Characterization of the Nero Siciliano Pig Fecal Microbiota after a Liquid Whey-Supplemented Diet
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Management and Experimental Design
2.2. Fecal Sample Collection and Next Generation Ssequencing
2.3. Bioinformatics and Statistical Analysis
3. Results
3.1. Growth Performance, Quality Check, and Taxonomic Composition of Pig Fecal Microbiota
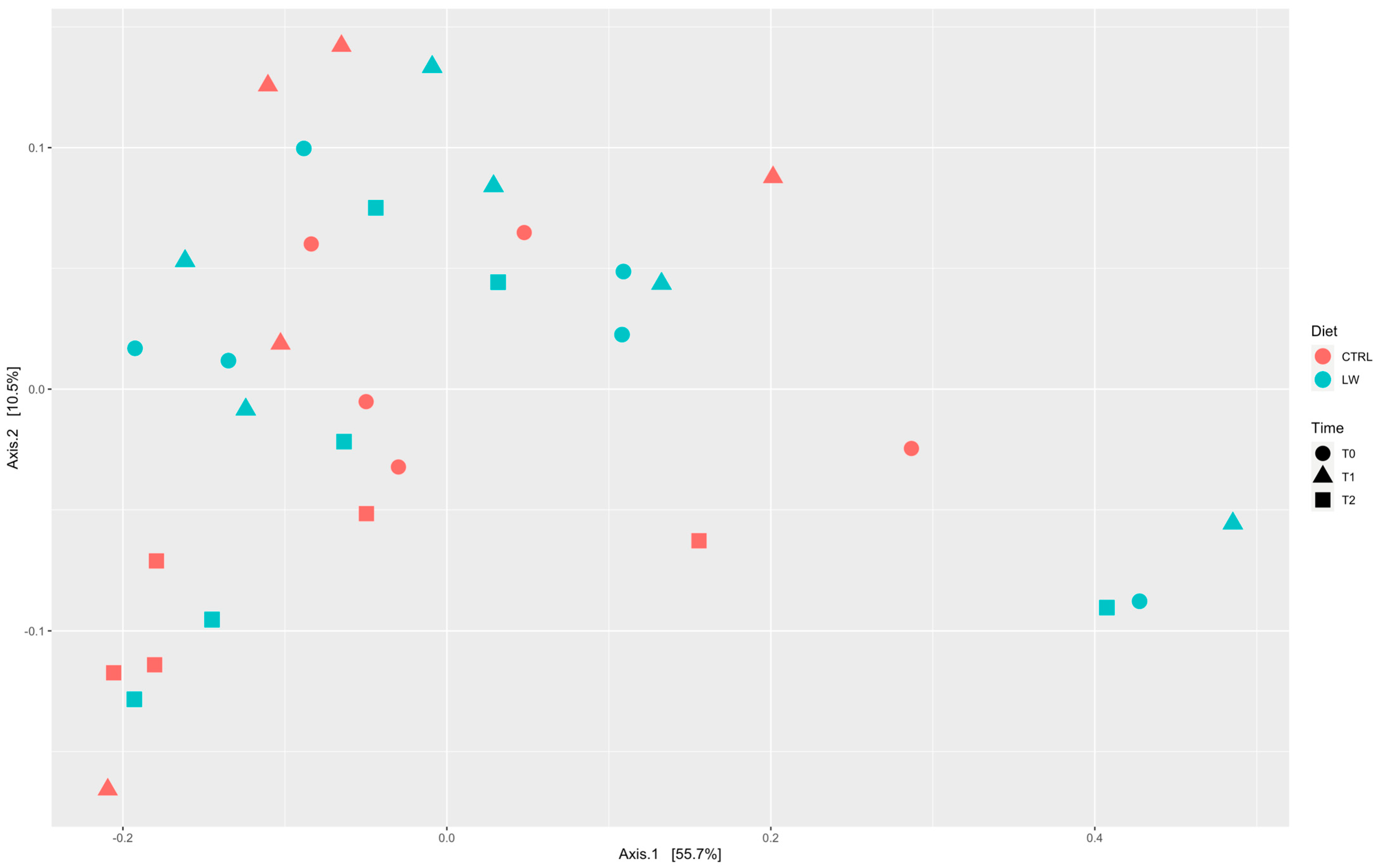
3.2. Fecal Microbiota Diversity Correlated with Feeding Condition and Time Point
3.3. Analysis of Microbial Genera Abundance at Different Time Points
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Egea, M.; Peñaranda, I.; Garrido, M.D.; Linares, M.B.; Sánchez, C.J.; Madrid, J.; Orengo, J.; Hernández, F.; Casañas, M.A.A.; Baños, A. Use of Mediterranean By-Products to Produce Entire Male Large White Pig: Meat and Fat Quality. Animals 2021, 11, 3128. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.H.; Hong, J.S.; Yoo, H.B.; Han, T.H.; Jeong, J.H.; Kim, Y.Y. Influence of various levels of milk by-products in weaner diets on growth performance, blood urea nitrogen, diarrhea incidence, and pork quality of weaning to finishing pigs. Asian-Australas. J. Anim. Sci. 2018, 31, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Pierce, K.M.; Callan, J.J.; McCarthy, P.; O’Doherty, J.V. The interaction between lactose level and crude protein concentration on piglet post-weaning performance, nitrogen metabolism, selected faecal microbial populations and faecal volatile fatty acid concentrations. Anim. Feed Sci. Technol. 2007, 132, 267–282. [Google Scholar] [CrossRef]
- Jang, K.; Purvis, J.M.; Kim, S.W. Dose-response and functional role of whey permeate as a source of lactose and milk oligosaccharides on intestinal health and growth of nursery pigs. J. Anim. Sci. 2021, 99, skab008. [Google Scholar] [CrossRef] [PubMed]
- González, S. The Biotechnological Utilization of Cheese Whey: A Review. Bioresour. Technol. 1996, 57, 1–11. [Google Scholar]
- De Wit, J.N. Nutritional and functional characteristics of whey proteins in food products. J. Dairy Sci. 1998, 81, 597–608. [Google Scholar] [CrossRef]
- Outinen, M.; Rantamäki, P.; Heino, A. Effect of milk pretreatment on the whey composition and whey powder functionality. J. Food Sci. 2010, 75, E1–E10. [Google Scholar] [CrossRef] [PubMed]
- Marshall, K. Therapeutic applications of whey protein. Altern. Med. Rev. 2004, 9, 136–156. [Google Scholar]
- Hering, N.A.; Andres, S.; Fromm, A.; van Tol, E.A.; Amasheh, M.; Mankertz, J.; Fromm, M.; Schulzke, J.D. Transforming growth factor-β, a whey protein component, strengthens the intestinal barrier by upregulating claudin-4 in HT-29/B6 cells. J. Nutr. 2011, 141, 783–789. [Google Scholar] [CrossRef]
- Xiao, K.; Jiao, L.; Cao, S.; Song, Z.; Hu, C.; Han, X. Whey protein concentrate enhances intestinal integrity and influences transforming growth factor-β1 and mitogen-activated protein kinase signalling pathways in piglets after lipopolysaccharide challenge. Br. J. Nutr. 2016, 115, 984–993. [Google Scholar] [CrossRef]
- Nielsen, C.H.; Hui, Y.; Nguyen, D.N.; Ahnfeldt, A.M.; Burrin, D.G.; Hartmann, B.; Heckmann, A.B.; Sangild, P.T.; Thymann, T.; Bering, S.B. Alpha-Lactalbumin enriched whey protein concentrate to improve gut, immunity and brain development in preterm pigs. Nutrients 2020, 12, 245. [Google Scholar] [CrossRef]
- D’Alessandro, E.; Arfuso, F.; Floridia, V.; Tardiolo, G.; Fazio, F.; Giannetto, C.; Piccione, G.; Zumbo, A. Different genotype and a liquid whey-supplemented diet influence the resilience of pigs through immune-modulation and anti-inflammatory response. Front. Vet. Sci. 2022, 9, 1046101. [Google Scholar] [CrossRef]
- Sprong, R.C.; Schonewille, A.J.; van der Meer, R. Dietary cheese whey protein protects rats against mild dextran sulfate sodium-induced colitis: Role of mucin and microbiota. J. Dairy Sci. 2010, 93, 1364–1371. [Google Scholar] [CrossRef] [PubMed]
- Playford, R.J.; Macdonald, C.E.; Johnson, W.S. Colostrum and milk-derived peptide growth factors for the treatment of gastrointestinal disorders. Am. J. Clin. Nutr. 2000, 72, 5–14. [Google Scholar] [CrossRef]
- Cho, I.; Blaser, M.J. The human microbiome: At the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Song, S.J.; Sanders, J.G.; Delsuc, F.; Metcalf, J.; Amato, K.; Taylor, M.W.; Mazel, F.; Lutz, H.L.; Winker, K.; Graves, G.R. Comparative Analyses of Vertebrate Gut Microbiomes Reveal Convergence between Birds and Bats. mBio 2020, 7, e02901-19. [Google Scholar] [CrossRef]
- Patil, Y.; Gooneratne, R.; Ju, X.H. Interactions between host and gut microbiota in domestic pigs: A review. Gut Microbes 2020, 11, 310–334. [Google Scholar] [CrossRef] [PubMed]
- McGlone, J.J. The Future of Pork Production in the World: Towards Sustainable, Welfare-Positive Systems. Animals 2013, 3, 401–415. [Google Scholar] [CrossRef] [PubMed]
- Pappas, G. Socio-economic, industrial and cultural parameters of pig-borne infections. Clin. Microbiol. Infect. 2013, 19, 605–610. [Google Scholar] [CrossRef]
- Cucchi, T.; Dai, L.; Balasse, M.; Zhao, C.; Gao, J.; Hu, Y.; Yuan, J.; Vigne, J.D. Social Complexification and Pig (Sus scrofa) Husbandry in Ancient China: A Combined Geometric Morphometric and Isotopic Approach. PLoS ONE 2016, 11, e0158523. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Colmenero, F.; Ventanas, J.; Toldrá, F. Nutritional composition of dry-cured ham and its role in a healthy diet. Meat Sci. 2010, 84, 585–593. [Google Scholar] [CrossRef]
- Pugliese, C.; Sirtori, F. Quality of meat and meat products produced from southern European pig breeds. Meat Sci. 2012, 90, 511–518. [Google Scholar] [CrossRef] [PubMed]
- Hryhorowicz, M.; Lipiński, D.; Hryhorowicz, S.; Nowak-Terpiłowska, A.; Ryczek, N.; Zeyland, J. Application of Genetically Engineered Pigs in Biomedical Research. Genes 2020, 11, 670. [Google Scholar] [CrossRef] [PubMed]
- Meurens, F.; Summerfield, A.; Nauwynck, H.; Saif, L.; Gerdts, V. The pig: A model for human infectious diseases. Trends Microbiol. 2012, 20, 50–57. [Google Scholar] [CrossRef]
- Zhang, Q.; Widmer, G.; Tzipori, S. A pig model of the human gastrointestinal tract. Gut Microbes 2013, 4, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L.; Estellé, J.; Kiilerich, P.; Ramayo-Caldas, Y.; Xia, Z.; Feng, Q.; Liang, S.; Pedersen, A.Ø.; Kjeldsen, N.J.; Liu, C. A reference gene catalogue of the pig gut microbiome. Nat. Microbiol. 2016, 1, 16161. [Google Scholar] [CrossRef]
- Yang, H.; Huang, X.; Fang, S.; He, M.; Zhao, Y.; Wu, Z.; Yang, M.; Zhang, Z.; Chen, C.; Huang, L. Un-raveling the Fecal Microbiota and Metagenomic Functional Capacity Associated with Feed Efficiency in Pigs. Front. Microbiol. 2017, 8, 1555. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, P.; Yan, Q.; Chen, L.; Li, T.; Zhang, W.; Li, H.; Chen, C.; Han, X.; Zhang, S. Characterization of the Pig Gut Microbiome and Antibiotic Resistome in Industrialized Feedlots in China. mSystems 2019, 4, e00206-19. [Google Scholar] [CrossRef] [PubMed]
- Ramayo-Caldas, Y.; Mach, N.; Lepage, P.; Levenez, F.; Denis, C.; Lemonnier, G.; Leplat, J.J.; Billon, Y.; Berri, M.; Dore, J. Phylogenetic network analysis applied to pig gut microbiota identifies an ecosystem structure linked with growth traits. ISME J. 2016, 10, 2973–2977. [Google Scholar] [CrossRef]
- Fouhse, J.M.; Zijlstra, R.T.; Willing, B.P. The role of gut microbiota in the health and disease of pigs. Anim. Front. 2016, 6, 30–36. [Google Scholar] [CrossRef]
- Canibe, N.; Højberg, O.; Højsgaard, S.; Jensen, B.B. Feed physical form and formic acid addition to the feed affect the gastrointestinal ecology and growth performance of growing pigs. J. Anim. Sci. 2005, 83, 1287–1302. [Google Scholar] [CrossRef] [PubMed]
- Mach, N.; Berri, M.; Estellé, J.; Levenez, F.; Lemonnier, G.; Denis, C.; Leplat, J.J.; Chevaleyre, C.; Billon, Y.; Doré, J. Early-life establishment of the swine gut microbiome and impact on host phenotypes: Role of early-life gut microbiome on pigs’ health. Environ. Microbiol. Rep. 2015, 7, 554–569. [Google Scholar] [CrossRef] [PubMed]
- Maltecca, C.; Bergamaschi, M.; Tiezzi, F. The interaction between microbiome and pig efficiency: A review. J. Anim. Breed Genet. 2020, 137, 4–13. [Google Scholar] [CrossRef] [PubMed]
- Feng, W.; Ao, H.; Peng, C. Gut microbiota, short-chain fatty acids, and herbal medicines. Front. Pharmacol. 2018, 9, 1354. [Google Scholar] [CrossRef]
- Muraca, M.; Putignani, L.; Fierabracci, A.; Teti, A.; Perilongo, G. Gut microbiota-derived outer membrane vesicles: Under-recognized major players in health and disease? Discov. Med. 2015, 19, 343–348. [Google Scholar]
- Zumbo, A.; Sutera, A.M.; Tardiolo, G.; D’Alessandro, E. Sicilian Black Pig: An Overview. Animals 2020, 10, 2326. [Google Scholar] [CrossRef]
- Chen, C.; D’Alessandro, E.; Murani, E.; Zheng, Y.; Giosa, D.; Yang, N.; Wang, X.; Gao, B.; Li, K.; Wimmers, K.; et al. SINE jumping contributes to large-scale polymorphisms in the pig genomes. Mob. DNA 2021, 12, 17. [Google Scholar] [CrossRef]
- Wang, X.; D′Alessandro, E.; Chi, C.; Moawad, A.S.; Zong, W.; Chen, C.; Song, C. Genetic Evaluation and Population Structure of Jiangsu Native Pigs in China Revealed by SINE Insertion Polymorphisms. Animals 2022, 12, 1345. [Google Scholar] [CrossRef]
- D’Alessandro, E.; Giosa, D.; Sapienza, I.; Giuffrè, L.; Aiese, C.R.; Romeo, O.; Zumbo, A. Whole genome SNPs discovery in Nero Siciliano pig. Genet. Mol. Biol. 2019, 42, 594–602. [Google Scholar] [CrossRef]
- D’Alessandro, E.; Sottile, G.; Sardinia, M.T.; Criscione, A.; Bordonaro, S.; Sutera, A.M.; Zumbo, A.; Portolano, B.; Mastrangelo, S. Genome-wide analyses reveal the regions involved in the phenotypic diversity in Sicilian pigs. Anim. Genet. 2020, 51, 101–105. [Google Scholar] [CrossRef]
- Chen, C.; Wang, X.; Zong, W.; D’Alessandro, E.; Giosa, D.; Guo, Y.; Mao, J.; Song, C. Genetic Diversity and Population Structures in Chinese Miniature Pigs Revealed by SINE Retrotransposon Insertion Polymorphisms, a New Type of Genetic Markers. Animals 2021, 11, 1136. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 1 October 2022).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Paytuví, A.; Battista, E.; Scippacercola, F.; Aiese Cigliano, R.; Sanseverino, W. GAIA: An integrated metagenomics suite. bioRxiv 2019, 804690. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-7. 2020. Available online: http://CRAN.R-project.org/package=vegan (accessed on 1 November 2022).
- Bray, J.R.; Curtis, J.T. An Ordination of the Upland Forest Communities of Southern Wisconsin. Ecol. Monogr. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Wilcoxon, F. Individual comparisons by ranking methods. Biometrics 1945, 1, 80–83. [Google Scholar] [CrossRef]
- Kruskal, W.H.; Wallis, W.A. Use of Ranks in One-Criterion Variance Analysis. J. Am. Stat. Ass. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Sattin, E.; Andreani, N.A.; Carraro, L.; Lucchini, R.; Fasolato, L.; Telatin, A.; Balzan, S.; Novelli, E.; Simionati, B.; Cardazzo, B. A Multi-Omics Approach to Evaluate the Quality of Milk Whey Used in Ricotta Cheese Production. Front. Microbiol. 2016, 7, 1272. [Google Scholar] [CrossRef]
- Giuffrè, L.; Giosa, D.; Galeano, G.; Cigliano, R.A.; Paytuví-Gallart, A.; Sutera, A.M.; Tardiolo, G.; Zumbo, Z.; Romeo, O.; D’Alessandro, E. Whole-metagenome shotgun sequencing of pig faecal microbiome. Ital. J. Anim. Sci. 2021, 20, 1147–1155. [Google Scholar] [CrossRef]
- Miragoli, F.; Patrone, V.; Prandini, A.; Sigolo, S.; Dell’Anno, M.; Rossi, L.; Senizza, A.; Morelli, L.; Callegari, M.L. Implications of Tributyrin on Gut Microbiota Shifts Related to Performances of Weaning Piglets. Microorganisms 2021, 9, 584. [Google Scholar] [CrossRef]
- Ives, A.R.; Carpenter, S.R. Stability and diversity of ecosystems. Science 2007, 317, 58–62. [Google Scholar] [CrossRef]
- Miller, A.W.; Oakeson, K.F.; Dale, C.; Dearing, M.D. Microbial Community Transplant Results in Increased and Long-Term Oxalate Degradation. Microb. Ecol. 2016, 72, 470–478. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, K.; Yokoi, T.; Sugiyama, M.; Osawa, R.; Mukai, T.; Okada, N. Roles of the Cell Surface Architecture of Bacteroides and Bifidobacterium in the Gut Colonization. Front. Microbiol. 2021, 12, 754819. [Google Scholar] [CrossRef]
- Han, G.G.; Lee, J.Y.; Jin, G.D.; Park, J.; Choi, Y.H.; Kang, S.K.; Chae, B.J.; Kim, E.B.; Choi, Y.J. Tracing of the fecal microbiota of commercial pigs at five growth stages from birth to shipment. Sci. Rep. 2018, 8, 6012. [Google Scholar] [CrossRef] [PubMed]
- Percy, D.H.; Ruhnke, H.L.; Soltys, M.A. A case of infectious cystitis and pyelonephritis of swine caused by Corynebacterium suis. Can. Vet. J. 1966, 7, 291–292. [Google Scholar]
- Bernard, K. The genus Corynebacterium and other medically relevant coryneform-like bacteria. J. Clin. Microbiol. 2012, 50, 3152–3158. [Google Scholar] [CrossRef]
- Kim, H.; Cho, J.H.; Song, M.; Cho, J.H.; Kim, S.; Kim, E.S.; Keum, G.B.; Kim, H.B.; Lee, J.H. Evaluating the Prevalence of Foodborne Pathogens in Livestock Using Metagenomics Approach. J. Microbiol. Biotechnol. 2021, 31, 1701–1708. [Google Scholar] [CrossRef]
- Chen, W.; Liu, F.; Ling, Z.; Tong, X.; Xiang, C. Human intestinal lumen and mucosa-associated microbiota in patients with colorectal cancer. PLoS ONE 2012, 7, e39743. [Google Scholar] [CrossRef]
- Berding, K.; Wang, M.; Monaco, M.H.; Alexander, L.S.; Mudd, A.T.; Chichlowski, M.; Waworuntu, R.V.; Berg, B.M.; Miller, M.J.; Dilger, R.N. Prebiotics and bioactive milk fractions affect gut development, microbiota, and neurotransmitter expression in piglets. J. Pediatr. Gastroenterol. Nutr. 2016, 63, 688–697. [Google Scholar] [CrossRef]
- Burrough, E.R.; Arruda, B.L.; Plummer, P.J. Comparison of the Luminal and Mucosa-Associated Microbiota in the Colon of Pigs with and without Swine Dysentery. Front. Vet. Sci. 2017, 4, 139. [Google Scholar] [CrossRef]
- Girolamo, F.D.; D’Amato, A.; Lante, I.; Signore, F.; Muraca, M.; Putignani, L. Farm animal serum proteomics and impact on human health. Int. J. Mol. Sci. 2014, 15, 15396–15411. [Google Scholar] [CrossRef] [PubMed]
- Greco, V.; Piras, C.; Pieroni, L.; Ronci, M.; Putignani, L.; Roncada, P.; Urbani, A. Applications of MALDI-TOF mass spectrometry in clinical proteomics. Expert Rev. Proteom. 2018, 15, 683–696. [Google Scholar] [CrossRef]

| Ingredients | g/Kg of DM |
|---|---|
| Corn | 550 |
| Broad bean | 125 |
| Peas beans | 110 |
| Sunflower meal (38% CP) | 80 |
| Wheat middling | 70 |
| Carob | 30 |
| Sugar cane molasses | 13 |
| Analytical components 1 | |
| CP | 17.4 |
| CFa | 5.7 |
| CFi | 4.5 |
| Ash | 5.3 |
| Calcium | 0.6 |
| Phosphorus | 0.5 |
| Sodium | 0.2 |
| Lysine | 1.2 |
| Methionine | 0.4 |
| Additive components | |
| Vitamin B1 | 1.0 mg |
| Vitamin B2 | 3.0 mg |
| Vitamin B6 | 1.5 mg |
| Vitamin B12 | 0.015 mg |
| Vitamin D3 | (1.000 UI) |
| Vitamin E | 20 mg |
| Vitamin K3 | 1.0 mg |
| Niacin | 15.0 mg |
| Calcium-D | 10.3 mg |
| Choline | 200 mg |
| Cu | 14.0 mg |
| Fe | 89.8 mg |
| I | 0.50 mg |
| Mn | 39.9 mg |
| Se | 0.15 mg |
| Zn | 99.6 mg |
| Biotin | 0.10 mg |
| DL-Methionine | 0.12 mg |
| Lysine | 500 mg |
| Phylum | Family | Genus | Log2 Fold Change * | p-Value | p-Adj |
|---|---|---|---|---|---|
| Actinobacteria | Bifidobacteriaceae | Bifidobacterium | 1.79 | 2.78 × 10−4 | 4.93 × 10−3 |
| Corynebacteriaceae | Corynebacterium | −1.35 | 4.81 × 10−3 | 4.02 × 10−2 | |
| Atopobiaceae | Olsenella | −2.34 | 1.21 × 10−4 | 2.87 × 10−3 | |
| Eggerthellaceae | Slackia | −1.78 | 2.23 × 10−3 | 2.43 × 10−2 | |
| Firmicutes | Lactobacillaceae | Lactobacillus | −1.11 | 6.28 × 10−3 | 4.69 × 10−2 |
| Clostridiales Family XIII. Incertae Sedis | Mogibacterium | −2.45 | 7.37 × 10−7 | 5.24 × 10−5 | |
| Lachnospiraceae | Blautia | −0.97 | 6.66 × 10−3 | 4.73 × 10−2 | |
| Cellulosilyticum | 1.97 | 3.74 × 10−3 | 3.31 × 10−2 | ||
| Coprococcus | −2.36 | 2.00 × 10−4 | 4.06 × 10−3 | ||
| Lachnospira | −1.62 | 2.64 × 10−3 | 2.68 × 10−2 | ||
| Lachnotalea | 1.79 | 1.63 × 10−3 | 2.19 × 10−2 | ||
| Ruminococcaceae | Anaeromassilibacillus | −2.05 | 9.01 × 10−4 | 1.42 × 10−2 | |
| Harryflintia | 0.07 | 6.77 × 10−8 | 9.62 × 10−6 | ||
| Ruminococcus | 1.41 | 4.77 × 10−5 | 1.35 × 10−3 | ||
| Saccharofermentans | 2.24 | 1.87 × 10−3 | 2.22 × 10−2 | ||
| Acidaminococcaceae | Phascolarctobacterium | 1.62 | 3.78 × 10−5 | 1.34 × 10−3 | |
| Proteobacteria | Oxalobacteraceae | Oxalobacter | 5.87 | 1.69 × 10−3 | 2.19 × 10−2 |
| Sutterellaceae | Duodenibacillus | −2.64 | 2.94 × 10−3 | 2.78 × 10−2 | |
| Parasutterella | 3.99 | 1.31 × 10−6 | 6.19 × 10−5 | ||
| Pasteurellaceae | Actinobacillus | −0.22 | 5.18 × 10−3 | 4.09 × 10−2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tardiolo, G.; Romeo, O.; Zumbo, A.; Di Marsico, M.; Sutera, A.M.; Cigliano, R.A.; Paytuví, A.; D’Alessandro, E. Characterization of the Nero Siciliano Pig Fecal Microbiota after a Liquid Whey-Supplemented Diet. Animals 2023, 13, 642. https://doi.org/10.3390/ani13040642
Tardiolo G, Romeo O, Zumbo A, Di Marsico M, Sutera AM, Cigliano RA, Paytuví A, D’Alessandro E. Characterization of the Nero Siciliano Pig Fecal Microbiota after a Liquid Whey-Supplemented Diet. Animals. 2023; 13(4):642. https://doi.org/10.3390/ani13040642
Chicago/Turabian StyleTardiolo, Giuseppe, Orazio Romeo, Alessandro Zumbo, Marco Di Marsico, Anna Maria Sutera, Riccardo Aiese Cigliano, Andreu Paytuví, and Enrico D’Alessandro. 2023. "Characterization of the Nero Siciliano Pig Fecal Microbiota after a Liquid Whey-Supplemented Diet" Animals 13, no. 4: 642. https://doi.org/10.3390/ani13040642
APA StyleTardiolo, G., Romeo, O., Zumbo, A., Di Marsico, M., Sutera, A. M., Cigliano, R. A., Paytuví, A., & D’Alessandro, E. (2023). Characterization of the Nero Siciliano Pig Fecal Microbiota after a Liquid Whey-Supplemented Diet. Animals, 13(4), 642. https://doi.org/10.3390/ani13040642






