Transcriptomic Analysis Reveals Regulatory Responses of Fatty Acid Positional Distribution in Triacylglycerols and Lipid Composition to Dietary n-3 HUFA in the Muscle of Trachinotus ovatus
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Fish, Diet Formulation, Feeding, and Sample Collection
2.2. Separation of Sample Lipids
2.3. Positional Distribution Analysis of Fatty Acid in TAG
2.4. Fatty Acid Analysis
2.5. RNA Extraction, cDNA Library Preparation, and Illumina Sequencing
2.6. De Novo Transcriptome Assembly, Sequence Annotation, and Differentially Expressed Gene Identification
2.7. Real-Time PCR
2.8. Statistical Analysis
3. Results
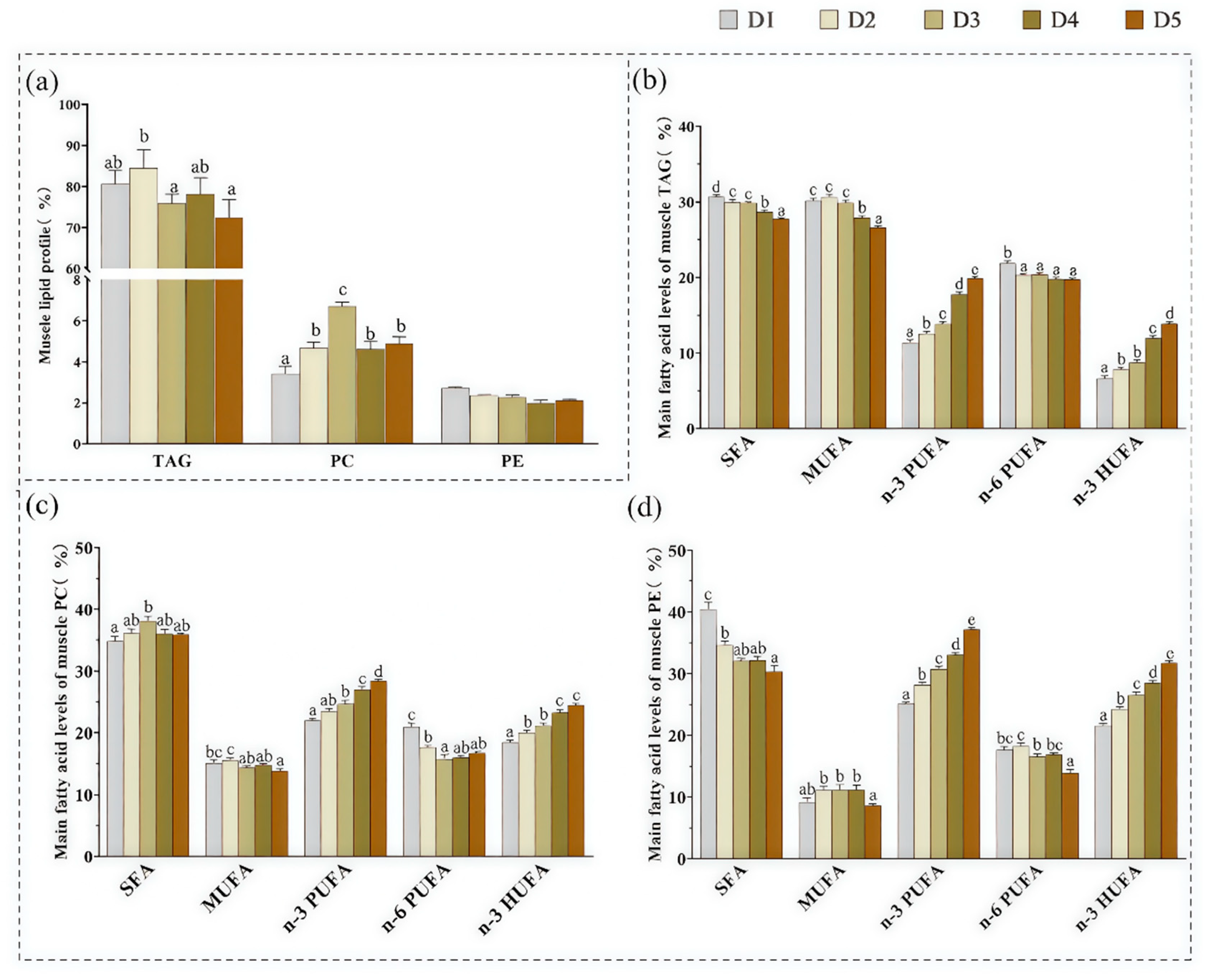
3.1. The Contents of TAG, PC, and PE in Muscle
3.2. Fatty Acid Composition of TAG, PC, and PE
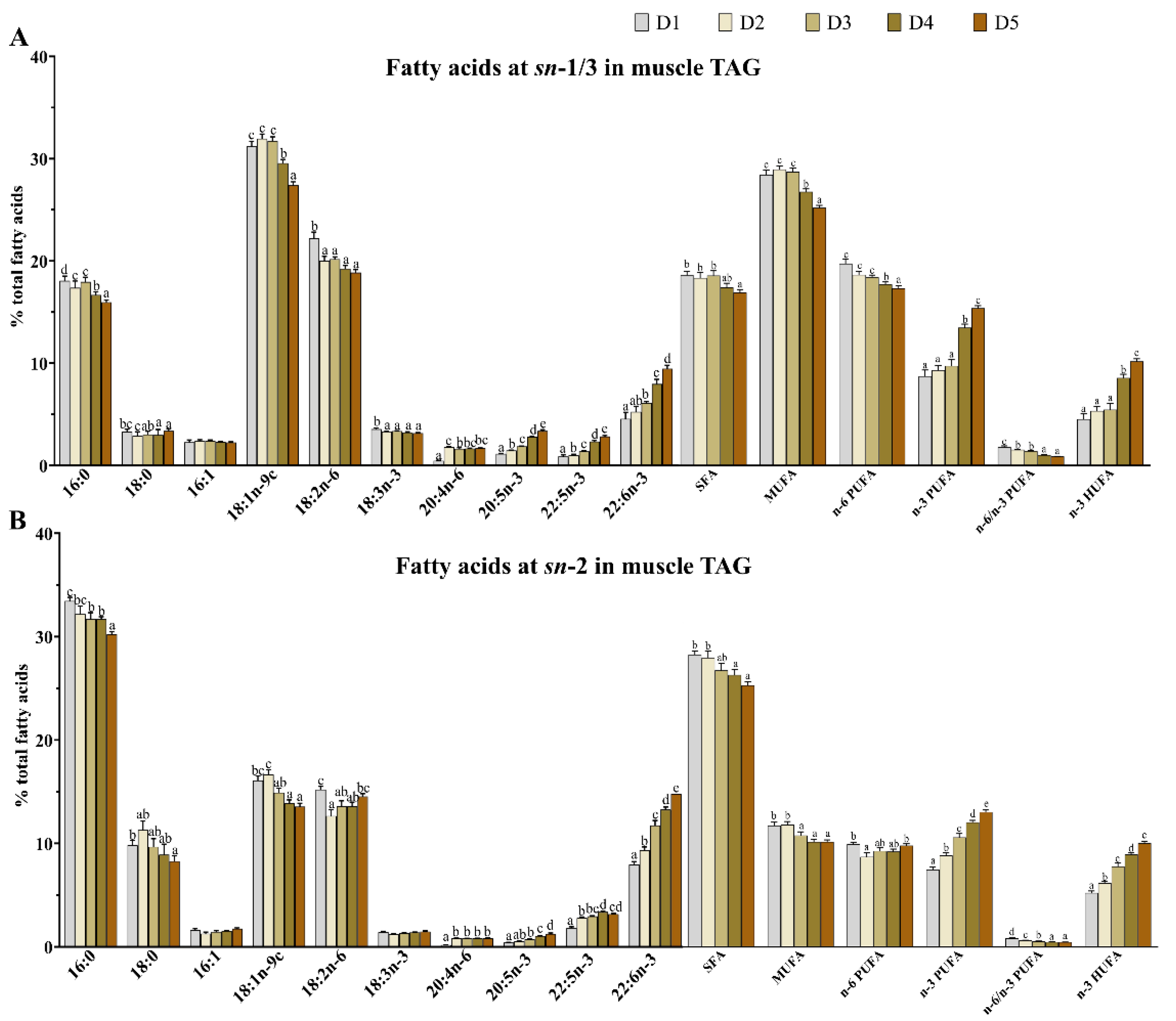
3.3. Fatty Acid Composition and Positional Distribution in Muscle TAG
3.4. Transcriptome Sequencing, Assembly, and Annotation
3.5. Function Enrichment Analysis of GO Terms and KEGG Pathway
3.6. Differentially Expressed Unigenes and Functional Enrichment
3.7. Expression of Genes Related to Fatty Acid Deposition and Positional Distribution
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Han, L.; Yu, J.; Chen, Y.; Cheng, D.; Wang, X.; Wang, C. Immunomodulatory activity of docosahexenoic acid on RAW264.7 cells activation through GPR120-mediated signaling pathway. J. Agric. Food Chem. 2018, 66, 926–934. [Google Scholar] [CrossRef]
- Serini, S.; Fasano, E.; Piccioni, E.; Cittadini, A.; Calviello, G. Differential anti-cancer effects of purified EPA and DHA and possible mechanisms involved. Curr. Med. Chem. 2011, 18, 4065–4075. [Google Scholar] [CrossRef]
- Jin, M.; Lu, Y.; Yuan, Y.; Li, Y.; Qiu, H.; Sun, P.; Ma, H.N.; Ding, L.Y.; Zhou, Q.C. Regulation of growth, antioxidant capacity, fatty acid profiles, hematological characteristics and expression of lipid related genes by different dietary n-3 highly unsaturated fatty acids in juvenile black seabream (Acanthopagrus schlegelii). Aquaculture 2017, 471, 55–65. [Google Scholar] [CrossRef]
- Vizcaíno-Ochoa, V.; Lazo, P.J.; Barón-Sevilla, B.; Drawbridge, M.A. The effect of dietary docosahexaenoic acid (DHA) on growth, survival and pigmentation of California halibut Paralichthys californicus larvae (Ayres, 1810). Aquaculture 2010, 302, 228–234. [Google Scholar] [CrossRef]
- Liu, Y.; Jiao, J.G.; Gao, S.; Ning, L.J.; Mchele Limbu, S.; Qiao, F.; Chen, L.Q.; Zhang, M.L.; Du, Z.Y. Dietary oils modify lipid molecules and nutritional value of fillet in Nile tilapia: A deep lipidomics analysis. Food Chem. 2019, 277, 515–523. [Google Scholar] [CrossRef]
- Ruizlopez, N.; Stubhaug, I.; Ipharraguerre, I.; Rimbach, G.; Menoyo, D. Positional distribution of fatty acids in triacylglycerols and phospholipids from fillets of Atlantic salmon (Salmo Salar) fed vegetable and fish oil blends. Mar. Drugs 2015, 13, 4255–4269. [Google Scholar] [CrossRef]
- Ramırez, M.; Amate, L.; Gil, A. Absorption and distribution of dietary fatty acids from different sources. Early Hum. Dev. 2001, 65, 95–101. [Google Scholar] [CrossRef]
- Christensen, J.H.; Korup, E.; Aarøe, J.; Toft, E.; Møller, J.; Rasmussen, K.; Dyerberg, J.; Schmidt, E.B. Fish consumption, n-3 fatty acids in cell membranes, and heart rate variability in survivors of myocardial infarction with left ventricular dysfunction. Am. J. Cardiol. 1997, 79, 1670–1673. [Google Scholar] [CrossRef] [PubMed]
- Yoshinaga, K.; Sasaki, K.; Watanabe, H.; Nagao, K.; Inoue, N.; Shirouchi, B.; Yanagita, T.; Nagai, T.; Mizobe, H.; Kojima, K. Differential effects of triacylglycerol positional isomers containing n-3 series highly unsaturated fatty acids on lipid metabolism in C57BL/6J mice. J. Nutr. Biochem. 2015, 26, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Alhazzaa, R.; Bridle, A.R.; Carter, C.G.; Nichols, P.D. Sesamin modulation of lipid class and fatty acid profile in early juvenile teleost, Lates calcarifer, fed different dietary oils. Food Chem. 2012, 134, 2057–2065. [Google Scholar] [CrossRef] [PubMed]
- Nanton, D.A.; Vegusdal, A.; Amb, R.; Ruyter, B.; Baeverfjord, G.; Torstensen, B.E. Muscle lipid storage pattern, composition, and adipocyte distribution in different parts of Atlantic salmon (Salmo salar) fed fish oil and vegetable oil. Aquaculture 2007, 265, 230–243. [Google Scholar] [CrossRef]
- Brockerhoff, H.; Yurkowski, M.; Hoyle, R.J.; Ackman, R.G. Fatty acid distribution in lipids of marine plankton. J. Fish. Res. Board Can. 1964, 21, 1379–1384. [Google Scholar] [CrossRef]
- Yamashita, A.; Sugiura, T.; Waku, K. Acyltransferases and transacylases involved in fatty acid remodeling of phospholipids and metabolism of bioactive lipids in mammalian cells. J. Biochem. 1997, 122, 1–16. [Google Scholar] [CrossRef]
- Lands, W.E. Metabolism of glycerolipids. 2. The enzymatic acylation of lysolecithin. J. Biol. Chem. 1960, 235, 2233–2237. [Google Scholar] [CrossRef] [PubMed]
- Coleman, R.A.; Lee, D.P. Enzymes of triacylglycerol synthesis and their regulation. Prog. Lipid Res. 2004, 43, 134–176. [Google Scholar] [CrossRef] [PubMed]
- Bell, J.G.; Dick, J.R.; Mcvicar, A.H.; Sargent, J.R.; Thompson, K.D. Dietary sunflower, linseed and fish oils affect phospholipid fatty acid composition, development of cardiac lesions, phospholipase activity and eicosanoid production in Atlantic salmon (Salmo salar). Prostaglandins Leukot. Essent. Fat. Acids 1993, 49, 665–673. [Google Scholar] [CrossRef]
- Ahmmed, M.K.; Carne, A.; Ahmmed, F.; Stewart, I.; Sabrina Tian, H.; Bekhit, A.E.A. Positional distribution of fatty acids and phospholipid composition in King salmon (Oncorhynchus tshawytscha) head, roe and skin using nuclear magnetic resonance spectroscopy. Food Chem. 2021, 363, 130302. [Google Scholar] [CrossRef] [PubMed]
- Ahmmed, M.K.; Ahmmed, F.; Stewart, I.; Carne, A.; Tian, H.S.; Bekhit, A.E.A. Omega-3 phospholipids in Pacific blue mackerel (Scomber australasicus) processing by-products. Food Chem. 2021, 353, 129451. [Google Scholar] [CrossRef]
- Zhang, M.; Chen, C.Y.; You, C.H.; Chen, B.J.; Wang, S.Q.; Li, Y.Y. Effects of different dietary ratios of docosahexaenoic to eicosapentaenoic acid (DHA/EPA) on the growth, non-specific immune indices, tissue fatty acid compositions and expression of genes related to LC-PUFA biosynthesis in juvenile golden pompano Trachinotus ovatus. Aquaculture 2019, 505, 488–495. [Google Scholar]
- Li, Y.; Zhao, J.; Dong, Y.; Yin, Z.; Li, Y.; Liu, Y.; You, C.; Monroig, O.; Tocher, D.R.; Wang, S. Sp1 is involved in vertebrate LC-PUFA biosynthesis by upregulating the expression of liver desaturase and elongase genes. Int. J. Mol. Sci. 2019, 20, 5066. [Google Scholar] [CrossRef]
- He, C.; Cao, J.; Jiang, X.; Wen, C.; Bai, X.; Li, C. Fatty acid profiles of triacylglycerols and phospholipids of sea-cage cultured trachinotus blochii: A comparative study of head, viscera, skin, bone, and muscle. J. Food Sci. 2019, 84, 650–658. [Google Scholar] [CrossRef]
- Ichihara, K.I.; Shibahara, A.; Yamamoto, K.; Nakayama, T. An improved method for rapid analysis of the fatty acids of glycerolipids. Lipids 1996, 31, 535–539. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Li, J.; Li, G.; Hu, J.; Tang, L.; Liu, R.; Fan, Y.; Deng, Z. Stereospecific analysis of triacylglycerol and phospholipid fractions of five wild freshwater fish from Poyang Lake. J. Agric. Food Chem. 2012, 60, 1857–1864. [Google Scholar] [CrossRef] [PubMed]
- Li, M.M.; Zhai, S.S.; Xie, Q.; Tian, L.; Li, X.C.; Zhang, J.M.; Ye, H.; Zhu, Y.W.; Yang, L.; Wang, W.C. Effects of dietary n-6: N-3 PUFA ratios on lipid levels and fatty acid profile of Cherry Valley ducks at 15-42 days of age. J. Agric. Food Chem. 2017, 65, 9995–10002. [Google Scholar] [CrossRef] [PubMed]
- Gerber, M. Omega-3 fatty acids and cancers: A systematic update review of epidemiological studies. Br. J. Nutr. 2012, 107, S228–S239. [Google Scholar] [CrossRef]
- Li, M.; Zhang, M.; Ma, Y.; Ye, R.; Chen, C.; You, C.; Wang, S.; Li, Y. Dietary supplementation with n-3 high unsaturated fatty acids decreases serum lipid levels and improves flesh quality in the marine teleost golden pompano Trachinotus ovatus. Aquaculture 2020, 516, 734632. [Google Scholar] [CrossRef]
- Hussain, S.M.; Sharif, A.; Bashir, F.; Ali, S.; Javid, A.; Hussain, A.I.; Ghafoor, A.; Alshehri, M.A.; Naeem, A.; Naeem, E.; et al. Polymerase Chain Reaction: A Toolbox for Molecular Discovery. Mol. Biotechnol. 2025; ahead of print. [Google Scholar] [CrossRef]
- Andersson, A.; Sjodin, A.; Hedman, A.; Olsson, R.; Vessby, B. Fatty acid profile of skeletal muscle phospholipids in trained and untrained young men. Am. J. Physiol. Endocrinol. Metab. 2000, 279, 744–751. [Google Scholar] [CrossRef]
- Dos Santos, J.; Burkow, I.C.; Jobling, M. Patterns of growth and lipid deposition in cod (Gadus morhua L.) fed natural prey and fish-based feeds. Aquaculture 1993, 110, 173–189. [Google Scholar] [CrossRef]
- Kubow, S. The influence of positional distribution of fatty acids in native, interesterified and structure-specific lipids on lipoprotein metabolism and atherogenesis. J. Nutr. Biochem. 1996, 7, 530–541. [Google Scholar] [CrossRef]
- Menoyo, D.; Lopez-Bote, C.J.; Bautista, J.M.; Obach, A. Growth, digestibility and fatty acid utilization in large Atlantic salmon (Salmo salar) fed varying levels of n-3 and saturated fatty acids. Aquaculture 2003, 225, 295–307. [Google Scholar] [CrossRef]
- Martin, J.; Bougnoux, P.; Antoine, J.; Lanson, M.; Couet, C. Triacylglycerol structure of human colostrum and mature milk. Lipids 1993, 28, 637–643. [Google Scholar] [CrossRef]
- Innis, S.M.; Dyer, R.; Nelson, C.M. Evidence that palmitic acid is absorbed as sn-2 monoacylglycerol from human milk by breast-fed infants. Lipids 1994, 29, 541–545. [Google Scholar] [CrossRef]
- Carnielli, V.P.; Luijendijk, I.H.; van Goudoever, J.B.; Sulkers, E.J.; Boerlage, A.A.; Degenhart, H.J.; Sauer, P.J. Feeding premature newborn infants palmitic acid in amounts and stereoisomeric position similar to that of human milk: Effects on fat and mineral balance. Am. J. Clin. Nutr. 1995, 61, 1037–1042. [Google Scholar] [CrossRef]
- Simonetti, M.S.; Blasi, F.; Bosi, A.; Maurizi, A.; Cossignani, L.; Damiani, P. Stereospecific analysis of triacylglycerol and phospholipid fractions of four freshwater fish species: Salmo trutta, Ictalurus punctatus, Ictalurus melas and Micropterus salmoides. Food Chem. 2008, 110, 199–206. [Google Scholar] [CrossRef]
- Wolk, A.; Bergström, R.; Hunter, D.; Willett, W.; Ljung, H.; Holmberg, L.; Bergkvist, L.; Bruce, A.; Adami, H.O. A prospective study of association of monounsaturated fat and other types of fat with risk of breast cancer. Arch. Intern. Med. 1998, 158, 41–45. [Google Scholar] [CrossRef]
- Stoneham, M.; Goldacre, M.; Seagroatt, V.; Gill, L. Olive oil, diet and colorectal cancer: An ecological study and a hypothesis. J. Epidemiol. Community Health 2000, 54, 756–760. [Google Scholar] [CrossRef]
- Psaltopoulou, T.; Naska, A.; Orfanos, P.; Trichopoulos, D.; Mountokalakis, T.; Trichopoulou, A. Olive oil, the Mediterranean diet, and arterial blood pressure: The Greek European Prospective Investigation into Cancer and Nutrition (EPIC) study. Am. J. Clin. Nutr. 2004, 80, 1012–1018. [Google Scholar] [CrossRef]
- Calder, P.C. Metabolism of polyunsaturated fatty acids by cells of the immune system. In Polyunsaturated Fatty Acid Metabolism; Elsevier: Amsterdam, The Netherlands, 2018; pp. 135–155. [Google Scholar] [CrossRef]
- Bracco, U. Effect of triglyceride structure on fat absorption. Am. J. Clin. Nutr. 1994, 60, 1002S–1009S. [Google Scholar] [CrossRef]
- Lien, E.L.; Boyle, F.G.; Yuhas, R.; Tomarelli, R.M.; Quinlan, P. The effect of triglyceride positional distribution on fatty acid absorption in rats. J. Pediatr. Gastroenterol. Nutr. 1997, 25, 167–174. [Google Scholar]
- Cao, J.S.; Li, J.L.; Li, D.M.; Tobin, J.F.; Gimeno, R.E. Molecular identification of microsomal acyl-CoA: Glycerol-3-phosphate acyltransferase, a key enzyme in de novo triacylglycerol synthesis. Proc. Natl. Acad. Sci. USA 2006, 103, 19695–19700. [Google Scholar] [CrossRef]
- Nagle, C.A.; Vergnes, L.; DeJong, H.; Wang, S.; Lewin, T.M.; Reue, K.; Coleman, R.A. Identification of a novel sn-glycerol-3-phosphate acyltransferase isoform, GPAT4, as the enzyme deficient in Agpat6−/− mice. J. Lipid Res. 2008, 49, 823–831. [Google Scholar] [CrossRef]
- Vergnes, L.; Beigneux, A.P.; Davis, R.; Watkins, S.M.; Young, S.G.; Reue, K. Agpat6 deficiency causes subdermal lipodystrophy and resistance to obesity. J. Lipid Res. 2006, 47, 745–754. [Google Scholar] [CrossRef]
- Li, J.; Romestaing, C.; Han, X.; Li, Y.; Hao, X.; Wu, Y.; Sun, C.; Liu, X.; Jefferson, L.S.; Xiong, J. Cardiolipin remodeling by ALCAT1 links oxidative stress and mitochondrial dysfunction to obesity. Cell Metab. 2010, 12, 154–165. [Google Scholar] [CrossRef]
- Imae, R.; Inoue, T.; Nakasaki, Y.; Uchida, Y.; Ohba, Y.; Kono, N.; Nakanishi, H.; Sasaki, T.; Mitani, S.; Arai, H. LYCAT, a homologue of C. elegans acl-8, acl-9, and acl-10, determines the fatty acid composition of phosphatidylinositol in mice. J. Lipid Res. 2012, 53, 335–347. [Google Scholar] [CrossRef] [PubMed]
- Hishikawa, D.; Valentine, W.J.; Iizuka Hishikawa, Y.; Shindou, H.; Shimizu, T. Metabolism and functions of docosahexaenoic acid-containing membrane glycerophospholipids. Febs. Lett. 2017, 591, 2730–2744. [Google Scholar] [CrossRef]
- Koichi, Y.; Hideo, S.; Daisuke, H.; Takao, S. Characterization of mouse lysophosphatidic acid acyltransferase 3: An enzyme with dual functions in the testis. J. Lipid Res. 2009, 50, 860–869. [Google Scholar] [CrossRef] [PubMed]
- Miki, E.; Shindou, H.; Shimizu, T. A novel lysophosphatidic acid acyltransferase enzyme (LPAAT4) with a possible role for incorporating docosahexaenoic acid into brain glycerophospholipids. Biochem. Biophys. Res. Commun. 2014, 443, 718–724. [Google Scholar] [CrossRef]




| Genes | Primers | Primer Sequences | Annealing Temperature (°C) | Annealing Time (s) | Accession No./Reference |
|---|---|---|---|---|---|
| gpat1 | gpat1-F | GAGTCCGACTACACCCAGAG | 60 | 30 | Genome sequences |
| gpat1-R | GTGGTCACCTTGCTCTCTCT | ||||
| gpat3 | gpat3-F | GTGGTGTTTGGCTGTAGGTG | 60 | 30 | Genome sequences |
| gpat3-R | GCTCCATCTCCTCCTCCATC | ||||
| gpat4 | gpat4-F | CATTGCCAAAGAGCCAACCT | 60 | 30 | Genome sequences |
| gpat4-R | GGTCAGCAGATTCCAGGACT | ||||
| agpat1 | agpat1-F | TAGGAGAAAGAGAGGCGGGA | 60 | 30 | Genome sequences |
| agpat1-R | TCGAGACAAAGGGGATGACG | ||||
| agpat5 | agpat5-F | AGCACCGATGTACCTTGTCA | 60 | 30 | Genome sequences |
| agpat5-R | TGGCCCTTCATGGTCTCAAT | ||||
| agpat8 | agpat8-F | CAGCCTGCCGAAATTTGAGT | 60 | 30 | Genome sequences |
| agpat8-R | CAAGATGAGGTGGCGTTCTG | ||||
| dgat1 | dgat1-F | TCCTCAACTGGTGTGTGGTT | 60 | 30 | Genome sequences |
| dgat1-R | TGGTACCCACAGCTAAACGT | ||||
| dgat2 | dgat2-F | TAAAACCCACAACCTGCTGC | 60 | 30 | Genome sequences |
| dgat2-R | CGGGACAGATACCTCCAGAC | ||||
| lpeat1 | lpeat1-F | CTGTGTGTTATGACCGAGCC | 60 | 30 | Genome sequences |
| lpeat1-R | GTAGTGGCGAACGGATTTCC | ||||
| lpgat1 | lpgat1-F | AGTGGATAGTGCTGTTCCCC | 60 | 30 | Genome sequences |
| lpgat1-R | TTACCCACTGCAGGCCTTTA | ||||
| lpcat1 | lpcat1-F | GCTTGGATGACTTCGCTCAG | 60 | 30 | Genome sequences |
| lpcat1-R | CCATCCTCCTCTGCCTCAAA | ||||
| lpcat2 | lpcat2-F | TTTGCCAGCAGAGTGAGAGA | 60 | 30 | Genome sequences |
| lpcat2-R | CGTTGTCCCACTTCAGTTGG | ||||
| lpcat3 | lpcat3-F | CGTTCTGGTATCGCTGTGTG | 60 | 30 | Genome sequences |
| lpcat3-R | GAGCCACACCTTCATGTTGG | ||||
| lpcat4 | lpcat4-F | GGCTGGTATTCCCTCCACTT | 60 | 30 | Genome sequences |
| lpcat4-R | CCCGCTTTGTCAAACCATCA | ||||
| β-actin | β-actin-F | TACGAGCTGCCTGACGGACA | 60 | 30 | KX987228 |
| β-actin-R | GGCTGTGATCTCCTTCTGC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, X.; Li, M.; Guan, J.; Cheng, Z.; Xie, D.; Li, Y. Transcriptomic Analysis Reveals Regulatory Responses of Fatty Acid Positional Distribution in Triacylglycerols and Lipid Composition to Dietary n-3 HUFA in the Muscle of Trachinotus ovatus. Animals 2025, 15, 2427. https://doi.org/10.3390/ani15162427
Gao X, Li M, Guan J, Cheng Z, Xie D, Li Y. Transcriptomic Analysis Reveals Regulatory Responses of Fatty Acid Positional Distribution in Triacylglycerols and Lipid Composition to Dietary n-3 HUFA in the Muscle of Trachinotus ovatus. Animals. 2025; 15(16):2427. https://doi.org/10.3390/ani15162427
Chicago/Turabian StyleGao, Xin, Mengmeng Li, Junfeng Guan, Zhiyi Cheng, Dizhi Xie, and Yuanyou Li. 2025. "Transcriptomic Analysis Reveals Regulatory Responses of Fatty Acid Positional Distribution in Triacylglycerols and Lipid Composition to Dietary n-3 HUFA in the Muscle of Trachinotus ovatus" Animals 15, no. 16: 2427. https://doi.org/10.3390/ani15162427
APA StyleGao, X., Li, M., Guan, J., Cheng, Z., Xie, D., & Li, Y. (2025). Transcriptomic Analysis Reveals Regulatory Responses of Fatty Acid Positional Distribution in Triacylglycerols and Lipid Composition to Dietary n-3 HUFA in the Muscle of Trachinotus ovatus. Animals, 15(16), 2427. https://doi.org/10.3390/ani15162427







