1. Introduction
The unified genetic evaluation of genotyped and non-genotyped animals has been of great interest. In an initial attempt, Misztal et al. [
1] suggested a unified pedigree (
A) and genomic (
G) relationship matrix (
), in which genomic relationships between genotyped animals replaced their pedigree relationship coefficients in
A. Denoting non-genotyped and genotyped animals with 1 and 2:
This relationship matrix did not condition the distributions of breeding values for genotyped and non-genotyped animals on each other, leading to incoherencies in the joint distribution of genetic values for genotyped and non-genotyped animals. Legarra et al. [
2] presented an augmented (
A and
G) relationship matrix in which the genetic values of non-genotyped animals were conditioned to the genetic values of genotyped animals. The resulting matrix was:
which can be simplified to any of the following:
In matrix
H, the genomic information in
G influences the relationships between non-genotyped and genotyped animals and among non-genotyped animals. Later, it was discovered that
can be indirectly obtained without forming and inverting
H [
3,
4].
Matrix
G is not always full-rank (e.g., when the number of genotyped animals is greater than the number of loci or when there are duplicated genotypes, such as for identical twins). To force
G to be positive-definite and avoid large diagonal values of
due to the bad numerical condition of
G, the first step of conditioning
G often involves blending it with
, which is always positive-definite (except in the existence of identical twins or clones [
5]) and of good numerical conditions (i.e.,
, 0 <
k < 1). Blending introduces residual polygenic effects (genetic effects not captured by genetic markers) to the evaluation model without explicitly modelling it, where the scalar
k is the ratio of the polygenic to the total additive genetic variance [
6].
It is theoretically true that no artificially inflated variance is introduced via the
H matrix [
2]. However, inflated genetic variances have been observed due to incompatibilities between
G and
[
6,
7,
8,
9]. Incompatible
G and
lead to incorrectly weighted pedigree and genomic information [
7,
8]. Besides different distributions of
G and
elements, incomplete and incorrect pedigree information, and genotyping and imputation errors, incompatibilities between
G and
can be due to the non-random selection of genotyped animals [
10], and the different bases and scales of the two matrices [
7]. Matrices
and
G regress data to different means. Matrix
regresses solutions towards pedigree founders, animals in the pedigree with unknown parents or genetic groups if considered in the pedigree. On the other hand,
G regresses solutions toward a founder population comprising genotyped animals [
5,
10] since the real allele frequencies in the founder population are unknown. The average genetic merit of genotyped animals can be different from founders, especially in the presence of selection. Different approaches (referred to as tuning) have been used for correcting the base difference between
G and
[
7,
11] and rebasing and scaling
G to improve its consistency with
[
10]. Those approaches were tested by Nilforooshan [
9] on New Zealand Romney sheep. Christensen [
8] and Gao et al. [
6] tuned
G by regressing its averages to the averages of
(Equations (
7) and (
8), respectively).
The
and
scalars obtained by solving either of the equations above are used for transforming
G into
. Another solution proposed to tackle the problem of inflated genomic evaluations (i.e., an increased variance of genomic predictions) as a result of incorrectly scaled genomic and pedigree information was scaling
in the form of
[
3,
12,
13]. Applying
is equivalent to transforming
G into
[
3,
9], which equals
. It is also equivalent to replacing
with
in Equation (
2) [
12].
Reducing
and
values toward 0 brings
G closer to
by bringing
closer to
. However, it is not easily quantifiable how
G and
are proportionally combined. With
and
deviating from each other and 1, there is a risk of distorting the conditional properties of
H, because the changes made in
are not reflected in other blocks of
. Whereas 1 –
k and
k are the commonly used blending coefficients of
G and
,
and
are the commonly used blending coefficients of
and
. i.e.,
Considering the above equation, there is no legitimate reason for
being out of the boundary of 0 and 1, and
being out of the boundary of –1 and 1. Martini et al. [
12] studied
ranging from 0.1 to 2, and
ranging from –1 to 1 by steps of 0.1, leading to 420 analyses. Dealing with two parameters increases the number of analyses and validation tests in a two-dimensional space. It is assuming that the
k coefficient has already been chosen and does not need to be validated. The most coherent approach for finding
k is by restricted maximum likelihood (REML), as proposed by Christensen and Lund [
4], rather than using empirical values by screening and validation.
Weighting
and
as
has been used until recently [
12,
13,
14,
15,
16,
17]. Several improvements have been made to ssGBLUP [
18] and the use of
is declining. For example, one of the factors leading to the need for an
considerably less than 1 was that inbreeding coefficients were considered in
but not in
[
19]. The aim of this study was to communicate the problems that might occur using
, and investigate the possible solutions for weighting the
components if the modifications in
G are not satisfactory and the weighting of the
components is still needed for the deflation/inflation of genomic breeding values.
3. Materials
Data were simulated for a species in a 1:1 sex ratio, litter size of 2, and generation overlap of 1. The pedigree, phenotypes, and genotypes were simulated using the R package pedSimulate [
21]. Initially, ten generations were simulated, starting with a base generation (F0) of 100 animals (50 of each sex). No non-random pre-mating mortality or selection was applied to F0. Genotypes were simulated on 5000 markers, and allele frequencies were sampled from a uniform distribution ranging from 0.1 to 0.9. Marker (allele substitution) effects were simulated from a gamma distribution with shape and rate parameters equal to 2. The distribution was rebased to have a mean of 0 and scaled to create a variance of (true) marker breeding values in F0,
= 9. Residual polygenic and environment (residual) effects were simulated from normal distributions with variances
= 1 and
= 30, respectively.
Following F0, half of the males were mated to half of the females, which were all randomly selected and mated. Where the numbers of mating animals per sex were not equal, the sex with the higher number of animals underwent random selection to match the number of animals of the opposite sex. These ten generations were followed by ten more generations, in which 50% of male candidates (to become sires of the next generation) were selected for their marker breeding value and mated to the same number of randomly selected females. Genotypes in each subsequent generation were obtained by combining sampled gametes from the parents’ genotypes.
Phenotypes were calculated as , where is the population mean, and g, a, and e are the vectors of effects corresponding to , , and . Genotypes before F8 and phenotypes for the last generation (F19) and before F7 were set to missing. Randomly, 5% of the known dams and 5% of the known sires (after F0) were set to missing. As such, missing pedigree and phenotype information, genomic pre-selection, and base and scale deviations between A and G were accommodated in the simulation. Data simulation was repeated ten times to reduce the possibility of observing the results specific to a dataset.
No fixed effect was simulated, and the data were analysed using the following mixed model equations:
where
Z is the matrix relating phenotypes to animals,
1 and
are the vectors of ones and predicted breeding values, and
is the mean estimate. Matrix
G was used in
and built according to method 1 of VanRaden [
5], where
,
W is the centred and scaled genotype matrix, and
p is the marker allele frequency. Markers with minor allele frequency below 0.02 were discarded before calculating
G. Then,
G was blended as
.
5. Discussion
Matrices
G and
indicate different means and variances for genotyped animals. This can cause differently scaled genomic and pedigree information in
[
3]. Usually,
G is blended and tuned (rebased and scaled) with
. If genomic breeding values are still inflated, a complementary weighting of
might be needed. A common practice is to weight using
. It was shown that some
combinations are likely to distort the properties of
H that provide conditionality between the breeding values of genotyped and non-genotyped animals. Other ways of weighting the components of
were presented that are unlikely to distort the conditional properties of
H.
Weighting
with
> 1 is equivalent to reducing h
2 and increasing inflation due to increased dispersion. It is equivalent to adding
to 1/h
2 or weighting the genetic variance by 1/
. Due to selection, h
2 can be lower than expected. The h
2 reduction is expected to be greater due to genomic selection. Change of genetic variance by genomic selection is propagated from
G throughout
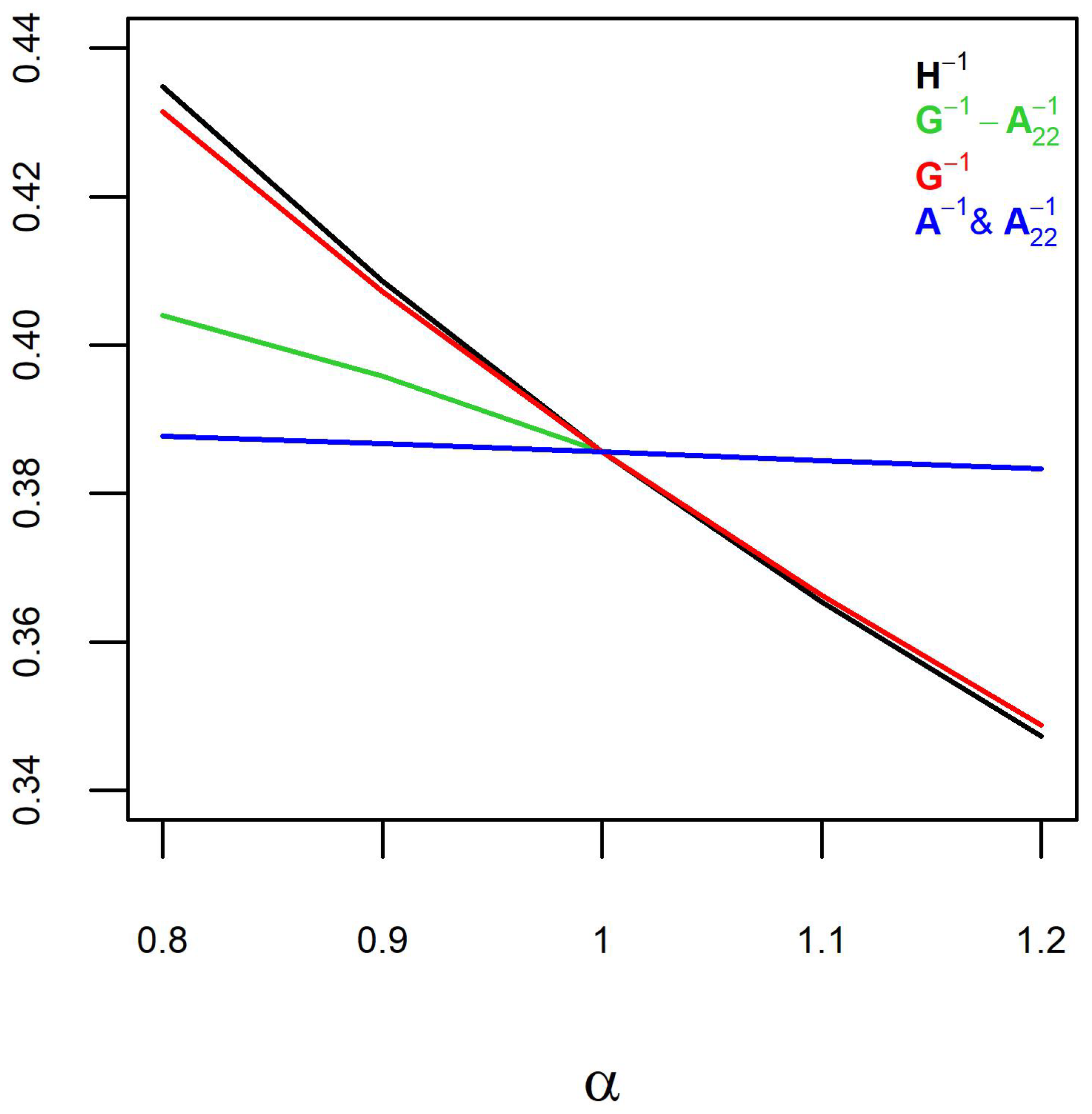
H. The predictive ability declined with increasing
(
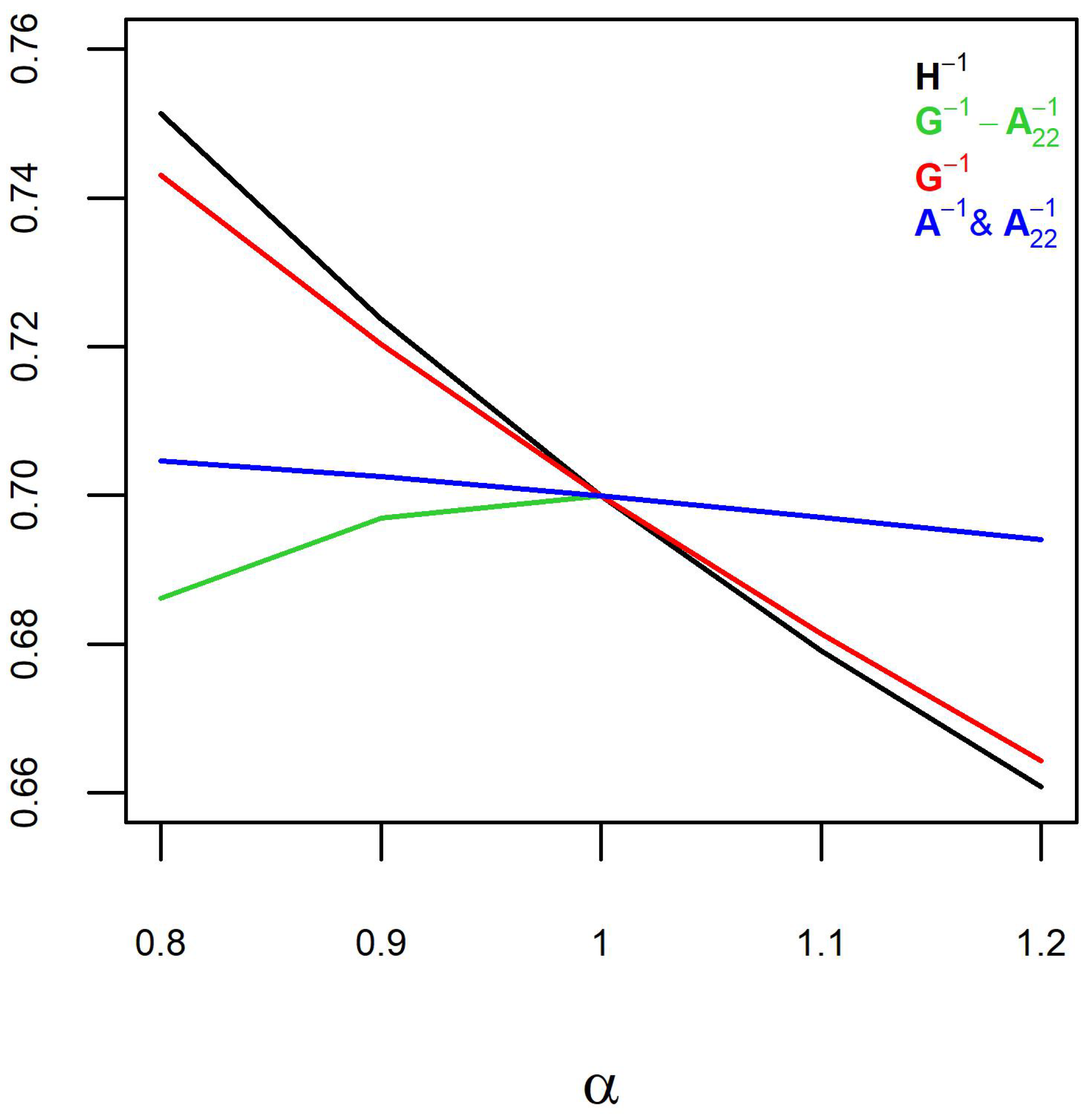
Figure 2), which might be concerning. However, predictive ability is a direct function of the slope of the regression line (
Figure 1). Therefore, the slope of the regression line (inflation) should be the main concern.
Weighting
(accompanied by weighting
) did not influence inflation and predictive ability. Predictive ability and the slope of the regression line decreased slightly (inflation increased slightly) over the increase in
. The reason for this is likely that
H is a genomic relationship matrix extended from
G for genotyped animals to non-genotyped animals via the
coefficients (Equations (
2)–(
5)). As such,
G is more influential in defining the variances in
H than
A. This was confirmed by similar trends for weighting
and
(
Figure 1 and
Figure 2). The slopes of the regression line (inflation) and predictive ability were slightly steeper for
than for
, and that was a result of the combined weighting of
,
and
. Weighting
by
< 1 increased the inflation but at a lower rate than weighting
or
with
> 1.
The inflation results are expected to be valid for other data as weighting or its components is equivalent to inversely weighting the genetic variance, regardless of the data. The exception is weighting . Whether weighting with a larger results in inflation or deflation depends on whether using instead of results in inflation or deflation. If using results in inflation, then weighting with a larger (more emphasis on than ) results in greater inflation. The predictive ability improved by weighting with decreasing from 1 to 0.8. Generally, predictive ability increases by the increase in the slope of the regression line. Notice that the predictive ability ignoring inflation can be misleading. Since the trends for prediction ability and the slope of the regression line were in opposite directions for weighting , it shows that the predictive ability benefited from blending and , mainly because the h2 was more compatible with a blended and than with .
This study does not completely rule out using
. However, weighting
components should meet specific conditions to avoid/minimise violating the conditional properties of
H. As such,
and
are better alternatives to
. By definition, none of these four options are better than the others. However, achieving good compatibility between the resulting
and h
2 without blending
and
at a high rate (low emphasis on genomic information) is important.
Concerning pedigree and genomic errors, regardless of the emphasis given to pedigree and genomic information, genotype errors propagate through non-genotyped animals, and pedigree errors incorrectly and insufficiently propagate genotype information through non-genotyped animals. Therefore, the correctness and the completeness of pedigree and genomic information are vital for accurate and unbiased ssGBLUP evaluations.
Future research may focus on changing genetic parameters over time or across populations in genomic predictions. It is possible to reduce inflation in genomic predictions for young animals by using smaller additive genetic variances. This can be done by replacing
with
. Considering no overall weight on
:
. Matrix
D is a diagonal matrix of positive values descending in function of the animal’s age. The researcher would need to decide the
range, where
d = diag(
D). With recent advances in ssGBLUP (mentioned by Misztal et al. [
18]), which improve the compatibility between
A and
G, conditioning
might become an interim solution from the past or be reduced to only weighting
.