Identification of Reference Genes for Expression Studies in the Whole-Blood from Three Cattle Breeds under Two States of Livestock Weather Safety
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Study Population
2.3. Weather Data
2.4. Samples, RNA Extraction, and cDNA Synthesis
2.5. Gene Selection and Primer Design
2.6. End-Point PCR and Quantitative Polymerase Chain Reaction (qPCR)
2.7. Analysis of Reference Gene Expression Stability
3. Results
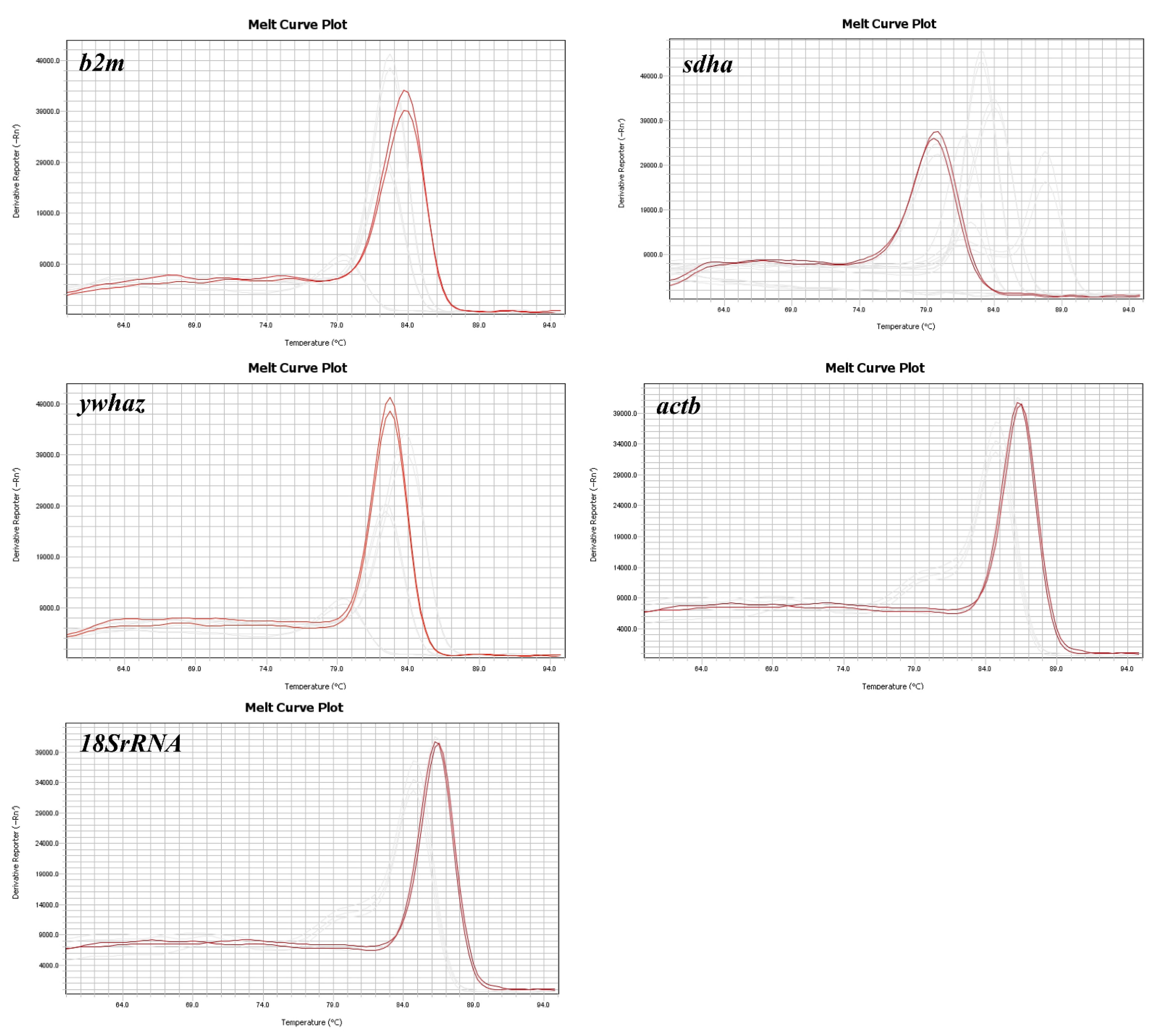
3.1. Primer Specificity
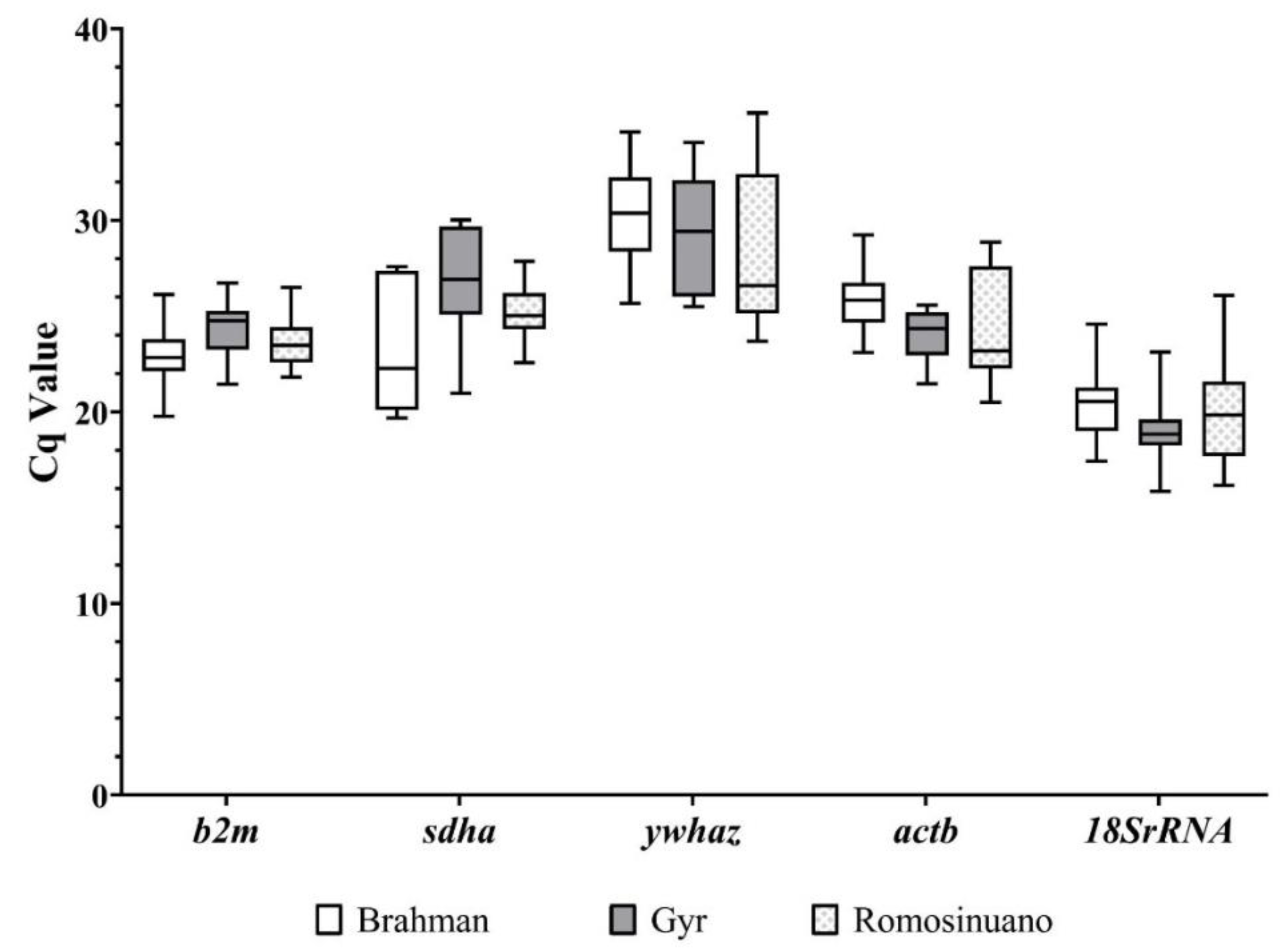
3.2. Expression Profiles of Reference Genes
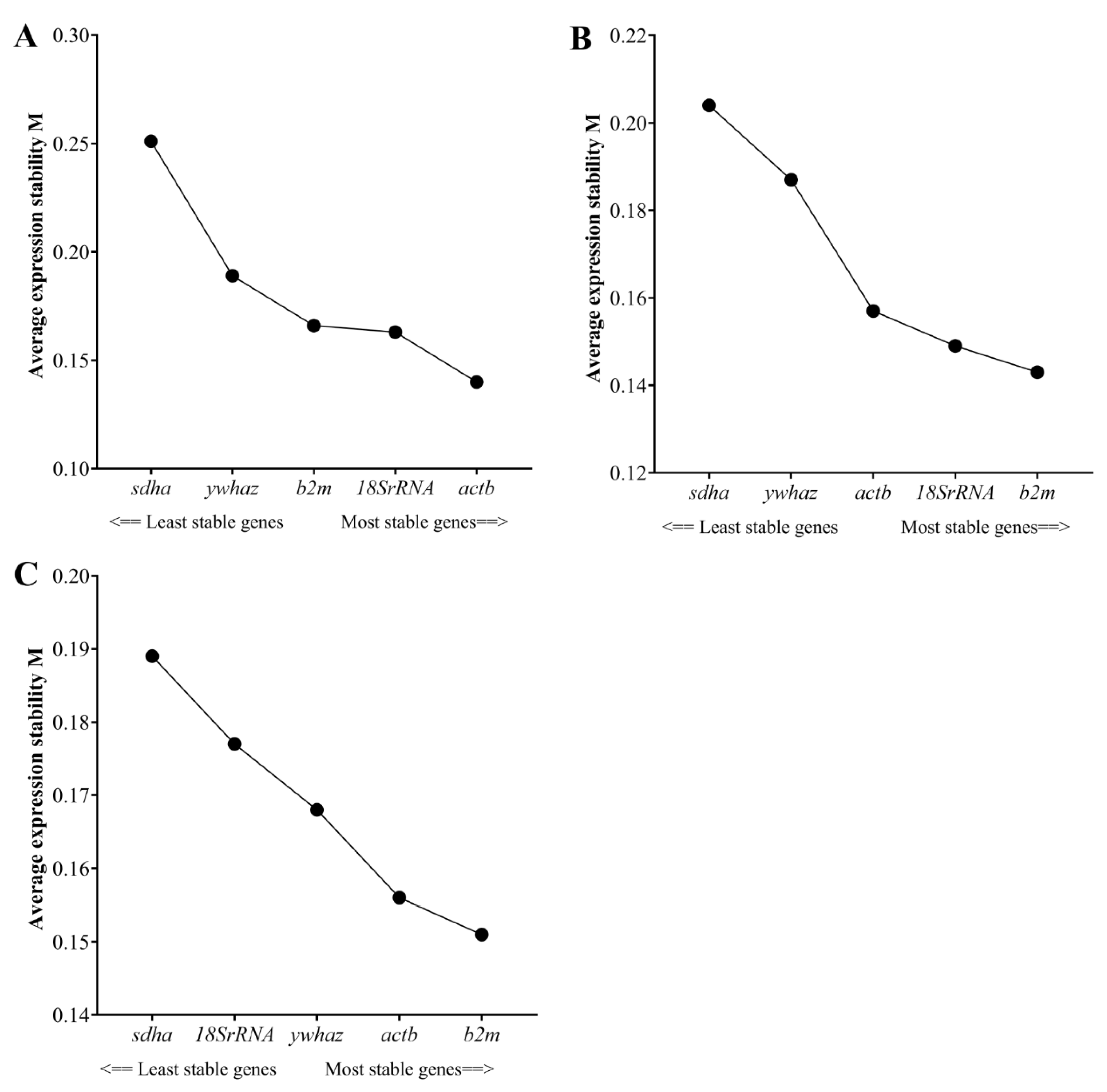
3.3. Reference Gene Stability: geNorm
3.4. Reference Gene Stability: NormFinder
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zeng, J.; Liu, S.; Zhao, Y.; Tan, X.; Aljohi, H.A.; Liu, W.; Hu, S. Identification and analysis of house-keeping and tissue-specific genes based on RNA-seq data sets across 15 mouse tissues. Gene 2016, 576, 560–570. [Google Scholar] [CrossRef]
- Summer, A.; Lora, I.; Formaggioni, P.; Gottardo, F. Impact of heat stress on milk and meat production. Anim. Front. 2019, 9, 39–46. [Google Scholar] [CrossRef]
- Vanselow, J.; Vernunft, A.; Koczan, D.; Spitschak, M.; Kuhla, B. Exposure of lactating dairy cows to acute pre-ovulatory heat stress affects granulosa cell-specific gene expression profiles in dominant follicles. PLoS ONE 2016, 11, e0160600. [Google Scholar] [CrossRef]
- Al-Katanani, Y.M.; Paula-Lopes, F.F.; Hansen, P.J. Effect of season and exposure to heat stress on oocyte competence in Holstein cows. J. Dairy Sci. 2002, 85, 390–396. [Google Scholar] [CrossRef]
- Cooke, R.F.; Daigle, C.L.; Moriel, P.; Smith, S.B.; Tedeschi, L.O.; Vendramini, J.M.B. Cattle adapted to tropical and subtropical environments: Social, nutritional, and carcass quality considerations. Anim. Sci. J. 2020, 98, skaa015. [Google Scholar] [CrossRef] [PubMed]
- Porto-Neto, L.R.; Reverter, A.; Prayaga, K.C.; Chan, E.K.F.; Johnston, D.J.; Hawken, R.J.; Fordyce, G.; Garcia, J.F.; Sonstegard, T.S.; Bolormaa, S.; et al. The genetic architecture of climatic adaptation of tropical cattle. PLoS ONE 2014, 9, e113284. [Google Scholar] [CrossRef] [PubMed]
- Herrera, J.P.; Rojas, M.D.C.; Estrada, E.G.; Tous, M.G. Behavioral biomarker of bovines of the dual purpose system. Rev. MVZ Córdoba 2017, 22, 5761–5776. [Google Scholar] [CrossRef]
- Bharati, J.; Dangi, S.S.; Bag, S.; Maurya, V.P.; Singh, G.; Kumar, P.; Sarkar, M. Expression dynamics of HSP90 and nitric oxide synthase (NOS) isoforms during heat stress acclimation in Tharparkar cattle. Int. J. Biometeorol. 2017, 61, 1461–1469. [Google Scholar] [CrossRef] [PubMed]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinforma. Biomath. 2013, 3, 71. [Google Scholar]
- Kozera, B.; Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 2013, 54, 391–406. [Google Scholar] [CrossRef]
- Hvid, H.; Ekstrøm, C.T.; Vienberg, S.; Oleksiewicz, M.B.; Klopfleisch, R. Identification of stable and oestrus cycle-independent housekeeping genes in the rat mammary gland and other tissues. Vet. J. 2011, 190, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Nolan, T. Pitfalls of quantitative real- time reverse-transcription polymerase chain reaction. J. Biomol. Tech. 2004, 15, 155. [Google Scholar] [PubMed]
- Adeola, F. Normalization of Gene Expression by Quantitative RT-PCR in Human Cell Line: Comparison of 12 Endogenous Reference Genes. Ethiop. J. Health Sci. 2018, 28, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 1–12. [Google Scholar] [CrossRef]
- Bonnet, M.; Bernard, L.; Bes, S.; Leroux, C. Selection of reference genes for quantitative real-time PCR normalisation in adipose tissue, muscle, liver and mammary gland from ruminants. Animal 2013, 7, 1344–1353. [Google Scholar] [CrossRef]
- Lisowski, P.; Pierzchała, M.; Gościk, J.; Pareek, C.S.; Zwierzchowski, L. Evaluation of reference genes for studies of gene expression in the bovine liver, kidney, pituitary, and thyroid. J. Appl. Genet. 2008, 49, 367–372. [Google Scholar] [CrossRef]
- Garner, J.B.; Chamberlain, A.J.; Vander Jagt, C.; Nguyen, T.T.T.; Mason, B.A.; Marett, L.C.; Leury, B.J.; Wales, W.J.; Hayes, B.J. Gene expression of the heat stress response in bovine peripheral white blood cells and milk somatic cells in vivo. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Adams, D.C.; Nielsen, M.K.; Schacht, W.H.; Clark, R.T. Designing and conducting experiments for range beef cows. J. Anim. Sci. 2000, 77, 510–528. [Google Scholar] [CrossRef]
- Clark, J.D.; Gebhart, G.F.; Gonder, J.C.; Keeling, M.E.; Kohn, D.F. The 1996 Guide for the Care and Use of Laboratory Animals. ILAR J. 1997, 38, 41–48. [Google Scholar] [CrossRef]
- Hahn, G.L.; Gaughan, J.B.; Mader, T.L.; Eigenberg, R.A. Chapter 5: Thermal Indices and Their Applications for Livestock Environments. In Livestock Energetics and Thermal Environmental Management, 1st ed.; DeShazer, J.A., Ed.; American Society of Agricultural and Biological Engineers: St. Joseph, MI, USA, 2009; pp. 113–130. [Google Scholar]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Choudhary, R.; Kumar, S.; Singh, S.V.; Sharma, A.K.; Goud, T.S.; Srivastava, A.K.; Kumar, A.; Mohanty, A.K.; Upadhyay, R.C. Validation of putative reference genes for gene expression studies in heat stressed and α-MSH treated melanocyte cells of Bos indicus using real-time quantitative PCR. Mol. Cell. Probes 2016, 30, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Xiao, P.; Chen, D.; Xu, L.; Zhang, B. miRDeepFinder: A miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Biol. 2012, 80, 75–84. [Google Scholar] [CrossRef]
- González-Bermúdez, L.; Anglada, T.; Genescà, A.; Martín, M.; Terradas, M. Identification of reference genes for RT-qPCR data normalisation in aging studies. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Hoogewijs, D.; Houthoofd, K.; Matthijssens, F.; Vandesompele, J.; Vanfleteren, J.R. Selection and validation of a set of reliable reference genes for quantitative sod gene expression analysis in C. elegans. BMC Mol. Biol. 2008, 9, 9. [Google Scholar] [CrossRef]
- Nolan, T.; Huggett, J.; Sanchez, E. Good Practice Guide for the Application of Quantitative PCR (qPCR). Natl. Meas. Syst. 2013. [Google Scholar] [CrossRef]
- Zhao, Y.; Luo, X.; Li, J.; Chang, T.; He, H.; Zhao, Y.; Yang, X.; Xu, Y. Stable Reference Gene Selection for RT-qPCR Analysis in Synechococcus elongatus PCC 7942 under Abiotic Stresses. Biomed Res. Int. 2019, 2019, 1–15. [Google Scholar] [CrossRef]
- Radonić, A.; Thulke, S.; Mackay, I.M.; Landt, O.; Siegert, W.; Nitsche, A. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 2004, 313, 856–862. [Google Scholar] [CrossRef] [PubMed]
- Bonefeld, B.E.; Elfving, B.; Wegener, G. Reference genes for normalization: A study of rat brain tissue. Synapse 2008, 62, 302–309. [Google Scholar] [CrossRef] [PubMed]
- Dowling, C.M.; Walsh, D.; Coffey, J.C.; Kiely, P.A. The importance of selecting the appropriate reference genes for quantitative real time PCR as illustrated using colon cancer cells and tissue [version 2; peer review: 2 approved]. F1000Research 2016, 5, 99. [Google Scholar] [CrossRef] [PubMed]
- Glare, E.M.; Divjak, M.; Bailey, M.J.; Walters, E.H. β-actin and GAPDH housekeeping gene expression in asthmatic airways is variable and not suitable for normalising mRNA levels. Thorax 2002, 57, 765–770. [Google Scholar] [CrossRef] [PubMed]
- Peng, S.; Liu, L.; Zhao, H.; Wang, H.; Li, H. Selection and validation of reference genes for quantitative real-time PCR normalization under ethanol stress conditions in Oenococcus oeni SD-2a. Front. Microbiol. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Nygard, A.B.; Jørgensen, C.B.; Cirera, S.; Fredholm, M. Selection of reference genes for gene expression studies in pig tissues using SYBR green qPCR. BMC Mol. Biol. 2007, 8, 67. [Google Scholar] [CrossRef]
- De Ketelaere, A.; Goossens, K.; Peelman, L.; Burvenich, C. Technical note: Validation of internal control genes for gene expression analysis in bovine polymorphonuclear leukocytes. J. Dairy Sci. 2006, 89, 4066–4069. [Google Scholar] [CrossRef]
- Christopher, H.H.M.; Olivier, A.E.S. Expression stability of reference genes in the skeletal muscles of beef cattle. Afr. J. Biotechnol. 2017, 16, 261–267. [Google Scholar] [CrossRef][Green Version]
- Habeeb, A.A.; Gad, A.E.; Atta, M.A. Temperature-Humidity Indices as Indicators to Heat Stress of Climatic Conditions with Relation to Production and Reproduction of Farm Animals. Int. J. Biotechnol. Recent Adv. 2018, 1, 35–50. [Google Scholar] [CrossRef]
- Lallo, C.H.O.; Cohen, J.; Rankine, D.; Taylor, M.; Cambell, J.; Stephenson, T. Characterizing heat stress on livestock using the temperature humidity index (THI)—prospects for a warmer Caribbean. Reg. Environ. Chang. 2018, 18, 2329–2340. [Google Scholar] [CrossRef]
- McDowell, R.E.; Hooven, N.W.; Camoens, J.K. Effect of Climate on Performance of Holsteins in First Lactation. J. Dairy Sci. 1976, 59, 965–971. [Google Scholar] [CrossRef]
- Brym, P.; Ruść, A.; Kamiński, S. Evaluation of reference genes for qRT-PCR gene expression studies in whole blood samples from healthy and leukemia-virus infected cattle. Vet. Immunol. Immunopathol. 2013, 153, 302–307. [Google Scholar] [CrossRef] [PubMed]
- Puech, C.; Dedieu, L.; Chantal, I.; Rodrigues, V. Design and evaluation of a unique SYBR Green real-time RT-PCR assay for quantification of five major cytokines in cattle, sheep and goats. BMC Vet. Res. 2015, 11, 65. [Google Scholar] [CrossRef] [PubMed]
- Kishore, A.; Sodhi, M.; Khate, K.; Kapila, N.; Kumari, P.; Mukesh, M. Selection of stable reference genes in heat stressed peripheral blood mononuclear cells of tropically adapted Indian cattle and buffaloes. Mol. Cell. Probes 2013, 27, 140–144. [Google Scholar] [CrossRef]
- Lovell, R.; Madden, L.; Carroll, S.; McNaughton, L. The time-profile of the PBMC HSP70 response to in vitro heat shock appears temperature-dependent. Amino Acids 2007, 33, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Lupberger, J.; Kreuzer, K.A.; Baskaynak, G.; Peters, U.R.; Le Coutre, P.; Schmidt, C.A. Quantitative analysis of beta-actin, beta-2-microglobulin and porphobilinogen deaminase mRNA and their comparison as control transcripts for RT-PCR. Mol. Cell. Probes 2002, 16, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Goidin, D.; Mamessier, A.; Staquet, M.J.; Schmitt, D.; Berthier-Vergnes, O. Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and β-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Anal. Biochem. 2001, 295, 17–21. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Zakrajsek, B.A. Effect of experimental treatment on housekeeping gene expression: Validation by real-time, quantitative RT-PCR. J. Biochem. Biophys. Methods 2000, 46, 69–81. [Google Scholar] [CrossRef]
- Selvey, S.; Thompson, E.W.; Matthaei, K.; Lea, R.A.; Irving, M.G.; Griffiths, L.R. β-Actin-An unsuitable internal control for RT-PCR. Mol. Cell. Probes 2001, 15, 307–311. [Google Scholar] [CrossRef]
- Mihi, B.; Rinaldi, M.; Geldhof, P. Effect of an Ostertagia ostertagi infection on the transcriptional stability of housekeeping genes in the bovine abomasum. Vet. Parasitol. 2011, 181, 354–359. [Google Scholar] [CrossRef]
- Bogdanović, M.D.; Dragićević, M.B.; Tanić, N.T.; Todorović, S.I.; Mišić, D.M.; Živković, S.T.; Tissier, A.; Simonović, A.D. Reverse Transcription of 18S rRNA with Poly(dT)18 and Other Homopolymers. Plant Mol. Biol. Rep. 2013, 31, 55–63. [Google Scholar] [CrossRef]
- Vorachek, W.R.; Bobe, G.; Hall, J.A. Reference gene selection for quantitative PCR studies in sheep neutrophils. Int. J. Mol. Sci. 2013, 14, 11484–11495. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef]
- Schmidtmann, C.; Schönherz, A.; Guldbrandtsen, B.; Marjanovic, J.; Calus, M.; Hinrichs, D.; Thaller, G. Assessing the genetic background and genomic relatedness of red cattle populations originating from Northern Europe. Genet. Sel. Evol. 2021, 53, 1–18. [Google Scholar] [CrossRef]
- Scharf, B.A. Comparison of thermoregulatory mechanisms in heat sensitive and tolerant breeds of bos taurus cattle. Master’s Thesis, University of Missouri-Columbia, Columbia, MO, USA, 2008. [Google Scholar]
- Paula-Lopes, F.F.; Lima, R.S.; Satrapa, R.A.; Barros, C.M. Physiology and endocrinology symposium: Influence of cattle genotype (Bos indicus vs. Bos taurus) on oocyte and preimplantation embryo resistance to increased temperature. J. Anim. Sci. 2013, 91, 1143–1153. [Google Scholar] [CrossRef]
- Silva, A.A.; Azevedo, A.L.S.; Verneque, R.S.; Gasparini, K.; Peixoto, M.G.C.D.; da Silva, M.V.G.B.; Lopes, P.S.; Guimarães, S.E.F.; Machado, M.A. Quantitative trait loci affecting milk production traits on bovine chromosome 6 in zebuine Gyr breed. J. Dairy Sci. 2011, 94, 971–980. [Google Scholar] [CrossRef] [PubMed]
- Fernández, J.C.; Pérez, J.E.; Herrera, N.; Martínez, R.; Bejarano, D.; Rocha, J.F. Research Article Genomic association study for age at first calving and calving interval in Romosinuano and Costeño con Cuernos cattle. Genet. Mol. Res. 2019, 18, 1–13. [Google Scholar] [CrossRef]
- Martínez, R.; Gallego, J.; Onofre, G.; Pérez, J.; Vasquez, R. Evaluación de la variabilidad y potencial genético de poblaciones de bovinos criollos colombianos. Anim. Genet. Resour. Inf. 2009, 44, 57–66. [Google Scholar] [CrossRef]
| Gene | Primer Sequence (5′–3′) | Primer Length (nt) | Tm (°C) | GC% | Annealing Temperature (°C) | Amplicon Size (bp) | Reference | |
|---|---|---|---|---|---|---|---|---|
| b2m | F | CTGCTATGTGTATGGGTTCC | 20 | 55.6 | 50 | 54 | 141 | [22] |
| R | GGAGTGAACTCAGCGTG | 17 | 54.8 | 58.8 | ||||
| sdha | F | TGCAGACCATCTACGGAGCGGA | 22 | 65.44 | 59.09 | 55 | 163 | This study |
| R | ACGTAGGAGAGCGTGTGCTTCCTCC | 25 | 67.96 | 60.00 | ||||
| ywhaz | F | AGCAGGCTGAGCGATATGAT | 20 | 59.02 | 50.00 | 55 | 180 | This study |
| R | TCTCAGCACCTTCCGTCTTT | 20 | 58.95 | 50.00 | ||||
| actb | F | GGGATGAGGCTCAGAGCAAGAGA | 23 | 63.65 | 56.52 | 60 | 118 | This study |
| R | AGCTCGTTGTAGAAGGTGTGGTGCC | 25 | 66.91 | 56.00 | ||||
| 18S rRNA | F | TAGAGGGACAAGTGGCGTTC | 20 | 59.39 | 55.00 | 55 | 104 | This study |
| R | CGCTGAGCCAGTCAGTGTAG | 20 | 60.46 | 60.00 | ||||
| Ranking | Brahman | Gyr | Romosinuano | |||
|---|---|---|---|---|---|---|
| Gene 1 | Stability Value | Gene 1 | Stability Value | Gene 1 | Stability Value | |
| 1 | actb | 0.009 | 18SrRNA | 0.009 | actb | 0.017 |
| 2 | 18SrRNA | 0.018 | b2m | 0.016 | b2m | 0.017 |
| 3 | b2m | 0.021 | ywhaz | 0.018 | ywhaz | 0.020 |
| 4 | ywhaz | 0.025 | actb | 0.021 | 18SrRNA | 0.022 |
| 5 | sdha | 0.043 | sdha | 0.032 | sdha | 0.029 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lozano-Villegas, K.J.; Rodríguez-Hernández, R.; Herrera-Sánchez, M.P.; Uribe-García, H.F.; Naranjo-Gómez, J.S.; Otero-Arroyo, R.J.; Rondón-Barragán, I.S. Identification of Reference Genes for Expression Studies in the Whole-Blood from Three Cattle Breeds under Two States of Livestock Weather Safety. Animals 2021, 11, 3073. https://doi.org/10.3390/ani11113073
Lozano-Villegas KJ, Rodríguez-Hernández R, Herrera-Sánchez MP, Uribe-García HF, Naranjo-Gómez JS, Otero-Arroyo RJ, Rondón-Barragán IS. Identification of Reference Genes for Expression Studies in the Whole-Blood from Three Cattle Breeds under Two States of Livestock Weather Safety. Animals. 2021; 11(11):3073. https://doi.org/10.3390/ani11113073
Chicago/Turabian StyleLozano-Villegas, Kelly J., Roy Rodríguez-Hernández, María P. Herrera-Sánchez, Heinner F. Uribe-García, Juan S. Naranjo-Gómez, Rafael J. Otero-Arroyo, and Iang S. Rondón-Barragán. 2021. "Identification of Reference Genes for Expression Studies in the Whole-Blood from Three Cattle Breeds under Two States of Livestock Weather Safety" Animals 11, no. 11: 3073. https://doi.org/10.3390/ani11113073
APA StyleLozano-Villegas, K. J., Rodríguez-Hernández, R., Herrera-Sánchez, M. P., Uribe-García, H. F., Naranjo-Gómez, J. S., Otero-Arroyo, R. J., & Rondón-Barragán, I. S. (2021). Identification of Reference Genes for Expression Studies in the Whole-Blood from Three Cattle Breeds under Two States of Livestock Weather Safety. Animals, 11(11), 3073. https://doi.org/10.3390/ani11113073











