Runs of Homozygosity and Quantitative Trait Locus/Association for Semen Parameters in Selected Chinese and South African Beef Cattle
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling, Genotyping and Quality Control
2.2. Estimation of Runs of Homozygosity (ROH)
2.3. QTL and Genomic Regions in ROH
2.4. Classification of Runs and Inbreeding Co-Efficient
3. Results
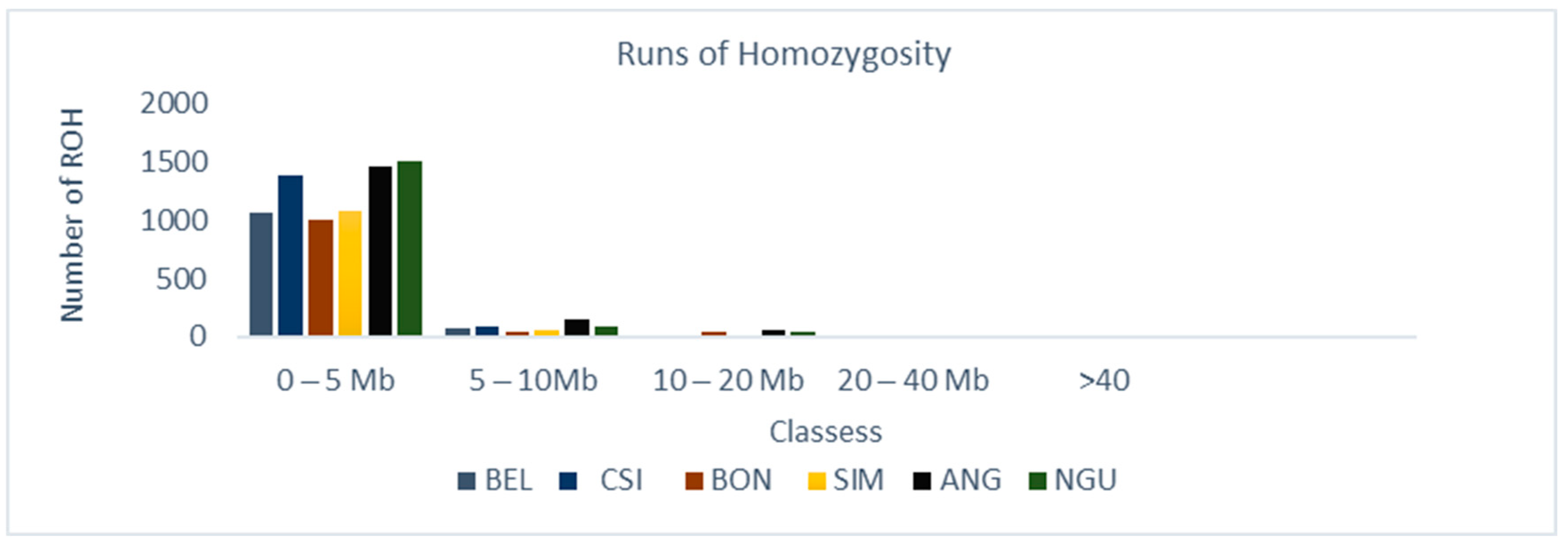
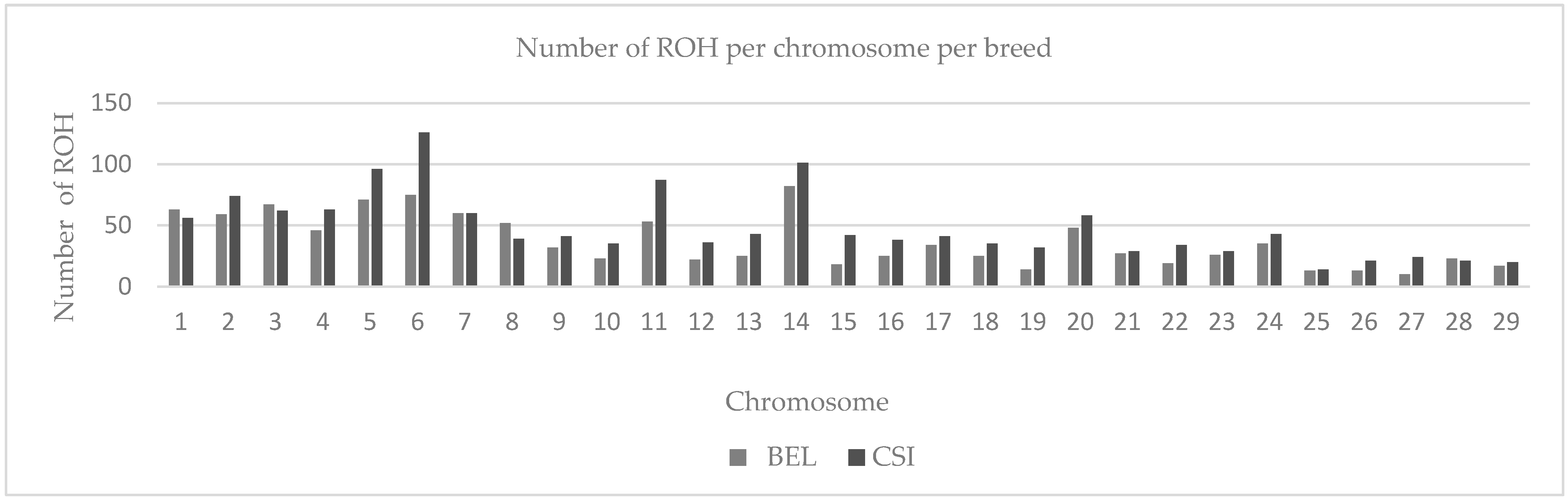
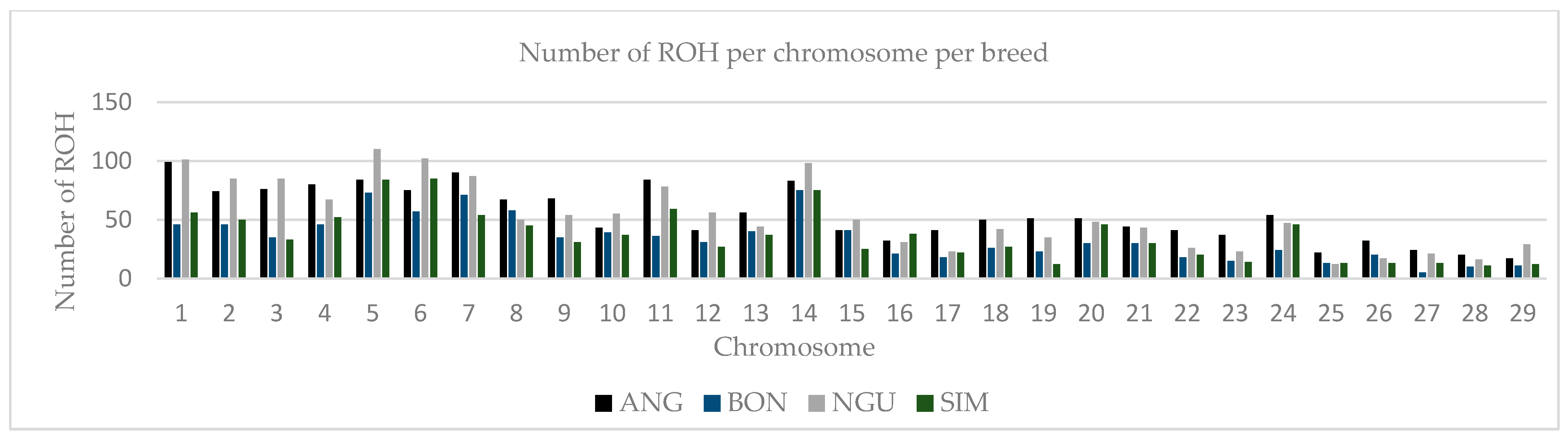
3.1. Distribution of Runs of Homozygosity (ROH)
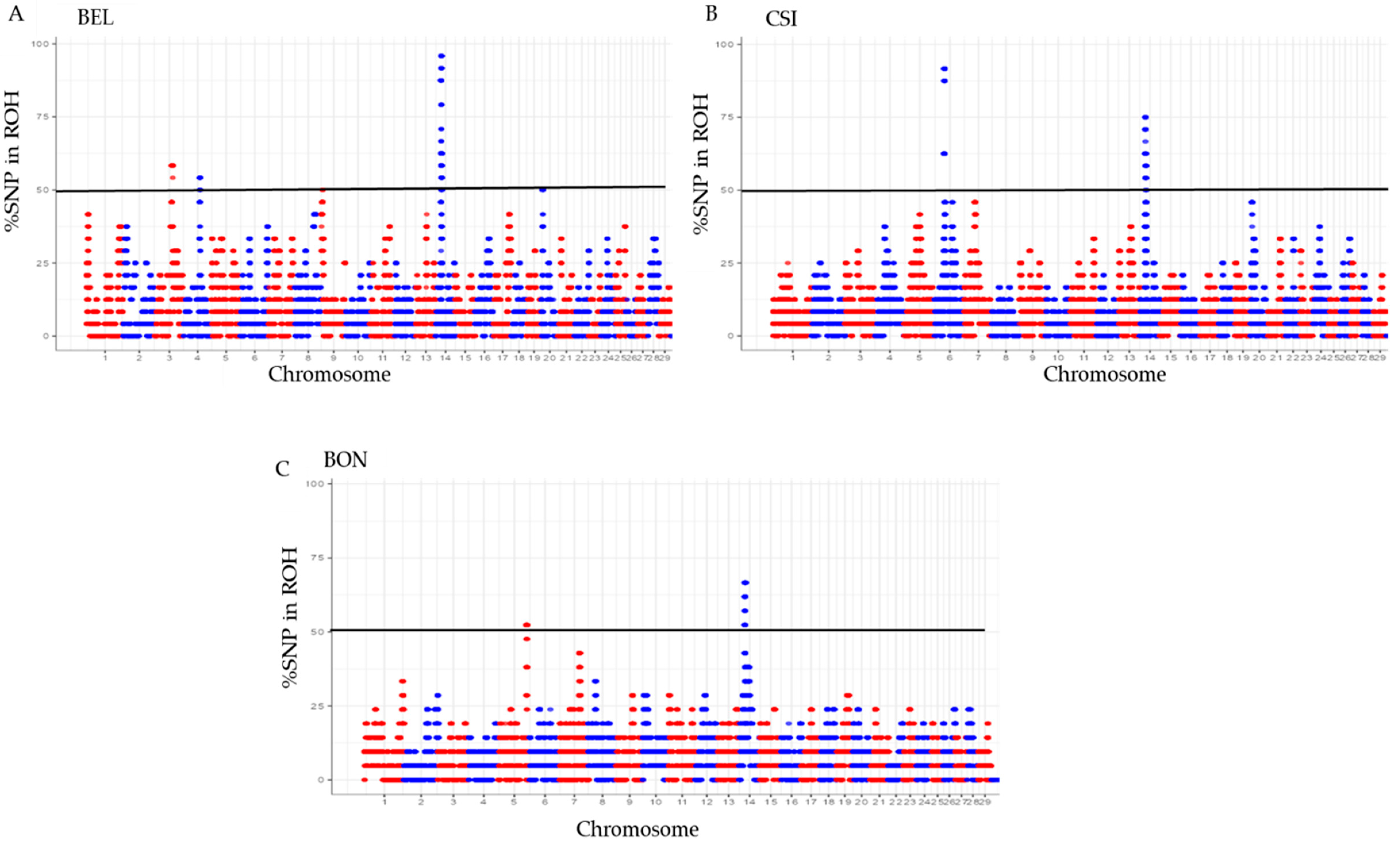
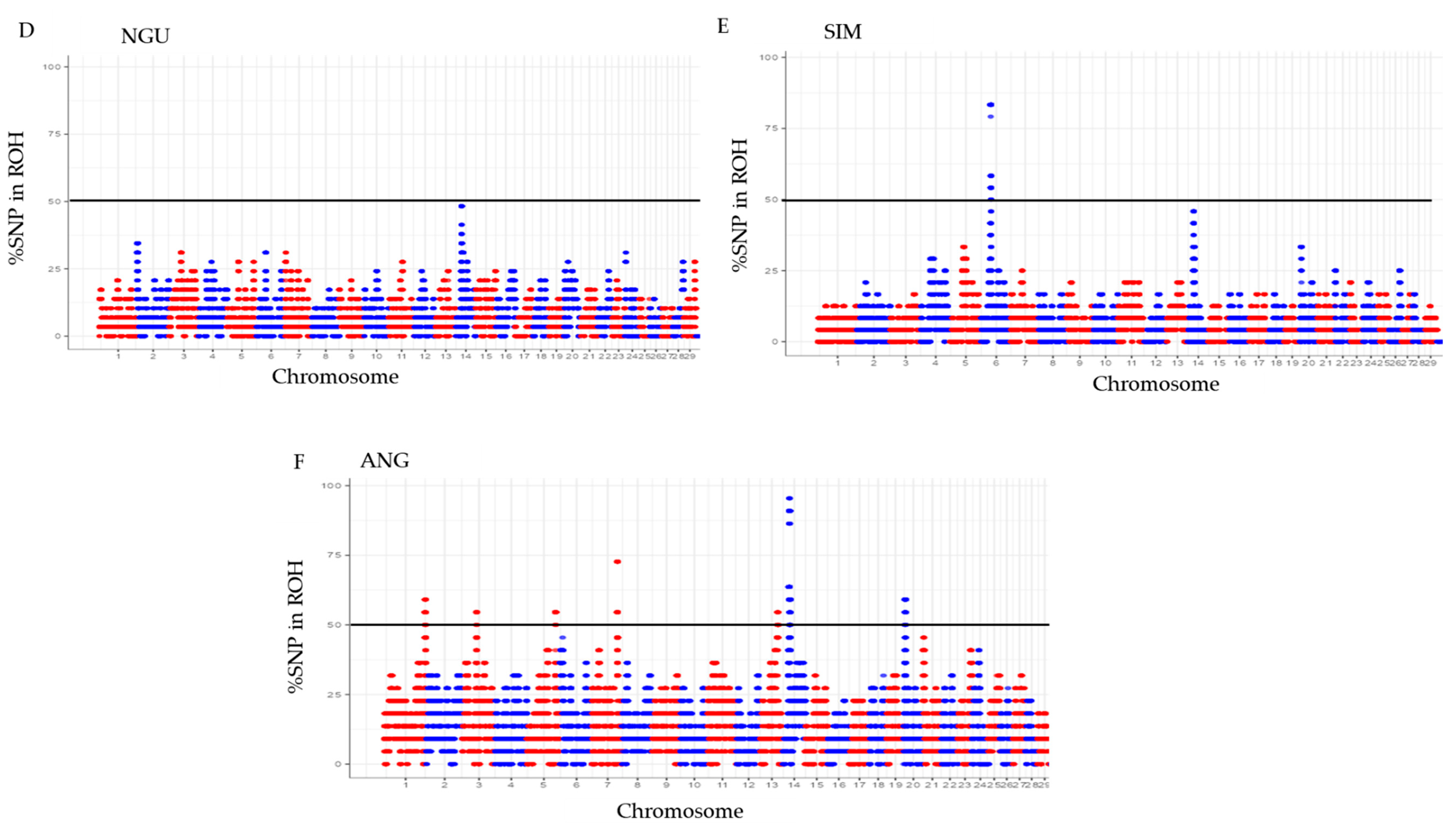
3.2. The proportion of SNPs in ROH
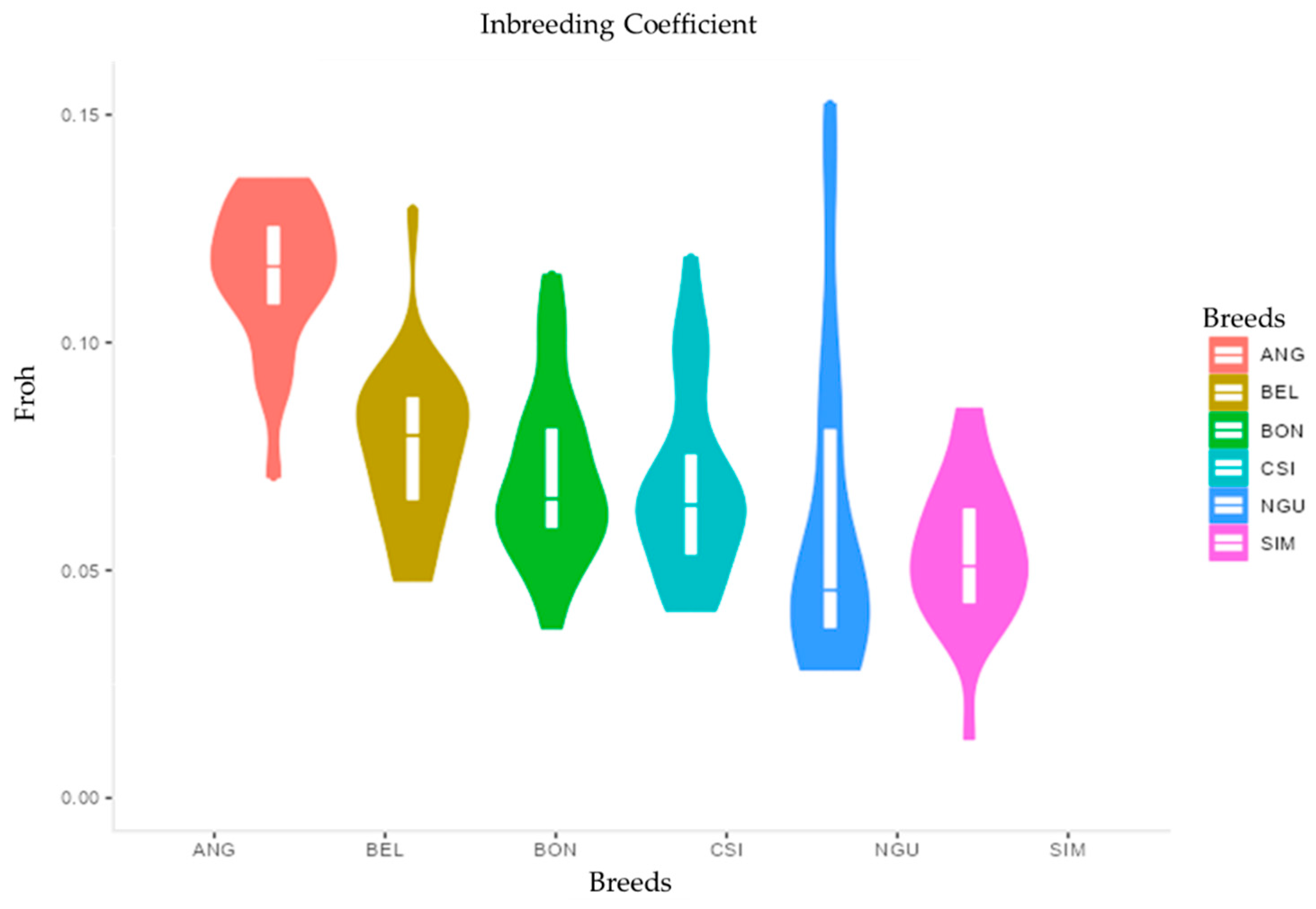
3.3. Inbreeding Coefficient FROH vs. FHOM and ROH Regions
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Scholtz, M.M.; Bester, J.; Mamabolo, J.M.; Ramsay, K.A. Results of the national cattle survey undertaken in South Africa, with emphasis on beef. Appl. Anim. Husb. Rural. Dev 2008, 1, 1–9. [Google Scholar]
- Mokolobate, M.C. Novelty Traits to Improve Cow-Calf Efficiency in Climate Smart Beef Production Systems. Ph.D. Thesis, University of the Free State, Bloemfontein, South Africa, 28 April 2015. [Google Scholar]
- Kolkman, I. Calving problems and calving ability in the phenotypically double muscled Belgian Blue breed. Ph.D. Thesis, Ghent University. Faculty of Veterinary Medicine, Ghent, Belgium, 6 June 2022. [Google Scholar]
- Peñagaricano, F.; Weigel, K.A.; Khatib, H. Genome-wide association study identifies candidate markers for bull fertility in Holstein dairy cattle. Anim. Genet. 2012, 43, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Liu, Y.; Niu, Q.; Zheng, X.; Zhang, T.; Wang, Z.; Xu, L.; Zhu, B.; Gao, X.; Zhang, L.; et al. Runs of homozygosity analysis reveals consensus homozygous regions affecting production traits in Chinese Simmental beef cattle. BMC Genom. 2021, 22, 678. [Google Scholar] [CrossRef] [PubMed]
- Peripolli, E.; Stafuzza, N.B.; Munari, D.P.; Lima, A.L.F.; Irgang, R.; Machado, M.A.; Panetto, J.C.D.C.; Ventura, R.V.; Baldi, F.; da Silva, M.V.G.B. Assessment of runs of homozygosity islands and estimates of genomic inbreeding in Gyr (Bos indicus) dairy cattle. BMC Genom. 2018, 19, 34. [Google Scholar] [CrossRef]
- Kirin, M.; McQuillan, R.; Franklin, C.S.; Campbell, H.; McKeigue, P.M.; Wilson, J.F. Genomic runs of homozygosity record population history and consanguinity. PLoS ONE 2010, 5, e13996. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gibson, J.; Morton, N.E.; Collins, A. Extended tracts of homozygosity in outbred human populations. Hum. Mol. Genet. 2006, 15, 789–795. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Broman, K.W.; Weber, J.L. Long homozygous chromosomal segments in reference families from the centre d’Etude du polymorphisme humain. Am. J. Hum. Genet. 1999, 65, 1493–1500. [Google Scholar] [CrossRef] [Green Version]
- Szmatoła, T.; Gurgul, A.; Jasielczuk, I.; Ząbek, T.; Ropka-Molik, K.; Litwińczuk, Z.; Bugno-Poniewierska, M. A comprehensive analysis of runs of homozygosity of eleven cattle breeds representing different production types. Animals 2019, 9, 1024. [Google Scholar] [CrossRef] [Green Version]
- Nani, J.P.; Peñagaricano, F. Whole-genome homozygosity mapping reveals candidate regions affecting bull fertility in US Holstein cattle. BMC Genom. 2020, 21, 338. [Google Scholar] [CrossRef]
- Bosse, M.; Megens, H.J.; Madsen, O.; Paudel, Y.; Frantz, L.A.; Schook, L.B.; Groenen, M.A. Regions of homozygosity in the porcine genome: Consequence of demography and the recombination landscape. PLoS Genet. 2012, 8, e1003100. [Google Scholar] [CrossRef] [Green Version]
- Curik, I.; Ferenčaković, M.; Sölkner, J. Inbreeding and runs of homozygosity: A possible solution to an old problem. Livest. Sci. 2014, 166, 26–34. [Google Scholar] [CrossRef]
- Purcell, S. PLINK (1.07). Documentation. pp. 1–293. Available online: http://zzz.bwh.harvard.edu/plink/dist/plink-doc-1.07.pdf (accessed on 6 June 2022).
- Childers, C.P.; Reese, J.T.; Sundaram, J.P.; Vile, D.C.; Dickens, C.M.; Childs, K.L.; Salih, H.; Bennett, A.K.; Hagen, D.E.; Adelson, D.L.; et al. Bovine Genome Database: Integrated tools for genome annotation and discovery. Nucleic Acids Res. 2011, 39 (Suppl. S1), D830–D834. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.L.; Park, C.A.; Wu, X.L.; Reecy, J.M. Animal QTLdb: An improved database tool for livestock animal QTL/association data dissemination in the post-genome era. Nucleic Acids Res. 2013, 41, D871–D879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kanehisa, M.; Goto, S. Comprehensive gene and pathway analysis of cervical cancer progression. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- McQuillan, R.; Leutenegger, A.L.; Abdel-Rahman, R.; Franklin, C.S.; Pericic, M.; Barac-Lauc, L.; Smolej-Narancic, N.; Janicijevic, B.; Polasek, O.; Tenesa, A.; et al. Runs of homozygosity in European populations. Am. J. Hum. Genet. 2008, 83, 359–372. [Google Scholar] [CrossRef] [Green Version]
- Ferencakovic, M.; Hamzic, E.; Gredler, B.; Curik, I.; Sölkner, J. Runs of homozygosity reveal genome-wide autozygosity in the Austrian Fleckvieh cattle. Agric. Conspec. Sci. 2011, 76, 325–329. [Google Scholar]
- Alin, A. Minitab. Wiley Interdiscip. Rev. Comput. Stat. 2010, 2, 723–727. [Google Scholar] [CrossRef]
- Ferenčaković, M.; Sölkner, J.; Kapš, M.; Curik, I. Genome-wide mapping and estimation of inbreeding depression of semen quality traits in a cattle population. J. Dairy Sci. 2017, 100, 4721–4730. [Google Scholar] [CrossRef]
- Margulis, S.W.; Walsh, A. The effects of inbreeding on testicular sperm concentration in Peromyscus polionotus. Reprod. Fertil. Dev. 2002, 14, 63–67. [Google Scholar] [CrossRef]
- Wildt, D.E.; Baas, E.J.; Chakraborty, P.K.; Wolfle, T.L.; Stewart, A.P. Influence of inbreeding on reproductive performance, ejaculate quality and testicular volume in the dog. Theriogenology 1982, 17, 445–452. [Google Scholar] [CrossRef]
- Aurich, C.; Achmann, R.; Aurich, J.E. Semen parameters and level of microsatellite heterozygosity in Noriker draught horse stallions. Theriogenology 2003, 60, 371–378. [Google Scholar] [CrossRef]
- Vickers, N.J. Animal communication: When i’m calling you, will you answer too? Curr. Biol. 2017, 27, R713–R715. [Google Scholar] [CrossRef]
- Doekes, H.P.; Veerkamp, R.F.; Bijma, P.; de Jong, G.; Hiemstra, S.J.; Windig, J.J. Inbreeding depression due to recent and ancient inbreeding in Dutch Holstein–Friesian dairy cattle. Genet. Sel. Evol. 2019, 51, 54. [Google Scholar] [CrossRef] [Green Version]
- Zhao, G.; Zhang, T.; Liu, Y.; Wang, Z.; Xu, L.; Zhu, B.; Gao, X.; Zhang, L.; Gao, H.; Liu, G.E.; et al. Genome-wide assessment of runs of homozygosity in Chinese wagyu beef cattle. Animals 2020, 10, 1425. [Google Scholar] [CrossRef]
- Ferenčaković, M. Molecular Dissection of Inbreeding Depression for Semen Quality Traits in Cattle. Ph.D. Thesis, University of Zagreb, Faculty of Agriculture, Zagreb, Croatia, 28 April 2015. [Google Scholar]
- Iacolina, L.; Stronen, A.V.; Pertoldi, C.; Tokarska, M.; Nørgaard, L.S.; Muñoz, J.; Kjærsgaard, A.; Ruiz-Gonzalez, A.; Kamiński, S.; Purfield, D.C. Novel graphical analyses of runs of homozygosity among species and livestock breeds. Int. J. Genom. 2016, 2016, 2152847. [Google Scholar] [CrossRef] [Green Version]
- Purfield, D.C.; Berry, D.P.; McParland, S.; Bradley, D.G. Runs of homozygosity and population history in cattle. BMC Genet. 2012, 13, 70. [Google Scholar] [CrossRef] [Green Version]
- Mastrangelo, S.; Di Gerlando, R.; Sardina, M.T.; Sutera, A.M.; Moscarelli, A.; Tolone, M.; Cortellari, M.; Marletta, D.; Crepaldi, P.; Portolano, B. Genome-wide patterns of homozygosity reveal the conservation status in five italian goat populations. Animals 2021, 11, 1510. [Google Scholar] [CrossRef]
- Macciotta, N.P.; Colli, L.; Cesarani, A.; Ajmone-Marsan, P.; Low, W.Y.; Tearle, R.; Williams, J.L. The distribution of runs of homozygosity in the genome of river and swamp buffaloes reveals a history of adaptation, migration and crossbred events. Genet. Sel. Evol. 2021, 53, 20. [Google Scholar] [CrossRef]
- Mastrangelo, S.; Tolone, M.; Sardina, M.T.; Sottile, G.; Sutera, A.M.; Di Gerlando, R.; Portolano, B. Genome-wide scan for runs of homozygosity identifies potential candidate genes associated with local adaptation in Valle del Belice sheep. Genet. Sel. Evol. 2017, 49, 84. [Google Scholar] [CrossRef] [Green Version]
- Tokarska, M.; Pertoldi, C.; Kowalczyk, R.; Perzanowski, K. Genetic status of the European bison Bison bonasus after extinction in the wild and subsequent recovery. Mammal. Rev. 2011, 41, 151–162. [Google Scholar] [CrossRef]
- Leroy, G. Inbreeding depression in livestock species: Review and meta-analysis. Anim. Genet. 2014, 45, 618–628. [Google Scholar] [CrossRef]
- Purfield, D.C.; McParland, S.; Wall, E.; Berry, D.P. The distribution of runs of homozygosity and selection signatures in six commercial meat sheep breeds. PLoS ONE 2017, 12, e0176780. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Yin, H.; Li, C.; Qin, C.; Cai, W.; Cao, M.; Zhang, S. Genetic effects of PDGFRB and MARCH1 identified in GWAS revealing strong associations with semen production traits in Chinese Holstein bulls. BMC Genet. 2017, 18, 63. [Google Scholar] [CrossRef] [Green Version]
- Tang, K.Q.; Yang, W.C.; Zhang, X.X.; Yang, L.G. Effects of polymorphisms in the bovine growth differentiation factor 9 gene on sperm quality in Holstein bulls. Genet. Mol. Res. 2013, 12, 2189–2195. [Google Scholar] [CrossRef]
- Hiltpold, M.; Niu, G.; Kadri, N.K.; Crysnanto, D.; Fang, Z.H.; Spengeler, M.; Schmitz-Hsu, F.; Fuerst, C.; Schwarzenbacher, H.; Seefried, F.R.; et al. Activation of cryptic splicing in bovine WDR19 is associated with reduced semen quality and male fertility. PLoS Genet. 2020, 16, e1008804. [Google Scholar] [CrossRef]
- Hering, D.M.; Oleński, K.; Ruść, A.; Kaminski, S. Genome-wide association study for semen volume and total number of sperm in Holstein-Friesian bulls. Anim. Reprod. Sci. 2014, 151, 126–130. [Google Scholar] [CrossRef]
- Druet, T.; Fritz, S.; Sellem, E.; Basso, B.; Gerard, O.; Salas-Cortes, L.; Humblot, P.; Druart, X.; Eggen, A. Estimation of genetic parameters and genome scan for 15 semen characteristics traits of Holstein bulls. J. Anim. Breed. Genet. 2009, 126, 269–277. [Google Scholar] [CrossRef]
| Breeds | Start SNP | End SNP | Start Position | End Position | BTA | nSNP |
|---|---|---|---|---|---|---|
| SIM | BovineHD0600010649 | BovineHD0600010935 | 38474338 | 39921321 | 6 | 258 |
| SIM | BovineHD0600010973 | ARS-BFGL-NGS-99026 | 40064733 | 72930338 | 6 | 1491 |
| CSI | BovineHD1400006792 | BovineHD1400006937 | 23394002 | 23917569 | 14 | 60 |
| CSI | BTB-01143619 | BovineHD1400007578 | 26196375 | 26302589 | 14 | 21 |
| CSI | BovineHD4100004545 | BovineHD0600010837 | 38290032 | 39418286 | 6 | 183 |
| BON | BovineHD1400006736 | BovineHD1400006801 | 23240328 | 23407192 | 14 | 55 |
| BON | BovineHD1400007096 | BovineHD1400007272 | 24448641 | 25069487 | 14 | 107 |
| BON | ARS-BFGL-NGS 100816 | BovineHD0500032567 | 111909943 | 112748475 | 5 | 39 |
| BEL | BovineHD1400001029 | BovineHD0300021345 | 7162753 | 73321455 | 3 | 43 |
| BEL | BTA-107777 | BovineHD0300022023 | 73921609 | 75752848 | 3 | 77 |
| BEL | BovineHD0500024941 | BovineHD0500026993 | 87870382 | 95025828 | 5 | 295 |
| BEL | BovineHD1400006790 | BovineHD1400007272 | 233392546 | 25069487 | 14 | 265 |
| BEL | BovineHD1400007377 | BovineHD1400007531 | 25607730 | 26108646 | 14 | 117 |
| BEL | BovineHD0400019201 | BovineHD0400019430 | 6990155 | 70569432 | 4 | 17 |
| ANG | BovineHD0100044669 | Hapmap23088-BTA-151194 | 153400885 | 154349918 | 1 | 47 |
| ANG | BovineHD1300018328 | BovineHD1300018406 | 64228423 | 64621429 | 13 | 10 |
| ANG | BovineHD1400006790 | BovineHD1400006916 | 23392546 | 23831754 | 14 | 31 |
| ANG | BovineHD1400007051 | BovineHD1400007272 | 24315353 | 25069487 | 14 | 55 |
| ANG | BovineHD1400007366 | BovineHD1400007408 | 25480962 | 25583674 | 14 | 31 |
| ANG | BovineHD1400007518 | BovineHD1400007583 | 26051609 | 26938603 | 14 | 49 |
| ANG | BovineHD1400007694 | BovineHD1400024442 | 26700286 | 26938603 | 14 | 52 |
| ANG | BovineHD2000001713 | BovineHD2000001840 | 5497761 | 5839847 | 20 | 81 |
| ANG | BovineHD0300015826 | BovineHD0300015865 | 52418548 | 52539507 | 3 | 32 |
| ANG | BovineHD0500030711 | BovineHD0500030764 | 106905471 | 106988256 | 5 | 47 |
| ANG | BovineHD0700027123 | BovineHD0700027253 | 92797461 | 93307177 | 7 | 53 |
| Breed | QTL TRAIT | BTA | Gene |
|---|---|---|---|
| SIM | Sperm motility, Sperm head abnormalities (SPHAB) and Sperm tail abnormalities | 6 | WD repeat domain 19 (WDR 19) |
| Semen volume, sperm motility and sperm count | 6 | MARCHF1 membrane associated ring-CH-type finger 1 (MARCH1) | |
| CSI | Sperm motility, Sperm head abnormalities (SPHAB) and Sperm tail abnormalities (SPTAB) | 6 | WD repeat domain 19 (WDR 19) |
| BON | Sperm motility (SPMOT) | 5 | WD repeat domain 19 (WDR 19) |
| Sperm motility (SPMOT) | 5 | Solute carrier organic anion transporter family member 1C1 (SLCOICI) | |
| BEL | Sperm motility (SPMOT) | 5 | Solute carrier organic anion transporter family member 1C1 (SLCO1C1) WD repeat domain 19 (WDR 19) |
| Sperm count and sperm motility | 4 | suppression of tumorigenicity 7 (ST7) inhibin subunit beta A (INHBA) | |
| ANG | Sperm motility and sperm count | 1 | DOP1 leucine zipper like protein B (DOP1B) |
| Sperm concentration | 1 | 1 cilia and flagella associated protein 91 CFAP9 | |
| Sperm motility | 1 | crystallin zeta like 1 (CRYZL1) ADAM metallopeptidase with thrombospondin type 1 motif 1 (ADAMTS1) | |
| Percentage normal sperm | 13 | - | |
| Sperm motility | 3 | - | |
| Sperm acrosome integrity rate, sperm motility and sperm count. | 7 | growth differentiation factor 9 GDF9 |
| Classes | Froh_Class_0 | Froh_Class_5 | Froh_Class_10 | Froh_Class_20 | Froh |
|---|---|---|---|---|---|
| Froh_Class_5 | 0.929 | ||||
| Froh_Class_10 | 0.711 | 0.864 | |||
| Froh_Class_20 | 0.567 | 0.743 | 0.881 | ||
| Froh_Class_40 | 0.400 | 0.527 | 0.599 | 0.665 | |
| Fhom | 0.603 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Modiba, M.C.; Nephawe, K.A.; Wang, J.; Hlongwane, N.; Hadebe, K.; Lu, W.; Mtileni, B. Runs of Homozygosity and Quantitative Trait Locus/Association for Semen Parameters in Selected Chinese and South African Beef Cattle. Animals 2022, 12, 1546. https://doi.org/10.3390/ani12121546
Modiba MC, Nephawe KA, Wang J, Hlongwane N, Hadebe K, Lu W, Mtileni B. Runs of Homozygosity and Quantitative Trait Locus/Association for Semen Parameters in Selected Chinese and South African Beef Cattle. Animals. 2022; 12(12):1546. https://doi.org/10.3390/ani12121546
Chicago/Turabian StyleModiba, Mamokoma Cathrine, Khathutshelo Agree Nephawe, Jun Wang, Nompilo Hlongwane, Khanyisile Hadebe, Wenfa Lu, and Bohani Mtileni. 2022. "Runs of Homozygosity and Quantitative Trait Locus/Association for Semen Parameters in Selected Chinese and South African Beef Cattle" Animals 12, no. 12: 1546. https://doi.org/10.3390/ani12121546
APA StyleModiba, M. C., Nephawe, K. A., Wang, J., Hlongwane, N., Hadebe, K., Lu, W., & Mtileni, B. (2022). Runs of Homozygosity and Quantitative Trait Locus/Association for Semen Parameters in Selected Chinese and South African Beef Cattle. Animals, 12(12), 1546. https://doi.org/10.3390/ani12121546







