Variation of Detailed Protein Composition of Cow Milk Predicted from a Large Database of Mid-Infrared Spectra
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. MIRS Calibration Models
2.3. Data Editing and Statistical Analyses
3. Results and Discussion
3.1. Descriptive Statistics
3.2. Breed Effect
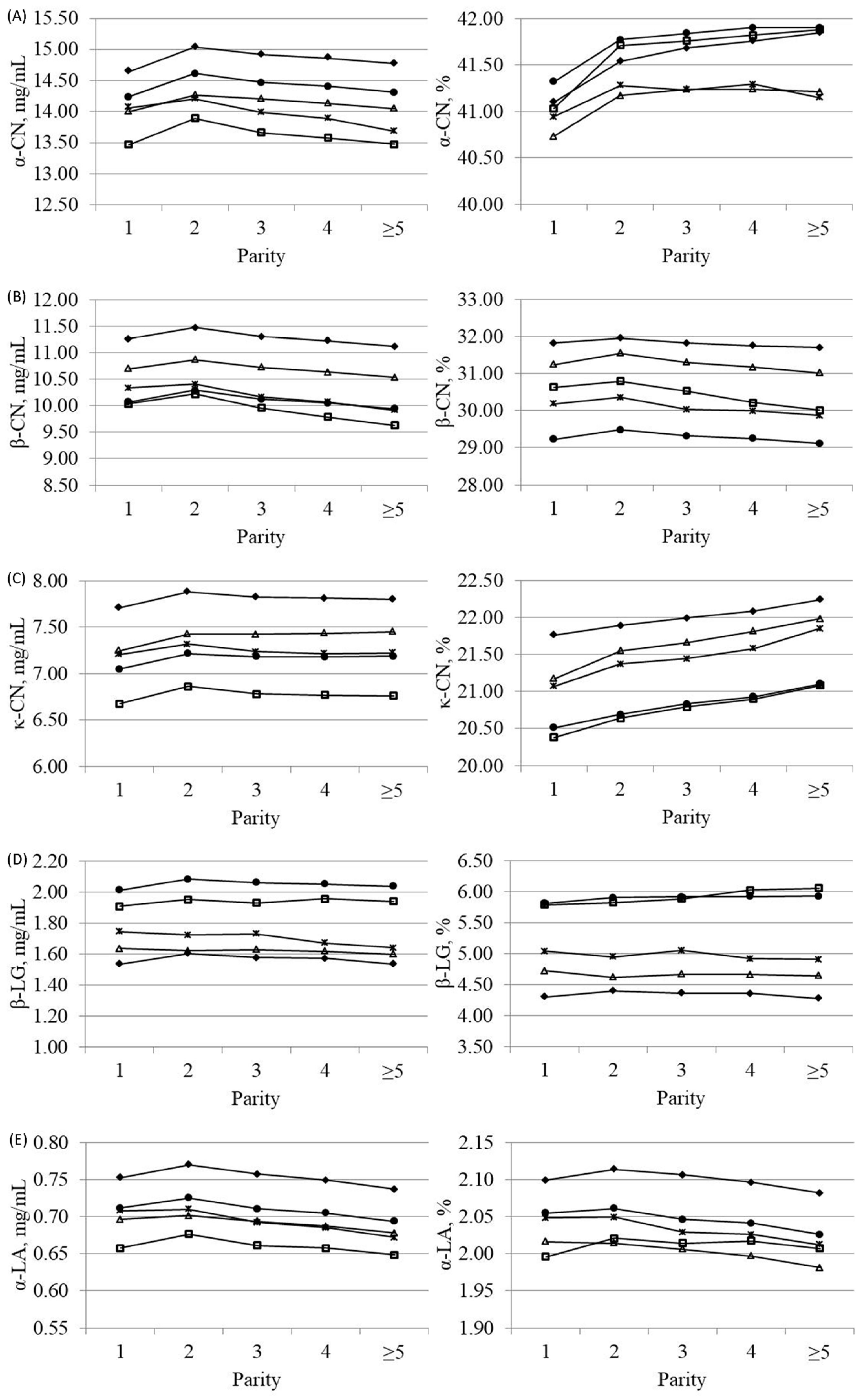
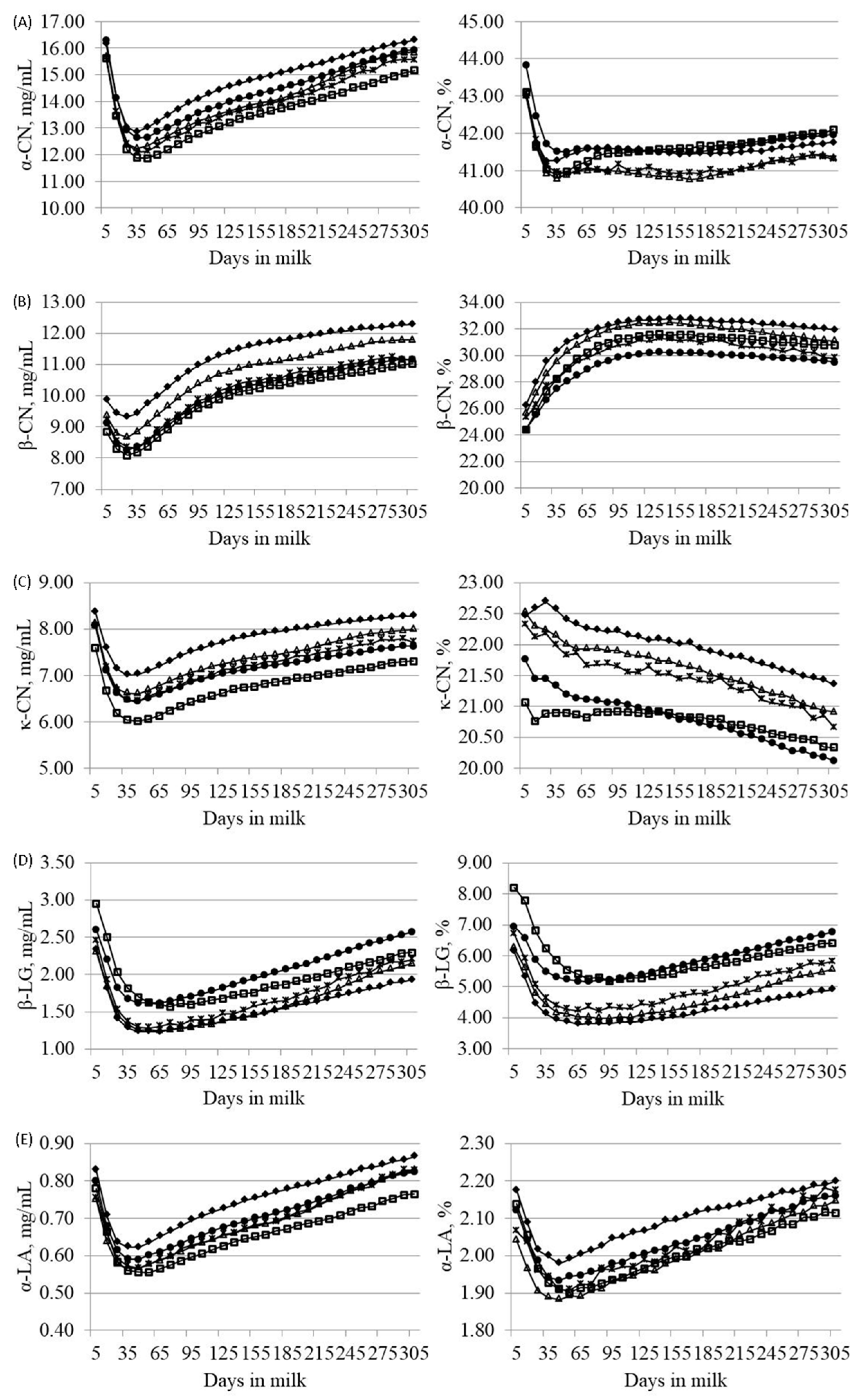
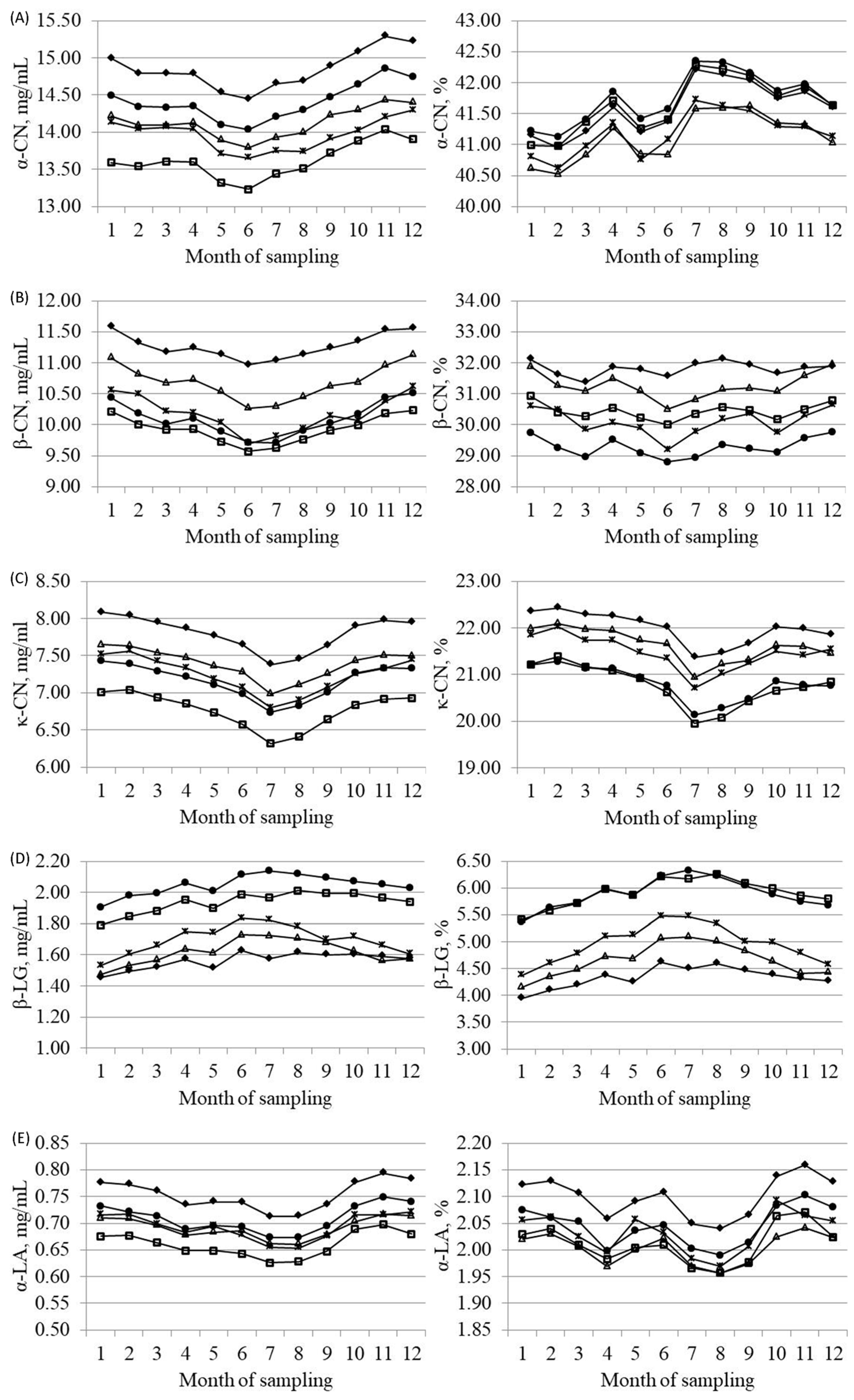
3.3. Effects of Parity, Lactation Stage and Season
3.4. Correlations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Singhal, S.; Baker, R.D.; Baker, S.S. A comparison of the nutritional value of cow’s milk and nondairy beverages. J. Pediatr. Gastroenterol. Nutr. 2017, 64, 799–805. [Google Scholar] [CrossRef]
- Mills, S.; Ross, R.P.; Hill, C.; Fitzgerald, G.F.; Stanton, C. Milk intelligence: Mining milk for bioactive substances associated with human health. Int. Dairy J. 2011, 21, 377–401. [Google Scholar] [CrossRef]
- Caroli, A.; Poli, A.; Ricotta, D.; Banfi, G.; Cocchi, D. Invited review: Dairy intake and bone health: A viewpoint from the state of the art. J. Dairy Sci. 2011, 94, 5249–5262. [Google Scholar] [CrossRef]
- Visentin, G.; McParland, S.; De Marchi, M.; McDermott, A.; Fenelon, M.A.; Penasa, M.; Berry, D.P. Processing characteristics of dairy cow milk are moderately heritable. J. Dairy Sci. 2017, 100, 6343–6355. [Google Scholar] [CrossRef]
- Jõudu, I.; Henno, M.; Kaart, T.; Püssa, T.; Kärt, O. The effect of milk protein contents on the rennet coagulation properties of milk from individual dairy cows. Int. Dairy J. 2008, 18, 964–967. [Google Scholar] [CrossRef]
- Marziali, A.S.; Ng-Kwai-Hang, K.F. Effects of milk composition and genetic polymorphism on coagulation properties of milk. J. Dairy Sci. 1986, 69, 1793–1798. [Google Scholar] [CrossRef]
- Tiezzi, F.; Valente, B.D.; Cassandro, M.; Maltecca, C. Causal relationships between milk quality and coagulation properties in Italian Holstein-Friesian dairy cattle. Genet. Sel. Evol. 2015, 47, 45. [Google Scholar] [CrossRef]
- Miglior, F.; Muir, B.L.; Van Doormaal, B.J. Selection indices in Holstein cattle of various countries. J. Dairy Sci. 2005, 88, 1255–1263. [Google Scholar] [CrossRef]
- Bonfatti, V.; Vicario, D.; Lugo, A.; Carnier, P. Genetic parameters of measures and population-wide infrared predictions of 92 traits describing the fine composition and technological properties of milk in Italian Simmental cattle. J. Dairy Sci. 2017, 100, 5526–5540. [Google Scholar] [CrossRef]
- Niero, G.; Penasa, M.; De Marchi, M.; Visentin, G.; Cassandro, M. Study of milk protein composition and coagulation properties of Burlina local cattle breed. Poljoprivreda 2015, 21, 101–104. [Google Scholar] [CrossRef]
- Niero, G.; Penasa, M.; Gottardo, P.; Cassandro, M.; De Marchi, M. Short communication: Selecting the most informative mid-infrared spectra wavenumbers to improve the accuracy of prediction models for detailed milk protein content. J. Dairy Sci. 2016, 99, 1853–1858. [Google Scholar] [CrossRef]
- Visentin, G.; De Marchi, M.; Berry, D.P.; McDermott, A.; Fenelon, M.A.; Penasa, M.; McParland, S. Factors associated with milk processing characteristics predicted by mid-infrared spectroscopy in a large database of dairy cows. J. Dairy Sci. 2017, 100, 3293–3304. [Google Scholar] [CrossRef] [PubMed]
- De Marchi, M.; Toffanin, V.; Cassandro, M.; Penasa, M. Invited review: Mid-infrared spectroscopy as phenotyping tool for milk traits. J. Dairy Sci. 2014, 97, 1171–1186. [Google Scholar] [CrossRef] [PubMed]
- Grelet, C.; Fernández Pierna, J.A.; Dardenne, P.; Baeten, V.; Dehareng, F. Standardization of milk mid-infrared spectra from a European dairy network. J. Dairy Sci. 2015, 98, 2150–2160. [Google Scholar] [CrossRef] [PubMed]
- McDermott, A.; De Marchi, M.; Berry, D.P.; Visentin, G.; Fenelon, M.A.; Lopez-Villalobos, N.; McParland, S. Cow and environmental factors associated with protein fractions and free amino acids predicted using mid-infrared spectroscopy in bovine milk. J. Dairy Sci. 2017, 100, 6272–6284. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.P.; Ferrand, M.; Gelé, M.; Pourchet, D.; Miranda, G.; Martin, P.; Brochard, M.; Boichard, D. Short communication: Genetic parameters for milk protein composition predicted using mid-infrared spectroscopy in the French Montbéliarde, Normande, and Holstein dairy cattle breeds. J. Dairy Sci. 2017, 100, 6371–6375. [Google Scholar] [CrossRef]
- Juhl, H.V. Method for compensating amplitude drift in a spectrometer and spectrometer performing said method. U.S. Patent No. US9606050B2, 2017. [Google Scholar]
- Maurmayr, A.; Cecchinato, A.; Grigoletto, L.; Bittante, G. Detection and quantification of αs1, αs2-, β-, κ-casein, α-lactalbumin, β-lactoglobulin and lactoferrin in bovine milk by reverse-phase high-performance liquid chromatography. Agric. Conspec. Sci. 2013, 78, 201–205. [Google Scholar]
- Gottardo, P.; De Marchi, M.; Cassandro, M.; Penasa, M. Technical note: Improving the accuracy of mid-infrared prediction models by selecting the most informative wavelengths. J. Dairy Sci. 2015, 98, 4168–4173. [Google Scholar] [CrossRef]
- De Marchi, M.; Dal Zotto, R.; Cassandro, M.; Bittante, G. Milk coagulation ability of five dairy cattle breeds. J. Dairy Sci. 2007, 90, 3986–3992. [Google Scholar] [CrossRef]
- Visentin, G.; Penasa, M.; Niero, G.; Cassandro, M.; De Marchi, M. Phenotypic characterisation of major mineral composition predicted by mid-infrared spectroscopy in cow milk. Ital. J. Anim. Sci. 2018, 17, 549–556. [Google Scholar] [CrossRef]
- Penasa, M.; Tiezzi, F.; Sturaro, A.; Cassandro, M.; De Marchi, M. A comparison of the predicted coagulation characteristics and composition of milk from multi-breed herds of Holstein-Friesian, Brown Swiss and Simmental cows. Int. Dairy J. 2014, 35, 6–10. [Google Scholar] [CrossRef]
- Bobe, G.; Beitz, D.C.; Freeman, A.E.; Lindberg, G.L. Separation and quantification of bovine milk proteins by reversed-phase high-performance liquid chromatography. J. Agric. Food Chem. 1998, 46, 458–463. [Google Scholar] [CrossRef] [PubMed]
- Niero, G.; Visentin, G.; Ton, S.; De Marchi, M.; Penasa, M.; Cassandro, M. Phenotypic characterisation of milk technological traits, protein fractions, and major mineral and fatty acid composition of Burlina cattle breed. Ital. J. Anim. Sci. 2016, 15, 576–583. [Google Scholar] [CrossRef]
- Walker, G.P.; Dunshea, F.R.; Doyle, P.T. Effects of nutrition and management on the production and composition of milk fat and protein: A review. Aust. J. Agric. Res. 2004, 55, 1009–1028. [Google Scholar] [CrossRef]
- Golkar, A.; Milani, J.M.; Vasiljevic, T. Altering allergenicity of cow’s milk by food processing for applications in infant formula. Crit. Rev. Food Sci. Nutr. 2019, 59, 159–172. [Google Scholar] [CrossRef]
- Cipolat-Gotet, C.; Cecchinato, A.; Malacarne, M.; Bittante, G.; Summer, A. Variations in milk protein fractions affect the efficiency of the cheese-making process. J. Dairy Sci. 2018, 101, 8788–8804. [Google Scholar] [CrossRef] [PubMed]
- Schopen, G.C.B.; Heck, J.M.L.; Bovenhuis, H.; Visker, M.H.P.W.; van Valenberg, H.J.F.; van Arendonk, J.A.M. Genetic parameters for major milk proteins in Dutch Holstein-Friesians. J. Dairy Sci. 2009, 92, 1182–1191. [Google Scholar] [CrossRef]
- Manuelian, C.L.; Penasa, M.; Visentin, G.; Cassandro, M.; De Marchi, M. Phenotypic analysis of milk coagulation properties and mineral content of Pinzgauer cattle breed. Arch. Anim. Breed. 2018, 61, 215–220. [Google Scholar] [CrossRef]
- Visentin, G.; Penasa, M.; Gottardo, P.; Niero, G.; Isaia, M.; Cassandro, M.; De Marchi, M. Milk coagulation properties of cattle breeds reared in Alpine area. Poljoprivreda 2015, 21, 237–240. [Google Scholar] [CrossRef]
- Maurmayr, A.; Pegolo, S.; Malchiodi, F.; Bittante, G.; Cecchinato, A. Milk protein composition in purebred Holsteins and in first/second-generation crossbred cows from Swedish Red, Montbeliarde and Brown Swiss bulls. Animal 2018, 12, 2214–2220. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Boeren, S.; Hageman, J.A.; van Hooijdonk, T.; Vervoort, J.; Hettinga, K. Perspective on calf and mammary gland development through changes in the bovine milk proteome over a complete lactation. J. Dairy Sci. 2015, 98, 5362–5373. [Google Scholar] [CrossRef]
- Bernabucci, U.; Basiricò, L.; Morera, P.; Dipasquale, D.; Vitali, A.; Cappelli, F.P.; Calamari, L. Effect of summer season on milk protein fractions in Holstein cows. J. Dairy Sci. 2015, 95, 1815–1827. [Google Scholar] [CrossRef]
- Niero, G.; Koczura, M.; De Marchi, M.; Currò, S.; Kreuzer, M.; Turille, G.; Berard, J. Are cheese-making properties of dual purpose cattle impaired by highland grazing? A case study using Aosta Red Pied cows. Ital. J. Anim. Sci. 2018, 17, 827–834. [Google Scholar] [CrossRef]
- Daniel, J.B.; Friggens, N.C.; Chapoutot, P.; Van Laar, H.; Sauvant, D. Milk yield and milk composition responses to change in predicted net energy and metabolizable protein: A meta-analysis. Animal 2016, 10, 1975–1985. [Google Scholar] [CrossRef]
- Mackle, T.R.; Bryant, A.M.; Petch, S.F.; Hill, J.P.; Auldist, M.J. Nutritional influences on the composition of milk from cows of different protein phenotypes in New Zealand. J. Dairy Sci. 1999, 82, 172–180. [Google Scholar] [CrossRef]
| Traits | Mean | SD | Range | CV (%) | σ2c (%) | σ2h (%) |
|---|---|---|---|---|---|---|
| Milk yield (kg/day) | 23.45 | 7.41 | 44.70 | 31.61 | 24.63 | 35.09 |
| Milk composition (%) | ||||||
| Fat | 4.03 | 0.65 | 4.49 | 16.16 | 25.58 | 7.65 |
| Crude protein | 3.46 | 0.38 | 2.45 | 11.06 | 40.62 | 17.48 |
| Casein | 2.72 | 0.30 | 1.90 | 10.93 | 41.78 | 17.40 |
| SCS | 2.48 | 1.78 | 11.18 | 72.02 | 29.52 | 8.88 |
| MUN (mg/dL) | 21.19 | 7.17 | 43.60 | 33.83 | 14.70 | 20.27 |
| Protein fractions (mg/mL) | ||||||
| α-casein | 14.30 | 1.78 | 10.88 | 12.43 | 35.67 | 18.12 |
| β-casein | 10.45 | 1.64 | 10.03 | 15.71 | 37.56 | 10.50 |
| κ-casein | 7.30 | 0.96 | 5.91 | 13.18 | 36.53 | 11.57 |
| β-lactoglobulin | 1.82 | 0.77 | 4.29 | 42.45 | 44.76 | 7.22 |
| α-lactalbumin | 0.70 | 0.15 | 0.22 | 21.43 | 22.67 | 6.98 |
| Protein fractions (% of crude protein) | ||||||
| α-casein | 41.36 | 1.90 | 35.80 | 4.59 | 21.46 | 9.58 |
| β-casein | 30.48 | 3.69 | 47.81 | 12.12 | 36.15 | 9.63 |
| κ-casein | 21.25 | 2.12 | 34.34 | 9.97 | 28.30 | 13.18 |
| β-lactoglobulin | 5.23 | 2.13 | 18.64 | 40.82 | 45.63 | 6.83 |
| α-lactalbumin | 2.02 | 0.31 | 3.25 | 15.47 | 8.11 | 2.62 |
| Traits | Brown Swiss | Holstein-Friesian | Simmental | Alpine Grey | Pinzgauer |
|---|---|---|---|---|---|
| Milk yield (kg/day) | 23.33 (0.13) a | 28.43 (0.26) b | 23.19 (0.14) a | 17.10 (0.17) c | 19.99 (0.55) d |
| Milk composition (%) | |||||
| Fat | 4.20 (0.01) a | 4.02 (0.01) b | 4.05 (0.01) b | 3.84 (0.01) c | 4.00 (0.03) b |
| Crude protein | 3.58 (0.01) a | 3.27 (0.01) b | 3.45 (0.01) c | 3.44 (0.01) d | 3.40 (0.02) d |
| Casein | 2.81 (0.01) a | 2.56 (0.01) b | 2.71 (0.01) c | 2.70 (0.01) c | 2.67 (0.02) c |
| SCS | 2.85 (0.02) a | 2.73 (0.04) ab | 2.45 (0.02) c | 2.62 (0.03) b | 2.79 (0.09) ab |
| MUN (mg/dL) | 21.64 (0.13) a | 19.04 (0.26) b | 20.22 (0.14) c | 21.74 (0.16) a | 20.19 (0.54) abc |
| Protein fractions (mg/mL) | |||||
| α-casein | 14.85 (0.03) a | 13.62 (0.05) b | 14.41 (0.03) c | 14.13 (0.03) d | 13.96 (0.11) bd |
| β-casein | 11.27 (0.02) a | 9.93 (0.04) b | 10.09 (0.02) c | 10.69 (0.02) d | 10.18 (0.08) bc |
| κ-casein | 7.81 (0.01) a | 6.77 (0.02) b | 7.16 (0.01) c | 7.40 (0.01) d | 7.24 (0.05) cd |
| β-lactoglobulin | 1.56 (0.03) a | 1.94 (0.02) b | 2.05 (0.01) c | 1.62 (0.01) d | 1.70 (0.04) d |
| α-lactalbumin | 0.75 (0.01) a | 0.66 (0.01) b | 0.71 (0.01) c | 0.69 (0.01) d | 0.69 (0.01) cd |
| Protein fractions (% of crude protein) | |||||
| α-casein | 41.59 (0.02) a | 41.64 (0.04) ab | 41.75 (0.02) b | 41.12 (0.03) c | 41.18 (0.09) c |
| β-casein | 31.81 (0.04) a | 30.44 (0.09) b | 29.28 (0.05) c | 31.25 (0.06) d | 30.08 (0.18) b |
| κ-casein | 21.99 (0.03) a | 20.76 (0.05) b | 20.81 (0.03) b | 21.63 (0.03) c | 21.46 (0.11) c |
| β-lactoglobulin | 4.34 (0.02) a | 5.91 (0.05) b | 5.89 (0.02) b | 4.66 (0.03) c | 4.97 (0.10) c |
| α-lactalbumin | 2.10 (0.01) a | 2.01 (0.01) bc | 2.05 (0.01) d | 2.00 (0.01) b | 2.03 (0.01) cd |
| Trait 1 | MY, kg/d | Fat, % | CP, % | Casein, % | SCS | MUN, mg/dL | α-CN, mg/mL | β-CN, mg/mL | κ-CN, mg/mL | β-LG, mg/mL | α-LA, mg/mL | α-CN, % | β-CN, % | κ-CN, % | β-LG, % |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Fat, % | −0.03 | ||||||||||||||
| CP, % | −0.18 | 0.14 | |||||||||||||
| Casein, % | −0.14 | 0.17 | 0.98 | ||||||||||||
| SCS | −0.11 | 0.08 | 0.10 | 0.07 | |||||||||||
| MUN, mg/dL | 0.02 | 0.04 | 0.02 | −0.02 | −0.02 | ||||||||||
| α-CN, mg/mL | −0.14 | 0.15 | 0.89 | 0.86 | 0.07 | 0.04 | |||||||||
| β-CN, mg/mL | −0.06 | −0.02 | 0.53 | 0.48 | −0.01 | 0.26 | 0.56 | ||||||||
| κ-CN, mg/mL | −0.06 | 0.20 | 0.58 | 0.55 | 0.09 | −0.15 | 0.52 | 0.56 | |||||||
| β-LG, mg/mL | −0.16 | −0.06 | 0.32 | 0.34 | 0.06 | −0.24 | 0.24 | −0.21 | −0.11 | ||||||
| α-LA, mg/mL | −0.08 | 0.41 | 0.46 | 0.48 | 0.07 | 0.02 | 0.42 | 0.18 | 0.30 | 0.10 | |||||
| α-CN, % | −0.02 | 0.11 | 0.23 | 0.20 | −0.01 | 0.07 | 0.64 | 0.30 | 0.14 | −0.04 | 0.13 | ||||
| β-CN, % | 0.04 | −0.11 | −0.05 | −0.10 | −0.07 | 0.31 | 0.05 | 0.81 | 0.27 | −0.49 | −0.11 | 0.20 | |||
| κ-CN, % | 0.07 | 0.15 | −0.12 | −0.14 | 0.04 | −0.20 | −0.11 | 0.24 | 0.72 | −0.41 | −0.01 | −0.04 | 0.37 | ||
| β-LG, % | −0.13 | −0.09 | 0.12 | 0.15 | 0.04 | −0.26 | 0.06 | −0.35 | −0.24 | 0.97 | 0.01 | −0.09 | −0.53 | −0.42 | |
| α-LA, % | −0.02 | 0.41 | 0.13 | 0.16 | 0.03 | 0.02 | 0.13 | −0.01 | 0.11 | −0.01 | 0.93 | 0.05 | −0.10 | 0.03 | −0.03 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Franzoi, M.; Niero, G.; Visentin, G.; Penasa, M.; Cassandro, M.; De Marchi, M. Variation of Detailed Protein Composition of Cow Milk Predicted from a Large Database of Mid-Infrared Spectra. Animals 2019, 9, 176. https://doi.org/10.3390/ani9040176
Franzoi M, Niero G, Visentin G, Penasa M, Cassandro M, De Marchi M. Variation of Detailed Protein Composition of Cow Milk Predicted from a Large Database of Mid-Infrared Spectra. Animals. 2019; 9(4):176. https://doi.org/10.3390/ani9040176
Chicago/Turabian StyleFranzoi, Marco, Giovanni Niero, Giulio Visentin, Mauro Penasa, Martino Cassandro, and Massimo De Marchi. 2019. "Variation of Detailed Protein Composition of Cow Milk Predicted from a Large Database of Mid-Infrared Spectra" Animals 9, no. 4: 176. https://doi.org/10.3390/ani9040176
APA StyleFranzoi, M., Niero, G., Visentin, G., Penasa, M., Cassandro, M., & De Marchi, M. (2019). Variation of Detailed Protein Composition of Cow Milk Predicted from a Large Database of Mid-Infrared Spectra. Animals, 9(4), 176. https://doi.org/10.3390/ani9040176






