Simple Summary
Canine malignant lymphoma is one of the most common neoplasias in dogs. Even though breed influences the prevalence and prognosis, most cases present resistance to traditional chemotherapy. Canine breeds such as Labrador and Golden Retriever often have a worse prognosis for this reason. This high resistance to anticancer drugs makes it necessary to search for other therapies and other therapeutic targets. In this sense, the epigenetic regulators of neoplasia could be a promising object of study in the search for new research advances.
Abstract
Canine malignant lymphoma is a common neoplasia in dogs, and some studies have used dogs as a research model for molecular mechanisms of lymphomas in humans. In two species, chemotherapy is the treatment of choice, but the resistance to conventional anticancer drugs is frequent. The knowledge of molecular mechanisms of development and progression of neoplasia has expanded in recent years, and the underlying epigenetic mechanisms are increasingly well known. These studies open up new ways of discovering therapeutic biomarkers. Histone deacetylases and demethylase inhibitors could be a future treatment for canine lymphoma, and the use of microRNAs as diagnosis and prognosis biomarkers is getting closer. This review summarises the epigenetic mechanisms underlying canine lymphoma and their possible application as treatment and biomarkers, both prognostic and diagnostic.
1. Introduction
Lymphomas are a common type of neoplasia in dogs, with prevalence estimated at around 100 cases per 100,000 dogs [1]. This prevalence depends on different variants, including canine breed. Labrador Retriever, Rottweiler and Boxer are breeds that present high prevalence of lymphomas [2,3,4]. Several clinical presentations and morphological subtypes have been studied, and the World Health Organization (WHO) classifies canine malignant lymphomas (CML) into different subtypes, regarding histopathology and immunohistochemistry, with two large groups according to origin: B-cell neoplasms and T-cell and putative natural killer cell neoplasms [5]. The most common clinical presentation includes multicentric, mediastinal, abdominal (gastrointestinal, hepatic, splenic and renal), cutaneous, ocular, central nervous system and pulmonary lymphoma [6]. Prognosis depends on type of lymphoma and clinical presentation, among other factors. In general, T-cell lymphomas present lower remission and survival time rates than B-cell lymphomas [5,7]. These forms are similar to human lymphomas, which have specific genetic abnormalities, including epigenetic changes, used in prognosis, diagnosis and treatment [8]. However, studies on this are very limited in CML. This review aims to summarise the molecular changes related to CML, epigenetics mechanisms and possible applications as biomarkers and treatments.
2. Canine Malignant Lymphomas (CML)
2.1. Aetiology of Lymphomas
CML is the most common haematopoietic neoplasm diagnosed in dogs and represents the most managed neoplasia in veterinary medical oncology [6]. Many studies of comparative oncology have investigated the epidemiology of CML, due to the similarities between CML and human non-Hodgkin’s lymphoma (NHL). CML shares similar characteristics of NHL patients, having similar clinical presentation, molecular and immunophenotypic composition, diagnosis, therapeutic protocols and treatment response [9,10]. These similarities make the dog one of the best models for the study of this human neoplasia. This disease consists of a heterogeneous group of lymphoid malignancies, so a multifactorial aetiology is considered, including age, sex, race/breed, autoimmune diseases, immunosuppression, and environmental factors.
In terms of age, CML presents most commonly in dogs from six to nine years old [3,11], whereas in humans, the average age of diagnosis for NHL is 67 years, with 57% of diagnostic cases found in those over 65 years of age [12]. Other factors related to NHL and CML are gender/sex. In fact, the influence of hormone status on the risk of NHL development has been extensively studied in human oncology, where the studies show a clear predominance of this disease in men [13,14] and poor prognosis, with 60% more likely to die of the disease [12]. In dogs, as in humans, several studies have reported a decreased risk in intact females [11,15]. Two mechanisms have been proposed to explain the reduced rate of NHL in females. The first is the antiproliferative effect of oestrogen on lymphoid cells through oestrogen receptor β signalling [16], and the second is related to interleukin 6 (IL6) serum levels. Low gene expression and IL6 protein production in peripheral blood mononuclear cells are related to low levels of 17β-oestradiol [17]. Considering that high serum levels of IL6 have been related to complete response and overall survival time [14,18], the elevated prevalence of NHL and poor prognosis in men could be explained by low levels of oestrogen.
In human oncology, race and ethnicity are considered risk factors for the development of NHL. Some studies have reported that white and non-Hispanic people are at higher risk of NHL than Asian/Pacific Islander, American Indian and black populations in the USA [12]. As in humans, canine breed is a relevant factor in CML. In fact, CML is over-represented in Labrador Retriever, Doberman, Rottweiler, Boxer, Bernese Mountain and Bullmastiff breeds [4]. For other breeds such as the Golden Retriever, the high prevalence has been reported only in the USA and Japan [19,20]. The genetic homogeneity of canine breeds facilitates the genetic studies related to lymphoma in dogs. A relation between canine breed and lymphoma immunophenotype has also been identified. Boxer, Shi-Tzu, Siberian Husky and Welsh Corgi have been linked to an increase of T-cell lymphoma, whereas Doberman, Rottweiler and German Shepherd have been related to B-cell lymphoma [4,21]. This breed predisposition suggests a genetic component in the pathogenesis of lymphoma. Specific mutations would explain the different prevalence of CML according to the canine breed. In one study, Modiano et al., (2005) identified patterns of chromosomal gains and losses as corresponding to the occurrence of B-cell and T-cell tumours in the Golden Retriever, a breed with a prevalence ratio of 1:1 B-cell/T-cell CML [3].
Other factors such as autoimmune disease and environmental characteristics have been correlated with NHL and CML. For example, several autoimmune diseases, such as Sjogren’s syndrome, systemic lupus erythematosus, celiac disease and scleroderma and immunosuppression due to infections or immunosuppressive therapy, have been associated with various subtypes of NHL [12]. In line with these results, some studies in veterinary medicine have reported high frequency of autoimmune diseases in dogs with CML [6]. Finally, the increased incidence of lymphoma in both humans and dogs suggests that environmental common risk factors might play a role in NHL and CML development. Some of the risks identified in the literature are exposure to chemicals and tobacco, living in industrial or farm areas and living near waste incinerators, radioactive or polluted sites [6,22]. Considering the environmental exposure of humans and their canine companions to be the same, the identification of canine genes that play a role in the pathogenesis of CML and its relationship with these risk factors may be useful for the identification of human cancer-associated genes and vice versa.
2.2. Diagnosis, Treatment and Prognosis of CML
The clinical signs associated with canine lymphoma are variable. In multicentric lymphoma, the most common form, there is usually generalised peripheral lymphadenopathy, painless, rubbery, hepatosplenomegaly and bone marrow involvement. Nonspecific signs such as anorexia, weight loss, vomiting, diarrhoea, emaciation, ascites, dyspnoea, polydipsia, polyuria and fever can appear [23]. However, the clinical features depend on the lymphoma presentation (Table 1).

Table 1.
Common clinical manifestations according to the type and presentation of the lymphoma.
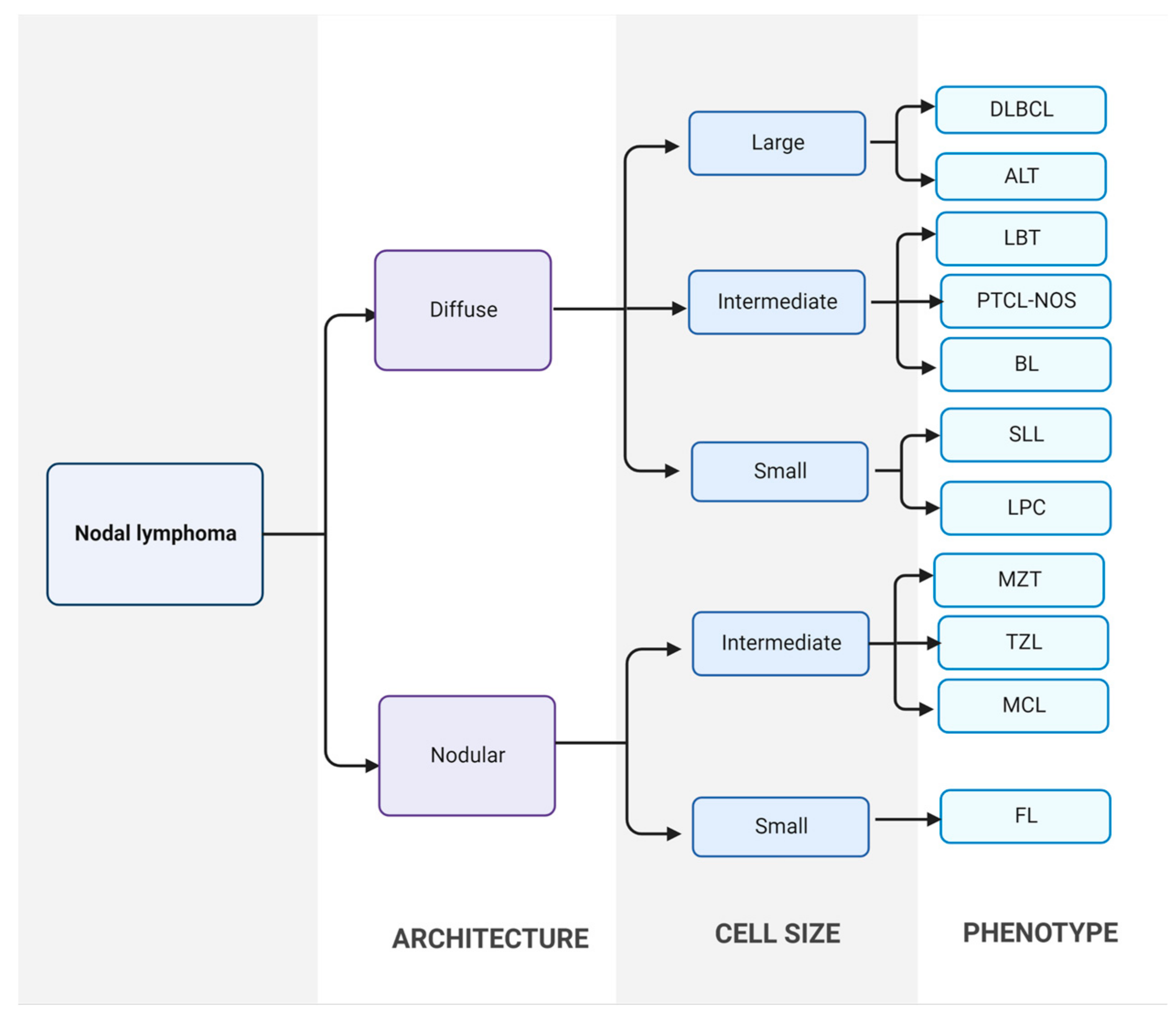
In clinical practice, the most common method for diagnosis of CML is by cytological examination of fine-needle aspirate from a neoplastic lymph node. Fine-needle aspiration is a cheap, quick, sensitive, and minimally invasive technique, making it the diagnostic method of choice for high-grade canine lymphomas. When the cytological examination is non-conclusive or needs to be confirmed, the histopathology study of an incisional, excisional or true-cut biopsy should be performed [9]. Then, lymphoma is classified based on histological and cytological tests, according to the WHO and Kiel classification, respectively (Figure 1) [5,27,28].
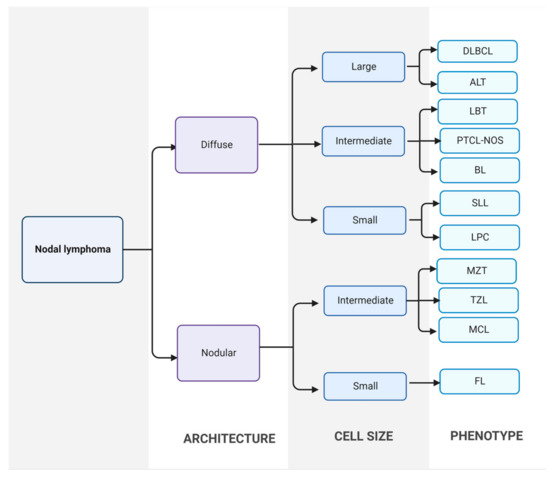
Figure 1.
Classification of canine nodal lymphoma. First, lymphomas are divided into diffuse or nodular forms using excisional lymph node sections. Then, neoplastic populations are divided into large, intermediate, and small cells. Finally, final diagnosis is established. DLBCL: diffuse large B-cell lymphoma; ALT: anaplastic large T-cell lymphoma; LBT: T-cell lymphoblastic lymphoma; PTCL-NOS: peripheral T-cell lymphoma; BL: B-cell lymphoma; SLL: small lymphocytic lymphoma; LPC: lymphoplasmacytic lymphoma; MZL: marginal zone lymphoma; TZL: T-zone lymphoma; MCL: mantle cell lymphoma; FL: follicular lymphoma [29].
In recent years, immunophenotyping has been carried out by immunohistochemistry, molecular methods to detect antigen receptor rearrangement (PARR) and flow cytometry to improve the first-choice treatment, according to this information. The latter method (flow cytometry), using fine-needle aspirates, is a minimally invasive method and assures rapid results. It is also an excellent tool for staging lymphoma, as it can identify neoplastic cell infiltration in different tissues, and one of the most recent and promising uses is the identification of minimal residual disease in peripheral blood, bone marrow and lymph nodes at the end of the chemotherapeutic protocol [30]. When flow cytometry or immunohistochemistry are not possible, the use of molecular techniques to detect antigen receptor rearrangement (PARR) can be used for diagnosing, staging and immunophenotyping in CML [31,32] in different samples, including fresh frozen tissue, formalin-fixed paraffin-embedded tissue, flow cytometry pellets and air-dried fine-needle aspirates [33]. This method is based on the fact that the lymphoma is a clonal lymphocyte expansion, so the neoplastic populations can be identified by detecting the presence of clonal populations of lymphocytes. This detection is carried out by amplification of the variable regions of immunoglobulin genes and T-cell receptor genes [34].
Multiagent protocols are the most used for the treatment of CML. The Wisconsin–Madison (CHOP) protocol, which combines cyclophosphamide, doxorubicin, vincristine and prednisolone, has the highest response rate and longest response durations [35]. Chemotherapy protocols follow two phases: the induction phase, which is a more intensive protocol that aims to induce complete remission, and in some cases, the clinician may choose to follow a less invasive maintenance protocol to maintain the remission status [6]. The recommendation to continue a maintenance chemotherapy protocol has been withdrawn, as it did not provide any treatment benefit after an induction protocol [35]. Another rescue protocol with good results is the LAP protocol, which combines lomustine, L-asparaginase and prednisone. The overall response with this rescue protocol has been estimated at around 77%, with 65% being a complete response [36].
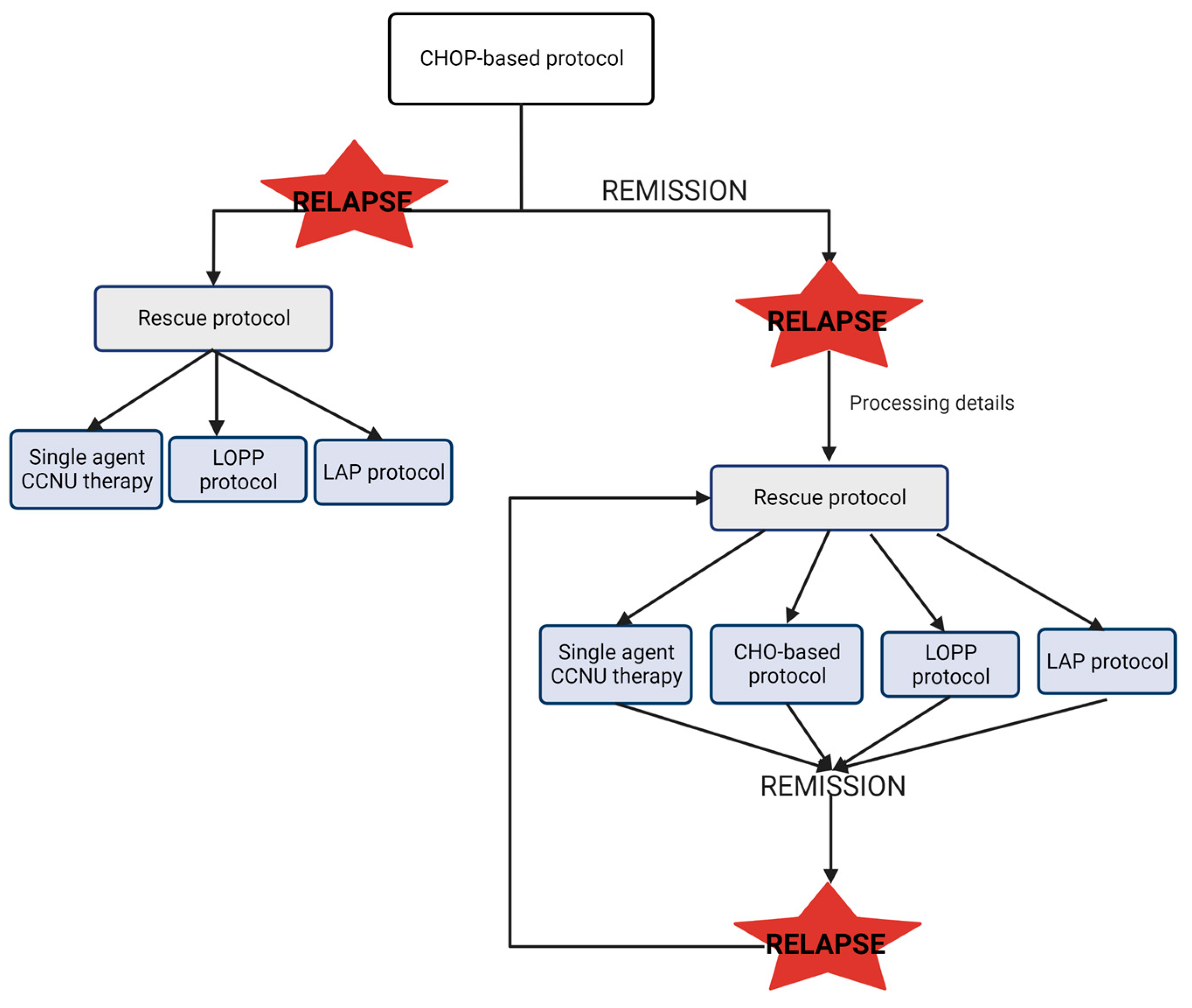
In the event of a relapse, different options are available, and the treatment choice will depend on the moment of relapse in relation to the original protocol and the individual clinician’s preference. The common protocol of treatment for CML is resumed in Figure 2. Briefly, if the relapse occurs during the first-line induction protocol, it may indicate drug resistance to the protocol used. In this case, the rescue protocol will require alternative drugs not included in the first protocol. On the contrary, if the relapse occurs following completion of the first-line protocol, sometimes the clinical decision is to repeat the first-line protocol or change to a different rescue protocol. In this scenario, the use of drugs from the original protocol is possible [37,38].
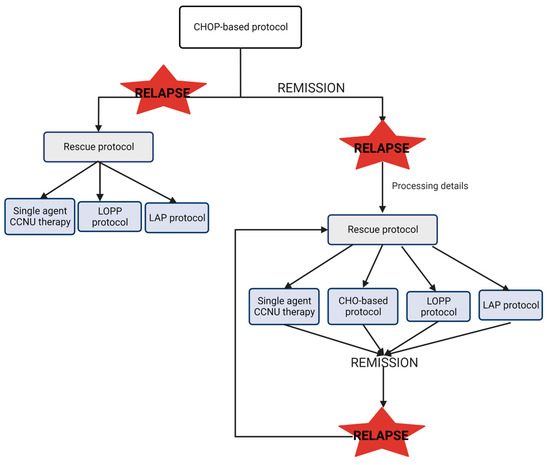
Figure 2.
Scheme of common treatment protocol in CML. CHOP: Wisconsin–Madison protocol; CCNU: lomustine; LOPP: vincristine procarbazine and prednisolone protocol. LAP: lomustine, L-asparaginase and prednisone protocol [35,36].
Unfortunately, the rescue protocol typically results in lower response rates and shorter response durations and in some cases presents more toxicity than the first-line protocol. Given the high prevalence of this neoplasm in dogs, its poor prognosis, and the high toxicity of chemotherapy drugs, mainly those used as rescue treatment, progress in new treatments and biomarkers, both prognostic and diagnostic, is increasingly important. The study of the molecular mechanisms that underlie the development of CML and the response to treatment is essential to improve both the diagnosis and the prognosis of patients.
3. Epigenetic Mechanisms and CML
3.1. Histone Modifications
Histone modifications, such as acetylation, phosphorylation and methylation, could change chromatic structure and affect transcription and gene expression [39]. Briefly, these modifications could activate or inactivate heterochromatin and euchromatin, respectively, adding methyl, acetyl, and phosphate groups.
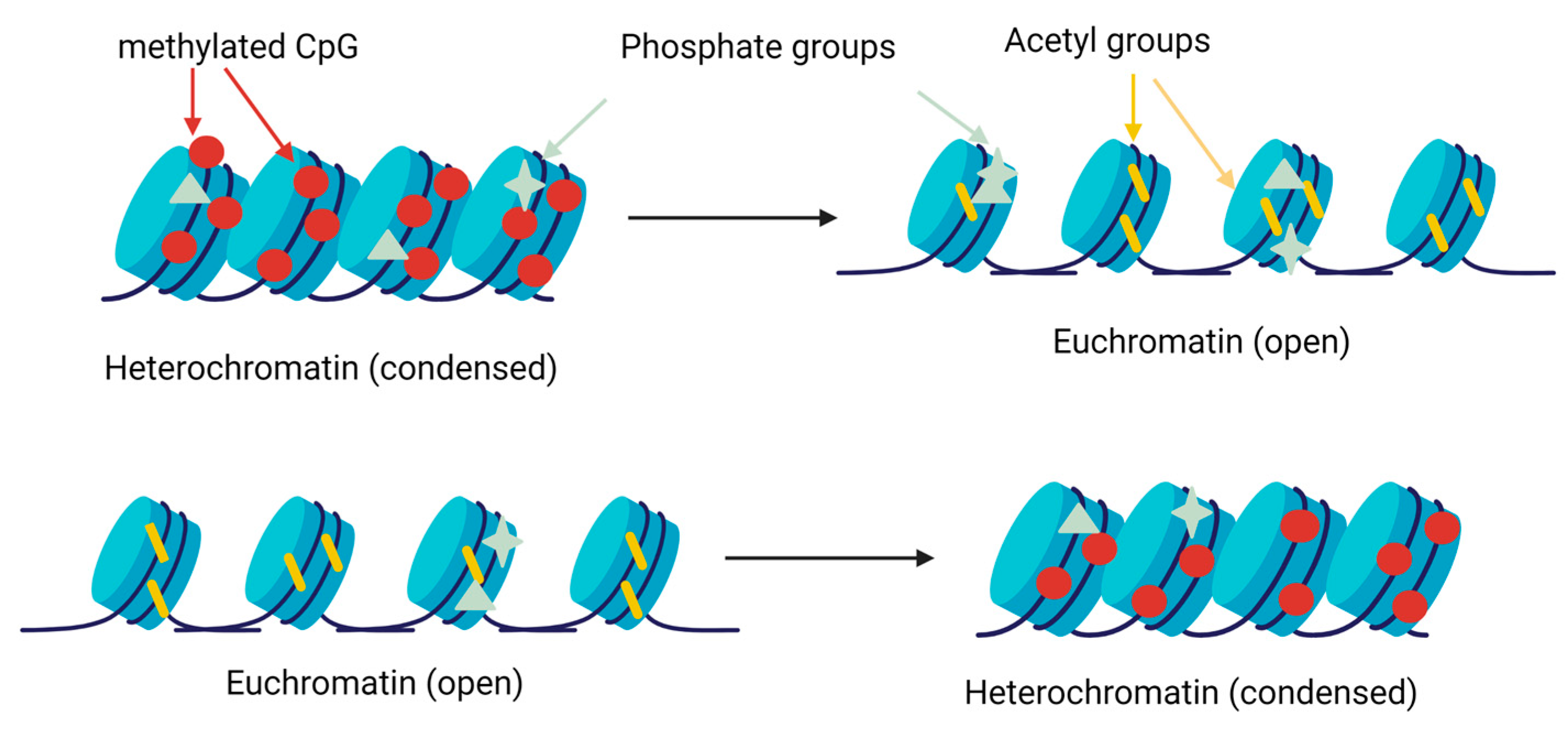
Histone methylation is carried out by specific methylases and demethylases on the residues of lysine and arginine [39,40]. Histone acetyltransferases (HAT) catalyse the transfer of an acetyl group of lysine, whereas histone deacetylases (HDAC) catalyse the reverse process [41]. Finally, histone kinases and phosphatase remove or add phosphate groups in the serine, threonine and tyrosine residues [42]. These changes modify the condensation of chromatin and facilitate or prevent gene expression (Figure 3).
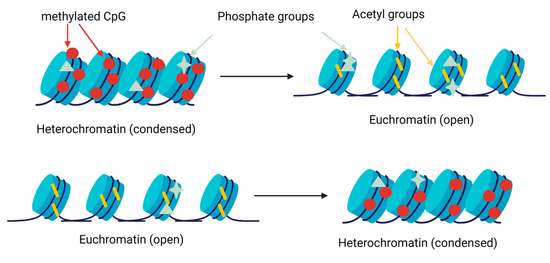
Figure 3.
Scheme of the gene expression regulation by histone modifications. Methyltransferases add methyl groups in lysine and arginine residues, favouring chromatin condensation and preventing gene expression. Acetyltransferases and deacetylases add or remove acetyl group in lysine residues, decreasing or increasing, respectively, the condensation of chromatin. The kinases and phosphatases add phosphate groups in different locations, changing the condensation of chromatin [43].
Histone modifications are related to regulation of different biological processes, including development, genomic imprinting and HOX gene expression [44]. Regarding tumour development and progression, several histone modifications have been found. For example, functional dysregulation of histone acetyltransferases has been associated with cancer development, including B-cell [45,46] and T-cell lymphomas [47,48]. In humans, histone modifier gene mutations have been associated with inferior progression-free survival time in patients with T-cell lymphomas [48], and it has been demonstrated that HDAC regulates the expression of BCL6, involved in cell survival and/or differentiation in B-cell lymphoma [49], and HDAC inhibitors have been tried as therapy in humans [50,51,52]. In CML, repression of p16 tumour suppressor gene is regulated by histone H3 acetylation in vitro [53].
3.2. DNA Methylation
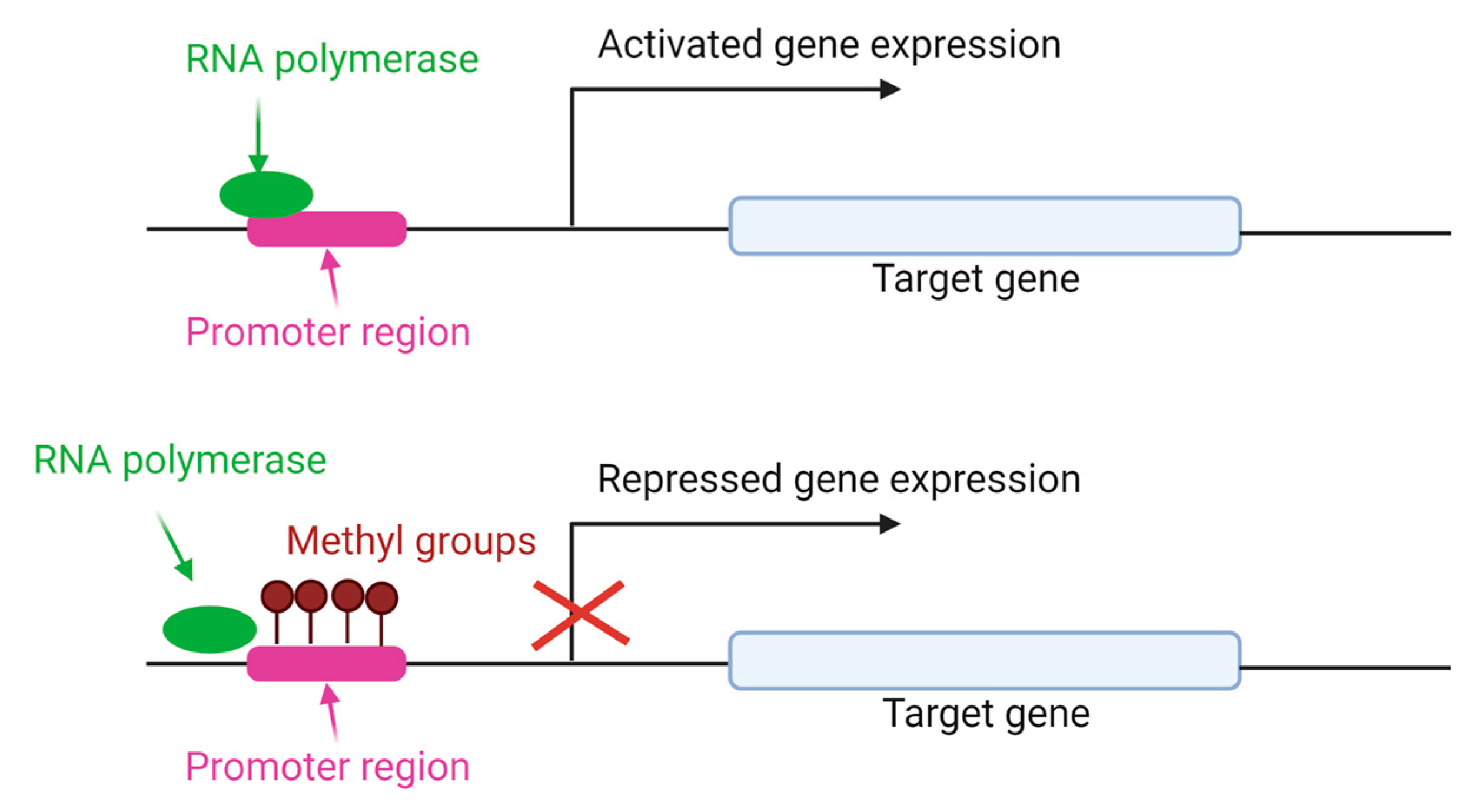
The addition of methyl groups to the cytosine CpG island of DNA is the most studied mechanism to regulate different processes, including chromosomal stability, genomic imprinting and X-chromosome inactivation [43]. DNA methylation is carried out primarily by three conserved enzymes in mammals: DNA methyltransferases (DNMT) 1, 3a and 3b [54], with different functions. DNMT1 maintains the methylation patterns during DNA replication and is expressed in somatic tissues and proliferating cells [55]. DNMT3a and 3b are de novo methyltransferases, and alternative transcripts with different functions are present in humans and mice [56]. As shown in Figure 4, DNMT adds methyl groups in the CpG island of promoter gene, preventing the RNA polymerase union and repressing the gene expression.
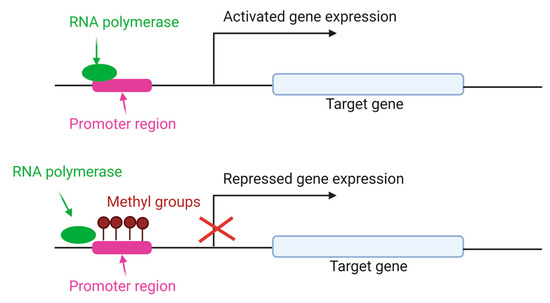
Figure 4.
Gene expression regulated by DNA methylation. DNMT enzymes add methyl groups in cytosine residues of CpG island. As a result, RNA polymerase is unable to bind to the DNA sequence of the gene promoter and the gene expression is repressed [57].
The relationship between DNA methylation aberrations and cancer has already been demonstrated in humans (see review [58]). Hypomethylation of several regions provokes the expression of oncogenes, whereas hypermethylation favours the development of cancer through the silencing of tumour suppressor genes. In dogs, the relationship between hypomethylation in genomic regions has been shown in canine leukaemia and lymphoma, where 30 and 69% of cases, respectively, were found. They are related to early phases of tumour transformation and progression [59]. In canine skin tumours, hypomethylation has been correlated to aggressiveness [60]. Hypomethylation has been found in canine osteosarcoma and lung cancer [61], whereas hypermethylation has been related to lymphoma [62], melanoma [63] and leukaemia [64], among other canine tumours, including canine mammary tumours. For example, hypomethylation of the intron region of the PAX6 motif of CDH2 and ADAM19 genes and hypermethylation at the PAX5 motifs in the intron regions of CDH5 and LRIG1 genes have been related to breast cancer [65].
Different GWAS (genome-wide association studies) have been carried out with relevant results. Specifically, in gastrointestinal lymphoma, 773 CpG islands are methylated, including promoter regions of 61 genes, including homeobox genes [66], whereas in high-grade B-cell lymphoma, Hsu et al., (2021) found 14 specific hypermethylated genes in samples of sick dogs but not in healthy dogs [67]. In canine diffuse large B-cell lymphoma, 1194 target loci are hypermethylated, including HOX, BMP and WNT genes, in promoter, 5′-UTRs upstream and exonic regions [68]. Hypomethylation in several regions and genes has been correlated with CML. Specifically, in dogs with NHL, higher DNA hypomethylation has been observed in circulating leukocytes [69].
In CML, not only methylated profiles have been studied. Different genes with relevant functions present methylation or non-methylation. In vitro studies have shown hypermethylation in promoter regions of TWIST2 (related to transcriptional regulation [70]) and TLX3 (orphan homeobox) genes in canine lymphoma cell lines [71]. Bryan et al., (2009) found abnormal hypermethylation of the CpG island in the DLC1 (tumour suppressor gene) promoter in canine NHL samples, although it is not associated with silencing expression gene or survival [62]. Other tumour suppressor genes are inhibited by hypermethylation in CML. For example, tissue factor pathway inhibitor-2 (TFPI-2) is hypermethylated in its promoter region, decreasing its expression [72], and death-associated protein kinase (DAPK) hypermethylation has been correlated to negative prognosis in canine B-cell lymphoma [73].
One of the most widely studied tumour suppressor genes in haematological neoplasia and others in different species is the p16 gene. The p16 protein inhibits the activity of cyclin-dependent kinase, as a negative control of the cell cycle to prevent phosphorylation of the retinoblastoma (pRb) protein. In fact, a high frequency of p16 mutations has been detected in many primary tumours and has been associated with progression of disease [74]. Related to CML, the inactivation of this gene by its CpG islands’ hypermethylation has been observed in canine lymphoid tumour cell lines [75], although results in vivo have demonstrated that p16 methylation status did not influence the prognosis [76]. Maylina et al., (2022) found a correlation between methylation of p16 gene, gene expression downregulation and hyperphosphorylation of pRb in canine lymphoma cell lines [77], and Fosmire et al., (2007) stated that inactivation of the p16 gene by methylation occurs in high-grade T-cell NHL but not in B-cell [78]. Another tumour suppressor gene, DAPK, has been found hypermethylated in canine nodal high-grade B-cell lymphomas [79,80]. Recently, silencing of tumour suppressor genes CIDEA, MAL and PCDH17 by hypermethylation in canine diffuse large B-cell lymphoma in vitro has been demonstrated [81].
Recent studies indicate that hypermethylome in canine B-cell lymphoma is conserved, unlike in humans [82], so these studies could improve the knowledge related to the molecular mechanism of CML and could open new lines of research to improve both treatments and prognostic and diagnostic biomarkers.
3.3. MicroRNAs (miRNAs)
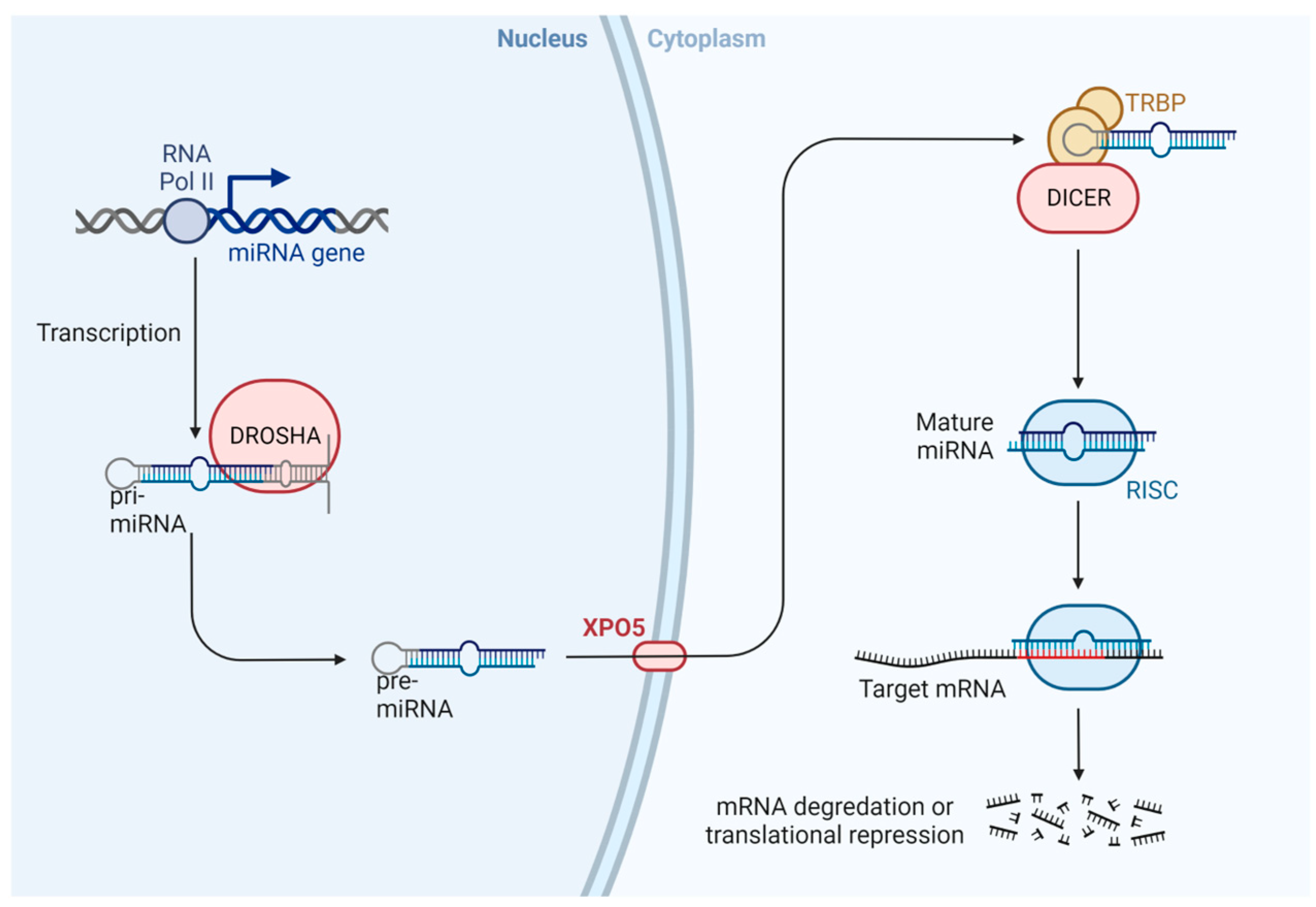
MicroRNAs are small non-coding RNA molecules of 20–24 base pairs, which are unable to code for proteins, with a crucial role in the regulation of gene expression and cellular processes [83,84]. The scheme of microRNAs is shown in Figure 5. Briefly, miRNA is transcribed by RNA polymerase II as the primary transcript (pri-miRNA) in the nucleus, and the enzyme Drosha cuts the pri-miRNA to form a pre-miRNA (70–90 nucleotides). Later, Exportin 5 (XPO5) transports pre-miRNA from the nucleus to the cytoplasm, where it is cut by the Dicer enzyme (regulated by XPO5), to form a mature and short double-stranded miRNA. One of these chains is degraded, and a single strand of miRNA is incorporated into the RISC complex (RNA-induced silencing complex). This complex is related to the silencing of mRNA and regulation post-transcriptional gene expression via inhibition of protein translation or destabilization of target transcripts [85,86,87,88,89,90,91].
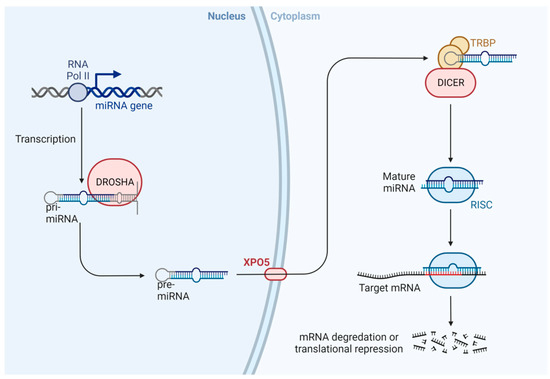
Figure 5.
Scheme of miRNA biogenesis. In the nucleus, miRNA is transcribed by RNA polymerase II as primary transcripts (pri-miRNA). The Drosha enzyme cuts this pri-miRNA to form a pre-miRNA, which is actively transported to the cytoplasm by the nuclear transport receptor exportin 5 (XPO5). In the cytoplasm, the pre-miRNA is cut by a second enzyme, Dicer, to form a mature and short double-stranded miRNA molecule. The miRNA duplex is incorporated into the RISC protein complex [92].
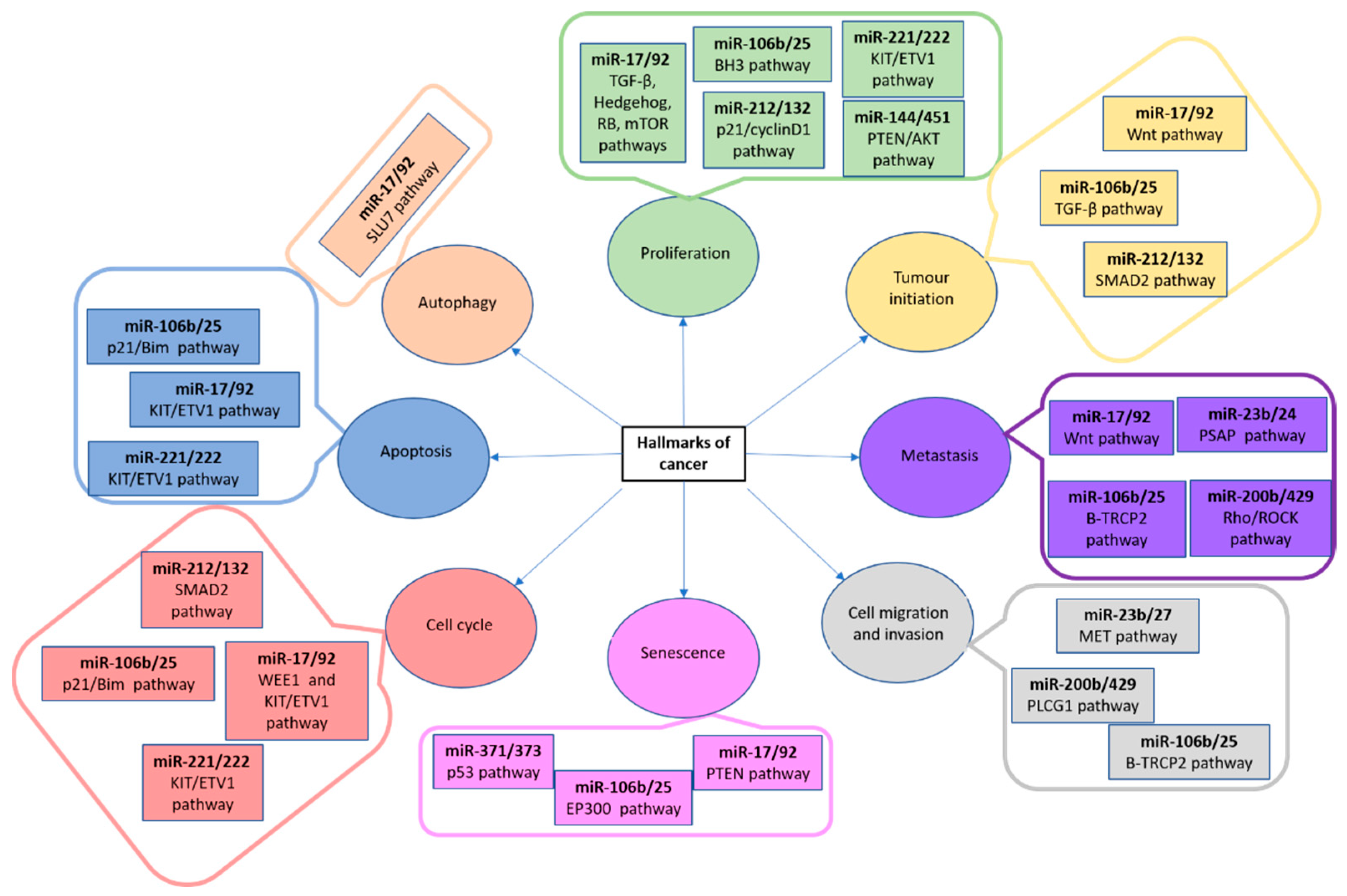
These small molecules of RNA are present both in intragenic and intergenic genome regions, are phylogenetically conserved [93] and have a crucial role in gene expression regulation and cellular processes, including neoplasia development [94,95]. In this sense, microRNAs are different target genes which have been related to proliferation, tumour initiation, metastasis, cell migration and invasion, senescence, cell cycle control, apoptosis and autophagy (Figure 6) [96,97,98,99,100,101,102].
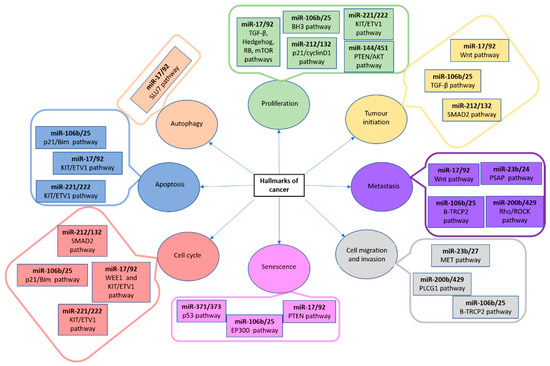
Figure 6.
MiRNA clusters, their pathways and associated with cancer hallmarks. AKT: AKT serine/threonine kinase; BH3: Bcl-2 homology 3 domain; BIM: Bcl-2-like protein 11; EP300: E1A-asosciated protein p300; ETV1: Ets variant gene 1; KIT: proto-oncogene tyrosine-protein kinase; MET: MET proto-oncogene receptor tyrosine kinase; mTOR: mammalian target of rapamycin complex 1; p21: cyclin-dependent kinase inhibitor 1A; PLCG1: phospholipase c gamma 1; PSAP: prosaposin; p53: tumour protein p53; PTEN: phosphatase and tensin homolog; RB1: RB transcriptional corepressor 1; RHO: ROCK Rho-associated protein kinase; SLU7: pre-mRNA splicing factor SLU7; SMAD2: mothers against dpp homolog 2; TRCP2: F-box and WD repeat domain containing 11; TGF: transforming growth factor; WEE1: Wee1A kinase; Wnt: wingless-type mmtv integration site family [57].
Several miRNAs have been related to lymphoma in humans. MiR-150 controls expression of c-Myc transcription factor [103], whereas miR-155 promotes lymphoma progression by the PI3K/AKT pathway regulation [104]. Other miRNAs related to B-cell lymphoma are miR-17, miR-18, miR-19, miR-21, miR-92 and miR-217 as oncomiRNAs and miR-181, miR-144, miR-27, miR-34a, miR-28, miR-145 and miR-146 as tumour suppressor miRNAs [105]. In T-cell lymphoma, several miRNAs have been associated with disease progression, such as miR-187, miR-17, miR-135, miR-155, miR-16, miR-29, miR-96, miR-146, miR-101, miR-30 and miR-21 [106]. Several trials with miRNAs inhibitors are being used as a treatment for lymphomas. Currently, phases I of miR-34 mimic (MRX34) [107] and anti-miR-155 (MRG-106) [108] have been finished for lymphoma treatment.
In dogs, some miRNAs have been studied. Craig et al., (2019) carried out an interesting study on canine multicentric lymphoma, where they demonstrated the relationship between altered expression of 16 miRNAs in lymph nodes and 15 miRNAs in plasma for B-cell lymphoma and 9 miRNAs in lymph nodes and 3 miRNAs in plasma for B-cell and T-cell lymphoma, respectively [109]. Specifically, these authors found a relationship between the miR-181b, -181a, -31, -99a, -146a, -30b and 130b with B-cell lymphoma diagnosis and miR-182, -143, -130b, -183, -450a, -450b, -181b and -23a with T-cell lymphoma diagnosis. Differential expression of miRNAs has been found in canine intestinal large B-cell lymphoma, so tumour suppressor miRNAs such as miR-194, miR-192, miR-141 and miR-203 are downregulated, whereas oncogenic miRNAs such as miR-106a are upregulated [110].
4. New Treatments and Biomarkers Based on Epigenetics in CML
4.1. Histone Deacetylase (HDAC) Inhibitors as Treatment
HDAC inhibitors (HDACi) are classified based on their chemical structure in four categories, such as carboxylate, benzamide, hydroxamate and cyclic peptide. In humans, hydroxamate-acid-based vorinostat (SAHA) was approved by the FDA in 2006 [111], whereas cyclin tetra peptide-based romidepsin and belistat were approved in 2011 [112] and 2014 [113] for the T-cell lymphoma, respectively. In 2015, Panobinostat was approved by the FDA and EMA for multiple myeloma treatments [114,115], and it is currently being studied in a phase II trial for lymphoma treatment [116], and mocetinostat is awaiting EMA approval for Hodgkin’s lymphoma [117].
The use and effectivity of HDACi in CML have been demonstrated in in vitro experiments. Dias et al., (2018) studied seven different HDACi in canine lymphoma cell lines, and their results indicated that Panobinostat is the most effective and with fewer toxicity compounds of the nine evaluated [118]. Two hydroxamate-based HDACi (Suberoylanilide hydroxamic acid-SHANA and Panobinostat) have shown demonstrated efficacy and low toxicity in dogs, and Zhang et al., (2016) evaluated the toxicity of HZ1006 (other hydroxamate-base HDACi) in dogs, with promising results [119]. However, the number of in vivo studies on the effectivity and toxicity of HDACi in dogs is very limited, so further studies of different HDACi as CML therapies are necessary to confirm the use of these molecules in this canine neoplasia.
4.2. Demethylation and Deacetylation Drugs as Treatment for Canine Lymphoma
Different demethylated agents have been tested for lymphoma treatment in humans. Inhibitors of lysine-specific demethylase 1 (LSD1) are some of the most studied. LSD1 regulates the transcription genes, repressing transcription by demethylation of histone H3 lysine 4 (H3K4) [120] and activating transcription by demethylation of histone 3 lysine 9 (H3K9) [121]. Different clinical trials are being carried out to evaluate the efficacy and toxicity of various LSD1 inhibitory molecules. Domatinostat (4SC-202) is an HDACi and LSD1 inhibitor, which induced cell death in T-cell lymphoma cell lines in vitro [122]. Hollebecque et al., (2022) showed the results of a phase I clinical trials with an oral LSD1 inhibitor (CC-90011) in NHL, indicating that this molecule presents a favourable tolerability profile, good clinical activity and durable responses [123]. The same research group has demonstrated that LSD1 expression in NHL is associated with chemoresistance [124], so the use of LSD1 inhibitors could be a new approach in drug-resistant lymphomas. In fact, LSD1 inhibitor GSK2879552 showed results in the phase I trial to relapsed/refractory myelodysplastic syndromes treatment [125]. Other LSD1 inhibitors are being tested. For example, ZY0511 is a potent LSD1 inhibitor, which induces the methylation of H3K4 and H3K9 in B-cell lymphoma, promoting apoptosis in both in vitro and in vivo xenograft experiments [126]. Regarding chemoresistance, high levels of inhibitor of histone 3 Lys 27 (KDM6B) are associated with poor prognosis and survival in patients with diffuse large B-cell lymphoma, and its inhibitor GSK-J4 induces apoptosis in cell lines with high levels of KDM6B [127]. These results suggest that these inhibitors could be the treatment of choice in lymphomas resistant to other conventional treatments. Other molecules related to lymphoma progression have been found, such as histone methylation inhibitors, including PKF118-310 (a KDM4A inhibitor) [128] and histone H3 Lys 27 demethylase [129], which could be used in future lymphoma treatments.
In CML, some methylation regulators have also been tested and could be a new possible treatment. For example, antimetabolite 6-thioguanine (6-TG) causes downregulation of DNA methyltransferase 1 in canine lymphoma cell lines, increasing cell survival [130]. Recently, Itoh et al., (2021) demonstrated that olsalazine reduces DNA methylation, including the methylation in the promoter regions of ADAM23, FES and CREB3L1 genes, inhibiting cell proliferation in canine lymphoid tumour cell lines [131]. Similar results have been found with azacytidine and decitabine, two hypomethylating drugs. However, in vivo studies in mice indicated that these drugs did not arrest tumour growth, although the methylation in the target gene promoter was reduced [132]. Sloan et al., (2021) showed an overexpression of protein arginine methyltransferase 5 (PRMT5) in primary B-cell canine lymphomas, arguing that PRMT5 inhibitors could be a candidate as a new therapy in CML [133].
Likewise, methylation has been related to chemoresistance in CML, so demethylated drugs are interesting as candidate therapy in refractory and/or chemo-resistant CML. Several studies have observed differences between drug-resistant cell methylation patterns. Thus, the CpG island in the upstream region of the exon 2 in the ABCB1 gene is hypermethylated (and presents low expression) in drug-sensitive canine lymphoma cell lines, whereas it is hypomethylated in drug-resistant canine lymphoma cell lines, and these results have been corroborated in in vivo studies [134]. Furthermore, O-methyl guanine DNA methyltransferase expression has been correlated to lomustine resistance in canine lymphoma cell lines [135], and canine cell lines resistant to doxorubicin show hypermethylation in gene promoter regions and low expression of 24 differentiated genes [67]. These results indicate that, as in humans, demethylating drugs could be interesting to use in CML with poor prognosis or resistant to chemotherapy.
Regarding deacetylation drugs, a multicentric study in canine lymphoma shows that treatment with sulforaphane, an isothiocyanate derived from some vegetables, is associated with change in proteome [136], although its results in clinical practice are as yet unknown. However, the effect of sulforaphane as an HDAC and DNMT inhibitor has been proven in human cancer prostate cells [137], acute lymphoblastic leukaemia cells [138] and multiple myeloma [139]. Other HDAC inhibitors have been studied as potential canine lymphoma treatments, such as class I HDAC inhibitors [140] or BET and SYK inhibitors [141,142].
4.3. microRNAs as Biomarkers and Treatment in CML
Alteration in miRNA expression has been related to different human diseases [143], including malignant tumours, where miRNAs have relevant roles in proliferation, apoptosis, invasion, metastasis and angiogenesis [144]. Several miRNAs have been related to CML and some of them have been postulated as possible biomarkers and/or therapeutic targets [145]. Two of the most studied miRNAs in CML are miR-155 and miR-17-5p. The former, miR-155, is downregulated in canine diffuse large B-cell lymphoma [146], whereas miR-17-5p is upregulated in lymph nodes of dogs with canine B-cell lymphomas [147]. Moreover, the molar ratio between these miRNAs in plasma is correlated with the WHO gradient classification [148], so this ratio could be used as a minimally invasive biomarker for diagnostic classification. miR-181 is an miRNA with tumour suppressor functions, and it is also downregulated in the lymph nodes of dogs with T-cell lymphomas [149]. Other miRNAs with tumour suppressor functions such as miR-203 and miR-218 present low expression in canine B- and T-cell lymphomas, as well as in humans, whereas miRNAs with oncogene upregulation function, such as miR-19a, miR-19b and miR17-5p, present high expression [147]. These results show that inhibitors of these miRNAs could be used in new treatments or as biomarkers in CML. Elshafie et al., (2021) observed an upregulation and downregulation of miR-34a and let-7 family miRNAs, respectively, with their results suggesting that the latter family of miRNAs alone can discriminate neoplastic canine diffuse B-cell lymphoma and non-neoplastic tissue in 97% of cases [146]. However, not only these miRNAs have been postulated as possible biomarkers in CML. The serum expression levels of let-7b, miR-223, miR-25 and miR-92a and mi-423a present reduced expression levels in dogs with lymphoma compared to healthy dogs. Specifically, miR-25 levels are related to histological grade in CML [150]. To determine the possible use of some miRNAs as biomarkers, Craig et al., (2019) analyzed the expression of 38 miRNAs both in plasma and lymph nodes in multicentric B- and T-cell lymphoma compared to healthy dogs. Their results showed altered expression in lymph nodes for 16 and 9 miRNAs in B- and T-cell lymphoma, respectively. Different miRNAs presented altered expression in plasma samples, without correlation with lymph node results. In fact, results in plasma show 15 and 3 miRNAs with altered expression in B- and T-cell lymphoma, respectively. In this study, the authors correlated expression levels of miRNAs with progression of diseases, correlating low expression of 10 miRNAs with progression-free survival and 3 with overall survival [109]. Table 2 summarises the miRNAs related to CML found in this study and others, the target gene of miRNAs, possible function and therapeutic applications.

Table 2.
MicroRNAs (miRNAs) related to canine malignant lymphoma (CML), target of these miRNAs, result of aberrant expression and possible uses.
Some altered expression miRNAs have been related to a specific clinical presentation of CML. High serum levels of miR-122 have been found in CML involving the liver [166], whereas canine intestinal T-cell lymphoma presents specific expression patterns, including downregulation of miR-194, miR-192, miR-141 and miR-203 and upregulation of the miR-106a cluster [110].
In recent years, a new approach has been studied. Plasma levels of circulating extracellular vesicles (EV) and the miRNAs present in these EV are being postulated as new possible biomarkers. Indeed, the plasma EV population is altered in human and canine diffuse large B-cell lymphomas [167] and related to progression of disease [168]. In addition, exosomes differ in their miRNAs content in cell lines resistant to chemotherapy; thus, the content of miR-151, miR-8908a-3p and miR-486 in plasma exosomes is different between vincristine-sensitive and resistant canine lymphoid tumour cell lines [169]. These results promote new lines of research in the search for biomarkers in malignant lymphoma in both humans and dogs.
5. Conclusions
The traditional treatment consists of radio- and chemotherapy, with poor results in many cases. The epigenetic mechanisms underlying canine lymphoma are increasingly well known, which implies that their possible application both as new treatments and/or biomarkers for prognosis and diagnosis is getting closer. In humans, therapies based on histone deacetylase and demethylase inhibitors have been approved in recent years. MiRNAs have been used as biomarkers in other tumours, so their use and application in the near future will probably improve CML prognosis and early diagnosis.
Author Contributions
Writing—original draft preparation, E.M.-A. and P.J.M.-G.; writing—review, editing and supervision, L.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding. Article Processing Change of publication was invited to Lola Llobat and Pablo Jesús Marín-García.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We are grateful to the Veterinary Medicine Faculty and Clinical Veterinary Hospital of Universidad Cardenal Herrera-CEU.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dobson, J.M.; Samuel, S.; Milstein, H.; Rogers, K.; Wood, J.L.N. Canine Neoplasia in the UK: Estimates of Incidence Rates from a Population of Insured Dogs. J. Small Anim. Pract. 2002, 43, 240–246. [Google Scholar] [CrossRef] [PubMed]
- Van Rooyen, L.J.; Hooijberg, E.; Reyers, F. Breed Prevalence of Canine Lymphoma in South Africa. J. S. Afr. Vet. Assoc. 2018, 89, e1–e11. [Google Scholar] [CrossRef] [PubMed]
- Modiano, J.F.; Breen, M.; Burnett, R.C.; Parker, H.G.; Inusah, S.; Thomas, R.; Avery, P.R.; Lindblad-Toh, K.; Ostrander, E.A.; Cutter, G.C.; et al. Distinct B-Cell and T-Cell Lymphoproliferative Disease Prevalence among Dog Breeds Indicates Heritable Risk. Cancer Res. 2005, 65, 5654–5661. [Google Scholar] [CrossRef] [PubMed]
- Comazzi, S.; Marelli, S.; Cozzi, M.; Rizzi, R.; Finotello, R.; Henriques, J.; Pastor, J.; Ponce, F.; Rohrer-Bley, C.; Rütgen, B.C.; et al. Breed-Associated Risks for Developing Canine Lymphoma Differ among Countries: An European Canine Lymphoma Network Study. BMC Vet. Res. 2018, 14, 232. [Google Scholar] [CrossRef] [PubMed]
- Valli, V.E.; San Myint, M.; Barthel, A.; Bienzle, D.; Caswell, J.; Colbatzky, F.; Durham, A.; Ehrhart, E.J.; Johnson, Y.; Jones, C.; et al. Classification of Canine Malignant Lymphomas According to the World Health Organization Criteria. Vet. Pathol. 2011, 48, 198–211. [Google Scholar] [CrossRef]
- Zandvliet, M. Canine Lymphoma: A Review. Vet. Q. 2016, 36, 76–104. [Google Scholar] [CrossRef]
- Ponce, F.; Marchal, T.; Magnol, J.P.; Turinelli, V.; Ledieu, D.; Bonnefont, C.; Pastor, M.; Delignette, M.L.; Fournel-Fleury, C. A Morphological Study of 608 Cases of Canine Malignant Lymphoma in France with a Focus on Comparative Similarities between Canine and Human Lymphoma Morphology. Vet. Pathol. 2010, 47, 414–433. [Google Scholar] [CrossRef]
- Kluin, P.; Schuuring, E. Molecular Cytogenetics of Lymphoma: Where Do We Stand in 2010? Histopathology 2011, 58, 128–144. [Google Scholar] [CrossRef]
- Teske, E. Canine Malignant Lymphoma: A Review and Comparison with Human Non-Hodgkin’s Lymphoma. Vet. Q. 1994, 16, 209–219. [Google Scholar] [CrossRef]
- Vail, D.M.; MacEwen, E.G. Spontaneously Occurring Tumors of Companion Animals as Models for Human Cancer. Cancer Investig. 2000, 18, 781–792. [Google Scholar] [CrossRef]
- Yau, P.; Dhand, N.K.; Thomson, P.C.; Taylor, R.M. Retrospective Study on the Occurrence of Canine Lymphoma and Associated Breed Risks in a Population of Dogs in NSW (2001–2009). Aust. Vet. J. 2017, 95, 149–155. [Google Scholar] [CrossRef]
- Thandra, K.C.; Barsouk, A.; Saginala, K.; Padala, S.A.; Barsouk, A.; Rawla, P. Epidemiology of Non-Hodgkin’s Lymphoma. Med. Sci. 2021, 9, 5. [Google Scholar] [CrossRef]
- Alexander, D.D.; Mink, P.J.; Adami, H.-O.; Chang, E.T.; Cole, P.; Mandel, J.S.; Trichopoulos, D. The Non-Hodgkin Lymphomas: A Review of the Epidemiologic Literature. Int. J. Cancer 2007, 120 (Suppl. S12), 1–39. [Google Scholar] [CrossRef]
- Horesh, N.; Horowitz, N.A. Does Gender Matter in Non-Hodgkin Lymphoma? Differences in Epidemiology, Clinical Behavior, and Therapy. Rambam. Maimonides Med. J. 2014, 5, e0038. [Google Scholar] [CrossRef]
- Villamil, J.A.; Henry, C.J.; Hahn, A.W.; Bryan, J.N.; Tyler, J.W.; Caldwell, C.W. Hormonal and Sex Impact on the Epidemiology of Canine Lymphoma. J. Cancer Epidemiol. 2009, 2009, 591753. [Google Scholar] [CrossRef]
- Pierdominici, M.; Maselli, A.; Colasanti, T.; Giammarioli, A.M.; Delunardo, F.; Vacirca, D.; Sanchez, M.; Giovannetti, A.; Malorni, W.; Ortona, E. Estrogen Receptor Profiles in Human Peripheral Blood Lymphocytes. Immunol. Lett. 2010, 132, 79–85. [Google Scholar] [CrossRef]
- Rachoń, D.; Myśliwska, J.; Suchecka-Rachoń, K.; Wieckiewicz, J.; Myśliwski, A. Effects of Oestrogen Deprivation on Interleukin-6 Production by Peripheral Blood Mononuclear Cells of Postmenopausal Women. J. Endocrinol. 2002, 172, 387–395. [Google Scholar] [CrossRef]
- Preti, H.A.; Cabanillas, F.; Talpaz, M.; Tucker, S.L.; Seymour, J.F.; Kurzrock, R. Prognostic Value of Serum Interleukin-6 in Diffuse Large-Cell Lymphoma. Ann. Intern. Med. 1997, 127, 186–194. [Google Scholar] [CrossRef]
- Mizutani, N.; Goto-Koshino, Y.; Takahashi, M.; Uchida, K.; Tsujimoto, H. Clinical and Histopathological Evaluation of 16 Dogs with T-Zone Lymphoma. J. Vet. Med. Sci. 2016, 78, 1237–1244. [Google Scholar] [CrossRef]
- Seelig, D.M.; Avery, P.; Webb, T.; Yoshimoto, J.; Bromberek, J.; Ehrhart, E.J.; Avery, A.C. Canine T-Zone Lymphoma: Unique Immunophenotypic Features, Outcome, and Population Characteristics. J. Vet. Intern. Med. 2014, 28, 878–886. [Google Scholar] [CrossRef]
- Jankowska, U.; Jagielski, D.; Czopowicz, M.; Sapierzyński, R. The Animal-Dependent Risk Factors in Canine T-Cell Lymphomas. Vet. Comp. Oncol. 2017, 15, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Pastor, M.; Chalvet-Monfray, K.; Marchal, T.; Keck, G.; Magnol, J.P.; Fournel-Fleury, C.; Ponce, F. Genetic and Environmental Risk Indicators in Canine Non-Hodgkin’s Lymphomas: Breed Associations and Geographic Distribution of 608 Cases Diagnosed throughout France over 1 Year. J. Vet. Intern. Med. 2009, 23, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Vail, D.M.; Thamm, D.H.; Liptak, J.M. Hematopoietic Tumors. In Withrow and MacEwen’s Small Animal Clinical Oncology; Elsevier Health Sciences: Amsterdam, The Netherlands, 2019; pp. 688–772. [Google Scholar] [CrossRef]
- Lane, J.; Price, J.; Moore, A.; Dandrieux, J.R.S.; Clifford, C.; Curran, K.; Choy, K.; Cannon, C. Low-Grade Gastrointestinal Lymphoma in Dogs: 20 Cases (2010 to 2016). J. Small Anim. Pract. 2018, 59, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Moore, E.L.; Vernau, W.; Rebhun, R.B.; Skorupski, K.A.; Burton, J.H. Patient Characteristics, Prognostic Factors and Outcome of Dogs with High-Grade Primary Mediastinal Lymphoma. Vet. Comp. Oncol. 2018, 16, E45–E51. [Google Scholar] [CrossRef]
- Fontaine, J.; Bovens, C.; Bettenay, S.; Mueller, R.S. Canine Cutaneous Epitheliotropic T-Cell Lymphoma: A Review. Vet. Comp. Oncol. 2009, 7, 1–14. [Google Scholar] [CrossRef]
- Ponce, F.; Magnol, J.-P.; Ledieu, D.; Marchal, T.; Turinelli, V.; Chalvet-Monfray, K.; Fournel-Fleury, C. Prognostic Significance of Morphological Subtypes in Canine Malignant Lymphomas during Chemotherapy. Vet. J. 2004, 167, 158–166. [Google Scholar] [CrossRef]
- Valli, V.E.; Kass, P.H.; San Myint, M.; Scott, F. Canine Lymphomas: Association of Classification Type, Disease Stage, Tumor Subtype, Mitotic Rate, and Treatment with Survival. Vet. Pathol. 2013, 50, 738–748. [Google Scholar] [CrossRef]
- Seelig, D.M.; Avery, A.C.; Ehrhart, E.J.; Linden, M.A. The Comparative Diagnostic Features of Canine and Human Lymphoma. Vet. Sci. 2016, 3, 11. [Google Scholar] [CrossRef]
- Riondato, F.; Comazzi, S. Flow Cytometry in the Diagnosis of Canine B-Cell Lymphoma. Front. Vet. Sci. 2021, 8, 600986. [Google Scholar] [CrossRef]
- Thalheim, L.; Williams, L.E.; Borst, L.B.; Fogle, J.E.; Suter, S.E. Lymphoma Immunophenotype of Dogs Determined by Immunohistochemistry, Flow Cytometry, and Polymerase Chain Reaction for Antigen Receptor Rearrangements. J. Vet. Intern. Med. 2013, 27, 1509–1516. [Google Scholar] [CrossRef]
- Waugh, E.M.; Gallagher, A.; Haining, H.; Johnston, P.E.J.; Marchesi, F.; Jarrett, R.F.; Morris, J.S. Optimisation and Validation of a PCR for Antigen Receptor Rearrangement (PARR) Assay to Detect Clonality in Canine Lymphoid Malignancies. Vet. Immunol. Immunopathol. 2016, 182, 115–124. [Google Scholar] [CrossRef]
- Ehrhart, E.J.; Wong, S.; Richter, K.; Zismann, V.; Grimes, C.; Hendricks, W.; Khanna, C. Polymerase Chain Reaction for Antigen Receptor Rearrangement: Benchmarking Performance of a Lymphoid Clonality Assay in Diverse Canine Sample Types. J. Vet. Intern. Med. 2019, 33, 1392–1402. [Google Scholar] [CrossRef]
- Burnett, R.C.; Vernau, W.; Modiano, J.F.; Olver, C.S.; Moore, P.F.; Avery, A.C. Diagnosis of Canine Lymphoid Neoplasia Using Clonal Rearrangements of Antigen Receptor Genes. Vet. Pathol. 2003, 40, 32–41. [Google Scholar] [CrossRef]
- Moore, A.S. Treatment of T Cell Lymphoma in Dogs. Vet. Rec. 2016, 179, 277. [Google Scholar] [CrossRef]
- Saba, C.F.; Hafeman, S.D.; Vail, D.M.; Thamm, D.H. Combination Chemotherapy with Continuous L-Asparaginase, Lomustine, and Prednisone for Relapsed Canine Lymphoma. J. Vet. Intern. Med. 2009, 23, 1058–1063. [Google Scholar] [CrossRef]
- Brown, P.M.; Tzannes, S.; Nguyen, S.; White, J.; Langova, V. LOPP Chemotherapy as a First-Line Treatment for Dogs with T-Cell Lymphoma. Vet. Comp. Oncol. 2018, 16, 108–113. [Google Scholar] [CrossRef]
- Lautscham, E.M.; Kessler, M.; Ernst, T.; Willimzig, L.; Neiger, R. Comparison of a CHOP-LAsp-Based Protocol with and without Maintenance for Canine Multicentric Lymphoma. Vet. Rec. 2017, 180, 303. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of Chromatin by Histone Modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef]
- Copeland, R.A.; Solomon, M.E.; Richon, V.M. Protein Methyltransferases as a Target Class for Drug Discovery. Nat. Rev. Drug Discov. 2009, 8, 724–732. [Google Scholar] [CrossRef]
- Xhemalce, B.; Dawson, M.A.; Bannister, A.J. Histone Modifications. In Reviews in Cell Biology and Molecular Medicine; American Cancer Society: Atlanta, GA, USA, 2011; ISBN 978-3-527-60090-8. [Google Scholar]
- Oki, M.; Aihara, H.; Ito, T. Role of Histone Phosphorylation in Chromatin Dynamics and Its Implications in Diseases. Subcell Biochem. 2007, 41, 319–336. [Google Scholar]
- Smith, Z.D.; Meissner, A. DNA Methylation: Roles in Mammalian Development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Jambhekar, A.; Dhall, A.; Shi, Y. Roles and Regulation of Histone Methylation in Animal Development. Nat. Rev. Mol. Cell Biol. 2019, 20, 625–641. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Serrano, M.; Winkler, R.; Santos, J.C.; Le Pannérer, M.-M.; Buschbeck, M.; Roué, G. Histone Modifications and Their Targeting in Lymphoid Malignancies. Int. J. Mol. Sci. 2021, 23, 253. [Google Scholar] [CrossRef] [PubMed]
- Shaknovich, R.; Melnick, A. Epigenetics and B-Cell Lymphoma. Curr. Opin. Hematol. 2011, 18, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Lai, P.; Wang, Y. Epigenetics of Cutaneous T-Cell Lymphoma: Biomarkers and Therapeutic Potentials. Cancer Biol. Med. 2021, 18, 34–51. [Google Scholar] [CrossRef]
- Ji, M.-M.; Huang, Y.-H.; Huang, J.-Y.; Wang, Z.-F.; Fu, D.; Liu, H.; Liu, F.; Leboeuf, C.; Wang, L.; Ye, J.; et al. Histone Modifier Gene Mutations in Peripheral T-Cell Lymphoma Not Otherwise Specified. Haematologica 2018, 103, 679–687. [Google Scholar] [CrossRef]
- Lemercier, C.; Brocard, M.-P.; Puvion-Dutilleul, F.; Kao, H.-Y.; Albagli, O.; Khochbin, S. Class II Histone Deacetylases Are Directly Recruited by BCL6 Transcriptional Repressor. J. Biol. Chem. 2002, 277, 22045–22052. [Google Scholar] [CrossRef]
- Yoshimitsu, M.; Ando, K.; Ishida, T.; Yoshida, S.; Choi, I.; Hidaka, M.; Takamatsu, Y.; Gillings, M.; Lee, G.T.; Onogi, H.; et al. Oral Histone Deacetylase Inhibitor HBI-8000 (Tucidinostat) in Japanese Patients with Relapsed or Refractory Non-Hodgkin’s Lymphoma: Phase I Safety and Efficacy. Jpn. J. Clin. Oncol. 2022, 52, 1014–1020. [Google Scholar] [CrossRef]
- Krämer, O.H.; Schneider, G. Single-Cell Profiling Guided Combination Therapy of c-Fos and Histone Deacetylase Inhibitors in Diffuse Large B-Cell Lymphoma. Clin. Transl. Med. 2022, 12, e858. [Google Scholar] [CrossRef]
- Zrimšek, M.; Kuchaříková, H.; Draganić, K.; Dobrovolná, P.; Heiss Spornberger, V.; Winkelmayer, L.; Hassler, M.R.; Lochmanová, G.; Zdráhal, Z.; Egger, G. Quantitative Acetylomics Uncover Acetylation-Mediated Pathway Changes Following Histone Deacetylase Inhibition in Anaplastic Large Cell Lymphoma. Cells 2022, 11, 2380. [Google Scholar] [CrossRef]
- Fujiwara-Igarashi, A.; Tomiyasu, H.; Igarashi, H.; Yu, Y.; Gogo-Koshino, Y.; Ohno, K.; Tsujimoto, H. Regulation of P16 Gene Expression by Histone H3 Acetylation in Canine Lymphoid Tumor Cell Lines. Jpn. J. Vet. Res. 2016, 64, 257–263. [Google Scholar]
- Okano, M.; Bell, D.W.; Haber, D.A.; Li, E. DNA Methyltransferases Dnmt3a and Dnmt3b Are Essential for de Novo Methylation and Mammalian Development. Cell 1999, 99, 247–257. [Google Scholar] [CrossRef]
- Deng, J.; Szyf, M. Multiple Isoforms of DNA Methyltransferase Are Encoded by the Vertebrate Cytosine DNA Methyltransferase Gene. J. Biol. Chem. 1998, 273, 22869–22872. [Google Scholar] [CrossRef][Green Version]
- Robertson, K.D.; Uzvolgyi, E.; Liang, G.; Talmadge, C.; Sumegi, J.; Gonzales, F.A.; Jones, P.A. The Human DNA Methyltransferases (DNMTs) 1, 3a and 3b: Coordinate MRNA Expression in Normal Tissues and Overexpression in Tumors. Nucleic Acids Res. 1999, 27, 2291–2298. [Google Scholar] [CrossRef]
- Llobat, L.; Gourbault, O. Role of MicroRNAs in Human Osteosarcoma: Future Perspectives. Biomedicines 2021, 9, 463. [Google Scholar] [CrossRef]
- Papanicolau-Sengos, A.; Aldape, K. DNA Methylation Profiling: An Emerging Paradigm for Cancer Diagnosis. Annu. Rev. Pathol. 2022, 17, 295–321. [Google Scholar] [CrossRef]
- Pelham, J.T.; Irwin, P.J.; Kay, P.H. Genomic Hypomethylation in Neoplastic Cells from Dogs with Malignant Lymphoproliferative Disorders. Res. Vet. Sci. 2003, 74, 101–104. [Google Scholar] [CrossRef]
- Morimoto, C.Y.; Tedardi, M.V.; da Fonseca, I.I.M.; Kimura, K.C.; Sanches, D.S.; Epiphanio, T.F.; de Francisco Strefezzi, R.; Dagli, M.L.Z. Evaluation of the Global DNA Methylation in Canine Mast Cell Tumour Samples by Immunostaining of 5-Methyl Cytosine. Vet. Comp. Oncol. 2017, 15, 1014–1018. [Google Scholar] [CrossRef]
- Herrera, C.L.; Kim, D.Y.; Kumar, S.R.; Bryan, J.N. Peroxisome Proliferator Activated Receptor γ Protein Expression Is Asymmetrically Distributed in Primary Lung Tumor and Metastatic to Lung Osteosarcoma Samples and Does Not Correlate with Gene Methylation. BMC Vet. Res. 2015, 11, 230. [Google Scholar] [CrossRef]
- Bryan, J.N.; Jabbes, M.; Berent, L.M.; Arthur, G.L.; Taylor, K.H.; Rissetto, K.C.; Henry, C.J.; Rahmatpanah, F.; Rankin, W.V.; Villamil, J.A.; et al. Hypermethylation of the DLC1 CpG Island Does Not Alter Gene Expression in Canine Lymphoma. BMC Genet. 2009, 10, 73. [Google Scholar] [CrossRef]
- Noguchi, S.; Mori, T.; Nakagawa, T.; Itamoto, K.; Haraguchi, T.; Mizuno, T. DNA Methylation Contributes toward Silencing of Antioncogenic MicroRNA-203 in Human and Canine Melanoma Cells. Melanoma Res. 2015, 25, 390–398. [Google Scholar] [CrossRef] [PubMed]
- Bronzini, I.; Aresu, L.; Paganin, M.; Marchioretto, L.; Comazzi, S.; Cian, F.; Riondato, F.; Marconato, L.; Martini, V.; Te Kronnie, G. DNA Methylation and Targeted Sequencing of Methyltransferases Family Genes in Canine Acute Myeloid Leukaemia, Modelling Human Myeloid Leukaemia. Vet. Comp. Oncol. 2017, 15, 910–918. [Google Scholar] [CrossRef] [PubMed]
- Nam, A.-R.; Lee, K.-H.; Hwang, H.-J.; Schabort, J.J.; An, J.-H.; Won, S.-H.; Cho, J.-Y. Alternative Methylation of Intron Motifs Is Associated with Cancer-Related Gene Expression in Both Canine Mammary Tumor and Human Breast Cancer. Clin. Epigenet. 2020, 12, 110. [Google Scholar] [CrossRef] [PubMed]
- Ohta, H.; Yamazaki, J.; Jelinek, J.; Ishizaki, T.; Kagawa, Y.; Yokoyama, N.; Nagata, N.; Sasaki, N.; Takiguchi, M. Genome-Wide DNA Methylation Analysis in Canine Gastrointestinal Lymphoma. J. Vet. Med. Sci. 2020, 82, 632–638. [Google Scholar] [CrossRef]
- Hsu, C.-H.; Tomiyasu, H.; Lee, J.-J.; Tung, C.-W.; Liao, C.-H.; Chuang, C.-H.; Huang, L.-Y.; Liao, K.-W.; Chou, C.-H.; Liao, A.T.C.; et al. Genome-Wide DNA Methylation Analysis Using MethylCap-Seq in Canine High-Grade B-Cell Lymphoma. J. Leukoc. Biol. 2021, 109, 1089–1103. [Google Scholar] [CrossRef]
- Ferraresso, S.; Aricò, A.; Sanavia, T.; Da Ros, S.; Milan, M.; Cascione, L.; Comazzi, S.; Martini, V.; Giantin, M.; Di Camillo, B.; et al. DNA Methylation Profiling Reveals Common Signatures of Tumorigenesis and Defines Epigenetic Prognostic Subtypes of Canine Diffuse Large B-Cell Lymphoma. Sci. Rep. 2017, 7, 11591. [Google Scholar] [CrossRef]
- Epiphanio, T.M.F.; de Azevedo Fernandes, N.C.C.; de Oliveira, T.F.; Lopes, P.A.; Réssio, R.A.; Gonçalves, S.; Scattone, N.V.; Tedardi, M.V.; Kulikowski, L.D.; Damasceno, J.; et al. Global DNA Methylation of Peripheral Blood Leukocytes from Dogs Bearing Multicentric Non-Hodgkin Lymphomas and Healthy Dogs: A Comparative Study. PLoS ONE 2019, 14, e0211898. [Google Scholar] [CrossRef]
- Chengxiao, Z.; Ze, Y. Biological Function and Molecular Mechanism of Twist2. Yi Chuan 2015, 37, 17–24. [Google Scholar] [CrossRef]
- Yamazaki, J.; Jelinek, J.; Hisamoto, S.; Tsukamoto, A.; Inaba, M. Dynamic Changes in DNA Methylation Patterns in Canine Lymphoma Cell Lines Demonstrated by Genome-Wide Quantitative DNA Methylation Analysis. Vet. J. 2018, 231, 48–54. [Google Scholar] [CrossRef]
- Ferraresso, S.; Bresolin, S.; Aricò, A.; Comazzi, S.; Gelain, M.E.; Riondato, F.; Bargelloni, L.; Marconato, L.; te Kronnie, G.; Aresu, L. Epigenetic Silencing of TFPI-2 in Canine Diffuse Large B-Cell Lymphoma. PLoS ONE 2014, 9, e92707. [Google Scholar] [CrossRef]
- Sato, M.; Mochizuki, H.; Goto-Koshino, Y.; Fujiwara-Igarashi, A.; Takahashi, M.; Ohno, K.; Tsujimoto, H. Prognostic Significance of Hypermethylation of Death-Associated Protein Kinase (DAPK) Gene CpG Island in Dogs with High-Grade B-Cell Lymphoma. Vet. Comp. Oncol. 2018, 16, 409–415. [Google Scholar] [CrossRef]
- Liggett, W.H.; Sidransky, D. Role of the P16 Tumor Suppressor Gene in Cancer. J. Clin. Oncol. 1998, 16, 1197–1206. [Google Scholar] [CrossRef]
- Fujiwara-Igarashi, A.; Goto-Koshino, Y.; Mochizuki, H.; Sato, M.; Fujino, Y.; Ohno, K.; Tsujimoto, H. Inhibition of P16 Tumor Suppressor Gene Expression via Promoter Hypermethylation in Canine Lymphoid Tumor Cells. Res. Vet. Sci. 2014, 97, 60–63. [Google Scholar] [CrossRef]
- Fujiwara-Igarashi, A.; Goto-Koshino, Y.; Sato, M.; Maeda, S.; Igarashi, H.; Takahashi, M.; Fujino, Y.; Ohno, K.; Tsujimoto, H. Prognostic Significance of the Expression Levels of the P16, P15, and P14 Genes in Dogs with High-Grade Lymphoma. Vet. J. 2014, 199, 236–244. [Google Scholar] [CrossRef]
- Maylina, L.; Kambayashi, S.; Baba, K.; Okuda, M. Simultaneous Analysis of the P16 Gene and Protein in Canine Lymphoma Cells and Their Correlation with PRb Phosphorylation. Vet. Sci. 2022, 9, 393. [Google Scholar] [CrossRef]
- Fosmire, S.P.; Thomas, R.; Jubala, C.M.; Wojcieszyn, J.W.; Valli, V.E.O.; Getzy, D.M.; Smith, T.L.; Gardner, L.A.; Ritt, M.G.; Bell, J.S.; et al. Inactivation of the P16 Cyclin-Dependent Kinase Inhibitor in High-Grade Canine Non-Hodgkin’s T-Cell Lymphoma. Vet. Pathol. 2007, 44, 467–478. [Google Scholar] [CrossRef]
- Sato, M.; Mochizuki, H.; Goto-Koshino, Y.; Fujiwara-Igarashi, A.; Takahashi, M.; Fujino, Y.; Ohno, K.; Tsujimoto, H. Hypermethylation of the Death-Associated Protein Kinase CpG Island in Canine B-Cell Lymphoid Tumors. Vet. Immunol. Immunopathol. 2014, 161, 222–231. [Google Scholar] [CrossRef]
- Aresu, L. Canine Lymphoma, More Than a Morphological Diagnosis: What We Have Learned about Diffuse Large B-Cell Lymphoma. Front. Vet. Sci. 2016, 3, 77. [Google Scholar] [CrossRef]
- Zorzan, E.; Elgendy, R.; Guerra, G.; Da Ros, S.; Gelain, M.E.; Bonsembiante, F.; Garaffo, G.; Vitale, N.; Piva, R.; Marconato, L.; et al. Hypermethylation-Mediated Silencing of CIDEA, MAL and PCDH17 Tumour Suppressor Genes in Canine DLBCL: From Multi-Omics Analyses to Mechanistic Studies. Int. J. Mol. Sci. 2022, 23, 4021. [Google Scholar] [CrossRef]
- Chu, S.; Avery, A.; Yoshimoto, J.; Bryan, J.N. Genome Wide Exploration of the Methylome in Aggressive B-Cell Lymphoma in Golden Retrievers Reveals a Conserved Hypermethylome. Epigenetics 2022, 17, 2022–2038. [Google Scholar] [CrossRef]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of Small RNAs in Animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Doench, J.G.; Sharp, P.A. Specificity of MicroRNA Target Selection in Translational Repression. Genes Dev. 2004, 18, 504–511. [Google Scholar] [CrossRef] [PubMed]
- Morlando, M.; Ballarino, M.; Gromak, N.; Pagano, F.; Bozzoni, I.; Proudfoot, N.J. Primary MicroRNA Transcripts Are Processed Co-Transcriptionally. Nat. Struct. Mol. Biol. 2008, 15, 902–909. [Google Scholar] [CrossRef] [PubMed]
- Bohnsack, M.T.; Czaplinski, K.; Gorlich, D. Exportin 5 Is a RanGTP-Dependent DsRNA-Binding Protein That Mediates Nuclear Export of Pre-MiRNAs. RNA 2004, 10, 185–191. [Google Scholar] [CrossRef]
- Lund, E.; Güttinger, S.; Calado, A.; Dahlberg, J.E.; Kutay, U. Nuclear Export of MicroRNA Precursors. Science 2004, 303, 95–98. [Google Scholar] [CrossRef]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 Mediates the Nuclear Export of Pre-MicroRNAs and Short Hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef]
- Hutvágner, G.; McLachlan, J.; Pasquinelli, A.E.; Bálint, E.; Tuschl, T.; Zamore, P.D. A Cellular Function for the RNA-Interference Enzyme Dicer in the Maturation of the Let-7 Small Temporal RNA. Science 2001, 293, 834–838. [Google Scholar] [CrossRef]
- Ketting, R.F.; Fischer, S.E.; Bernstein, E.; Sijen, T.; Hannon, G.J.; Plasterk, R.H. Dicer Functions in RNA Interference and in Synthesis of Small RNA Involved in Developmental Timing in C. Elegans. Genes Dev. 2001, 15, 2654–2659. [Google Scholar] [CrossRef]
- Altuvia, Y.; Landgraf, P.; Lithwick, G.; Elefant, N.; Pfeffer, S.; Aravin, A.; Brownstein, M.J.; Tuschl, T.; Margalit, H. Clustering and Conservation Patterns of Human MicroRNAs. Nucleic Acids Res. 2005, 33, 2697–2706. [Google Scholar] [CrossRef]
- Gourbault, O.; Llobat, L. MicroRNAs as Biomarkers in Canine Osteosarcoma: A New Future? Vet. Sci. 2020, 7, 146. [Google Scholar] [CrossRef]
- Friedman, R.C.; Farh, K.K.-H.; Burge, C.B.; Bartel, D.P. Most Mammalian MRNAs Are Conserved Targets of MicroRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef]
- Heneghan, H.M.; Miller, N.; Kerin, M.J. MiRNAs as Biomarkers and Therapeutic Targets in Cancer. Curr. Opin. Pharmacol. 2010, 10, 543–550. [Google Scholar] [CrossRef]
- Macfarlane, L.-A.; Murphy, P.R. MicroRNA: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef]
- Conkrite, K.; Sundby, M.; Mukai, S.; Thomson, J.M.; Mu, D.; Hammond, S.M.; MacPherson, D. MiR-17∼92 Cooperates with RB Pathway Mutations to Promote Retinoblastoma. Genes Dev. 2011, 25, 1734–1745. [Google Scholar] [CrossRef]
- Jiang, H.; Wang, P.; Wang, Q.; Wang, B.; Mu, J.; Zhuang, X.; Zhang, L.; Yan, J.; Miller, D.; Zhang, H.-G. Quantitatively Controlling Expression of MiR-17∼92 Determines Colon Tumor Progression in a Mouse Tumor Model. Am. J. Pathol. 2014, 184, 1355–1368. [Google Scholar] [CrossRef]
- Hong, L.; Lai, M.; Chen, M.; Xie, C.; Liao, R.; Kang, Y.J.; Xiao, C.; Hu, W.-Y.; Han, J.; Sun, P. The MiR-17-92 Cluster of MicroRNAs Confers Tumorigenicity by Inhibiting Oncogene-Induced Senescence. Cancer Res. 2010, 70, 8547–8557. [Google Scholar] [CrossRef]
- Brockway, S.; Zeleznik-Le, N.J. WEE1 Is a Validated Target of the MicroRNA MiR-17-92 Cluster in Leukemia. Cancer Genet. 2015, 208, 279–287. [Google Scholar] [CrossRef][Green Version]
- Urtasun, R.; Elizalde, M.; Azkona, M.; Latasa, M.U.; García-Irigoyen, O.; Uriarte, I.; Fernández-Barrena, M.G.; Vicent, S.; Alonso, M.M.; Muntané, J.; et al. Splicing Regulator SLU7 Preserves Survival of Hepatocellular Carcinoma Cells and Other Solid Tumors via Oncogenic MiR-17-92 Cluster Expression. Oncogene 2016, 35, 4719–4729. [Google Scholar] [CrossRef]
- Smith, A.L.; Iwanaga, R.; Drasin, D.J.; Micalizzi, D.S.; Vartuli, R.L.; Tan, A.-C.; Ford, H.L. The MiR-106b-25 Cluster Targets Smad7, Activates TGF-β Signaling, and Induces EMT and Tumor Initiating Cell Characteristics Downstream of Six1 in Human Breast Cancer. Oncogene 2012, 31, 5162–5171. [Google Scholar] [CrossRef]
- Savita, U.; Karunagaran, D. MicroRNA-106b-25 Cluster Targets β-TRCP2, Increases the Expression of Snail and Enhances Cell Migration and Invasion in H1299 (Non Small Cell Lung Cancer) Cells. Biochem. Biophys. Res. Commun. 2013, 434, 841–847. [Google Scholar] [CrossRef]
- Xiao, C.; Calado, D.P.; Galler, G.; Thai, T.-H.; Patterson, H.C.; Wang, J.; Rajewsky, N.; Bender, T.P.; Rajewsky, K. MiR-150 Controls B-Cell Differentiation by Targeting the Transcription Factor c-Myb. Cell 2016, 165, 1027. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-D.; Li, X.-M.; Gu, J.-W.; Sun, X.-C. MiR-155 Regulates Lymphoma Cell Proliferation and Apoptosis through Targeting SOCS3/JAK-STAT3 Signaling Pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 7577. [Google Scholar] [CrossRef] [PubMed]
- Fuertes, T.; Ramiro, A.R.; de Yebenes, V.G. MiRNA-Based Therapies in B-Cell Non-Hodgkin Lymphoma. Trends Immunol. 2020, 41, 932–947. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhang, M. Epigenetic Alterations and Advancement of Treatment in Peripheral T-Cell Lymphoma. Clin. Epigenet. 2020, 12, 169. [Google Scholar] [CrossRef]
- Mirna Therapeutics, Inc. A Multicenter Phase I Study of MRX34, MicroRNA MiR-RX34 Liposomal Injection; Mirna Therapeutics, Inc.: Austin, TX, USA, 2016. Available online: clinicaltrials.gov (accessed on 27 November 2022).
- miRagen Therapeutics, Inc. A Phase 1 Dose-Ranging Study to Investigate the Safety, Tolerability, and Pharmacokinetics of MRG-106 Following Local Intratumoral, Subcutaneous, and Intravenous Administration in Subjects with Various Lymphomas and Leukemias; miRagen Therapeutics, Inc.: Waltham, MA, USA, 2020. Available online: clinicaltrials.gov (accessed on 27 November 2022).
- Craig, K.K.L.; Wood, G.A.; Keller, S.M.; Mutsaers, A.J.; Wood, R.D. MicroRNA Profiling in Canine Multicentric Lymphoma. PLoS ONE 2019, 14, e0226357. [Google Scholar] [CrossRef]
- Joos, D.; Leipig-Rudolph, M.; Weber, K. Tumour-Specific MicroRNA Expression Pattern in Canine Intestinal T-Cell-Lymphomas. Vet. Comp. Oncol. 2020, 18, 502–508. [Google Scholar] [CrossRef]
- Olsen, E.A.; Kim, Y.H.; Kuzel, T.M.; Pacheco, T.R.; Foss, F.M.; Parker, S.; Frankel, S.R.; Chen, C.; Ricker, J.L.; Arduino, J.M.; et al. Phase IIb Multicenter Trial of Vorinostat in Patients with Persistent, Progressive, or Treatment Refractory Cutaneous T-Cell Lymphoma. J. Clin. Oncol. 2007, 25, 3109–3115. [Google Scholar] [CrossRef]
- Piekarz, R.L.; Frye, R.; Turner, M.; Wright, J.J.; Allen, S.L.; Kirschbaum, M.H.; Zain, J.; Prince, H.M.; Leonard, J.P.; Geskin, L.J.; et al. Phase II Multi-Institutional Trial of the Histone Deacetylase Inhibitor Romidepsin as Monotherapy for Patients with Cutaneous T-Cell Lymphoma. J. Clin. Oncol. 2009, 27, 5410–5417. [Google Scholar] [CrossRef]
- Foss, F.; Advani, R.; Duvic, M.; Hymes, K.B.; Intragumtornchai, T.; Lekhakula, A.; Shpilberg, O.; Lerner, A.; Belt, R.J.; Jacobsen, E.D.; et al. A Phase II Trial of Belinostat (PXD101) in Patients with Relapsed or Refractory Peripheral or Cutaneous T-Cell Lymphoma. Br. J. Haematol. 2015, 168, 811–819. [Google Scholar] [CrossRef]
- EMA Farydak. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/farydak (accessed on 27 November 2022).
- Drug Approval Package. Available online: https://www.accessdata.fda.gov/drugsatfda_docs/nda/2015/205353Orig1s000TOC.cfm (accessed on 27 November 2022).
- Panobinostat May Be Active in Select Patients With Refractory DLBCL—Cancer Therapy Advisor. Available online: https://www.cancertherapyadvisor.com/home/cancer-topics/lymphoma/panobinostat-may-be-active-in-select-patients-with-refractory-dlbcl/ (accessed on 27 November 2022).
- Batlevi, C.L.; Crump, M.; Andreadis, C.; Rizzieri, D.; Assouline, S.E.; Fox, S.; van der Jagt, R.H.C.; Copeland, A.; Potvin, D.; Chao, R.; et al. A Phase 2 Study of Mocetinostat, a Histone Deacetylase Inhibitor, in Relapsed or Refractory Lymphoma. Br. J. Haematol. 2017, 178, 434–441. [Google Scholar] [CrossRef]
- Dias, J.N.R.; Aguiar, S.I.; Pereira, D.M.; André, A.S.; Gano, L.; Correia, J.D.G.; Carrapiço, B.; Rütgen, B.; Malhó, R.; Peleteiro, C.; et al. The Histone Deacetylase Inhibitor Panobinostat Is a Potent Antitumor Agent in Canine Diffuse Large B-Cell Lymphoma. Oncotarget 2018, 9, 28586–28598. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, X.; Yuan, B.; Ren, L.; Zhang, T.; Lu, G. Subchronic Toxicities of HZ1006, a Hydroxamate-Based Histone Deacetylase Inhibitor, in Beagle Dogs and Sprague-Dawley Rats. Int. J. Environ. Res. Public Health 2016, 13, 1190. [Google Scholar] [CrossRef]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A.; Shi, Y. Histone Demethylation Mediated by the Nuclear Amine Oxidase Homolog LSD1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef]
- Metzger, J.; Nolte, A.; Uhde, A.-K.; Hewicker-Trautwein, M.; Distl, O. Whole Genome Sequencing Identifies Missense Mutation in MTBP in Shar-Pei Affected with Autoinflammatory Disease (SPAID). BMC Genom. 2017, 18, 348. [Google Scholar] [CrossRef]
- Wobser, M.; Weber, A.; Glunz, A.; Tauch, S.; Seitz, K.; Butelmann, T.; Hesbacher, S.; Goebeler, M.; Bartz, R.; Kohlhof, H.; et al. Elucidating the Mechanism of Action of Domatinostat (4SC-202) in Cutaneous T Cell Lymphoma Cells. J. Hematol. Oncol. 2019, 12, 30. [Google Scholar] [CrossRef]
- Hollebecque, A.; Salvagni, S.; Plummer, R.; Niccoli, P.; Capdevila, J.; Curigliano, G.; Moreno, V.; de Braud, F.; de Villambrosia, S.G.; Martin-Romano, P.; et al. Clinical Activity of CC-90011, an Oral, Potent, and Reversible LSD1 Inhibitor, in Advanced Malignancies. Cancer 2022, 128, 3185–3195. [Google Scholar] [CrossRef]
- Hollebecque, A.; Salvagni, S.; Plummer, R.; Isambert, N.; Niccoli, P.; Capdevila, J.; Curigliano, G.; Moreno, V.; Martin-Romano, P.; Baudin, E.; et al. Phase I Study of Lysine-Specific Demethylase 1 Inhibitor, CC-90011, in Patients with Advanced Solid Tumors and Relapsed/Refractory Non-Hodgkin Lymphoma. Clin. Cancer Res. 2021, 27, 438–446. [Google Scholar] [CrossRef]
- Roboz, G.J.; Yee, K.; Verma, A.; Borthakur, G.; de la Fuente Burguera, A.; Sanz, G.; Mohammad, H.P.; Kruger, R.G.; Karpinich, N.O.; Ferron-Brady, G.; et al. Phase I Trials of the Lysine-Specific Demethylase 1 Inhibitor, GSK2879552, as Mono- and Combination-Therapy in Relapsed/Refractory Acute Myeloid Leukemia or High-Risk Myelodysplastic Syndromes. Leuk. Lymphoma 2022, 63, 463–467. [Google Scholar] [CrossRef]
- Liu, H.; Wei, J.; Sang, N.; Zhong, X.; Zhou, X.; Yang, X.; Zhang, J.; Zuo, Z.; Zhou, Y.; Yang, S.; et al. The Novel LSD1 Inhibitor ZY0511 Suppresses Diffuse Large B-Cell Lymphoma Proliferation by Inducing Apoptosis and Autophagy. Med. Oncol. 2021, 38, 124. [Google Scholar] [CrossRef]
- Mathur, R.; Sehgal, L.; Havranek, O.; Köhrer, S.; Khashab, T.; Jain, N.; Burger, J.A.; Neelapu, S.S.; Davis, R.E.; Samaniego, F. Inhibition of Demethylase KDM6B Sensitizes Diffuse Large B-Cell Lymphoma to Chemotherapeutic Drugs. Haematologica 2017, 102, 373–380. [Google Scholar] [CrossRef]
- Franci, G.; Sarno, F.; Nebbioso, A.; Altucci, L. Identification and Characterization of PKF118-310 as a KDM4A Inhibitor. Epigenetics 2017, 12, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Shen, L.; Stupack, D.G.; Bai, N.; Xun, J.; Ren, G.; Han, J.; Li, L.; Luo, Y.; Xiang, R.; et al. JMJD3 Promotes Survival of Diffuse Large B-Cell Lymphoma Subtypes via Distinct Mechanisms. Oncotarget 2016, 7, 29387–29399. [Google Scholar] [CrossRef] [PubMed]
- Flesner, B.K.; Kumar, S.R.; Bryan, J.N. 6-Thioguanine and Zebularine down-Regulate DNMT1 and Globally Demethylate Canine Malignant Lymphoid Cells. BMC Vet. Res. 2014, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Itoh, S.; Yamazaki, J.; Iwahana, M.; Tsukamoto, A. Olsalazine Inhibits Cell Proliferation and DNA Methylation in Canine Lymphoid Tumor Cell Lines. Pol. J. Vet. Sci. 2021, 24, 515–523. [Google Scholar] [CrossRef]
- Da Ros, S.; Aresu, L.; Ferraresso, S.; Zorzan, E.; Gaudio, E.; Bertoni, F.; Dacasto, M.; Giantin, M. Validation of Epigenetic Mechanisms Regulating Gene Expression in Canine B-Cell Lymphoma: An in Vitro and in Vivo Approach. PLoS ONE 2018, 13, e0208709. [Google Scholar] [CrossRef]
- Sloan, S.L.; Renaldo, K.A.; Long, M.; Chung, J.-H.; Courtney, L.E.; Shilo, K.; Youssef, Y.; Schlotter, S.; Brown, F.; Klamer, B.G.; et al. Validation of Protein Arginine Methyltransferase 5 (PRMT5) as a Candidate Therapeutic Target in the Spontaneous Canine Model of Non-Hodgkin Lymphoma. PLoS ONE 2021, 16, e0250839. [Google Scholar] [CrossRef]
- Tomiyasu, H.; Fujiwara-Igarashi, A.; Goto-Koshino, Y.; Fujino, Y.; Ohno, K.; Tsujimoto, H. Evaluation of DNA Methylation Profiles of the CpG Island of the ABCB1 Gene in Dogs with Lymphoma. Am. J. Vet. Res. 2014, 75, 835–841. [Google Scholar] [CrossRef]
- Kambayashi, S.; Minami, K.; Ogawa, Y.; Hamaji, T.; Hwang, C.C.; Igase, M.; Hiraoka, H.; Miyama, T.S.; Noguchi, S.; Baba, K.; et al. Expression of O(6)-Methylguanine-DNA Methyltransferase Causes Lomustine Resistance in Canine Lymphoma Cells. Can. J. Vet. Res. 2015, 79, 201–209. [Google Scholar]
- Parachini-Winter, C.; Bracha, S.; Ramsey, S.A.; Yang, L.; Ho, E.; Leeper, H.J.; Curran, K.M. Prospective Evaluation of the Lymph Node Proteome in Dogs with Multicentric Lymphoma Supplemented with Sulforaphane. J. Vet. Intern. Med. 2020, 34, 2036–2047. [Google Scholar] [CrossRef]
- Clarke, J.D.; Hsu, A.; Yu, Z.; Dashwood, R.H.; Ho, E. Differential Effects of Sulforaphane on Histone Deacetylases, Cell Cycle Arrest and Apoptosis in Normal Prostate Cells versus Hyperplastic and Cancerous Prostate Cells. Mol. Nutr. Food. Res. 2011, 55, 999–1009. [Google Scholar] [CrossRef]
- Suppipat, K.; Park, C.S.; Shen, Y.; Zhu, X.; Lacorazza, H.D. Sulforaphane Induces Cell Cycle Arrest and Apoptosis in Acute Lymphoblastic Leukemia Cells. PLoS ONE 2012, 7, e51251. [Google Scholar] [CrossRef]
- Jakubikova, J.; Cervi, D.; Ooi, M.; Kim, K.; Nahar, S.; Klippel, S.; Cholujova, D.; Leiba, M.; Daley, J.F.; Delmore, J.; et al. Anti-Tumor Activity and Signaling Events Triggered by the Isothiocyanates, Sulforaphane and Phenethyl Isothiocyanate, in Multiple Myeloma. Haematologica 2011, 96, 1170–1179. [Google Scholar] [CrossRef]
- Cui, H.; Hong, Q.; Wei, R.; Li, H.; Wan, C.; Chen, X.; Zhao, S.; Bu, H.; Zhang, B.; Yang, D.; et al. Design and Synthesis of HDAC Inhibitors to Enhance the Therapeutic Effect of Diffuse Large B-Cell Lymphoma by Improving Metabolic Stability and Pharmacokinetic Characteristics. Eur. J. Med. Chem. 2022, 229, 114049. [Google Scholar] [CrossRef]
- Sender, S.; Sultan, A.W.; Palmer, D.; Koczan, D.; Sekora, A.; Beck, J.; Schuetz, E.; Taher, L.; Brenig, B.; Fuellen, G.; et al. Evaluation of the Synergistic Potential of Simultaneous Pan- or Isoform-Specific BET and SYK Inhibition in B-Cell Lymphoma: An In Vitro Approach. Cancers 2022, 14, 4691. [Google Scholar] [CrossRef]
- Kong, W.; Sender, S.; Perez, S.V.; Sekora, A.; Ruetgen, B.; Junghanss, C.; Nolte, I.; Murua Escobar, H. Pan- and Isoform-Specific Inhibition of the Bromodomain and Extra-Terminal Proteins and Evaluation of Synergistic Potential With Entospletinib in Canine Lymphoma. Anticancer Res. 2020, 40, 3781–3792. [Google Scholar] [CrossRef]
- Hammond, S.M. An Overview of MicroRNAs. Adv. Drug Deliv. Rev. 2015, 87, 3–14. [Google Scholar] [CrossRef]
- Lee, Y.S.; Dutta, A. MicroRNAs in Cancer. Annu. Rev. Pathol. 2009, 4, 199–227. [Google Scholar] [CrossRef]
- Bryan, J.N. The Current State of Clinical Application of Serum Biomarkers for Canine Lymphoma. Front. Vet. Sci. 2016, 3, 87. [Google Scholar] [CrossRef]
- Elshafie, N.O.; do Nascimento, N.C.; Lichti, N.I.; Kasinski, A.L.; Childress, M.O.; Santos, A.P.D. MicroRNA Biomarkers in Canine Diffuse Large B-Cell Lymphoma. Vet. Pathol. 2021, 58, 34–41. [Google Scholar] [CrossRef]
- Uhl, E.; Krimer, P.; Schliekelman, P.; Tompkins, S.M.; Suter, S. Identification of Altered MicroRNA Expression in Canine Lymphoid Cell Lines and Cases of B- and T-Cell Lymphomas. Genes Chromosomes Cancer 2011, 50, 950–967. [Google Scholar] [CrossRef]
- Albonico, F.; Mortarino, M.; Avallone, G.; Gioia, G.; Comazzi, S.; Roccabianca, P. The Expression Ratio of MiR-17-5p and MiR-155 Correlates with Grading in Canine Splenic Lymphoma. Vet. Immunol. Immunopathol. 2013, 155, 117–123. [Google Scholar] [CrossRef] [PubMed]
- Mortarino, M.; Gioia, G.; Gelain, M.E.; Albonico, F.; Roccabianca, P.; Ferri, E.; Comazzi, S. Identification of Suitable Endogenous Controls and Differentially Expressed MicroRNAs in Canine Fresh-Frozen and FFPE Lymphoma Samples. Leuk. Res. 2010, 34, 1070–1077. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara-Igarashi, A.; Igarashi, H.; Mizutani, N.; Goto-Koshino, Y.; Takahashi, M.; Ohno, K.; Tsujimoto, H. Expression Profile of Circulating Serum MicroRNAs in Dogs with Lymphoma. Vet. J. 2015, 205, 317–321. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Ding, L.; Hu, Q.; Xia, J.; Sun, J.; Wang, X.; Xiong, H.; Gurbani, D.; Li, L.; Liu, Y.; et al. MicroRNA-218 Functions as a Tumor Suppressor in Lung Cancer by Targeting IL-6/STAT3 and Negatively Correlates with Poor Prognosis. Mol. Cancer 2017, 16, 141. [Google Scholar] [CrossRef] [PubMed]
- Tagawa, H.; Ikeda, S.; Sawada, K. Role of MicroRNA in the Pathogenesis of Malignant Lymphoma. Cancer Sci. 2013, 104, 801–809. [Google Scholar] [CrossRef]
- Chen, A.-H.; Qin, Y.-E.; Tang, W.-F.; Tao, J.; Song, H.-M.; Zuo, M. MiR-34a and MiR-206 Act as Novel Prognostic and Therapy Biomarkers in Cervical Cancer. Cancer Cell Int. 2017, 17, 63. [Google Scholar] [CrossRef]
- Nie, K.; Zhang, T.; Allawi, H.; Gomez, M.; Liu, Y.; Chadburn, A.; Wang, Y.L.; Knowles, D.M.; Tam, W. Epigenetic Down-Regulation of the Tumor Suppressor Gene PRDM1/Blimp-1 in Diffuse Large B-Cell Lymphomas: A Potential Role of the MicroRNA Let-7. Am. J. Pathol. 2010, 177, 1470–1479. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, J. MiR-223 Suppresses Proliferation and Promotes Apoptosis of Diffuse Large B-Cell Lymphoma Cells Through Lmo2 and MAPK Signaling Pathway. J. BUON 2021, 26, 580–586. [Google Scholar]
- Wu, T.; Hu, H.; Zhang, T.; Jiang, L.; Li, X.; Liu, S.; Zheng, C.; Yan, G.; Chen, W.; Ning, Y.; et al. MiR-25 Promotes Cell Proliferation, Migration, and Invasion of Non-Small-Cell Lung Cancer by Targeting the LATS2/YAP Signaling Pathway. Oxid. Med. Cell Longev. 2019, 2019, 9719723. [Google Scholar] [CrossRef]
- Elliott, E.K.; Hopkins, L.N.; Hensen, R.; Sutherland, H.G.; Haupt, L.M.; Griffiths, L.R. Epigenetic Regulation of MiR-92a and TET2 and Their Association in Non-Hodgkin Lymphoma. Front. Genet. 2021, 12, 768913. [Google Scholar] [CrossRef]
- Mazzoccoli, L.; Robaina, M.C.; Bacchi, C.E.; Soares Lima, S.C.; Klumb, C.E. MiR-29 Promoter and Enhancer Methylation Identified by Pyrosequencing in Burkitt Lymhoma Cells: Interplay between MYC and MiR-29 Regulation. Oncol. Rep. 2019, 42, 775–784. [Google Scholar] [CrossRef]
- Thompson, M.A.; Edmonds, M.D.; Liang, S.; McClintock-Treep, S.; Wang, X.; Li, S.; Eischen, C.M. MiR-31 and MiR-17-5p Levels Change during Transformation of Follicular Lymphoma. Hum. Pathol. 2016, 50, 118–126. [Google Scholar] [CrossRef]
- Gao, P.; Tchernyshyov, I.; Chang, T.-C.; Lee, Y.-S.; Kita, K.; Ochi, T.; Zeller, K.I.; De Marzo, A.M.; Van Eyk, J.E.; Mendell, J.T.; et al. C-Myc Suppression of MiR-23a/b Enhances Mitochondrial Glutaminase Expression and Glutamine Metabolism. Nature 2009, 458, 762–765. [Google Scholar] [CrossRef]
- Niu, F.; Kazimierska, M.; Nolte, I.M.; Terpstra, M.M.; de Jong, D.; Koerts, J.; van der Sluis, T.; Rutgers, B.; O’Connell, R.M.; Kok, K.; et al. The MiR-26b-5p/KPNA2 Axis Is an Important Regulator of Burkitt Lymphoma Cell Growth. Cancers 2020, 12, 1464. [Google Scholar] [CrossRef]
- Si, X.; Zhang, X.; Hao, X.; Li, Y.; Chen, Z.; Ding, Y.; Shi, H.; Bai, J.; Gao, Y.; Cheng, T.; et al. Upregulation of MiR-99a Is Associated with Poor Prognosis of Acute Myeloid Leukemia and Promotes Myeloid Leukemia Cell Expansion. Oncotarget 2016, 7, 78095–78109. [Google Scholar] [CrossRef]
- Kim, S.-W.; Ramasamy, K.; Bouamar, H.; Lin, A.-P.; Jiang, D.; Aguiar, R.C.T. MicroRNAs MiR-125a and MiR-125b Constitutively Activate the NF-ΚB Pathway by Targeting the Tumor Necrosis Factor Alpha-Induced Protein 3 (TNFAIP3, A20). Proc. Natl. Acad. Sci. USA 2012, 109, 7865–7870. [Google Scholar] [CrossRef]
- Zhao, H.; Cheng, Y.; Dong, S.; Du, J.; Gao, F.; Sun, D.; Cui, J.; Ni, J.; Cai, J. Down Regulation of MiR-143 Promotes Radiation—Induced Thymic Lymphoma by Targeting B7H1. Toxicol. Lett. 2017, 280, 116–124. [Google Scholar] [CrossRef]
- Lone, W.; Bouska, A.; Sharma, S.; Amador, C.; Saumyaranjan, M.; Herek, T.A.; Heavican, T.B.; Yu, J.; Lim, S.T.; Ong, C.K.; et al. Genome-Wide MiRNA Expression Profiling of Molecular Subgroups of Peripheral T-Cell Lymphoma. Clin. Cancer Res. 2021, 27, 6039–6053. [Google Scholar] [CrossRef]
- Ramadan, E.S.; Kubesy, A.A.; Baraka, T.A.; Torad, F.A.; Salem, S.I.; Salem, N.Y. Expression of Blood Hepatocyte-Derived MicroRNA-122 in Canine Multicentric Lymphoma with Hepatic Involvement. Vet. Res. Commun. 2019, 43, 231–238. [Google Scholar] [CrossRef]
- Kulka, M.; Brennan, K.; Mc Gee, M. Investigation of Canine Extracellular Vesicles in Diffuse Large B-Cell Lymphomas. PLoS ONE 2022, 17, e0274261. [Google Scholar] [CrossRef]
- Garnica, T.K.; Lesbon, J.C.C.; Ávila, A.C.F.C.M.; Rochetti, A.L.; Matiz, O.R.S.; Ribeiro, R.C.S.; Zoppa, A.; Nishiya, A.T.; Costa, M.T.; de Nardi, A.B.; et al. Liquid Biopsy Based on Small Extracellular Vesicles Predicts Chemotherapy Response of Canine Multicentric Lymphomas. Sci. Rep. 2020, 10, 20371. [Google Scholar] [CrossRef] [PubMed]
- Asada, H.; Tomiyasu, H.; Uchikai, T.; Ishihara, G.; Goto-Koshino, Y.; Ohno, K.; Tsujimoto, H. Comprehensive Analysis of MiRNA and Protein Profiles within Exosomes Derived from Canine Lymphoid Tumour Cell Lines. PLoS ONE 2019, 14, e0208567. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).