Dose-Dependent Porcine Deltacoronavirus Infection Reveals Linkage Between Infectious Dose and Immune Response
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Virus
2.2. Reagents and Antibodies
2.3. Plasmid Construction
2.4. Virus Infection
2.5. RNA Extraction and Quantitative Real-Time PCR (RT-qPCR)
2.6. Western Blotting
2.7. Plaque Assay
2.8. Indirect Immunofluorescence
2.9. Transcriptome Sequencing Analysis
2.10. Dual Luciferase Assay
2.11. Statistical Analysis
3. Result
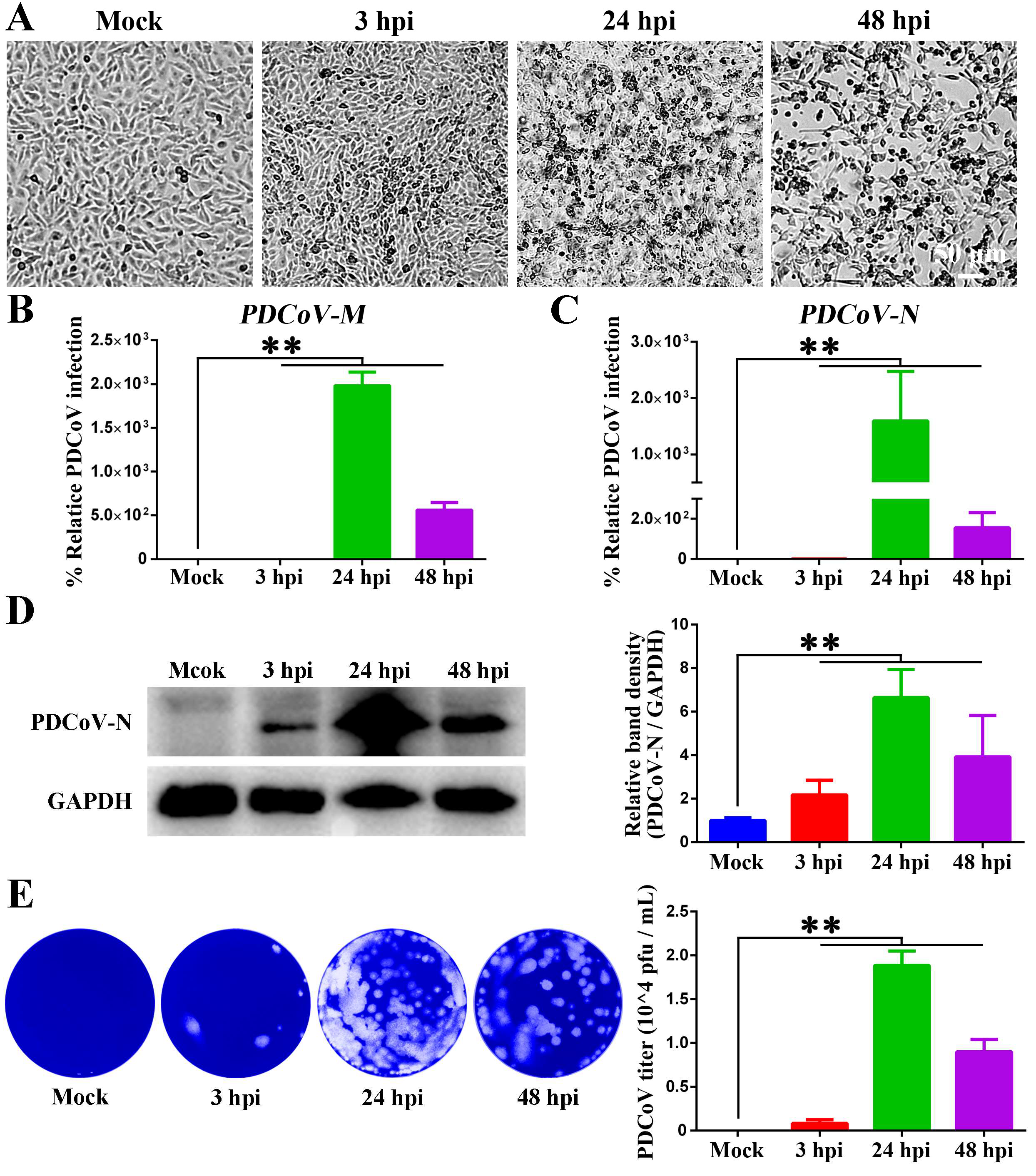
3.1. Proliferation Kinetics of PDCoV in IPEC-J2 Cells
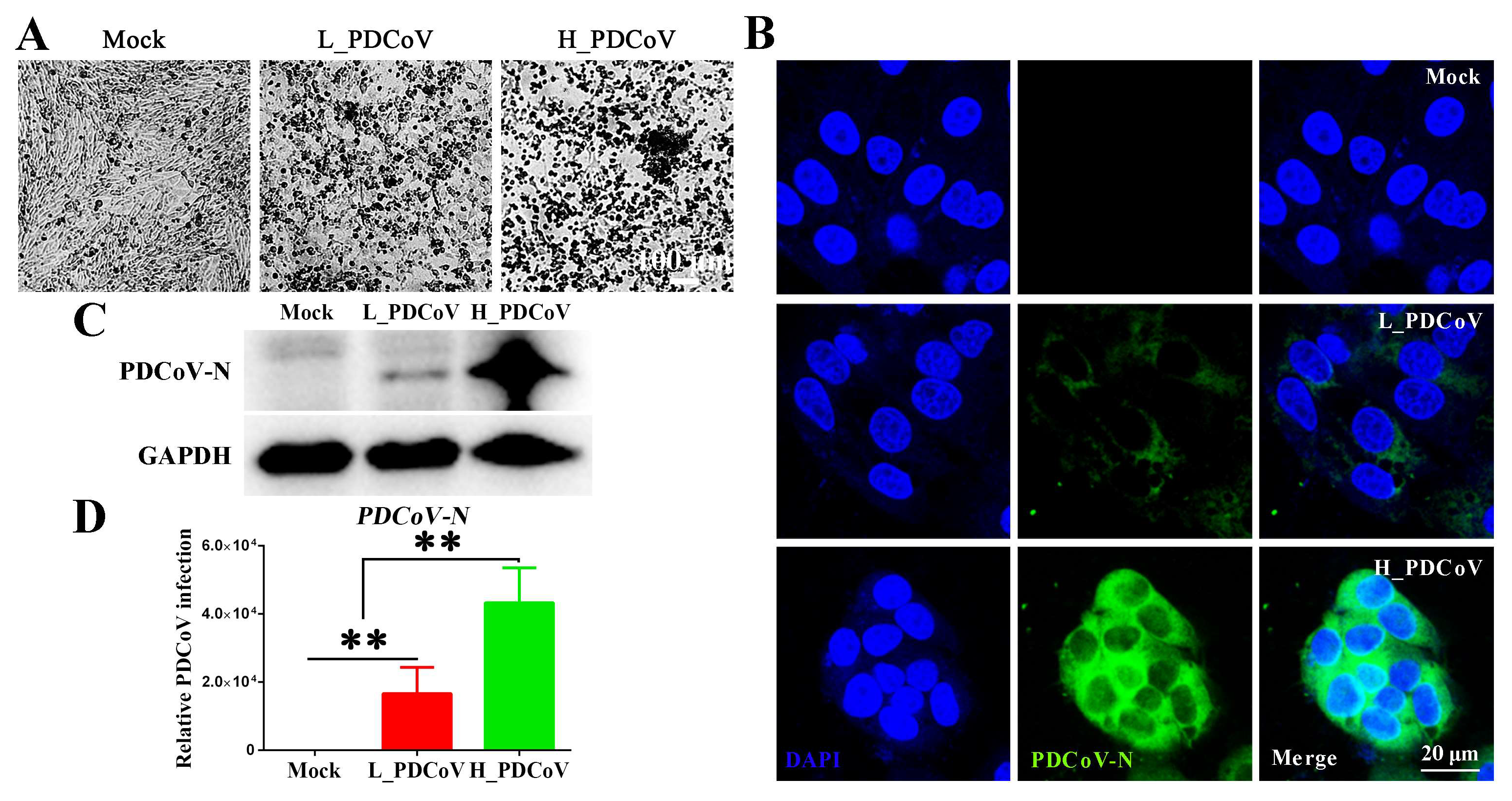
3.2. The Outcome of PDCoV Infection Depends on the Dose of Infection
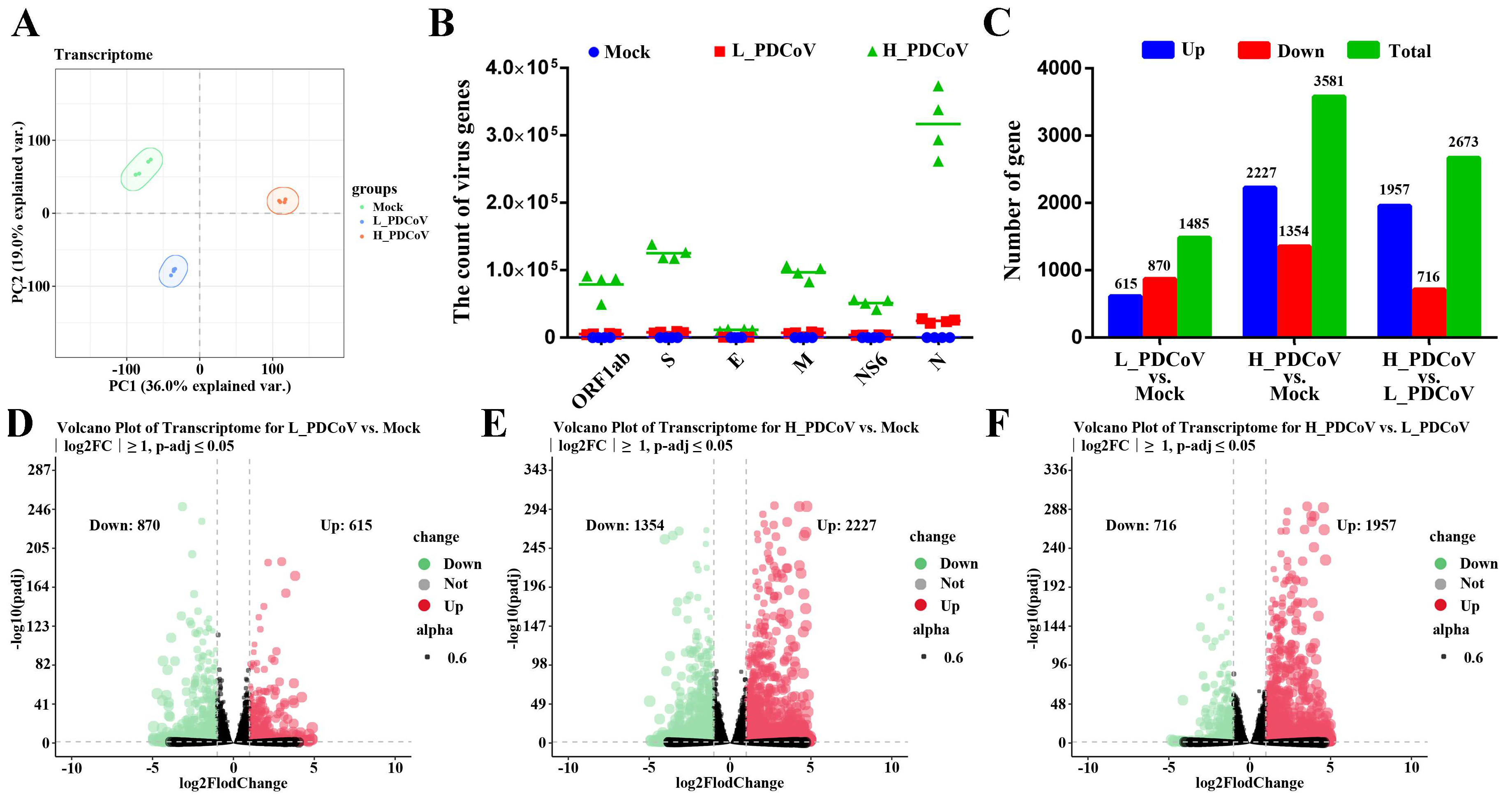
3.3. Transcriptional Profiles of Differently Dosed PDCoV-Infected IPEC-J2 Cells
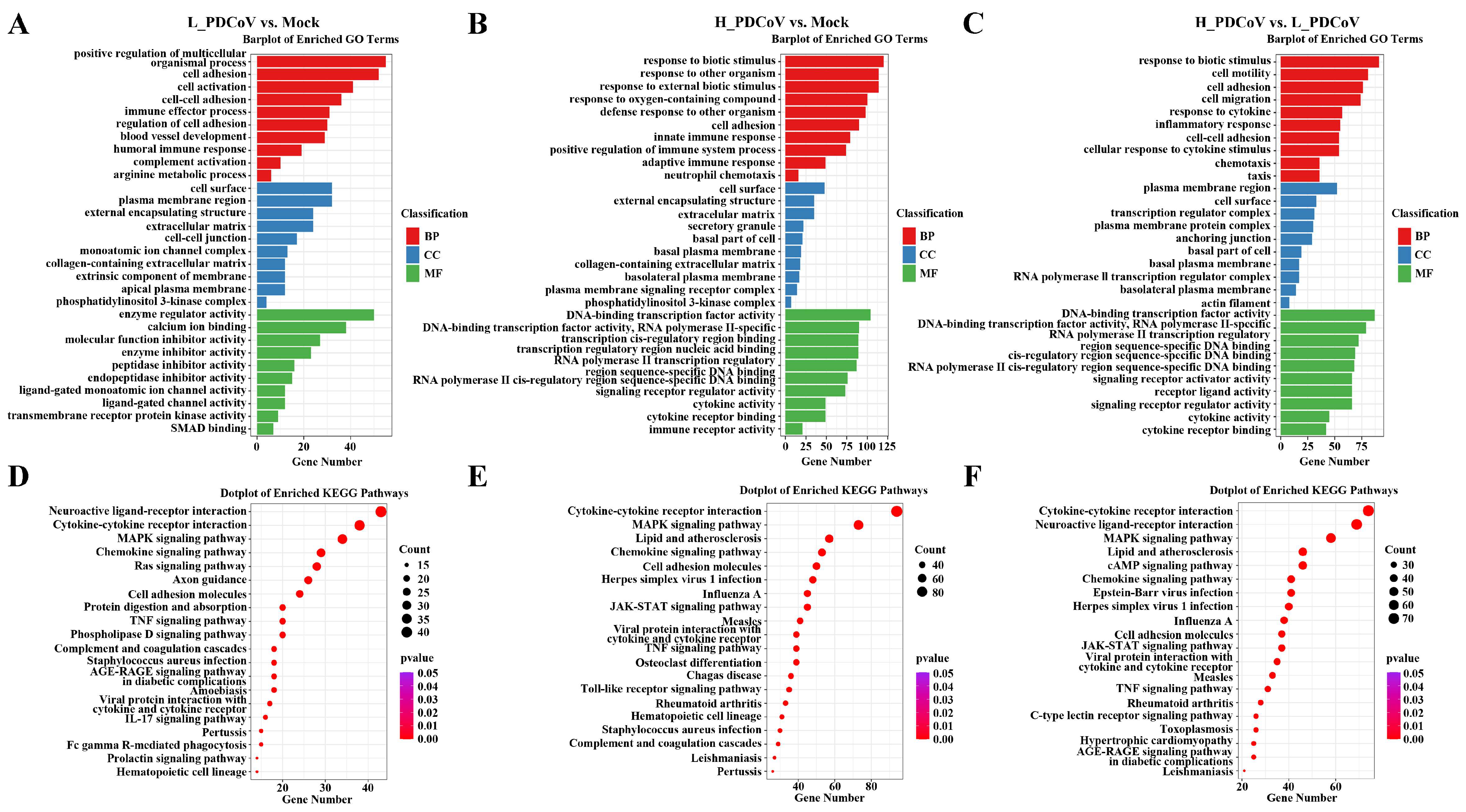
3.4. The Dose-Dependent Effects of PDCoV Infection on Host Gene Function and Signaling Pathways
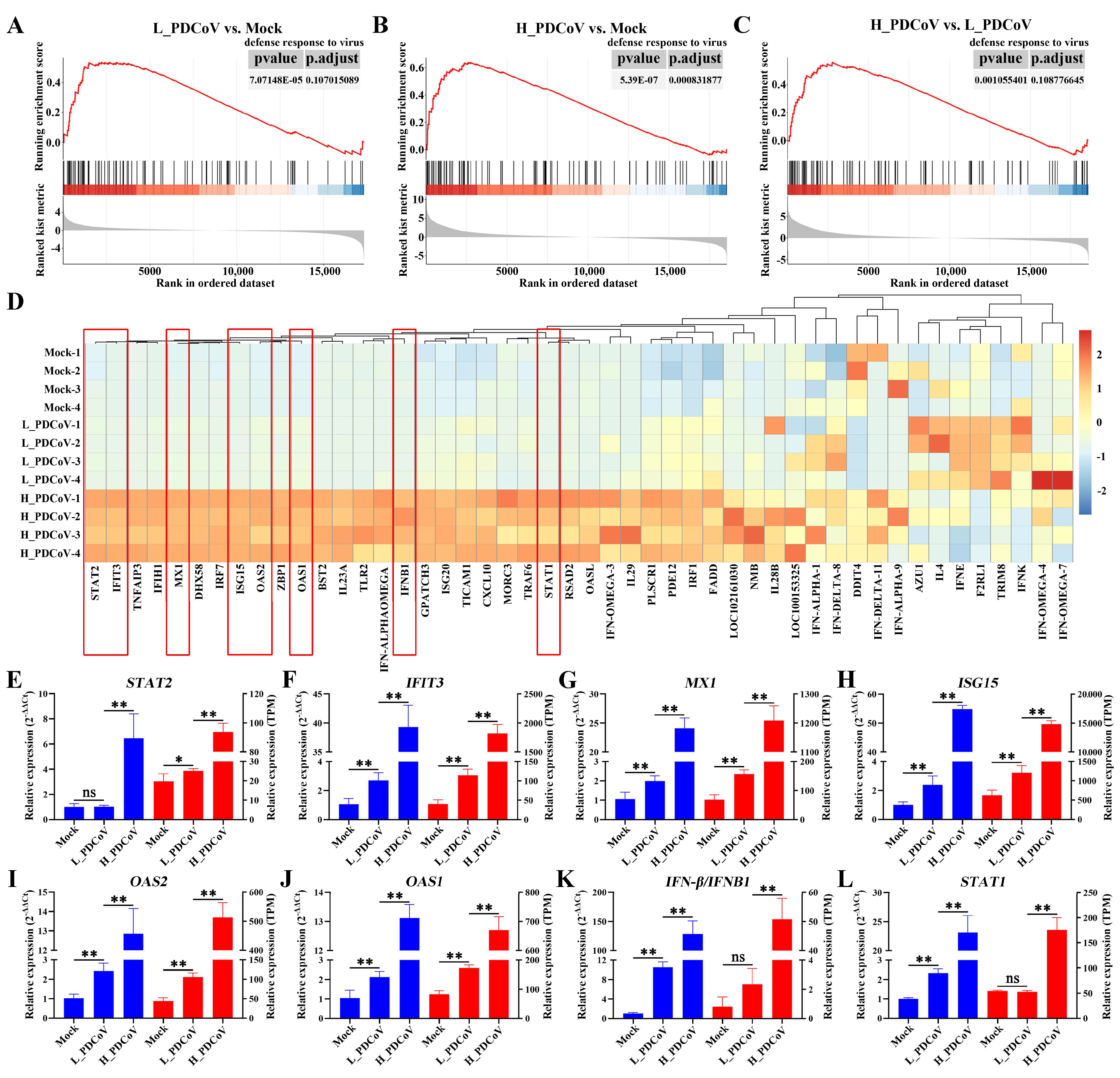
3.5. Assessment and Real-Time qPCR Verification of Genes Related to Defense Response to Virus
3.6. Interactions of STAT1, ISG15, and MX2 with PDCoV in Antiviral Responses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lau, C.C.Y.; Tsang, A.K.L.; Lau, J.H.N.; Bai, R.; Teng, J.L.L.; Tsang, C.C.C.; Wang, M.; et al. Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef]
- Zhang, J. Porcine deltacoronavirus: Overview of infection dynamics, diagnostic methods, prevalence and genetic evolution. Virus Res. 2016, 226, 71–84. [Google Scholar] [CrossRef]
- Wang, Z.; Qu, K.; Li, J.; Wang, Y.; Wang, L.; Yu, Y. Prevalence and potential risk factors of PDCoV in pigs based on publications during 2015–2021 in China: Comprehensive literature review and meta-analysis. Microb. Pathog. 2023, 179, 106118. [Google Scholar] [CrossRef]
- Kong, F.; Jia, H.; Xiao, Q.; Fang, L.; Wang, Q. Prevention and Control of Swine Enteric Coronaviruses in China: A Review of Vaccine Development and Application. Vaccines 2023, 12, 11. [Google Scholar] [CrossRef]
- Janetanakit, T.; Lumyai, M.; Bunpapong, N.; Boonyapisitsopa, S.; Chaiyawong, S.; Nonthabenjawan, N.; Kesdaengsakonwut, S.; Amonsin, A. Porcine Deltacoronavirus, Thailand, 2015. Emerg. Infect. Dis. 2016, 22, 757–759. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Shibahara, T.; Imai, N.; Yamamoto, T.; Ohashi, S. Genetic characterization and pathogenicity of Japanese porcine deltacoronavirus. Infect. Genet. Evol. 2018, 61, 176–182. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Gauger, P.; Stafne, M.; Thomas, J.; Arruda, P.; Burrough, E.; Madson, D.; Brodie, J.; Magstadt, D.; Derscheid, R.; et al. Pathogenicity and pathogenesis of a United States porcine deltacoronavirus cell culture isolate in 5-day-old neonatal piglets. Virology 2015, 482, 51–59. [Google Scholar] [CrossRef]
- Niederwerder, M.C.; Hesse, R.A. Swine enteric coronavirus disease: A review of 4 years with porcine epidemic diarrhoea virus and porcine deltacoronavirus in the United States and Canada. Transbound. Emerg. Dis. 2018, 65, 660–675. [Google Scholar] [CrossRef]
- Jung, K.; Hu, H.; Saif, L.J. Porcine deltacoronavirus infection: Etiology, cell culture for virus isolation and propagation, molecular epidemiology and pathogenesis. Virus Res. 2016, 226, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Ding, Q.; Yuan, J.; Han, F.; Wei, Z.; Hu, H. Susceptibility to mice and potential evolutionary characteristics of porcine deltacoronavirus. J. Med. Virol. 2022, 94, 5723–5738. [Google Scholar] [CrossRef]
- Boley, P.A.; Alhamo, M.A.; Lossie, G.; Yadav, K.K.; Vasquez-Lee, M.; Saif, L.J.; Kenney, S.P. Porcine Deltacoronavirus Infection and Transmission in Poultry, United States1. Emerg. Infect. Dis. 2020, 26, 255–265. [Google Scholar] [CrossRef]
- Jung, K.; Vasquez-Lee, M.; Saif, L.J. Replicative capacity of porcine deltacoronavirus and porcine epidemic diarrhea virus in primary bovine mesenchymal cells. Vet. Microbiol. 2020, 244, 108660. [Google Scholar] [CrossRef]
- Lednicky, J.A.; Tagliamonte, M.S.; White, S.K.; Elbadry, M.A.; Alam, M.M.; Stephenson, C.J.; Bonny, T.S.; Loeb, J.C.; Telisma, T.; Chavannes, S.; et al. Independent infections of porcine deltacoronavirus among Haitian children. Nature 2021, 600, 133–137. [Google Scholar] [CrossRef]
- McNab, F.; Mayer-Barber, K.; Sher, A.; Wack, A.; O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 2015, 15, 87–103. [Google Scholar] [CrossRef] [PubMed]
- Han, A.; Czajkowski, L.M.; Donaldson, A.; Baus, H.A.; Reed, S.M.; Athota, R.S.; Bristol, T.; Rosas, L.A.; Cervantes-Medina, A.; Taubenberger, J.K.; et al. A Dose-finding Study of a Wild-type Influenza A(H3N2) Virus in a Healthy Volunteer Human Challenge Model. Clin. Infect. Dis. 2019, 69, 2082–2090. [Google Scholar] [CrossRef] [PubMed]
- Memoli, M.J.; Czajkowski, L.; Reed, S.; Athota, R.; Bristol, T.; Proudfoot, K.; Fargis, S.; Stein, M.; Dunfee, R.L.; Shaw, P.A.; et al. Validation of the wild-type influenza A human challenge model H1N1pdMIST: An A(H1N1)pdm09 dose-finding investigational new drug study. Clin. Infect. Dis. 2015, 60, 693–702. [Google Scholar] [CrossRef]
- Chu, C.M.; Poon, L.L.; Cheng, V.C.; Chan, K.S.; Hung, I.F.; Wong, M.M.; Chan, K.H.; Leung, W.S.; Tang, B.S.; Chan, V.L.; et al. Initial viral load and the outcomes of SARS. Can. Med Assoc. J. 2004, 171, 1349–1352. [Google Scholar] [CrossRef] [PubMed]
- Bradburne, A.F.; Bynoe, M.L.; Tyrrell, D.A. Effects of a “new” human respiratory virus in volunteers. Br. Med. J. 1967, 3, 767–769. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Tsang, C.-C.; Lau, C.C.Y.; Wong, P.-C.; Chow, F.W.N.; Fong, J.Y.H.; Yuen, K.-Y. Coronavirus HKU15 in respiratory tract of pigs and first discovery of coronavirus quasispecies in 5′-untranslated region. Emerg. Microbes Infect. 2017, 6, 1–7. [Google Scholar] [CrossRef]
- Lammerding, A.M.; Paoli, G.M. Quantitative risk assessment: An emerging tool for emerging foodborne pathogens. Emerg. Infect. Dis. 1997, 3, 483–487. [Google Scholar] [CrossRef]
- Wong, G.; Liu, W.; Liu, Y.; Zhou, B.; Bi, Y.; Gao, G.F. MERS, SARS, and Ebola: The Role of Super-Spreaders in Infectious Disease. Cell Host Microbe 2015, 18, 398–401. [Google Scholar] [CrossRef]
- Marois, I.; Cloutier, A.; Garneau, É.; Richter, M.V. Initial infectious dose dictates the innate, adaptive, and memory responses to influenza in the respiratory tract. J. Leukoc. Biol. 2012, 92, 107–121. [Google Scholar] [CrossRef]
- Imai, M.; Iwatsuki-Horimoto, K.; Hatta, M.; Loeber, S.; Halfmann, P.J.; Nakajima, N.; Watanabe, T.; Ujie, M.; Takahashi, K.; Ito, M.; et al. Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proc. Natl. Acad. Sci. USA 2020, 117, 16587–16595. [Google Scholar] [CrossRef]
- Kojima, Y.; Nakakubo, S.; Kamada, K.; Yamashita, Y.; Takei, N.; Nakamura, J.; Matsumoto, M.; Horii, H.; Sato, K.; Shima, H.; et al. Combination therapy with remdesivir and immunomodulators improves respiratory status in COVID-19: A retrospective study. J. Med. Virol. 2022, 94, 5702–5712. [Google Scholar] [CrossRef]
- Kawai, T.; Akira, S. The role of pattern-recognition receptors in innate immunity: Update on Toll-like receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, S.; Takeuchi, K.; Chihara, K.; Honjoh, C.; Kato, Y.; Yoshiki, H.; Hotta, H.; Sada, K. STAT1 is essential for the inhibition of hepatitis C virus replication by interferon-λ but not by interferon-α. Sci. Rep. 2016, 6, 38336. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Gao, Q.; Zhang, W.; Zheng, J.; Chen, R.; Han, X.; Mao, J.; Shan, Y.; Shi, F.; He, F.; et al. Porcine epidemic diarrhea virus antagonizes host IFN-λ-mediated responses by tilting transcription factor STAT1 toward acetylation over phosphorylation to block its activation. mBio 2023, 14, e0340822. [Google Scholar] [CrossRef]
- Guo, L.; Luo, X.; Li, R.; Xu, Y.; Zhang, J.; Ge, J.; Bu, Z.; Feng, L.; Wang, Y. Porcine epidemic diarrhea virus infection inhibits interferon signaling by targeted degradation of STAT1. J. Virol. 2016, 90, 8281–8292. [Google Scholar] [CrossRef]
- Ma, X.Z.; Bartczak, A.; Zhang, J.; He, W.; Shalev, I.; Smil, D.; Chen, L.; Phillips, J.; Feld, J.J.; Selzner, N.; et al. Protein interferon-stimulated gene 15 conjugation delays but does not overcome coronavirus proliferation in a model of fulminant hepatitis. J. Virol. 2014, 88, 6195–6204. [Google Scholar] [CrossRef] [PubMed][Green Version]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bao, X.; Xia, L.; Bao, W.; Sun, M.; Zhang, S. Dose-Dependent Porcine Deltacoronavirus Infection Reveals Linkage Between Infectious Dose and Immune Response. Animals 2025, 15, 2536. https://doi.org/10.3390/ani15172536
Bao X, Xia L, Bao W, Sun M, Zhang S. Dose-Dependent Porcine Deltacoronavirus Infection Reveals Linkage Between Infectious Dose and Immune Response. Animals. 2025; 15(17):2536. https://doi.org/10.3390/ani15172536
Chicago/Turabian StyleBao, Xiaocheng, Liangxin Xia, Wenbin Bao, Ming’an Sun, and Shuai Zhang. 2025. "Dose-Dependent Porcine Deltacoronavirus Infection Reveals Linkage Between Infectious Dose and Immune Response" Animals 15, no. 17: 2536. https://doi.org/10.3390/ani15172536
APA StyleBao, X., Xia, L., Bao, W., Sun, M., & Zhang, S. (2025). Dose-Dependent Porcine Deltacoronavirus Infection Reveals Linkage Between Infectious Dose and Immune Response. Animals, 15(17), 2536. https://doi.org/10.3390/ani15172536






