Are Copy Number Variations within the FecB Gene Significantly Associated with Morphometric Traits in Goats?
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. CNV Genotyping
2.3. Association Analysis
2.4. Cluster Analysis, LD Block, and Transcription Factor Binding Prediction
3. Results
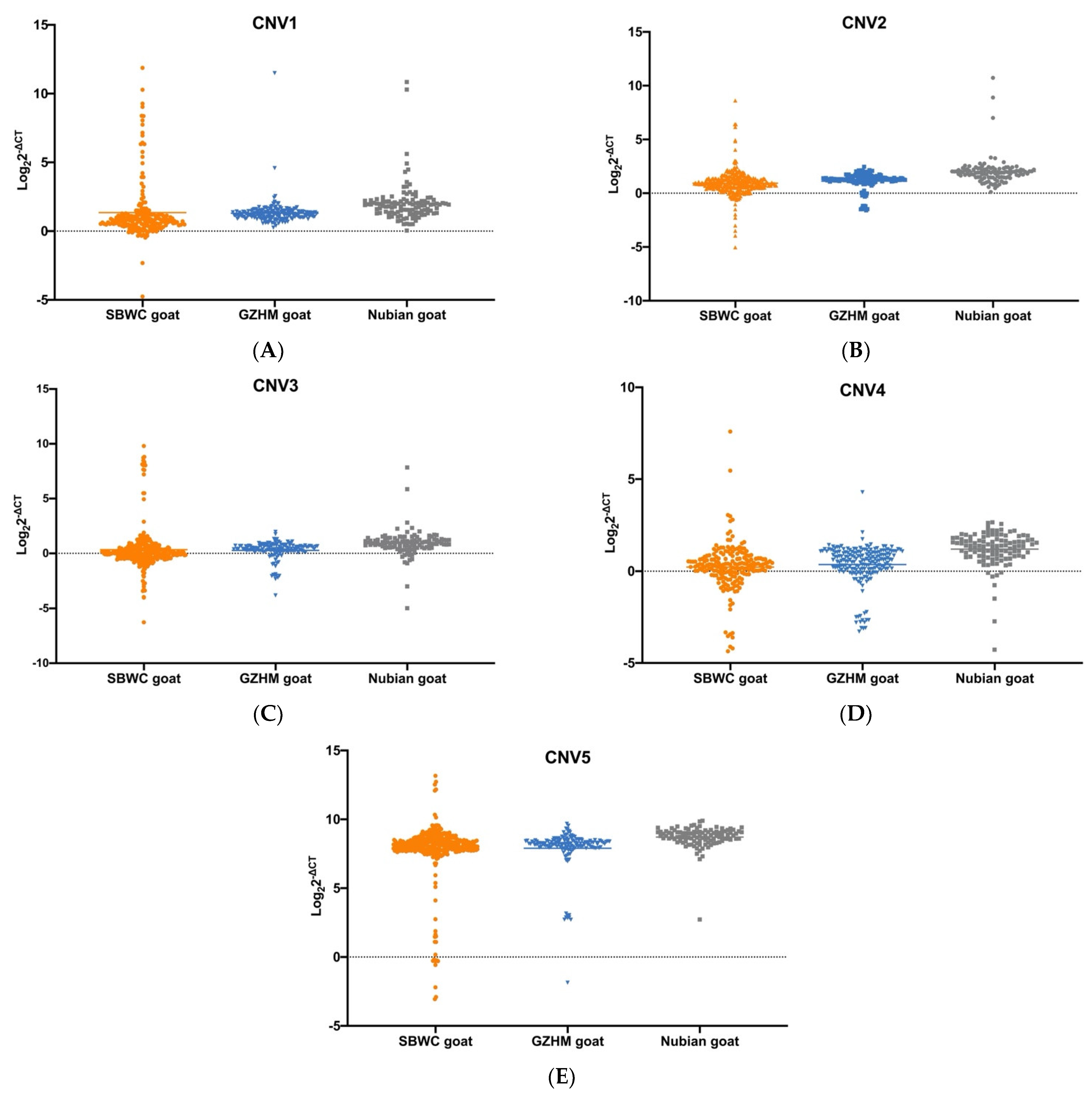
3.1. Detection and Frequency Distribution Analysis
3.2. Association Analysis
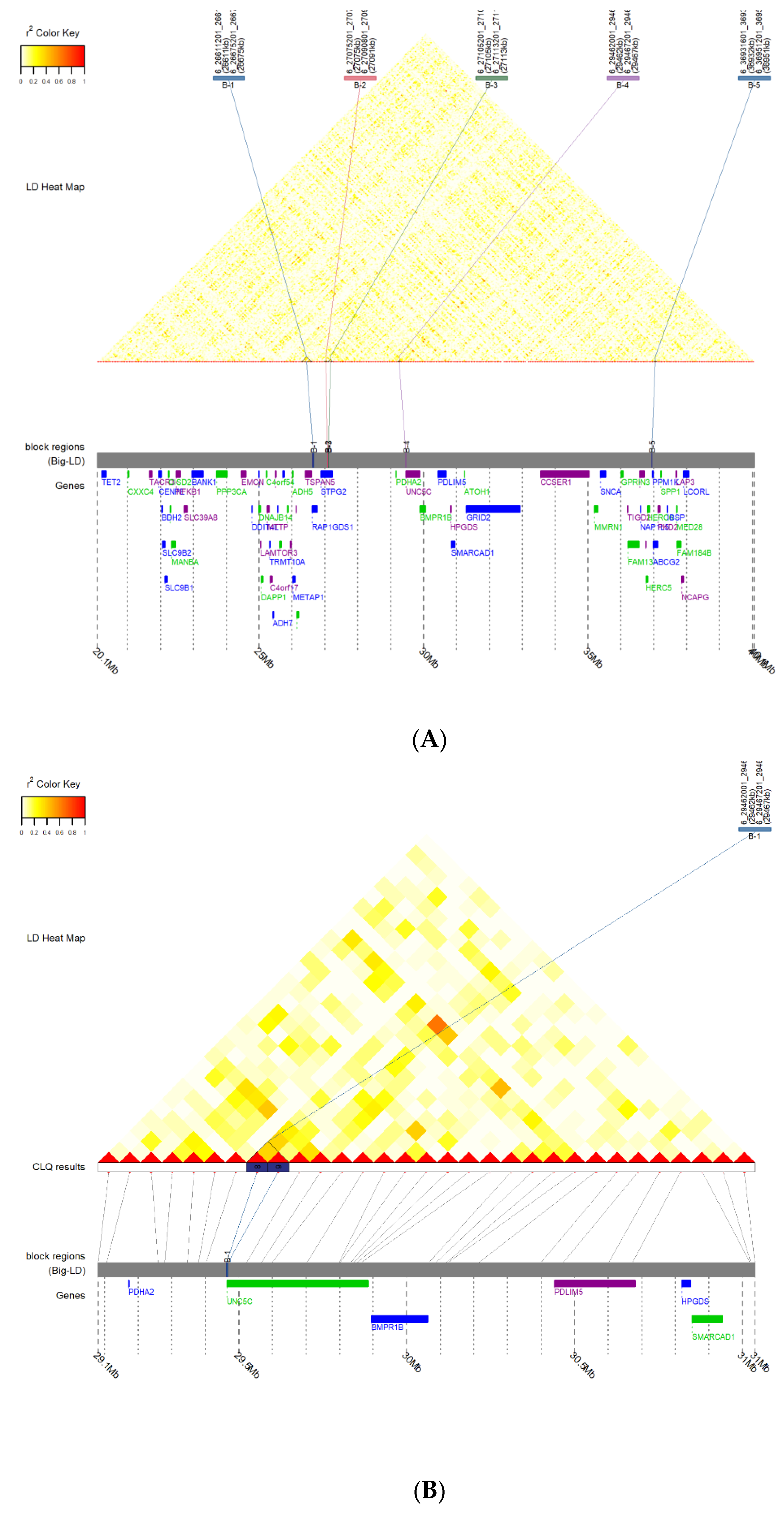
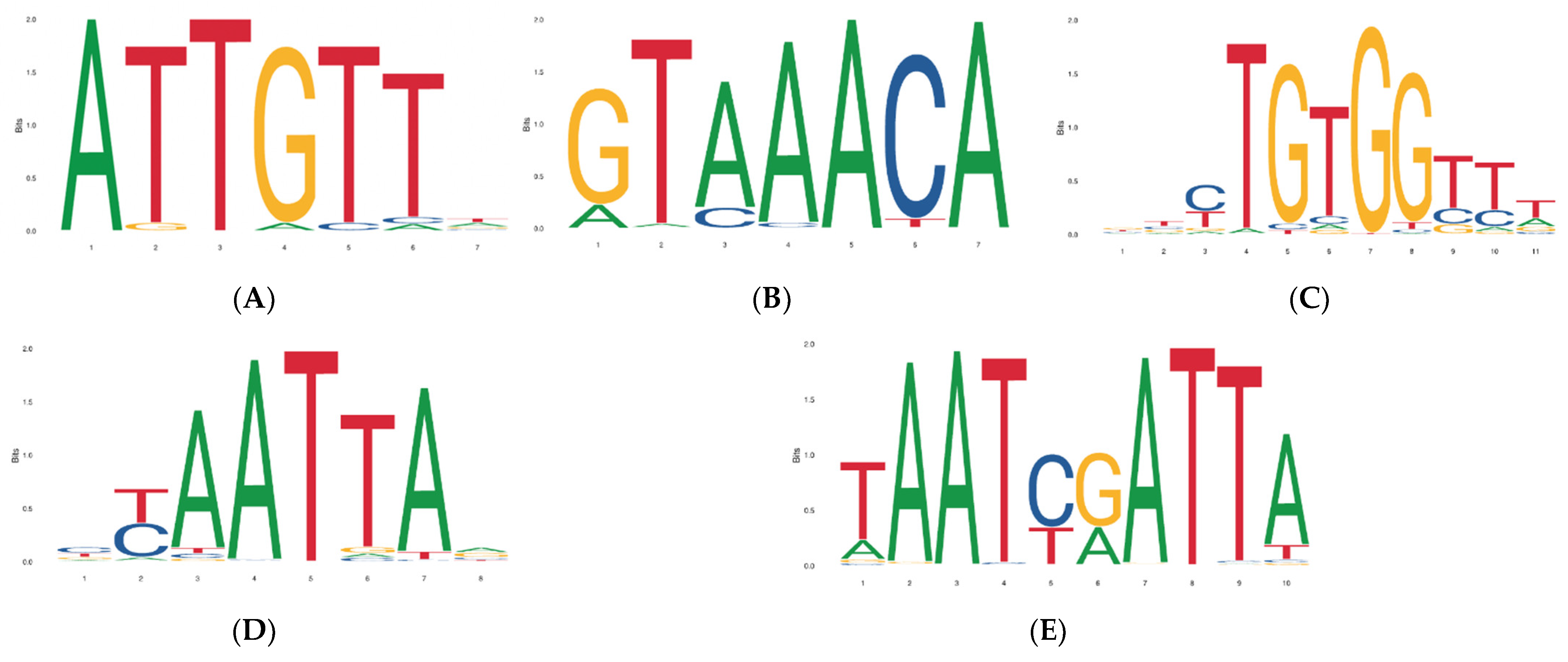
3.3. Prediction of LD Block and Transcription Factor Binding
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Montgomery, G.W.; Crawford, A.M.; Penty, J.M.; Dodds, K.G.; Ede, A.J.; Henry, H.M.; Pierson, C.A.; Lord, E.A.; Galloway, S.M.; Schmack, A.E.; et al. The ovine booroola fecundity gene (FecB) is linked to markers from a region of human chromosome 4q. Nat. Genet. 1993, 4, 410–414. [Google Scholar] [CrossRef] [PubMed]
- Campbell, B. The FecB (Booroola) gene acts at the ovary: In vivo evidence. Reproduction 2003, 126, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Hua, G.H.; Yang, L.G. A review of research progress of FecB gene in Chinese breeds of sheep. Anim. Reprod. Sci. 2009, 116, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Jin, J.; Chen, Q.; Yuan, Z.; Li, H.; Bian, J.; Gui, L. eukaryotic expression, co-ip and ms identify bmpr-1b protein-protein interaction network. Biol. Res. 2020, 53, 24. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Guo, X.; He, X.; Liu, Q.; Di, R.; Hu, W.; Cao, X.; Zhang, X.; Zhang, J.; Chu, M. Effects of FecB mutation on estrus, ovulation, and endocrine characteristics in small tail han sheep. Front. Vet. Sci. 2021, 8, 709737. [Google Scholar] [CrossRef] [PubMed]
- McNatty, K.P.; Hudson, N.L.; Lun, S.; Heath, D.A.; Shaw, L.; Condell, L.; Phillips, D.J.; Clarke, I.J. Gonadotrophin-releasing hormone and the control of ovulation rate by the fec(b) gene in booroola ewes. J. Reprod. Fertil. 1993, 98, 97–105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qi, M.Y.; Xu, L.Q.; Zhang, J.N.; Li, M.O.; Lu, M.H.; Yao, Y.C. Effect of the booroola fecundity (FecB) gene on the reproductive performance of ewes under assisted reproduction. Theriogenology 2020, 142, 246–250. [Google Scholar] [CrossRef]
- Mulsant, P.; Lecerf, F.; Fabre, S.; Schibler, L.; Monget, P.; Lanneluc, I.; Pisselet, C.; Riquet, J.; Monniaux, D.; Callebaut, I.; et al. mutation in bone morphogenetic protein receptor-ib is associated with increased ovulation rate in booroola m√©rino ewes. Proc. Natl. Acad. Sci. USA 2001, 98, 5104–5109. [Google Scholar] [CrossRef] [Green Version]
- Souza, C.J.H.; MacDougall, C.; Campbell, B.K.; McNeilly, A.S.; Baird, D.T. The booroola (FecB) phenotype is associated with a mutation in the bone morphogenetic receptor type 1 b (bmpr1b) gene. J. Endocrinol. 2001, 169, R1–R6. [Google Scholar] [CrossRef] [Green Version]
- Wilson, T.; Wu, X.Y.; Juengel, J.L.; Ross, I.K.; Lumsden, J.M.; Lord, E.A.; Dodds, K.G.; Walling, G.A.; McEwan, J.C.; O’Connell, A.R.; et al. highly prolific booroola sheep have a mutation in the intracellular kinase domain of bone morphogenetic protein ib receptor (alk-6) that is expressed in both oocytes and granulosa cells. Biol. Reprod. 2001, 64, 1225–1235. [Google Scholar] [CrossRef]
- Davis, G.H.; Montgomery, G.W.; Allison, A.J.; Kelly, R.W.; Bray, A.R. Segregation of a major gene influencing fecundity in progeny of booroola sheep. N. Z. J. Agric. Res. 1982, 25, 525–529. [Google Scholar] [CrossRef]
- McNatty, K.P.; Hudson, N.L.; Shaw, L.; Condell, L.A.; Ball, K.; Seah, S.L.; Clarke, I.J. GnRH-induced gonadotrophin secretion in ovariectomized booroola ewes with hypothalamic-pituitary disconnection. J. Reprod. Fertil. 1991, 91, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Wang, J. Study on BMPR-IB, GDF9, RXRG Genes and Reproductive Hormones for Lambing Traits in Three Sheep Breeds; Gansu Agricultural University: Lanzhou, China, 2015. [Google Scholar]
- Wang, W.; Liu, S.; Li, F.; Pan, X.; Li, C.; Zhang, X.; Ma, Y.; La, Y.; Xi, R.; Li, T. Polymorphisms of the ovine bmpr-ib, bmp-15 and fshr and their associations with litter size in two chinese indigenous sheep breeds. Int. J. Mol. Sci. 2015, 16, 11385–11397. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, J.; Hu, W.; Di, R.; Liu, Q.; Wang, X.; Zhang, X.; Zhang, J.; Chu, M. expression analysis of the prolific candidate genes, bmpr1b, bmp15, and gdf9 in small tail han ewes with three fecundity (FecB gene) genotypes. Animals 2018, 8, 166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bi, Y.; Feng, W.; Kang, Y.; Wang, K.; Yang, Y.; Qu, L.; Chen, H.; Lan, X.; Pan, C. Detection of mrna expression and copy number variations within the goat FecB gene associated with litter size. Front. Vet. Sci. 2021, 8, 758705. [Google Scholar] [CrossRef]
- Pourali, S.; Mirhoseini, S.Z.; Tufarelli, V.; Hossein-Zadeh, N.G.; Badbarin, S.; Colonna, M.A.; Seidavi, A.; Selvaggi, M. Investigating the polymorphism of bone morphogenetic protein receptor-1b (bmpr1b) gene in markhoz goat breed. Animals 2020, 10, 1582. [Google Scholar] [CrossRef]
- Yue, C.; Bai, W.L.; Zheng, Y.Y.; Hui, T.Y.; Sun, J.M.; Guo, D.; Guo, S.L.; Wang, Z.Y. Correlation Analysis of candidate gene snp for high-yield in liaoning cashmere goats with litter size and cashmere performance. Anim. Biotechnol. 2021, 32, 43–50. [Google Scholar] [CrossRef]
- Ahlawat, S.; Sharmaa, R.; Roy, M.; Mandakmale, S.; Prakash, V.; Tantia, M.S. Genotyping of novel snps in bmpr1b, bmp15, and gdf9 genes for association with prolificacy in seven Indian goat breeds. Anim. Biotechnol. 2016, 27, 199–207. [Google Scholar] [CrossRef]
- Kang, Y.; Bi, Y.; Luo, B.; Wang, K.; Bai, Y.; He, L.; Wang, R.; Pan, C.; Yan, H.; Liu, J.; et al. Correlation and regression analysis between litter size and morphometric traits of Shaanbei white cashmere goat. Acta Ecol. Anim. Domastici 2022, 43, 16–21. (In Chinese) [Google Scholar]
- Wang, K.; Liu, X.; Qi, T.; Hui, Y.; Yan, H.; Qu, L.; Lan, X.; Pan, C. Whole-genome sequencing to identify candidate genes for litter size and to uncover the variant function in goats (Capra hircus). Genomics 2021, 113, 142–150. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, S.; Tang, Q.; Jiang, E.; Wang, K.; Lan, X.; Pan, C. Goat sperm associated antigen 17 protein gene (spag17): Small and large fragment genetic variation detection, association analysis, and mrna expression in gonads. Genomics 2020, 112, 5115–5121. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Yan, H.; Li, J.; Xu, H.; Wang, K.; Zhu, H.; Chen, H.; Qu, L.; Lan, X. A novel 14-bp duplicated deletion within goat ghr gene is significantly associated with morphometric traits and litter size. Anim. Genet. 2017, 48, 499–500. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, S.; Brossard, M.; Roshandel, D.; Paterson, A.; Bull, S.; Yoo, Y. Gpart: Human genome partitioning and visualization of high-density SNP data by identifying haplotype blocks. Bioinformatics 2019, 35, 4419–4421. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zheng, Z.; Cai, Y.; Chen, T.; Li, C.; Fu, W.; Jiang, Y. CNVcaller: Highly efficient and widely applicable software for detecting copy number variations in large populations. Gigascience 2017, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yang, Q.; Wang, K.; Zhang, S.; Pan, C.; Chen, H.; Qu, L.; Yan, H.; Lan, X. A novel 12-bp indel polymorphism within the gdf9 gene is significantly associated with litter size and morphometric traits in goats. Anim. Genet. 2017, 48, 735–736. [Google Scholar] [CrossRef]
- Bai, Y.; Zhang, T.; Liu, N.; Wang, C.; Guo, Z.; Pan, C.; Zhu, H.; Lan, X. Investigation of copy number variations (CNVs) of the goat PPP3CA gene and their effect on litter size and semen quality. Animals 2022, 12, 445. [Google Scholar] [CrossRef]
- Liedvogel, M.; Ökesson, S.; Bensch, S. The genetics of migration on the move. Trends Ecol. Evol. 2011, 26, 561–569. [Google Scholar] [CrossRef]
- Harringmeyer, O.S.; Woolfolk, M.L.; Hoekstra, H.E. Fishing for the genetic basis of migratory behavior. Cell 2021, 184, 303–305. [Google Scholar] [CrossRef]
- Zan, Y.; Sheng, Z.; Lillie, M.; Rönnegård, L.; Honaker, C.F.; Siegel, P.B.; Carlborg, Ö. Artificial selection response due to polygenic adaptation from a multilocus, multiallelic genetic architecture. Mol. Biol. Evol. 2017, 34, 2678–2689. [Google Scholar] [CrossRef]
- Gosden, T.P.; Reddiex, A.J.; Chenoweth, S.F. Artificial selection reveals sex differences in the genetic basis of sexual attractiveness. Proc. Natl. Acad. Sci. USA 2018, 115, 5498–5503. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dekkers, J.C.M.; Hospital, F. The Use of molecular genetics in the improvement of agricultural populations. Nat. Rev. Genet. 2002, 3, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Zheng, P.; Hu, Y.; Wei, F. Genome-scale analysis of demographic history and adaptive selection. Protein Cell 2014, 5, 99–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nei, M.; Niimura, Y.; Nozawa, M. The Evolution of animal chemosensory receptor gene repertoires: Roles of chance and necessity. Nat. Rev. Genet. 2008, 9, 951–963. [Google Scholar] [CrossRef] [PubMed]
- Childs, G.V. Growth hormone cells as co-gonadotropes: Partners in the regulation of the reproductive system. Trends Endocrinol. Metab. 2000, 11, 168–175. [Google Scholar] [CrossRef]
- Liang, C.; Han, M.; Zhou, Z.; Liu, Y.; He, X.; Jiang, Y.; Ouyang, Y.; Hong, Q.; Chu, M. Hypothalamic transcriptome analysis reveals the crucial micrornas and mrnas affecting litter size in goats. Front. Vet. Sci. 2021, 8, 747100. [Google Scholar] [CrossRef]
- Long, F.; Shi, H.; Li, P.; Guo, S.; Ma, Y.; Wei, S.; Li, Y.; Gao, F.; Wang, M.; Duan, R.; et al. A SMOC2 variant inhibits BMP signaling by competitively binding to BMPR1B and cause growth plate defects. Bone 2021, 142, 115686. [Google Scholar] [CrossRef]
- Song, H.; Park, K.H. Regulation and function of sox9 during cartilage development and regeneration. Semin. Cancer Biol. 2020, 67, 12–23. [Google Scholar] [CrossRef]
- Fahrner, J.A.; Lin, W.Y.; Riddle, R.C.; Boukas, L.; DeLeon, V.B.; Chopra, S.; Lad, S.E.; Luperchio, T.R.; Hansen, K.D.; Bjornsson, H.T. Precocious chondrocyte differentiation disrupts skeletal growth in kabuki syndrome mice. JCI Insight 2019, 4, e129380. [Google Scholar] [CrossRef] [Green Version]
- Tang, C.Y.; Wu, M.; Zhao, D.; Edwards, D.; McVicar, A.; Luo, Y.; Zhu, G.; Wang, Y.; de Zhou, H.; Chen, W.; et al. Runx1 is a central regulator of osteogenesis for bone homeostasis by orchestrating bmp and wnt signaling pathways. PLoS Genet. 2021, 17, e1009233. [Google Scholar] [CrossRef]
- Cobb, J.; Dierich, A.; Huss-Garcia, Y.; Duboule, D.A. Mouse model for human short-stature syndromes identifies shox2 as an upstream regulator of runx2 during long-bone development. Proc. Natl. Acad. Sci. USA 2006, 103, 4511–4555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buckingham, M.; Relaix, F. The role of pax genes in the development of tissues and organs: Pax3 and pax7 regulate muscle progenitor cell functions. Annu. Rev. Cell Dev. Biol. 2007, 23, 645–673. [Google Scholar] [CrossRef] [PubMed]
- Buckingham, M.; Rigby, P.W.J. Gene regulatory networks and transcriptional mechanisms that control myogenesis. Dev. Cell 2014, 28, 225–238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Epstein, J.A.; Shapiro, D.N.; Cheng, J.; Lam, P.Y.P.; Maas, R.L. Pax3 modulates expression of the c-met receptor during limb muscle development. Proc. Natl. Acad. Sci. USA 1996, 93, 4213–4218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.X.; Rudnicki, M.A. Satellite cells, the engines of muscle repair. Nat. Rev. Mol. Cell Biol. 2012, 13, 127–133. [Google Scholar] [CrossRef]
- Sanchez, A.M.J.; Candau, R.B.; Bernardi, H. FoxO transcription factors: Their roles in the maintenance of skeletal muscle homeostasis. Cell. Mol. Life Sci. 2014, 71, 1657–1671. [Google Scholar] [CrossRef]
- Saxena, A.; Sharma, V.; Muthuirulan, P.; Neufeld, S.J.; Tran, M.P.; Gutierrez, H.L.; Chen, K.D.; Erberich, J.M.; Birmingham, A.; Capellini, T.D.; et al. Interspecies transcriptomics identify genes that underlie disproportionate foot growth in jerboas. Curr. Biol. 2021, 32, 289–303. [Google Scholar] [CrossRef]
- Jiang, E.; Kang, Z.; Wang, X.; Liu, Y.; Liu, X.; Wang, Z.; Li, X.; Lan, X. Detection of insertioans/deletions (indels) within the goat runx2 gene and their association with litter size and morphometric traits. Anim. Biotechnol. 2021, 32, 169–177. [Google Scholar] [CrossRef]
- Zhang, Y.; Cui, W.; Yang, H.; Wang, M.; Yan, H.; Zhu, H.; Liu, J.; Qu, L.; Lan, X.; Pan, C. A novel missense mutation (l280v) within pou1f1 gene strongly affects litter size and morphometric traits in goat. Theriogenology 2019, 135, 198–203. [Google Scholar] [CrossRef]


| Locus | CNV1 | CNV2 | CNV3 | CNV4 | CNV5 |
|---|---|---|---|---|---|
| Start | 30,065,201 | 30,086,001 | 30,116,801 | 30,121,201 | 30,180,401 |
| End | 30,067,200 | 30,088,400 | 30,118,800 | 30,123,200 | 30,182,000 |
| Length (bp) | 2000 | 2400 | 2000 | 2000 | 1600 |
| Breed | Locus | Size | Frequency | ||
|---|---|---|---|---|---|
| Loss | Normal | Gain | |||
| SBWC goat | CNV1 | n = 215 | 0.1163 (n = 25) | 0.1256 (n = 27) | 0.7581 (n = 163) |
| CNV2 | n = 318 | 0.1384 (n = 44) | 0.1101 (n = 35) | 0.7515 (n = 239) | |
| CNV3 | n = 318 | 0.1667 (n = 53) | 0.3019 (n = 96) | 0.5314 (n = 169) | |
| CNV4 | n = 213 | 0.2113 (n = 45) | 0.3286 (n = 70) | 0.4601 (n = 98) | |
| CNV5 | n = 302 | 0.0232 (n = 7) | 0.0232 (n = 7) | 0.9536 (n = 288) | |
| GZHM goat | CNV1 | n = 203 | 0 (n = 0) | 0.0049 (n = 1) | 0.9951 (n = 202) |
| CNV2 | n = 158 | 0.0696 (n = 11) | 0.0570 (n = 9) | 0.8734 (n = 138) | |
| CNV3 | n = 150 | 0.1133 (n = 17) | 0.2800 (n = 42) | 0.6067 (n = 91) | |
| CNV4 | n = 199 | 0.1206 (n = 24) | 0.2663 (n = 53) | 0.6131 (n = 122) | |
| CNV5 | n = 158 | 0.0127 (n = 2) | 0 (n = 0) | 0.9873 (n = 156) | |
| Nubian goat | CNV1 | n = 120 | 0 (n = 0) | 0.0083 (n = 1) | 0.9917 (n = 119) |
| CNV2 | n = 120 | 0 (n = 0) | 0.0083 (n = 1) | 0.9917 (n = 119) | |
| CNV3 | n = 116 | 0.1121 (n = 13) | 0.0603 (n = 7) | 0.8276 (n = 96) | |
| CNV4 | n = 120 | 0.0417 (n = 5) | 0.0333 (n = 4) | 0.9250 (n = 111) | |
| CNV5 | n = 120 | 0.0083 (n = 1) | 0 (n = 0) | 0.9917 (n = 119) | |
| Locus | Trait | Genotype (LSM ± SE) | p-Value | ||
|---|---|---|---|---|---|
| Loss | Normal | Gain | |||
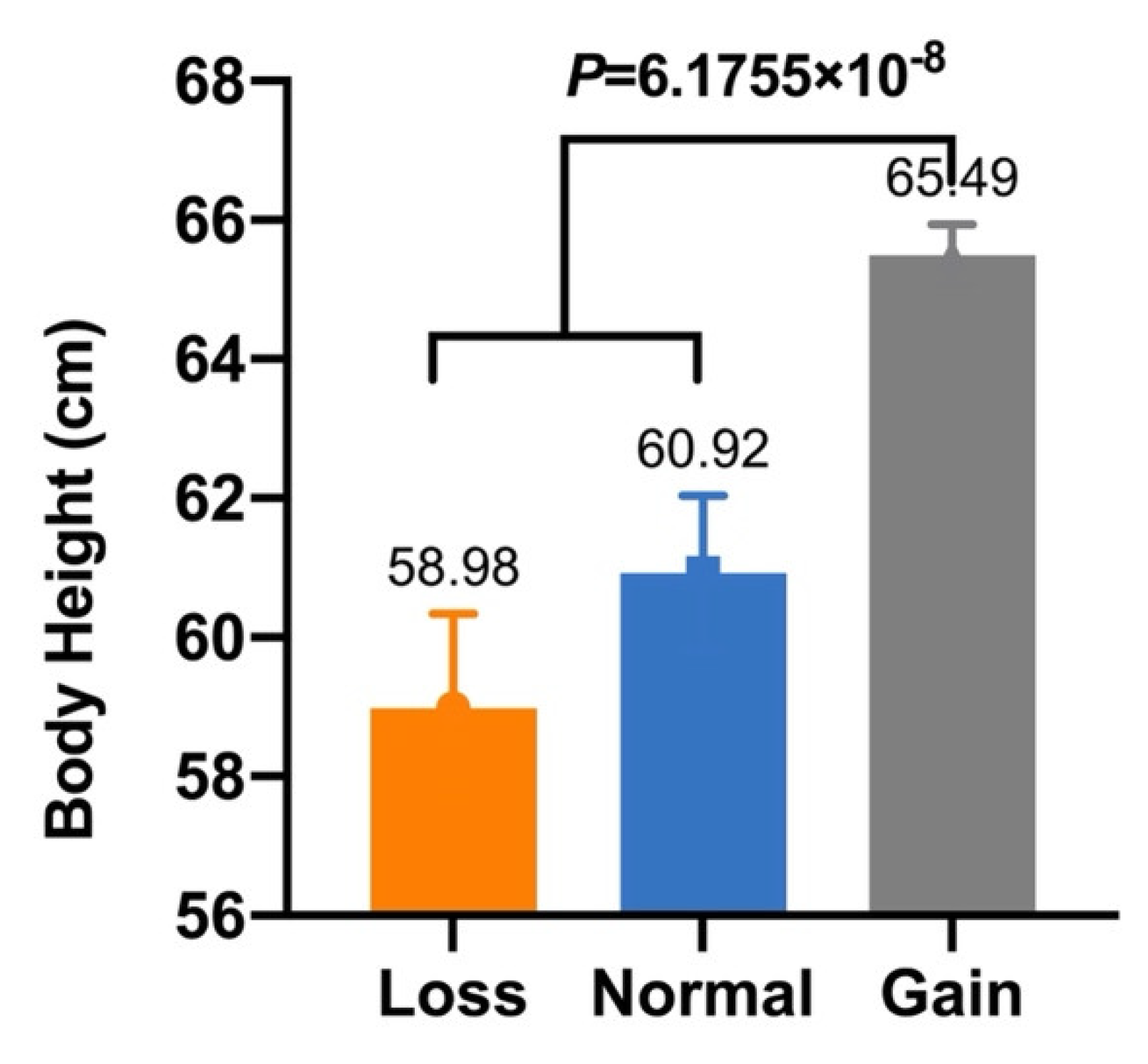
| CNV1 | Body Height (cm) | 58.98 ± 1.36 B (n = 25) | 60.92 ± 1.12 B (n = 27) | 65.49 ± 0.45 A (n = 163) | 6.1755 × 10−8 |
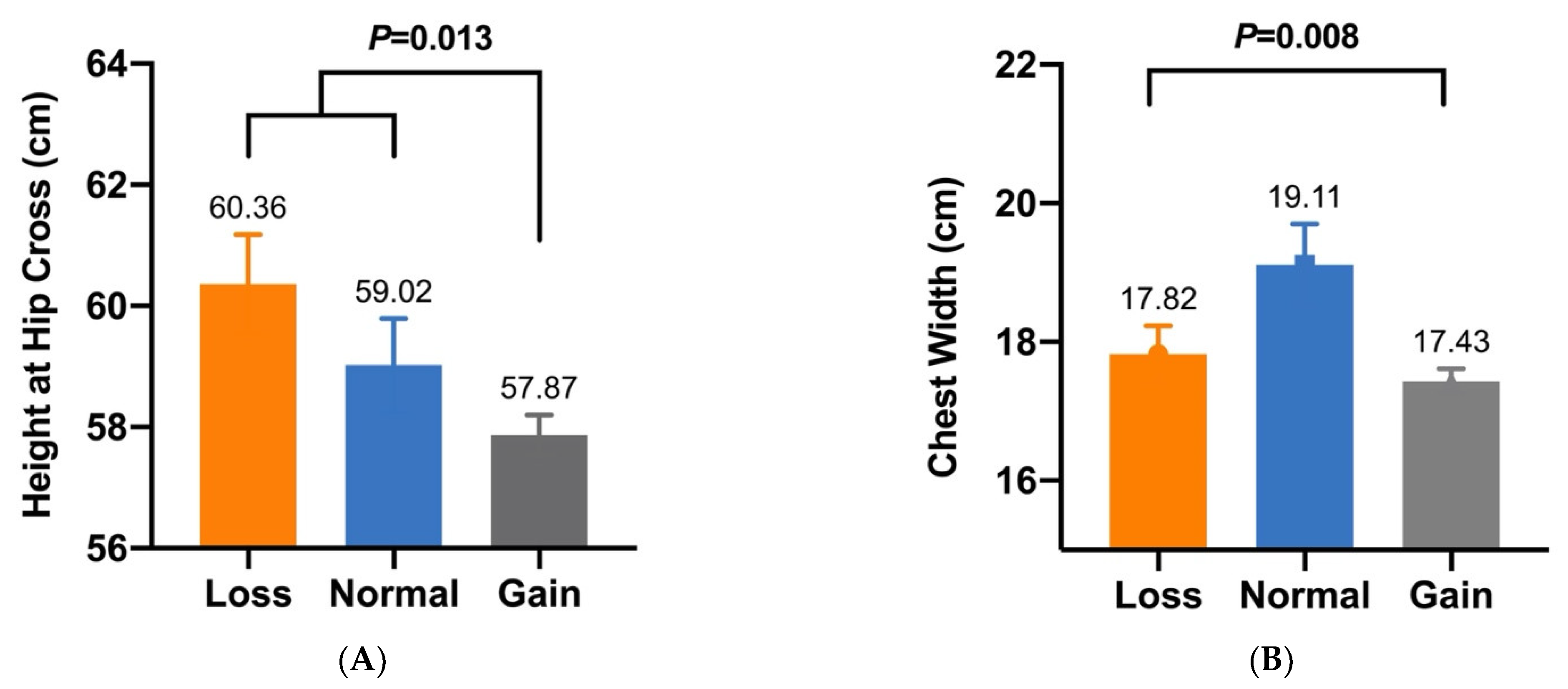
| CNV2 | Height at Hip Cross (cm) | 60.36 ± 0.78 a (n = 44) | 59.02 ± 0.77 a (n = 35) | 57.87 ± 0.33 b (n = 239) | 0.013 |
| CNV2 | Chest Width (cm) | 17.82 ± 0.41 B (n = 44) | 19.11 ± 0.59 A (n = 35) | 17.43 ± 0.18 B (n = 239) | 0.008 |
| CNV3 | Cannon Bone Circumference (cm) | 7.81 ± 0.11 A (n = 53) | 8.01 ± 0.08 A (n = 96) | 7.37 ± 0.07 B (n = 169) | 2.5303 × 10−7 |
| CNV3 | Chest Depth (cm) | 26.82 ± 0.36 a (n = 53) | 27.33 ± 0.26 a (n = 96) | 26.32 ± 0.23 b (n = 169) | 0.023 |
| CNV4 | Body Length (cm) | 62.70 ± 1.14 b (n = 45) | 64.99 ± 0.67 a (n = 70) | 65.88 ± 0.46 a (n = 98) | 0.037 |
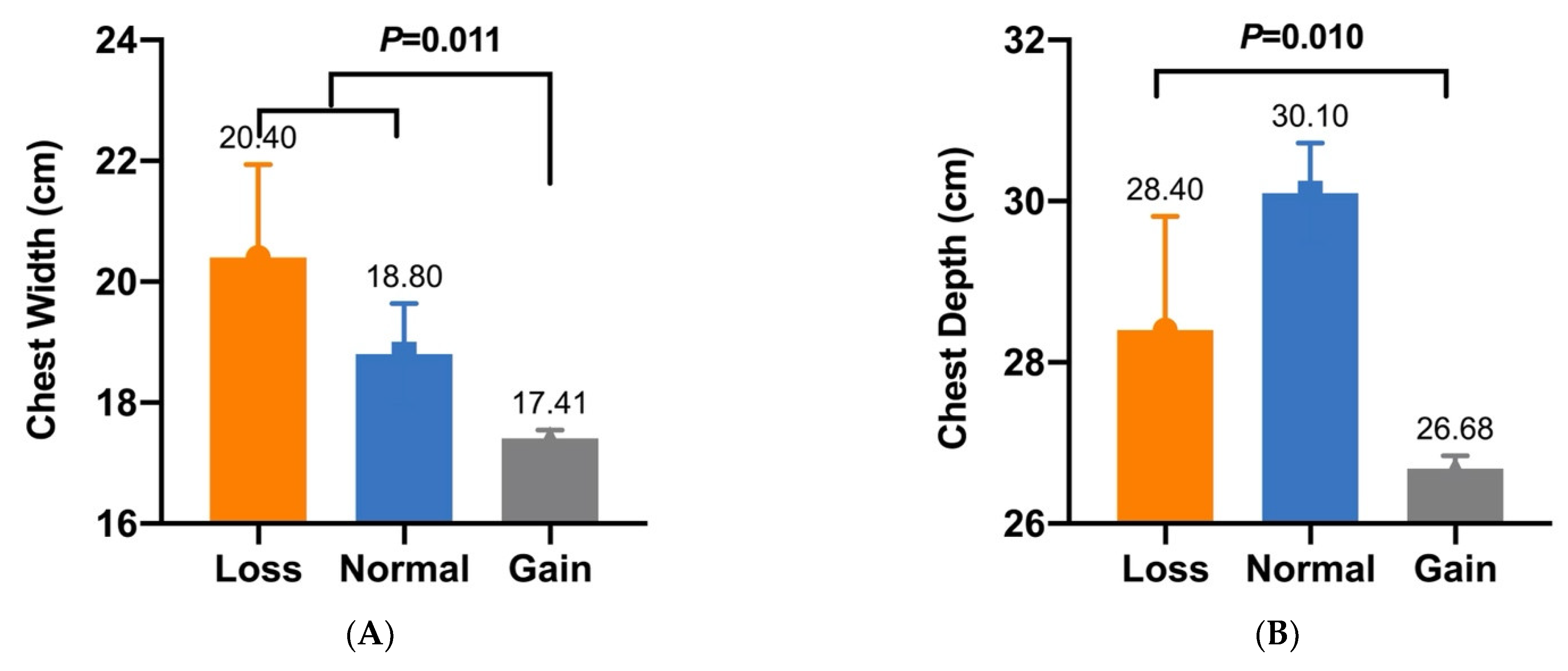
| CNV5 | Chest Width (cm) | 20.40 ± 1.54 a (n = 7) | 18.80 ± 0.84 a (n = 7) | 17.41 ± 0.14 b (n = 288) | 0.011 |
| CNV5 | Chest Depth (cm) | 28.40 ± 1.41 B (n = 7) | 30.10 ± 0.62 A (n = 7) | 26.68 ± 0.16 B (n = 288) | 0.010 |
| Locus | Trait | Genotype (LSM ± SE) | p-Value | ||
|---|---|---|---|---|---|
| Loss | Normal | Gain | |||
| CNV1 | Body Weight (kg) | - | 47.50 ± 0 (n = 1) | 32.74 ± 0.89 (n = 152) | - |
| Body Height (cm) | - | 65.00 ± 0 (n = 1) | 63.46 ± 0.45 (n = 181) | - | |
| Body Length (cm) | - | 70.00 ± 0 (n = 1) | 67.61 ± 0.50 (n = 181) | - | |
| Chest Depth (cm) | - | 34.00 ± 0 (n = 1) | 32.29 ± 0.26 (n = 181) | - | |
| Chest Width (cm) | - | 22.00 ± 0 (n = 1) | 22.27 ± 0.36 (n = 181) | - | |
| Heart Girth (cm) | - | 84.00 ± 0 (n = 1) | 76.20 ± 0.54 (n = 181) | - | |
| Cannon Bone Circumference (cm) | - | 8.00 ± 0 (n = 1) | 7.90 ± 0.03 (n = 181) | - | |
| CNV2 | Body Weight (kg) | 32.5 ± 0.62 (n = 10) | 40.19 ± 0.14 (n = 9) | 32.22 ± 0.97 (n = 126) | 0.120 |
| Body Height (cm) | 66.33 ± 1.40 (n = 9) | 64.11 ± 0.56 (n = 9) | 63.91 ± 0.53 (n = 123) | 0.495 | |
| Body Length (cm) | 70.22 ± 1.23 (n = 9) | 70.22 ± 1.68 (n = 9) | 68.17 ± 0.66 (n = 123) | 0.523 | |
| Chest Depth (cm) | 34.44 ± 0.50 (n = 9) | 33.66 ± 0.64 (n = 9) | 32.44 ± 0.33 (n = 123) | 0.186 | |
| Chest Width (cm) | 22.67 ± 0.50 (n = 9) | 23.67 ± 0.74 (n = 9) | 22.02 ± 0.19 (n = 123) | 0.054 | |
| Heart Girth (cm) | 81.00 ± 1.44 (n = 9) | 80.67 ± 0.69 (n = 9) | 76.66 ± 0.67 (n = 123) | 0.082 | |
| Cannon Bone Circumference (cm) | 7.88 ± 0.16 (n = 9) | 8.00 ± 0.16 (n = 9) | 7.87 ± 0.04 (n = 123) | 0.564 | |
| CNV3 | Body Weight (kg) | 31.41 ± 0.95 (n = 17) | 32.55 ± 1.66 (n = 35) | 32.83 ± 1.28 (n = 85) | 0.895 |
| Body Height (cm) | 64.42 ± 0.76 (n = 14) | 64.30 ± 0.81 (n = 39) | 63.83 ± 0.75 (n = 80) | 0.926 | |
| Body Length (cm) | 69.00 ± 0.85 (n = 14) | 67.85 ± 1.04 (n = 39) | 68.46 ± 0.87 (n = 80) | 0.775 | |
| Chest Depth (cm) | 33.78 ± 0.54 (n = 14) | 32.28 ± 0.54 (n = 39) | 32.73 ± 0.44 (n = 80) | 0.166 | |
| Chest Width (cm) | 22.71 ± 0.59 (n = 14) | 22.12 ± 0.33 (n = 39) | 22.13 ± 0.24 (n = 80) | 0.642 | |
| Heart Girth (cm) | 78.57 ± 1.62 (n = 14) | 76.07 ± 1.07 (n = 39) | 77.53 ± 0.91 (n = 80) | 0.483 | |
| Cannon Bone Circumference (cm) | 7.85 ± 0.14 (n = 14) | 7.87 ± 0.07 (n = 39) | 7.86 ± 0.05 (n = 80) | 0.994 | |
| CNV4 | Body Weight (kg) | 33.06 ± 1.95 (n = 20) | 33.53 ± 1.88 (n = 34) | 32.71 ± 1.18 (n = 96) | 0.933 |
| Body Height (cm) | 63.26 ± 1.01 (n = 23) | 63.86 ± 0.80 (n = 49) | 63.41 ± 0.62 (n = 107) | 0.894 | |
| Body Length (cm) | 68.00 ± 1.06 (n = 23) | 67.80 ± 0.98 (n = 49) | 67.55 ± 0.68 (n = 107) | 0.950 | |
| Chest Depth (cm) | 32.04 ± 0.64 (n = 23) | 32.59 ± 0.50 (n = 49) | 32.26 ± 0.35 (n = 107) | 0.797 | |
| Chest Width (cm) | 22.04 ± 0.40 (n = 23) | 21.88 ± 0.26 (n = 49) | 22.51 ± 0.60 (n = 107) | 0.738 | |
| Heart Girth (cm) | 76.26 ± 1.33 (n = 23) | 76.20 ± 1.12 (n = 49) | 76.27 ± 0.72 (n = 107) | 0.999 | |
| Cannon Bone Circumference (cm) | 7.87 ± 0.09 (n = 23) | 7.84 ± 0.08 (n = 49) | 7.88 ± 0.04 (n = 107) | 0.764 | |
| CNV5 | Body Weight (kg) | 29.30 ± 5.30 (n = 2) | - | 32.95 ± 0.93 (n = 143) | - |
| Body Height (cm) | 54.00 ± 0 (n = 1) | - | 64.19 ± 0.49 (n = 140) | - | |
| Body Length (cm) | 60.00 ± 0 (n = 1) | - | 68.49 ± 0.59 (n = 140) | - | |
| Chest Depth (cm) | 27.00 ± 0 (n = 1) | - | 32.78 ± 0.30 (n = 140) | - | |
| Chest Width (cm) | 20.00 ± 0 (n = 1) | - | 22.23 ± 0.18 (n = 140) | - | |
| Heart Girth (cm) | 66.00 ± 0 (n = 1) | - | 77.32 ± 0.62 (n = 140) | - | |
| Cannon Bone Circumference (cm) | 8.00 ± 0 (n = 1) | - | 7.87 ± 0.04 (n = 140) | - | |
| Locus | Trait | Genotype (LSM ± SE) | p-Value | ||
|---|---|---|---|---|---|
| Loss | Normal | Gain | |||
| CNV1 | Body Weight (kg) | - | 47.55 ± 0.55 (n = 2) | 50.29 ± 0.98 (n = 118) | - |
| Body Height (cm) | - | 75.15 ± 0.35 (n = 2) | 72.23 ± 0.50 (n = 118) | - | |
| Body Length (cm) | - | 70.90 ± 1.30 (n = 2) | 65.05 ± 0.51 (n = 118) | - | |
| Heart Girth (cm) | - | 89.50 ± 1.90 (n = 2) | 87.34 ± 0.78 (n = 118) | - | |
| Chest Width (cm) | - | 21.15 ± 0.25 (n = 2) | 21.45 ± 0.39 (n = 118) | - | |
| Chest Depth (cm) | - | 32.50 ± 2.00 (n = 2) | 34.91 ± 0.44 (n = 118) | - | |
| Cannon Bone Circumference (cm) | - | 10.50 ± 0.80 (n = 2) | 10.41 ± 0.71 (n = 118) | - | |
| CNV2 | Body Weight (kg) | 64.50 ± 0 (n = 1) | 44.52 ± 3.55 (n = 4) | 50.38 ± 1.00 (n = 112) | - |
| Body Height (cm) | 73.00 ± 0 (n = 1) | 70.75 ± 1.02 (n = 4) | 72.18 ± 0.50 (n = 112) | - | |
| Body Length (cm) | 68.50 ± 0 (n = 1) | 68.30 ± 1.86 (n = 4) | 64.95 ± 0.53 (n = 112) | - | |
| Heart Girth (cm) | 97.30 ± 0 (n = 1) | 88.58 ± 1.16 (n = 4) | 87.11 ± 0.80 (n = 112) | - | |
| Chest Width (cm) | 29.10 ± 0 (n = 1) | 21.60 ± 1.35 (n = 4) | 21.30 ± 0.41 (n = 112) | - | |
| Chest Depth (cm) | 41.00 ± 0 (n = 1) | 32.80 ± 3.80 (n = 4) | 34.74 ± 0.43 (n = 112) | - | |
| Cannon Bone Circumference (cm) | 10.50 ± 0 (n = 1) | 9.88 ± 0.55 (n = 4) | 10.42 ± 0.75 (n = 112) | - | |
| CNV3 | Body Weight (kg) | 46.21 ± 1.46 (n = 13) | 52.53 ± 0.43 (n = 7) | 50.70 ± 1.02 (n = 96) | 0.309 |
| Body Height (cm) | 69.77 ± 1.90 (n = 13) | 69.75 ± 2.55 (n = 7) | 72.56 ± 0.50 (n = 96) | 0.106 | |
| Body Length (cm) | 63.95 ± 1.96 (n = 13) | 61.94 ± 0.78 (n = 7) | 65.45 ± 0.54 (n = 96) | 0.216 | |
| Heart Girth (cm) | 32.33 ± 1.14 (n = 13) | 32.56 ± 1.58 (n = 7) | 35.13 ± 0.41 (n = 96) | 0.417 | |
| Chest Width (cm) | 21.12 ± 1.24 (n = 13) | 18.75 ± 1.16 (n = 7) | 21.56 ± 0.44 (n = 96) | 0.251 | |
| Chest Depth (cm) | 85.43 ± 1.50 (n = 13) | 83.96 ± 1.87 (n = 7) | 97.58 ± 0.84 (n = 96) | 0.060 | |
| Cannon Bone Circumference (cm) | 9.51 ± 0.25 (n = 13) | 9.41 ± 0.20 (n = 7) | 10.60 ± 0.87 (n = 96) | 0.845 | |
| CNV4 | Body Weight (kg) | 50.43 ± 3.16 (n = 6) | 47.77 ± 3.71 (n = 3) | 50.30 ± 1.01 (n = 111) | - |
| Body Height (cm) | 69.77 ± 2.44 (n = 6) | 69.80 ± 1.61 (n = 3) | 72.48 ± 0.49 (n = 111) | - | |
| Body Length (cm) | 65.88 ± 2.80 (n = 6) | 67.80 ± 2.66 (n = 3) | 65.04 ± 0.53 (n = 111) | - | |
| Heart Girth (cm) | 88.18 ± 4.07 (n = 6) | 84.33 ± 5.58 (n = 3) | 87.38 ± 0.77 (n = 111) | - | |
| Chest Width (cm) | 21.91 ± 2.01 (n = 6) | 18.37 ± 3.91 (n = 3) | 21.50 ± 0.40 (n = 111) | - | |
| Chest Depth (cm) | 34.53 ± 2.78 (n = 6) | 32.07 ± 1.57 (n = 3) | 34.96 ± 0.44 (n = 111) | - | |
| Cannon Bone Circumference | 9.67 ± 0.25 (n = 6) | 10.30 ± 1.15 (n = 3) | 10.46 ± 0.76 (n = 111) | - | |
| CNV5 | Body Weight (kg) | 67.10 ± 0 (n = 1) | - | 50.01 ± 1.00 (n = 108) | - |
| Body Height (cm) | 71.20 ± 0 (n = 1) | - | 72.05 ± 0.50 (n = 108) | - | |
| Body Length (cm) | 67.50 ± 0 (n = 1) | - | 65.04 ± 0.53 (n = 108) | - | |
| Heart Girth (cm) | 90.80 ± 0 (n = 1) | - | 87.04 ± 0.80 (n = 108) | - | |
| Chest Width (cm) | 20.70 ± 0 (n = 1) | - | 21.27 ± 0.41 (n = 108) | - | |
| Chest Depth (cm) | 34.30 ± 0 (n = 1) | - | 34.61 ± 0.42 (n = 108) | - | |
| Cannon Bone Circumference (cm) | 9.00 ± 0 (n = 1) | - | 10.46 ± 0.78 (n = 108) | - | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bi, Y.; Wang, Z.; Wang, Q.; Liu, H.; Guo, Z.; Pan, C.; Chen, H.; Zhu, H.; Wu, L.; Lan, X. Are Copy Number Variations within the FecB Gene Significantly Associated with Morphometric Traits in Goats? Animals 2022, 12, 1547. https://doi.org/10.3390/ani12121547
Bi Y, Wang Z, Wang Q, Liu H, Guo Z, Pan C, Chen H, Zhu H, Wu L, Lan X. Are Copy Number Variations within the FecB Gene Significantly Associated with Morphometric Traits in Goats? Animals. 2022; 12(12):1547. https://doi.org/10.3390/ani12121547
Chicago/Turabian StyleBi, Yi, Zhiying Wang, Qian Wang, Hongfei Liu, Zhengang Guo, Chuanying Pan, Hong Chen, Haijing Zhu, Lian Wu, and Xianyong Lan. 2022. "Are Copy Number Variations within the FecB Gene Significantly Associated with Morphometric Traits in Goats?" Animals 12, no. 12: 1547. https://doi.org/10.3390/ani12121547
APA StyleBi, Y., Wang, Z., Wang, Q., Liu, H., Guo, Z., Pan, C., Chen, H., Zhu, H., Wu, L., & Lan, X. (2022). Are Copy Number Variations within the FecB Gene Significantly Associated with Morphometric Traits in Goats? Animals, 12(12), 1547. https://doi.org/10.3390/ani12121547






