Establishment of the Primary Avian Gonadal Somatic Cell Lines for Cytogenetic Studies
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Specimens
2.2. Primary Cell Culture Establishment
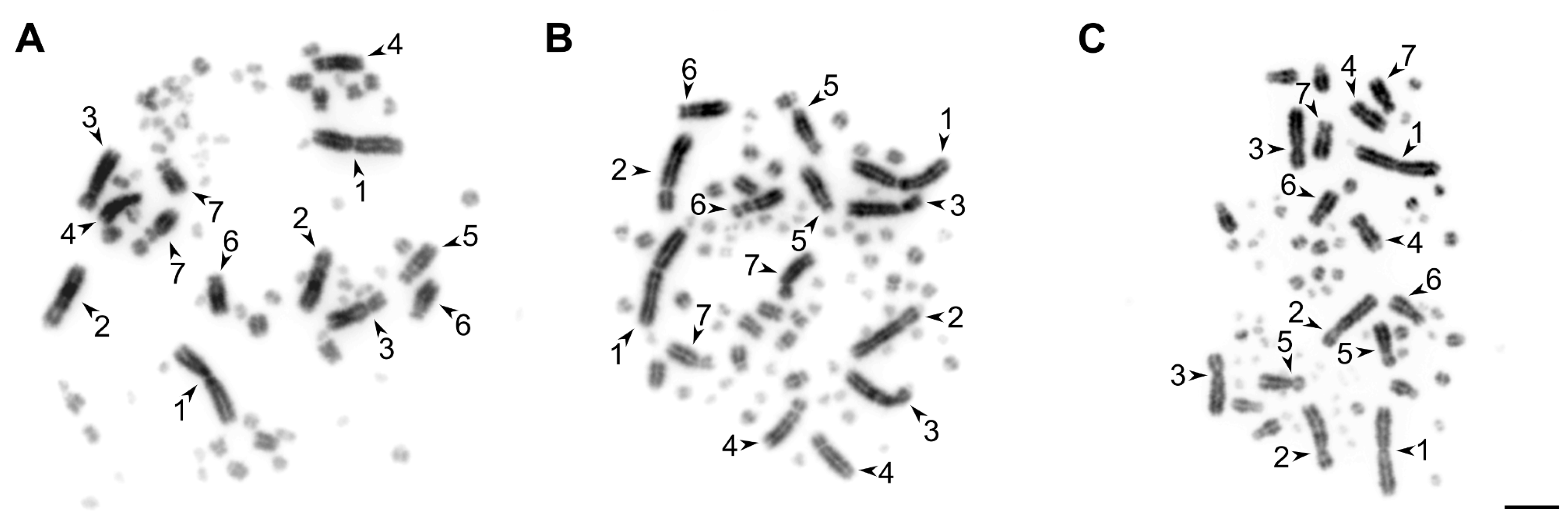
2.3. Mitotic Metaphase Chromosome Preparation
2.4. Microscopic Image Analysis
2.5. Measuring the Doubling Time of the Cell Population
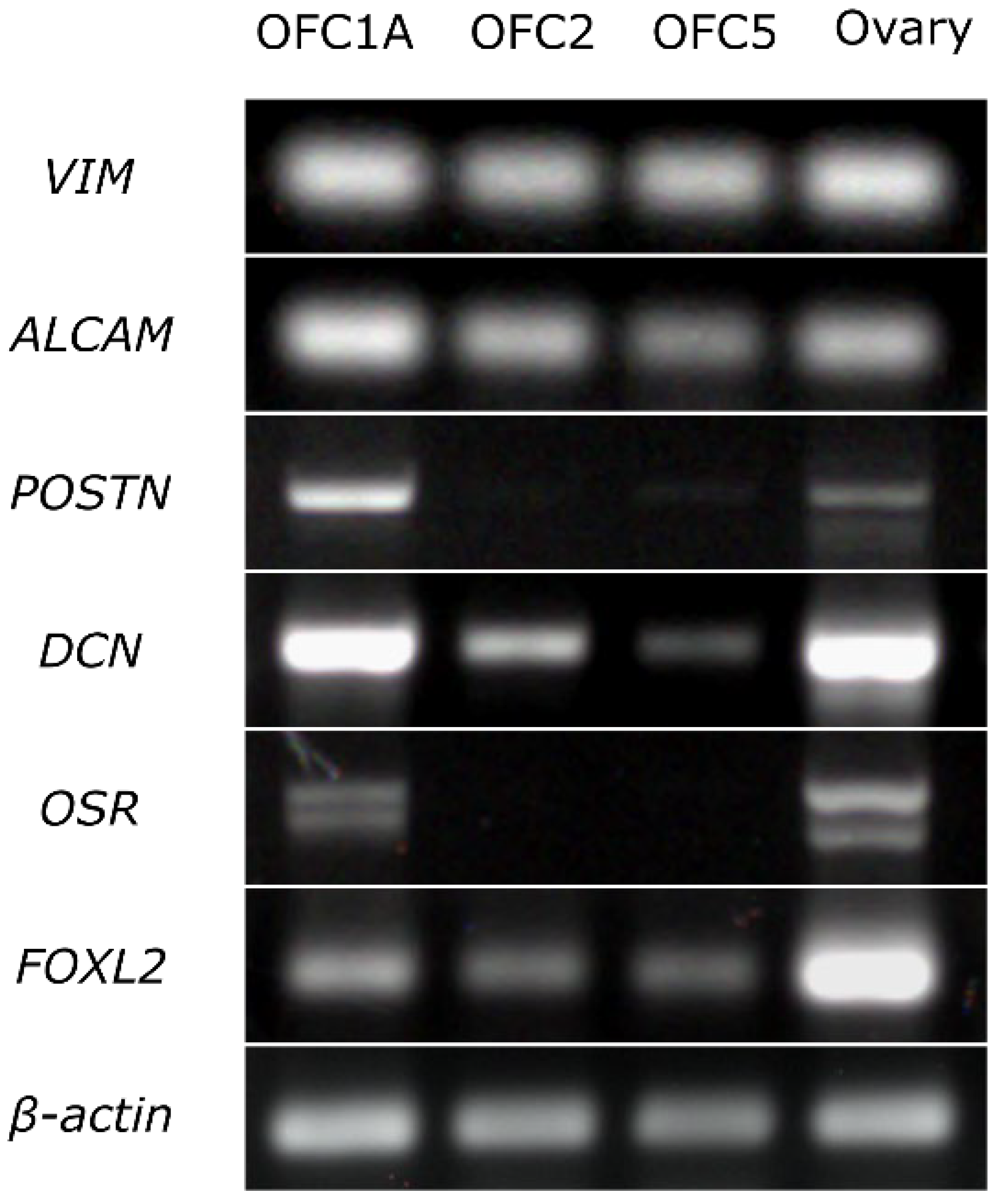
2.6. RT-PCR Analysis of Gene Expression
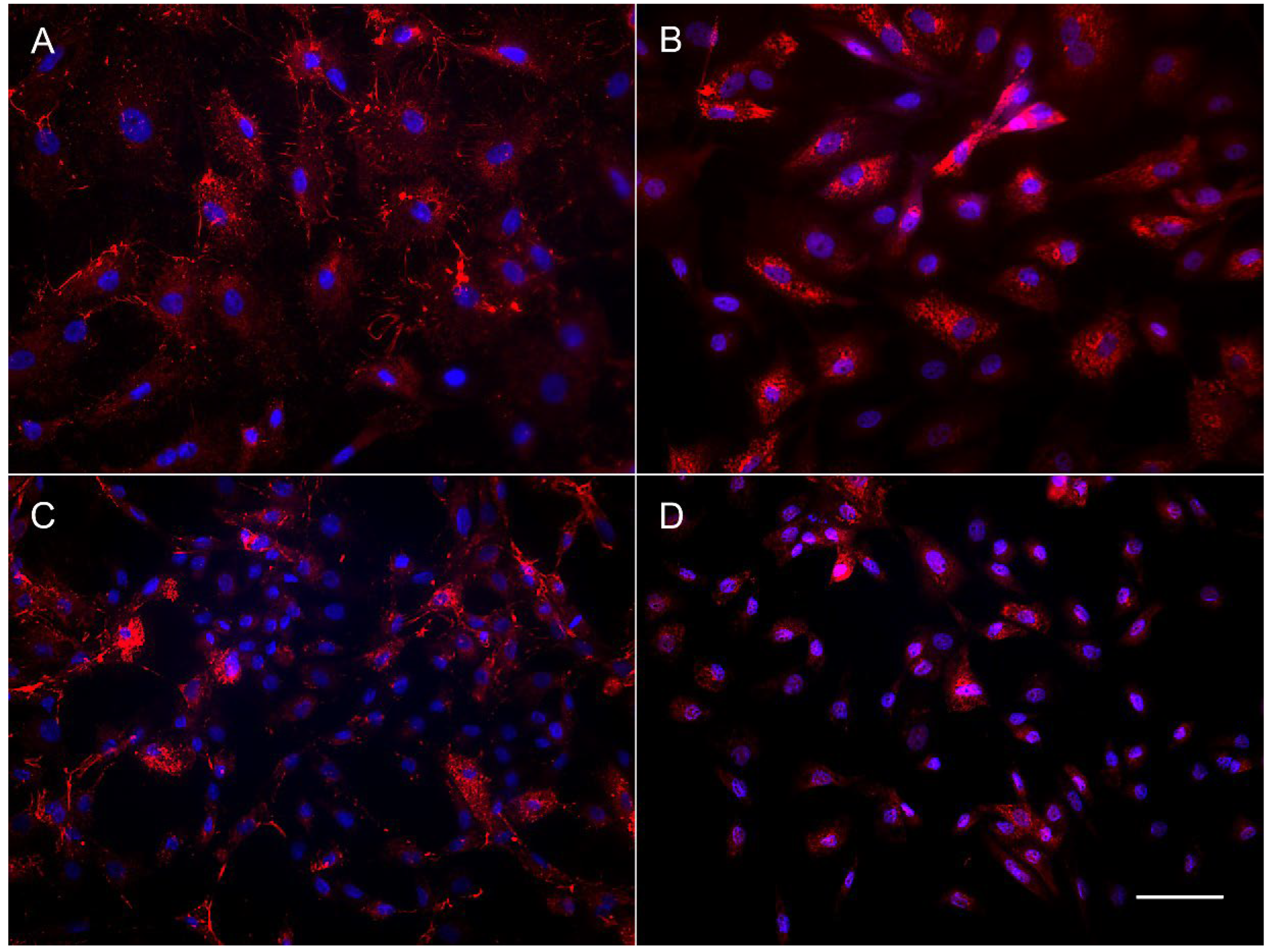
2.7. Immunocytochemical Analysis
2.8. Transduction with Lentivirus Vectors Expressing GFP
2.9. Electroporation with Episomal Vectors Expressing GFP
3. Results and Discussion
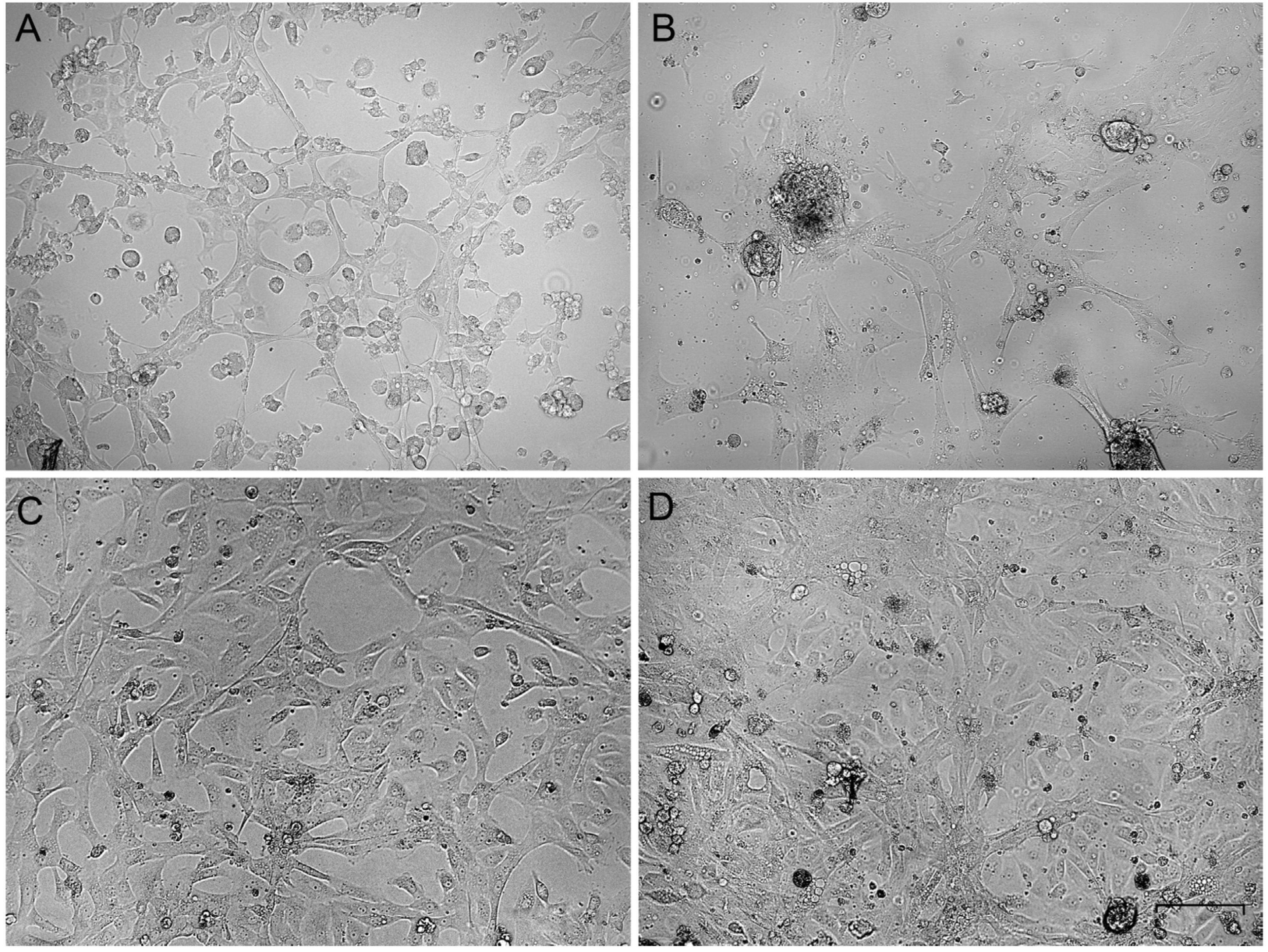
3.1. Establishment of the Primary Cell Cultures from the Passerine Gonads
3.2. Growth Rate and Cellular Composition of the Ovary-Derived Cell Lines
3.3. Transfection and Electroporation with GFP Reporter Gene
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, G.; Li, C.; Li, Q.; Li, B.; Larkin, D.M.; Lee, C.; Storz, J.F.; Antunes, A.; Greenwold, M.J.; Meredith, R.W.; et al. Comparative genomics reveals insights into avian genome evolution and adaptation. Science 2014, 346, 1311–1320. [Google Scholar] [CrossRef] [PubMed]
- Stryjewski, K.F.; Sorenson, M.D. Mosaic genome evolution in a recent and rapid avian radiation. Nat. Ecol. Evol. 2017, 1, 1912–1922. [Google Scholar] [CrossRef] [PubMed]
- Damas, J.; Kim, J.; Farré, M.; Griffin, D.K.; Larkin, D.M. Reconstruction of avian ancestral karyotypes reveals differences in the evolutionary history of macro- and microchromosomes. Genome Biol. 2018, 19, 155. [Google Scholar] [CrossRef]
- Bravo, G.A.; Schmitt, C.J.; Edwards, S.V. What have we learned from the first 500 avian genomes? Annu. Rev. Ecol. Evol. Syst. 2021, 52, 611–639. [Google Scholar] [CrossRef]
- Lohr, V.; Hädicke, O.; Genzel, Y.; Jordan, I.; Büntemeyer, H.; Klamt, S.; Reichl, U. The avian cell line AGE1.CR.pIX characterized by metabolic flux analysis. BMC Biotechnol. 2014, 14, 72. [Google Scholar] [CrossRef]
- Biegler, M.T.; Fedrigo, O.; Collier, P.; Mountcastle, J.; Haase, B.; Tilgner, H.U.; Jarvis, E.D. Induction of an immortalized songbird cell line allows for gene characterization and knockout by CRISPR-Cas9. Sci. Rep. 2022, 12, 4369. [Google Scholar] [CrossRef]
- Wang, W.; Said, A.; Wang, B.; Qu, G.; Xu, Q.; Liu, B.; Shen, Z. Establishment and evaluation of the goose embryo epithelial (GEE) cell line as a new model for propagation of avian viruses. PLoS ONE 2018, 13, e0193876. [Google Scholar] [CrossRef]
- Guan, W.; He, X.; Li, L.; Liang, H.; Zhao, Q.; Pu, Y.; Ma, Y.H. Establishment and biological characterization of fibroblast cell line from the Langshan chicken. Cell Prolif. 2010, 43, 157–163. [Google Scholar] [CrossRef]
- Niikura, M.; Narita, T.; Mikami, T. Establishment and characterization of a thymidine kinase deficient avian fibroblast cell line derived from a Japanese quail cell line, QT35. J. Vet. Med. Sci. 1991, 53, 439–446. [Google Scholar] [CrossRef][Green Version]
- Sasaki, M.; Ikeuchi, T.; Makino, S. A feather pulp culture technique for avian chromosomes, with notes on the chromosomes of the peafowl and the ostrich. Experientia 1968, 24, 1292–1293. [Google Scholar] [CrossRef]
- Barcellos, S.A.; Kretschmer, R.; de Souza, M.S.; Costa, A.L.; Degrandi, T.M.; dos Santos, M.S.; de Oliveira, E.H.C.; Cioffi, M.B.; Gunski, R.J.; Garnero, A.D.V. Karyotype evolution and distinct evolutionary history of the W chromosomes in swallows (Aves, Passeriformes). Cytogenet. Genome Res. 2019, 158, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Kulak, M.; Takki, O.; Galkina, S. Cell culture establishment from zebra finch embryonic fibroblast. Adv. Anim. Vet. Sci. 2020, 9, 344–347. [Google Scholar] [CrossRef]
- Kretschmer, R.; de Oliveira Furo, I.; Gunski, R.J.; del Valle Garnero, A.; Pereira, J.C.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; de Oliveira, E.H.C.; de Freitas, T.R.O. Comparative chromosome painting in Columbidae (Columbiformes) reinforces divergence in Passerea and Columbea. Chromosom. Res. 2018, 26, 211–223. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Duggavathi, R.; Zadworny, D. Regulatory mechanisms underlying the expression of prolactin receptor in chicken granulosa cells. PLoS ONE 2017, 12, e0170409. [Google Scholar] [CrossRef]
- D’Herde, K.; Leybaert, L. Intracellular tree calcium related to apoptotic cell death in quail granulosa cell sheets kept in serum-free culture. Cell Death Differ. 1997, 4, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Xiao, H.; Pi, J.; Zhang, H.; Pan, A.; Pu, Y.; Liang, Z.; Shen, J.; Du, J. EGFR promotes the proliferation of quail follicular granulosa cells through the MAPK/extracellular signal-regulated kinase (ERK) signaling pathway. Cell Cycle 2019, 18, 2742–2756. [Google Scholar] [CrossRef] [PubMed]
- Asem, E.K.; Feng, S.; Stingley-Salazar, S.R.; Turek, J.J.; Peter, A.T.; Robinson, J.P. Basal lamina of avian ovarian follicle: Influence on morphology of granulosa cells in-vitro. Comp. Biochem. Physiol. C Pharmacol. Toxicol. Endocrinol. 2000, 125, 189–201. [Google Scholar] [CrossRef]
- Deng, Y.; Gan, X.; Chen, D.; Huang, H.; Yuan, J.; Qiu, J.; Hu, S.; Hu, J.; Wang, J. Comparison of growth characteristics of in vitro cultured granulosa cells from geese follicles at different developmental stages. Biosci. Rep. 2018, 38, BSR20171361. [Google Scholar] [CrossRef]
- Gan, X.; Wang, Y.; Gao, S.; Chen, X.; Hu, S.; Wang, J.; Hu, J.; Li, L.; Han, C. Co-culture model reveals the characteristics of theca cells and the effect of granulosa cells on theca cells at different stages of follicular development. Reprod. Domest. Anim. 2021, 56, 58–73. [Google Scholar] [CrossRef]
- ARRIVE. ARRIVE Guidelines: Animal Research: Reporting of In Vivo Experiments. Available online: https://arriveguidelines.org (accessed on 1 June 2022).
- Matveeva, N.M.; Pristyazhnyuk, I.E.; Temirova, S.A.; Menzorov, A.G.; Vasilkova, A.; Shilov, A.G.; Smith, A.; Serov, O.L. Unequal segregation of parental chromosomes in embryonic stem cell hybrids. Mol. Reprod. Dev. 2005, 71, 305–314. [Google Scholar] [CrossRef]
- Menzorov, A.G.; Matveeva, N.M.; Markakis, M.N.; Fishman, V.S.; Christensen, K.; Khabarova, A.A.; Pristyazhnyuk, I.E.; Kizilova, E.A.; Cirera, S.; Anistoroaei, R.; et al. Comparison of American mink embryonic stem and induced pluripotent stem cell transcriptomes. BMC Genom. 2015, 16, S6. [Google Scholar] [CrossRef] [PubMed]
- Pristyazhnyuk, I.E.; Menzorov, A.G. Generation of American mink induced pluripotent stem cells: A protocol. Vavilovskii Z. Genet. Sel. 2017, 21, 701–709. [Google Scholar] [CrossRef]
- Remley, V.A.; Jin, J.; Sarkar, S.; Moses, L.; Prochazkova, M.; Cai, Y.; Shao, L.; Liu, H.; Fuksenko, T.; Jin, P.; et al. High efficiency closed-system gene transfer using automated spinoculation. J. Transl. Med. 2021, 19, 474. [Google Scholar] [CrossRef] [PubMed]
- Millington, M.; Arndt, A.; Boyd, M.; Applegate, T.; Shen, S. Towards a clinically relevant lentiviral transduction protocol for primary human CD34+ hematopoietic stem/progenitor cells. PLoS ONE 2009, 4, e6461. [Google Scholar] [CrossRef]
- Bulatova, N.S. A cytotaxonomic study of three related families of birds: Fringillidae, Emberizidae, Ploceidae. J. Zool. Syst. Evol. Res. 1973, 11, 233–239. [Google Scholar] [CrossRef]
- Cardoso, C.A.; Motta, L.C.B.; Oliveira, V.C.D.; Martins, D.D.S. Somatic feather follicle cell culture of the gallus domesticus species for creating a wild bird genetic resource bank. Anim. Reprod. 2020, 17, e20200044. [Google Scholar] [CrossRef]
- Biederman, B.M.; Lin, C.C. A leukocyte culture and chromosome preparation technique for avian species. In Vitro 1982, 18, 415–418. [Google Scholar] [CrossRef]
- Harper, J.M.; Wang, M.; Galecki, A.T.; Ro, J.; Williams, J.B.; Miller, R.A. Fibroblasts from long-lived bird species are resistant to multiple forms of stress. J. Exp. Biol. 2011, 214, 1902–1910. [Google Scholar] [CrossRef]
- Jung, K.M.; Kim, Y.M.; Keyte, A.L.; Biegler, M.T.; Rengaraj, D.; Lee, H.J.; Mello, C.V.; Velho, T.A.F.; Fedrigo, O.; Haase, B.; et al. Identification and characterization of primordial germ cells in a vocal learning Neoaves species, the zebra finch. FASEB J. 2019, 33, 13825–13836. [Google Scholar] [CrossRef]
- Shi, Q.-Q.; Sun, M.; Zhang, Z.-T.; Zhang, Y.-N.; Elsayed, A.K.; Zhang, L.; Huang, X.-M.; Li, B.-C. A screen of suitable inducers for germline differentiation of chicken embryonic stem cells. Anim. Reprod. Sci. 2014, 147, 74–85. [Google Scholar] [CrossRef]
- Crispo, M.; Vilariño, M.; dos Santos-Neto, P.C.; Núñez-Olivera, R.; Cuadro, F.; Barrera, N.; Mulet, A.P.; Nguyen, T.H.; Anegón, I.; Menchaca, A. Embryo development, fetal growth and postnatal phenotype of eGFP lambs generated by lentiviral transgenesis. Transgen. Res. 2015, 24, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Macdonald, J.; Glover, J.D.; Taylor, L.; Sang, H.M.; McGrew, M.J. Characterisation and germline transmission of cultured avian primordial germ cells. PLoS ONE 2010, 5, e15518. [Google Scholar] [CrossRef] [PubMed]
- Bednarczyk, M.; Kozłowska, I.; Łakota, P.; Szczerba, A.; Stadnicka, K.; Kuwana, T. Generation of transgenic chickens by the non-viral, cell-based method: Effectiveness of some elements of this strategy. J. Appl. Genet. 2018, 59, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Hruska-Plochan, M.; Miyanohara, A. Lentiviral-mediated transgenesis in songbirds. In Methods in Molecular Biology; Humana Press: New York, NY, USA, 2017; Volume 1650, pp. 149–165. [Google Scholar]
- Agate, R.J.; Scott, B.B.; Haripal, B.; Lois, C.; Nottebohm, F. Transgenic songbirds offer an opportunity to develop a genetic model for vocal learning. Proc. Natl. Acad. Sci. USA 2009, 106, 17963–17967. [Google Scholar] [CrossRef] [PubMed]
- Carsience, R.S.; Clark, M.E.; Gibbins, A.M.V.; Etches, R.J. Germline chimeric chickens from dispersed donor blastodermal cells and compromised recipient embryos. Development 1993, 117, 669–675. [Google Scholar] [CrossRef]
- Zhang, Z.; Sun, P.; Yu, F.; Yan, L.; Yuan, F.; Zhang, W.; Wang, T.; Wan, Z.; Shao, Q.; Li, Z. Transgenic quail production by microinjection of lentiviral vector into the early embryo blood vessels. PLoS ONE 2012, 7, e50817. [Google Scholar] [CrossRef]
- Chojnacka-Puchta, L.; Kasperczyk, K.; Płucienniczak, G.; Sawicka, D.; Bednarczyk, M. Primordial germ cells (PGCs) as a tool for creating transgenic chickens. Pol. J. Vet. Sci. 2012, 15, 181–188. [Google Scholar] [CrossRef]
| Source of Material | Advantages | Disadvantages | References |
|---|---|---|---|
| Bone marrow | Quick and simple protocol | Need to sacrifice birds; poor chromosome morphology | [26] |
| Feather pulp cell culture | No need to sacrifice birds; good chromosome morphology | Long procedure, fraught with failures; could not be applied to the small birds | [10,27] |
| Leukocyte cells culture | No need to sacrifice birds; good chromosome morphology | Long procedure, fraught with failures; could not be applied to the small birds | [28] |
| Skin fibroblast cells culture | No need to sacrifice birds; good chromosome morphology | Traumatic for small birds | [29] |
| Fetal fibroblasts cell culture | Relatively rapid procedure; good chromosome morphology | Difficult to obtain the material, which is available during the breeding season only | [11,12,13] |
| Gonads cell culture | Rapid procedure; good chromosome morphology; high and stable cell proliferation | Need to sacrifice birds | This paper |
| Cell Line | % of Lentivirus Containing Supernatant | Without Spinoculation | With Spinoculation | ||||
|---|---|---|---|---|---|---|---|
| % of GFP-Positive Cells | 105 TU/mL | MOI | % of GFP-Positive Cells | 105 TU/mL | MOI | ||
| OFC1A | 10 | 16 | 19.2 | 0.18 | 9.9 | 118.8 | 0.11 |
| 50 | 9.8 | 23.5 | 0.11 | 37.0 | 88.8 | 0.46 | |
| Zebra finch fibroblasts | 10 | 11.8 | 141.6 | 0.13 | 78.7 | 944.4 | 1.55 |
| 50 | 50.6 | 121.4 | 0.71 | 42.6 | 102.24 | 0.56 | |
| Protocol | Pulse Voltage | Pulse Width | Pulse No | % of GFP-Positive Cells | Number of Cells Per Field-of-View | |
|---|---|---|---|---|---|---|
| OFC1A | 7 | 1200 | 30 | 1 | 79.0 ± 1.9 | 34.7 ± 1.4 |
| 15 | 1300 | 20 | 2 | 70.7 ± 4.1 | 19.5 ± 0.9 | |
| 22 | 1400 | 10 | 3 | 53.5 ± 5.3 | 123.7 ± 5.9 | |
| Great tit fibroblasts | 7 | 1200 | 30 | 1 | 54.5 ± 0.8 | 24.0 ± 0.7 |
| 15 | 1300 | 20 | 2 | 66.3 ± 1.0 | 23.9 ± 0.4 | |
| 22 | 1600 | 10 | 3 | 59.1 ± 1.1 | 18.4 ± 0.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pristyazhnyuk, I.E.; Malinovskaya, L.P.; Borodin, P.M. Establishment of the Primary Avian Gonadal Somatic Cell Lines for Cytogenetic Studies. Animals 2022, 12, 1724. https://doi.org/10.3390/ani12131724
Pristyazhnyuk IE, Malinovskaya LP, Borodin PM. Establishment of the Primary Avian Gonadal Somatic Cell Lines for Cytogenetic Studies. Animals. 2022; 12(13):1724. https://doi.org/10.3390/ani12131724
Chicago/Turabian StylePristyazhnyuk, Inna E., Lyubov P. Malinovskaya, and Pavel M. Borodin. 2022. "Establishment of the Primary Avian Gonadal Somatic Cell Lines for Cytogenetic Studies" Animals 12, no. 13: 1724. https://doi.org/10.3390/ani12131724
APA StylePristyazhnyuk, I. E., Malinovskaya, L. P., & Borodin, P. M. (2022). Establishment of the Primary Avian Gonadal Somatic Cell Lines for Cytogenetic Studies. Animals, 12(13), 1724. https://doi.org/10.3390/ani12131724







