Associations between Bovine β-Defensin 4 Genotypes and Production Traits of Polish Holstein-Friesian Dairy Cattle
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Statistical Analysis
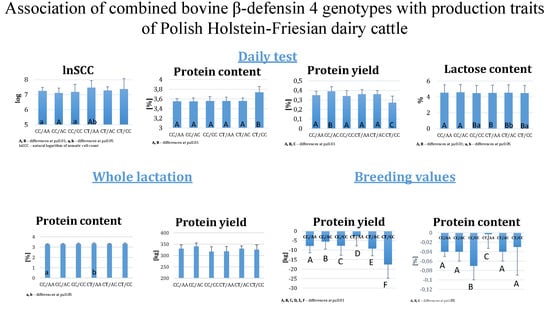
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Davies, D.; Meade, K.G.; Herath, S.; Eckersall, P.D.; Gonzalez, D.; White, J.O.; Conlan, D. Toll-like receptor and antimicrobial peptide expression in the bovine endometrium. Reprod. Biol. Endocrin. 2008, 6, 53. [Google Scholar] [CrossRef] [PubMed]
- Mainau, E.; Cuevas, A.; Ruiz-de-la-Torre, J.L.; Abbeloos, E.; Manteca, X. Effect of meloxicam administration after calving on milk production, acute phase proteins, and behavior in dairy cows. J. Vet. Behav. 2014, 9, 357–363. [Google Scholar] [CrossRef]
- Sender, G.; Korwin-Kossakowska, A.; Pawlik, A.; Hameed, K.G.A.; Oprządek, J. Genetic Basis of Mastitis Resistance in Dairy Cattle—A Review. Ann. Anim. Sci. 2013, 13, 663–673. [Google Scholar] [CrossRef]
- Togashi, K.; Lin, C.Y. Theoretical efficiency of multiple-trait quantitative trait loci-assisted selection. J. Anim. Breed. Genet. 2010, 127, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Song, H.; Zhu, X.; Xing, S.; Zhang, M.; Zhang, H.; Wang, X.; Yang, Z.; Ding, X.; Karrow, N.A.; et al. Toll-like receptor 4 gene polymorphisms influence milk production traits in Chinese Holstein cows. J. Dairy. Res. 2018, 85, 407–411. [Google Scholar] [CrossRef] [PubMed]
- Rangel, A.H.N.; Zaros, L.G.; Lima, T.C.; Borba, L.H.F.; Novaes, L.P.; Mota, L.F.M.; Silva, M.S. Polymorphism in the Beta Casein Gene and analysis of milk characteristics in Gir and Guzerá dairy cattle. Genet. Mol. Res. 2017, 16, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bojarojć-Nosowicz, B.; Oleński, K.; Sitarz, K.; Brym, P.; Zabolewicz, T.; Kamiński, S. Expression and polymorphism of ADAM32 gene and its association with somatic cell count in Holstein-Friesian cows. Anim. Sci. Pap. Rep. 2017, 35, 5–15. [Google Scholar]
- Bhattarai, D.; Chen, X.; Rehman, Z.; Hao, X.; Ullah, F.; Dad, R.; Talpur, H.S.; Kadariya, I.; Cui, L.; Fan, M.; et al. Association of MAP4K4 gene single nucleotide polymorphism with mastitis and milk traits in Chinese Holstein cattle. J. Dairy. Res. 2017, 84, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Stone, R.T.; Casas, E.; Smith, T.P.; Keele, J.W.; Harhay, G.; Bennet, G.L.; Koohmaraie, M.; Wheeler, T.L.; Shackelford, S.D.; Snelling, W.M. Identification of genetic markers for fat deposition and meat tenderness on bovine chromosome 5: Development of Low-density single nucleotide polymorphism map. J. Anim. Sci. 2005, 83, 2280–2288. [Google Scholar] [CrossRef]
- Wakchaure, R.; Ganguly, S.; Praveen, P.K.; Kumar, A.; Sharma, S.; Mahajn, T. Marker Assisted Selection (MAS) in Animal Breeding: A Review. J. Drug. Metab. Toxicol. 2015, 6, 127. [Google Scholar] [CrossRef]
- Goddard, M. Genomic selection: Prediction of accuracy and maximisation of long term response. Genetica 2009, 136, 245–257. [Google Scholar] [CrossRef]
- Schaeffer, L.R. Strategy for applying genome-wide selection in dairy cattle. J. Anim. Sci. 2006, 123, 218–223. [Google Scholar] [CrossRef] [PubMed]
- Meuwissen, T. Genomic Selection: The future of marker assisted selection and animal breeding. In Proceedings of the Electronic Forum on Biotechnology in Food and Agriculture: Conference 10, Workshop “Marker Assisted Selection: A Fast Track to Increase Genetic Gain in Plant and Animal Breeding?”, Session II: MAS in Animals, Turin, Italy, 17–18 October 2003. [Google Scholar]
- Matukumalli, L.K.; Lawley, C.T.; Schnabel, R.D.; Taylor, J.F.; Allan, M.F.; Heaton, M.P.; O’Connell, J.; Moore, S.S.; Smith, T.P.; Sonstegard, T.S. Development and Characterization of a High Density SNP Genotyping Assay for Cattle. PloS ONE 2009, 4, e5350. [Google Scholar] [CrossRef] [PubMed]
- Dadi, H.; Kim, J.J.; Yoon, H.; Kim, K.S. Evaluation of Single Nucleotide Polymorphisms (SNPs) Genotyped by the Illumina Bovine SNP50K in Cattle Focusing on Hanwoo Breed. Asian-Australasian J. Anim. Sci. 2012, 25, 28–32. [Google Scholar] [CrossRef]
- Meuwissen, T.; Hayes, B.; Goddard, M. Genomic selection: A paradigm shift in animal breeding. Anim. Front. 2016, 6, 6–14. [Google Scholar] [CrossRef]
- Bagnicka, E.; Strzałkowska, N.; Flisikowski, K.; Szreder, T.; Jóźwik, A.; Prusak, B.; Krzyżewski, J.; Zwierzchowski, L. The polymorphism in the β4-defensin gene and its association with production and somatic cell count in Holstein-Friesian cows. J. Anim. Sci. 2007, 124, 150–156. [Google Scholar]
- Bagnicka, E.; Strzałkowska, N.; Jóźwik, A.; Krzyżewski, J.; Horbańczuk, J.; Zwierzchowski, L. Expression and polymorphism of defensins in farm animals. Acta Biochim. Pol. 2010, 57, 487–497. [Google Scholar] [CrossRef]
- Jarczak, J.; Kościuczuk, E.M.; Lisowski, P.; Strzałkowska, N.; Jóźwik, A.; Horbańczuk, J.; Krzyżewski, J.; Zwierzchowski, L.; Bagnicka, E. Defensins: Natural component of human innate immunity. Hum. Immunol. 2013, 74, 1069–1079. [Google Scholar] [CrossRef]
- Zhu, S.; Gao, B. Evolutionary origin of β-defensins. Dev. Comp. Immunol. 2013, 39, 79–84. [Google Scholar] [CrossRef]
- Gurao, A.; Kashyap, S.K.; Singh, R. β-defensins: An innate defense for bovine mastitis. Vet. World 2017, 10, 990. [Google Scholar] [CrossRef]
- Roosen, S.; Exner, K.; Paul, S.; Schroder, J.M.; Kalm, E.; Looft, C. Bovine betadefensins: Identification and characterization of novel bovine beta-defensin genes and their expression in mammary gland tissue. Mamm. Genome. 2004, 15, 834–842. [Google Scholar] [CrossRef] [PubMed]
- O’brien, S.J.; Womack, J.E.; Lyons, L.A.; Moore, K.J.; Jenkins, N.A.; Copeland, N.G. Anchored reference loci for comparative genome mapping in mammals. Nat. Genet. 1993, 3, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Ashwell, M.S.; Da, Y.; Vanraden, P.M.; Rexroad, J.R.; Miller, R.H. Detection of putative loci affecting conformational type traits in an elite population of United States Holsteins using microsatellite markers. J. Anim. Sci. 1998, 81, 1120–1125. [Google Scholar] [CrossRef]
- Kościuszczuk, E.; Lisowski, P.; Jarczak, J.; Krzyżewski, J.; Zwierzchowski, L.; Bagnicka, E. Expression patterns of β-defensin and cathelicidin genes in parenchyma of bovine mammary gland infected with coagulase-positive or coagulase-negative Staphylococci. BMC Vet. Res. 2014, 10, 246. [Google Scholar]
- Ryniewicz, Z.; Zwierzchowski, L.; Bagnicka, E.; Krzyżewski, J.; Strzałkowska, N. Preliminary investigations on the polymorphism of defensin genes in cattle—relation with milk somatic cell count. Anim. Sci. Pap. Rep. 2002, 20, 125–131. [Google Scholar]
- Ryniewicz, Z.; Zwierzchowski, L.; Bagnicka, E.; Flisikowski, K.; Maj, A.; Krzyżewski, J.; Strzałkowska, N. Association of the polymorphism at defensin gene loci with dairy production traits and milk somatic cell count in Black-and-White cows. Anim. Sci. Pap. Rep. 2003, 21, 209–222. [Google Scholar]
- Wojdak-Maksymiec, K.; Strabel, T.; Szyda, J.; Mikołajczyk, K. Clinical Mastitis and Combined Defensin Polymorphism in Dairy Cattle. J. Vet. Med. 2012, 11, 2230–2237. [Google Scholar]
- Bagnicka, E.; Strzałkowska, N.; Szreder, T.; Prusak, B.; Joźwik, A.; Kosciuczuk, E.; Krzyżewski, J.; Zwierzchowski, L. A/C polymorphism in the β-4 defensin gene and its association with phenotypic and breeding values of milk production traits in Polish- Fresian cows. Anim. Sci. Pap. Rep. 2008, 26, 239–250. [Google Scholar]
- Krzyżewski, J.; Bagnicka, E.; Strzałkowska, N.; Jóźwik, A.; Pyzel, B.; Zwierzchowski, L. Association between the polymorphism of bovine B4-defensin gene and milk traits in Holstein-Friesian cows as computed for standard (305 days) and the whole lactation. Anim. Sci. Pap. Rep. 2008, 26, 191–198. [Google Scholar]
- Strzetelski, J.; Śliwiński, B. Wartość pokarmowa francuskich i krajowych pasz dla przeżuwaczy red; Strzetelski, J., Ed.; Research Institute of Animal Production: Cracow, Poland, 2009; pp. 21–90. (In Polish) [Google Scholar]
- Madsen, P.; Jensen, J. A User’s Guide to DMU. A Package for Analysing Multivariate Mixed Models. 2000, Version 6. Release 4. Available online: http://dmu.agrsci.dk/DMU/Doc/Current/dmuv6_guide.5.2.pdf (accessed on 15 November 2018).
- Brotherstone, S.; White, I.M.S.; Meyer, K. Genetic modeling of dairy milk yield using orthogonal polynomials and parametric curves. Anim. Sci. 2000, 70, 407–415. [Google Scholar] [CrossRef]
- Sjaunja, L.O.; Baevre, L.; Junkkarinen, L.; Pedersen, J.; Setaelae, J. A nordic proposal for an energy corrected milk (ECM) formula. In Proceedings of the 27th Session of International. Committee of Recording and Productivity of Milk Animal, Paris, France, 2–6 July 1990; pp. 156–157. [Google Scholar]
- Gaunt, T.R.; Rodríguez, S.; Day, I.N. Cubic exact solutions for the estimation of pairwise haplotype frequencies: Implications for linkage disequilibrium analyses and a web tool ‘CubeX’. BMC Bioinformatics 2007, 8, 428. [Google Scholar] [CrossRef] [PubMed]
- Wiechula, B.E.; Tustanowski, J.P.; Martirosian, G. Peptydy antydrobnoustrojowe. Wiad. Lek. 2006, 59, 542–547. [Google Scholar] [PubMed]
- Meredith, B.K.; Berry, D.P.; Kearney, F.; Finlay, E.K.; Fahey, A.G.; Bradley, D.G.; Lynn, D.J. A genome-wide association study for somatic cell score using the Illumina high-density bovine beadchip identifies several novel QTL potentially related to mastitis susceptibility. Front. Genet. 2013, 6, 229. [Google Scholar] [CrossRef] [PubMed]
- Tetens, J.; Friedrich, J.J.; Hartmann, A.; Schwerin, M.; Kalm, E.; Thaller, G. Thespatial expression pattern of antimicrobial peptides across the healthy bovine Udder. J. Anim. Sci. 2010, 93, 775–783. [Google Scholar]
- Jurevic, R.J.; Bai, M.; Chadwick, R.B.; White, T.C.; Dale, B.A. Single-nucleotide polymorphisms (SNPs) in human β-Defensin 1: High-Throughput SNP Assays and Association with Candidia Carriage in Type I Diabetics and Nondiabetic Controls. J. Clin. Microbiol. 2003, 41, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Ryan, L.K.; Diamond, G. Modulation of Human β-Defensin-1 Production by Viruses. Viruses 2017, 9, 153. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Chou, Y.Y.; Chang, T.L. Defensins in Viral Infections. J. Innate. Immun. 2009, 1, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, I.; Hasegawa, K.; Nakata, K.; Yasuda, K.; Tokunaga, K.; Keicho, N. Genetic Variants of Human β-defensin-l and Chronic Obstructive Pulmonary Disease. Biochem. Bioph. Res. Co. 2002, 291, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Levy, H.; Raby, B.A.; Lake, S.; Tantisira, K.G.; Kwiatkowski, D.; Lazarus, R.; Silverman, E.; Richter, B.; Klimecki, W.T.; Vercelli, D.; et al. Association of β-defensin gene polymorphisms with asthma. J. Allergy Clin. Immun. 2005, 115, 252–258. [Google Scholar] [CrossRef]
- Braida, L.; Boniotto, M.; Pontillo, A.; Tovo, P.A.; Amoroso, A.; Crovella, S. A single- nucleotide polymorphism in the human beta-defensin 1 gene is associated with HIV-1 infection in Italian children. AIDS 2004, 18, 1598–1600. [Google Scholar] [CrossRef]
- Milanese, M.; Segat, L.; Pontillo, A.; Arraes, L.C.; de Lima Filho, J.L.; Crovella, S. DEFB1 gene polymorphisms and increased risk of HIV-1 infection in Brasilian children. AIDS 2006, 20, 1673–1675. [Google Scholar] [CrossRef] [PubMed]
- Urech, E.; Puhan, Z.; Schallibaum, M. Changes in Milk Protein Fraction as Affected by Subclinical Mastitis. J. Anim. Sci. 1999, 82, 2402–2411. [Google Scholar] [CrossRef]
- Ekine, C.C.; Rowe, S.J.; Bishop, S.C.; de Koning, D.J. Why breeding values estimated using familial data should not be used for genome-wide association studies. G3-Genes Genom. Genet. 2014, 4, 341–347. [Google Scholar] [CrossRef] [PubMed]
- Hayes, B.J.; Chamberlain, A.J.; McPartlan, H.; Macleond, I.; Sethuraman, L.; Goddard, M.E. Accuracy of marker-assisted selection with single markers and marker haplotypes in cattle. Genet. Res. 2007, 89, 215–220. [Google Scholar] [CrossRef] [PubMed]
| Genotype | N | Estimate (SE) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Milk kg | ECM | Fat kg | Protein kg | Fat % | Protein % | Lactose % | lnSCC | Dry Matter % | ||
| CC/AA | 4761 | 29.49 A (1.15) | 31.01 A (1.17) | 1.25 A (0.08) | 0.35 A (0.04) | 4.39 A,a (0.14) | 3.55 A (0.06) | 4.54 A (0.05) | 7.25 a (0.23) | 13.23 A (0.17) |
| CC/AC | 625 | 30.72 B (1.44) | 32.66 B (1.42) | 1.32 B (0.09) | 0.39 B (0.05) | 4.42 a (0.17) | 3.55 A (0.07) | 4.57 A (0.06) | 7.11 A (0.33) | 13.31 a (0.21) |
| CC/CC | 163 | 29.20 A (2.00) | 31.91 A (1.92) | 1.31 B (0.10) | 0.34 A (0.06) | 4.56 B (0.23) | 3.56 A (0.09) | 4.46 B,a (0.08) | 7.18 a (0.51) | 13.34 (0.30) |
| CT/AA | 145 | 30.09 A,B (1.82) | 30.00 A (1.77) | 1.21 A (0.09) | 0.36 A (0.05) | 4.44 (0.22) | 3.56 A (0.08) | 4.49 B (0.07) | 7.47 B,b (0.46) | 13.31 (0.27) |
| CT/AC | 1875 | 29.38 A (1.22) | 30.72 A (1.23) | 1.24 A (0.08) | 0.36 A (0.04) | 4.36 A,a (0.16) | 3.56 A (0.06) | 4.52 B,b (0.05) | 7.28 (0.25) | 13.19 A,b (0.19) |
| CT/CC | 61 | 24.44 C (2.64) | 25.04 C (2.54) | 1.04 C (0.12) | 0.27 C (0.07) | 4.61 b (0.32) | 3.74 B (0.12) | 4.46 B,a (0.10) | 7.37 (0.70) | 13.54 B (0.41) |
| Genotype | N | Estimate (SE) | |||||
|---|---|---|---|---|---|---|---|
| Milk kg | ECM kg | Fat kg | Protein kg | Fat % | Protein % | ||
| CC/AA | 382 | 10,994.14 a (395.72) | 12,235 a (503) | 423.58 (14.58) | 329.93 (17.07) | 3.90 A (0.11) | 3.25 a (0.05) |
| CC/AC | 57 | 11,379.85 b (482.32) | 12,682 b (571) | 362.02 (17.33) | 340.44 (14.74) | 3.93 a (0.14) | 3.27 (0.06) |
| CC/CC | 14 | 10,487.42 a (704.34) | 1231 (752) | 379.35 (24.66) | 317.39 (21.32) | 4.17 B,b (0.21) | 3.28 (0.09) |
| CT/AA | 12 | 10,487.94 a (710.65) | 11,428 a (765) | 379.93 (25.24) | 317.78 (22.46) | 3.91 (0.21) | 3.31 b (0.09) |
| CT/AC | 151 | 10,895.82 a (421.97) | 12,076 a (528) | 370.04 (15.35) | 339.70 (13.08) | 3.90 A (0.12) | 3.28 (0.05) |
| CT/CC | 13 | 10,355.52 a (714.44) | 11,594 a (764) | 379.88 (25.19) | 324.94 (22.61) | 4.06 (0.21) | 3.31 b (0.09) |
| Genotype | N | Estimate (SE) | ||||
|---|---|---|---|---|---|---|
| Milk kg | Fat kg | Protein kg | Fat % | Protein % | ||
| CC/AA | 1278 | 73.47 A (114.00) | −11.29 A (4.78) | −7.79 A (3.71) | −0.08 A (0.03) | −0.04 A (0.01) |
| CC/AC | 182 | 156.67 B (129.53) | −8.44 B,b (5.22) | −5.50 B (4.09) | −0.09 B (0.05) | −0.04 A (0.02) |
| CC/CC | 40 | 162.23 B (165.20) | −7.09 B (6.32) | −7.69 C (5.02) | −0.05 C (0.07) | −0.07 B (0.03) |
| CT/AA | 37 | 178.16 B (170.57) | −5.82 B,b (6.47) | −2.69 D (5.16) | −0.04 D (0.08) | −0.003 C (0.03) |
| CT/AC | 491 | 41.94 A (119.23) | −13.79 C (4.92) | −9.17 E (3.83) | −0.10 E (0.04) | −0.04 A (0.02) |
| CT/CC | 10 | −264.03 C (250.70) | −29.24 D (9.01) | −17.50 F (7.30) | −0.18 F (0.13) | −0.03 A (0.06) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brodowska, P.; Zwierzchowski, L.; Marczak, S.; Jarmuż, W.; Bagnicka, E. Associations between Bovine β-Defensin 4 Genotypes and Production Traits of Polish Holstein-Friesian Dairy Cattle. Animals 2019, 9, 723. https://doi.org/10.3390/ani9100723
Brodowska P, Zwierzchowski L, Marczak S, Jarmuż W, Bagnicka E. Associations between Bovine β-Defensin 4 Genotypes and Production Traits of Polish Holstein-Friesian Dairy Cattle. Animals. 2019; 9(10):723. https://doi.org/10.3390/ani9100723
Chicago/Turabian StyleBrodowska, Paulina, Lech Zwierzchowski, Sylwester Marczak, Wiesław Jarmuż, and Emilia Bagnicka. 2019. "Associations between Bovine β-Defensin 4 Genotypes and Production Traits of Polish Holstein-Friesian Dairy Cattle" Animals 9, no. 10: 723. https://doi.org/10.3390/ani9100723
APA StyleBrodowska, P., Zwierzchowski, L., Marczak, S., Jarmuż, W., & Bagnicka, E. (2019). Associations between Bovine β-Defensin 4 Genotypes and Production Traits of Polish Holstein-Friesian Dairy Cattle. Animals, 9(10), 723. https://doi.org/10.3390/ani9100723