lettre, G.; Berndt, S.; Weedon, M.N.; Rivadeneira, F.; Willer, C.J.; Jackson, A.U.; Vedantam, S.; Raychaudhuri, S.; et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010, 467, 832–838. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rubin, C.J.; Megens, H.J.; Martinez, B.A.; Maqbool, K.; Andersson, L. Strong signatures of selection in the domestic pig genome. Proc. Natl. Acad. Sci. USA 2012, 109, 19529–19536. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borchers, N.; Reinsch, N.; Kalm, E. The number of ribs and vertebrae in a Pietrain cross:variation, heritability and effects on performance traits. J. Anim. Breed. Genet. 2004, 121, 392–403. [Google Scholar] [CrossRef]
- Qiu-Fei, J.; Long, H.; Yuan, F.; Juan, Z.; Yu, W.; Liang, W.U.; Gu, Y.L. Polymorphism in UCP3 Gene and Its Association with Growth Traits in Hybrid Simmental Cattle (Bos taurus). J. Agric. Biotechnol. 2018, 26, 642–651. [Google Scholar]
- Zhang, L.; Zhang, A.L.; Zhang, L.Z.; Zhang, Q.; Zhang, C.L.; Lan, X.Y.; Chen, H. Association analysis between variants in bovine NPY gene and growth traits in Nanyang cattle (Bos taurus). Gen. Comp. Endocr. 2011, 170, 189–192. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zan, L.; Xin, Y.; Tian, W.; Li, L.; Wang, H. ZBTB38 gene polymorphism associated with body measurement traits in native Chinese cattle breeds. Gene 2013, 513, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Duijvesteijn, N.; Veltmaat, J.M.; Knol, E.F.; Harlizius, B. High-resolution association mapping of number of teats in pigs reveals regions controlling vertebral development. BMC Genomics 2014, 15, 542. [Google Scholar] [CrossRef]
- Burgos, C.; Latorre, P.; Altarriba, J.; Carrodeguas, J.A.; Varona, L.; López-Buesa, P. Allelic frequencies of NR6A1 and VRTN, two genes that affect vertebrae number in diverse pig breeds: A study of the effects of the VRTN insertion on phenotypic traits of a Duroc × Landrace-Large White cross. Meat Sci. 2015, 100, 150–155. [Google Scholar] [CrossRef]
- Zhang, X.; Li, C.; Li, X.; Liu, Z.; Ni, W.; Cao, Y.; Yao, Y.; Islamov, E.; Wei, J.; Hou, X.; et al. Association analysis of polymorphism in the NR6A1 gene with the lumbar vertebrae number traits in sheep. Genes Genom. 2019, 1, 1–7. [Google Scholar] [CrossRef]
- Gu, P.; LeMenuet, D.; Chung, A.C.; Mancini, M.; Wheeler, D.A.; Cooney, A.J. Orphan Nuclear Recepor GCNF Is Required for the Repression of Pluripotenncy Genes during Retinoic Acid-Induced Embryonic Stem Cell Differentiation. Mol. Cell. Biol. 2005, 25, 8507–8519. [Google Scholar] [CrossRef] [PubMed]
- Noriko, S.; Mitsumasa, K.; Ken-ichi, A. The orphan nulear receptor GCNF recruits DNA methyltransferase for Otc-3/4 silencing. BBRC 2006, 344, 845–851. [Google Scholar]
- Mikawa, S.; Hayashi, T.; Nii, M.; Shimanuki, S.; Morozumi, T.; Awata, T. Two quantitative trait loci on Sus scrofa chromosomes 1 and 7 affecting the number of vertebrae. J. Anim. Sci. 2005, 83, 2247–2254. [Google Scholar] [CrossRef] [PubMed]
- Mikawa, S.; Morozumi, T.; Shimanuki, S.I.; Hayashi, T.; Uenishi, H.; Domukai, M.; Naohiko, O.; Takashi, A. Fine mapping of a swine quantitative trait locus for number of vertebrae and analysis of an orphan nuclear receptor, germ cell nuclear factor (NR6A1). Gent. Res. 2007, 17, 586. [Google Scholar] [CrossRef] [PubMed]
- Klomtong, P.; Chaweewan, K.; Phasuk, Y.; Duangjinda, M. MC1R, KIT, IGF2, and NR6A1 as markers for genetic differentiation in Thai native, wild boars, and Duroc and Chinese Meishan pigs. Genet. Mol. Res. 2015, 14, 12723–12732. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Meng, W.; Jiao, Y.; Xiang, Z.; Sanping, X.; Bang, L. Association of polymorphisms in NR6A1, PLAG 1 and VRTN with the number of vertebrae in Chinese Tongcheng × Large White crossbred pigs. Anim. Genet. 2018, 49. [Google Scholar] [CrossRef]
- Yang, G.; Ren, J.; Zhang, Z.; Huang, L. Genetic evidence for the introgression of Western NR6A1 haplotype into Chinese Licha breed associated with increased vertebral number. Anim. Genet. 2009, 40, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Braat, A.K.; Zandbergen, M.A.; De Vries, E.; Goos, H.J. Cloning and expression of the zebrafish germ cell nuclear factor. Mol. Reprod. Dev. 1999, 53, 369–375. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratories: New York, NY, USA, 1989; pp. 179–187. [Google Scholar]
- Anglani, F.; Piccl, L.; Camporese, C.; Zacchello, F.R. Heteroduplex formation in polymerase chain reaction. Am. J. Hum. Genet. 1990, 47, 169–170. [Google Scholar]
- Wang, H.; Wang, X.; Xu, X.; Kyba, M.; Cooney, A.J. Germ Cell Nuclear Factor (GCNF) Represses Oct4 Expression and Globally Modulates Gene Expression in Human Embryonic Stem (hES) Cells. J. Biol. Chem. 2016, 291, 8644–8652. [Google Scholar] [CrossRef] [Green Version]
- Gu, P.; Xu, X.; Le Menuet, D.; Chung, A.C.; Cooney, A.J. Differential Recruitment of Methyl CpG-Binding Domain Factors and DNA Methyltransferases by the Orphan Receptor Germ Cell Nuclear Factor Initiates the Repression and Silencing of Oct4. Stem Cell. 2011, 29, 1041–1051. [Google Scholar] [CrossRef] [PubMed]
- Akamatsu, W.; Deveale, B.; Okano, H.; Cooney, A.J.; van der Kooy, D. Suppression of Oct4 by germ cell nuclear factor restricts pluripotency and promotes neural stem cell development in the early neural lineage. J. Neurosci. 2009, 29, 2113. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, X.; Archer, T.K.; Zwaka, T.P.; Cooney, A.J. GCNF-Dependent Activation of Cyclin D1 Expression via Repression of Mir302a During ESC Differentiation. Stem Cell. 2014, 32, 1527–1537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, W.; Zhao, D.; Han, S.; Tian, Z.; Zhang, L. CYP3A4*1G regulates CYP3A4 intron 10 enhancer and promoter activity in an allelicdependent manner. Int. J. Clin. Pharm. Th. 2015, 53, 647–657. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Sadee, W. CYP3A4 intronic SNP rs35599367 (CYP3A4*22) alters RNA splicing. Pharnacogenet. Genom. 2016, 26, 40–43. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Locus | Breeds | N | Genotypic Frequencies | Allelic Frequencies | HWE | Population Parameters | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| II | ID | DD | I | D | p | Ho | He | Ne | PIC | |||
| NR6A1 (NW-014637488) | DZ | 346 | 0.84 (n = 291) | 0.15 (n = 53) | 0.01 (n = 2) | 0.92 | 0.08 | 0.81 | 0.849 | 0.151 | 1.178 | 0.14 |
| GZ | 62 | 0.82 (n = 51) | 0.16 (n = 10) | 0.02 (n = 1) | 0.90 | 0.10 | 0.54 | 0.825 | 0.175 | 1.212 | 0.16 | |
| Locus | Types | Breeds | DZ | GZ |
|---|---|---|---|---|
| NR6A1 (NW-014637488) | Genotypic frequencies | DZ | – | χ2 = 0.81 |
| GZ | (p > 0.05) | – | ||
| Allelic frequencies | DZ | – | χ2 = 0.06 | |
| GZ | (p > 0.05) | – |
| Locus | Breeds | Growth Traits | Observed Genotypes (LSM a ± SE) | p | ||
|---|---|---|---|---|---|---|
| II | ID | DD | ||||
| NR6A1 (NW-014637488) | DZ | Body height | 136.56 ± 6.94 a (n = 291) | 134.98 ± 5.11 b (n = 53) | 133.00 ± 7.07 (n = 2) | 0.019 |
| Body length | 135.78 ± 8.17 a (n = 291) | 135.32 ± 6.38 b (n = 53) | 133.00 ± 5.65 (n = 2) | 0.025 | ||
| Chest circumference | 148.66 ± 8.89 a (n = 291) | 148.11 ± 6.90 b (n = 53) | 157.75 ± 10.96 (n = 2) | 0.048 | ||
| Chest depth | 55.26 ± 3.86 a (n = 291) | 54.83 ± 2.61 b (n = 53) | 56.00 ± 4.24 (n = 2) | 0.002 | ||
| GZ | Body height | 135.68 ± 6.77 a (n = 51) | 133.10 ± 4.53 b (n = 10) | 0 | 0.048 | |
| Body length | 128.77 ± 7.07 a (n = 51) | 125.70 ± 5.27 a (n = 10) | 0 | 0.467 | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fang, X.; Lai, Z.; Liu, J.; Zhang, C.; Li, S.; Wu, F.; Zhou, Z.; Lei, C.; Dang, R. A Novel 13 bp Deletion within the NR6A1 Gene Is Significantly Associated with Growth Traits in Donkeys. Animals 2019, 9, 681. https://doi.org/10.3390/ani9090681
Fang X, Lai Z, Liu J, Zhang C, Li S, Wu F, Zhou Z, Lei C, Dang R. A Novel 13 bp Deletion within the NR6A1 Gene Is Significantly Associated with Growth Traits in Donkeys. Animals. 2019; 9(9):681. https://doi.org/10.3390/ani9090681
Chicago/Turabian StyleFang, Xiya, Zhenyu Lai, Jie Liu, Chunlan Zhang, Shipeng Li, Fei Wu, Zihui Zhou, Chuzhao Lei, and Ruihua Dang. 2019. "A Novel 13 bp Deletion within the NR6A1 Gene Is Significantly Associated with Growth Traits in Donkeys" Animals 9, no. 9: 681. https://doi.org/10.3390/ani9090681
APA StyleFang, X., Lai, Z., Liu, J., Zhang, C., Li, S., Wu, F., Zhou, Z., Lei, C., & Dang, R. (2019). A Novel 13 bp Deletion within the NR6A1 Gene Is Significantly Associated with Growth Traits in Donkeys. Animals, 9(9), 681. https://doi.org/10.3390/ani9090681





