Gut Microbiome Composition is Associated with Age and Memory Performance in Pet Dogs
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Subjects
2.3. 16S rRNA Gene Amplicon Sequencing
2.4. Assessing the Cognitive Performance: Memory Test
2.5. Statistical Analysis and Ordination
3. Results
3.1. Gut Microbiome Composition
3.2. Gut Microbiota Composition Associations with Age and Memory Performance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Clarke, G.; Grenham, S.; Scully, P.; Fitzgerald, P.; Moloney, R.D.; Shanahan, F.; Dinan, T.G.; Cryan, J.F. The microbiome-gut-brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatry 2013, 18, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Desbonnet, L.; Clarke, G.; Shanahan, F.; Dinan, T.G.; Cryan, J.F. Microbiota is essential for social development in the mouse. Mol. Psychiatry 2014, 19, 146–148. [Google Scholar] [CrossRef] [PubMed]
- Stilling, R.M.; Ryan, F.J.; Hoban, A.E.; Shanahan, F.; Clarke, G.; Claesson, M.J.; Dinan, T.G.; Cryan, J.F. Microbes & neurodevelopment—Absence of microbiota during early life increases activity-related transcriptional pathways in the amygdala. Brain. Behav. Immun. 2015, 50, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Ambrosini, Y.M.; Borcherding, D.; Kanthasamy, A.; Kim, H.J.; Willette, A.A.; Jergens, A.; Allenspach, K.; Mochel, J.P. The Gut-Brain Axis in Neurodegenerative Diseases and Relevance of the Canine Model: A Review. Front. Aging Neurosci. 2019, 11, 130. [Google Scholar] [CrossRef] [PubMed]
- Barrett, E.; Ross, R.P.; O’Toole, P.W.; Fitzgerald, G.F.; Stanton, C. γ-Aminobutyric acid production by culturable bacteria from the human intestine. J. Appl. Microbiol. 2012, 113, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Strandwitz, P. Neurotransmitter modulation by the gut microbiota. Brain Res. 2018, 1693, 128–133. [Google Scholar] [CrossRef]
- Foster, J.A.; McVey Neufeld, K.-A. Gut–brain axis: How the microbiome influences anxiety and depression. Trends Neurosci. 2013, 36, 305–312. [Google Scholar] [CrossRef]
- Topál, J.; Miklósi, Á.; Gácsi, M.; Dóka, A.; Pongrácz, P.; Kubinyi, E.; Virányi, Z.; Csányi, V. The Dog as a Model for Understanding Human Social Behavior. In Advances in the Study of Behavior; Brockmann, H.J., Roper, T.J., Naguib, M., Wynne-Edwards, K.E., Mitani, J.C., Simmons, L.W., Eds.; Academic Press: Burlington, VT, USA, 2009; Volume 39, pp. 71–116. ISBN 9780123744746. [Google Scholar]
- Sutter, N.B.; Ostrander, E.A. Dog star rising: The canine genetic system. Nat. Rev. Genet. 2004, 5, 900–910. [Google Scholar] [CrossRef]
- Sándor, S.; Kubinyi, E. Genetic pathways of aging and their relevance in the dog as a natural model of human aging. Front. Genet. 2019, 10, 948. [Google Scholar] [CrossRef]
- Kubinyi, E.; Iotchev, I.B. A Preliminary Study toward a Rapid Assessment of Age-Related Behavioral Differences in Family Dogs. Animals 2020, 10, 1222. [Google Scholar] [CrossRef]
- Szabó, D.; Miklósi, Á.; Kubinyi, E. Owner reported sensory impairments affect behavioural signs associated with cognitive decline in dogs. Behav. Processes 2018, 157, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Jónás, D.; Sándor, S.; Tátrai, K.; Egyed, B.; Kubinyi, E. A Preliminary Study to Investigate the Genetic Background of Longevity Based on Whole-Genome Sequence Data of Two Methuselah Dogs. Front. Genet. 2020, 11, 315. [Google Scholar] [CrossRef] [PubMed]
- Chapagain, D.; Range, F.; Huber, L.; Virányi, Z. Cognitive Aging in Dogs. Gerontology 2018, 64, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Urfer, S.R.; Kaeberlein, T.L.; Mailheau, S.; Bergman, P.J.; Creevy, K.E.; Promislow, D.E.L.; Kaeberlein, M. A randomized controlled trial to establish effects of short-term rapamycin treatment in 24 middle-aged companion dogs. GeroScience 2017. [Google Scholar] [CrossRef]
- Wallis, L.J.; Szabó, D.; Erdélyi-Belle, B.; Kubinyi, E. Demographic Change Across the Lifespan of Pet Dogs and Their Impact on Health Status. Front. Vet. Sci. 2018, 5, 200. [Google Scholar] [CrossRef]
- Mongillo, P.; Araujo, J.A.; Pitteri, E.; Carnier, P.; Adamelli, S.; Regolin, L.; Marinelli, L. Spatial reversal learning is impaired by age in pet dogs. Age (Omaha) 2013, 35, 2273–2282. [Google Scholar] [CrossRef]
- Mongillo, P.; Pitteri, E.; Carnier, P.; Gabai, G.; Adamelli, S.; Marinelli, L. Does the attachment system towards owners change in aged dogs? Physiol. Behav. 2013, 120, 64–69. [Google Scholar] [CrossRef]
- Turcsán, B.; Tátrai, K.; Petró, E.; Topál, J.; Balogh, L.; Egyed, B.; Kubinyi, E. Comparison of Behavior and Genetic Structure in Populations of Family and Kenneled Beagles. Front. Vet. Sci. 2020, 7, 183. [Google Scholar] [CrossRef]
- Coelho, L.P.; Kultima, J.R.; Costea, P.I.; Fournier, C.; Pan, Y.; Czarnecki-Maulden, G.; Hayward, M.R.; Forslund, S.K.; Schmidt, T.S.B.; Descombes, P.; et al. Similarity of the dog and human gut microbiomes in gene content and response to diet. Microbiome 2018, 6, 72. [Google Scholar] [CrossRef]
- Mizukami, K.; Uchiyama, J.; Igarashi, H.; Murakami, H.; Osumi, T.; Shima, A.; Ishiahra, G.; Nasukawa, T.; Une, Y.; Sakaguchi, M. Age-related analysis of the gut microbiome in a purebred dog colony. FEMS Microbiol. Lett. 2019, 366. [Google Scholar] [CrossRef]
- Pilla, R.; Suchodolski, J.S. The Role of the Canine Gut Microbiome and Metabolome in Health and Gastrointestinal Disease. Front. Vet. Sci. 2020, 6, 498. [Google Scholar] [CrossRef]
- Heath, S.E.; Barabas, S.; Craze, P.G. Nutritional supplementation in cases of canine cognitive dysfunction—A clinical trial. Appl. Anim. Behav. Sci. 2007, 105, 284–296. [Google Scholar] [CrossRef]
- Masouka, H.; Shimada, K.; Kiyosue-Yasuda, T.; Kiyosue, M.; Oishi, Y.; Kimura, S.; Yamada, A.; Hirayama, K. Transition of the intestinal microbiota of dogs with age. Biosci. Microbiota Food Heal. 2017, 36, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.H.; Kim, Y.M.; Zhou, Y.X.; Hu, D.; Zhong, C.; Chang, H.; Brislawn, C.; Langley, S.; Wang, Y.; Peisl, B.Y.L.; et al. Genetic and metabolic links between the murine microbiome and memory. Microbiome 2020, 8, 53. [Google Scholar] [CrossRef] [PubMed]
- Omatsu, T.; Omura, M.; Katayama, Y.; Kimura, T.; Okumura, M.; Okumura, A.; Murata, Y.; Mizutani, T. Molecular diversity of the faecal microbiota of Toy Poodles in Japan. J. Vet. Med. Sci. 2018, 80, 749–754. [Google Scholar] [CrossRef]
- Iotchev, I.B.; Szabó, D.; Kis, A.; Kubinyi, E. Possible association between spindle frequency and reversal learning in aged family dogs. Sci. Rep. 2020. [Google Scholar] [CrossRef]
- Benedek, K.; Bálint, J.; Máthé, I.; Mara, G.; Felföldi, T.; Szabó, A.; Fazakas, C.; Albert, C.; Buchkowski, R.W.; Schmitz, O.J.; et al. Linking intraspecific variation in plant chemical defence with arthropod and soil bacterial community structure and N allocation. Plant Soil 2019, 444, 383–397. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef]
- Piotti, P.; Szabó, D.; Wallis, L.; Bognár, Z.; Stiegmann, B.S.; Egerer, A.; Marty, P.; Kubinyi, E. The effect of age on visuo-spatial short-term memory in family dogs. Pet Behav. Sci. 2017, 17. [Google Scholar] [CrossRef]
- Kubinyi, E.; Turcsán, B.; Miklósi, Á. Dog and owner demographic characteristics and dog personality trait associations. Behav. Processes 2009, 81, 392–401. [Google Scholar] [CrossRef] [PubMed]
- Middelbos, I.S.; Vester Boler, B.M.; Qu, A.; White, B.A.; Swanson, K.S.; Fahey, G.C. Phylogenetic Characterization of Fecal Microbial Communities of Dogs Fed Diets with or without Supplemental Dietary Fiber Using 454 Pyrosequencing. PLoS ONE 2010, 5, e9768. [Google Scholar] [CrossRef] [PubMed]
- Swanson, K.S.; Dowd, S.E.; Suchodolski, J.S.; Middelbos, I.S.; Vester, B.M.; Barry, K.A.; Nelson, K.E.; Torralba, M.; Henrissat, B.; Coutinho, P.M.; et al. Phylogenetic and gene-centric metagenomics of the canine intestinal microbiome reveals similarities with humans and mice. ISME J. 2011, 5, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.; Unterer, S.; Suchodolski, J.S.; Honneffer, J.B.; Guard, B.C.; Lidbury, J.A.; Steiner, J.M.; Fritz, J.; Kölle, P. The fecal microbiome and metabolome differs between dogs fed Bones and Raw Food (BARF) diets and dogs fed commercial diets. PLoS ONE 2018, 13. [Google Scholar] [CrossRef] [PubMed]
- Jarett, J.K.; Carlson, A.; Rossoni Serao, M.; Strickland, J.; Serfilippi, L.; Ganz, H.H. Diets with and without edible cricket support a similar level of diversity in the gut microbiome of dogs. PeerJ 2019, 2019, e7661. [Google Scholar] [CrossRef]
- Kerr, K.R.; Beloshapka, A.N.; Swanson, K.S. 2011 and 2012 early careers acihevement awards: Use of genomic biology to study companion animal intestinal microbiota. J. Anim. Sci. 2013, 91, 2504–2511. [Google Scholar] [CrossRef]
- da Fonseca Ferreira, M.; Salavati Schmitz, S.; Schoenebeck, J.J.; Clements, D.N.; Campbell, S.M.; Gaylor, D.E.; Mellanby, R.J.; Gow, A.G.; Salavati, M. Lactulose drives a reversible reduction and qualitative modulation of the faecal microbiota diversity in healthy dogs. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Biagi, E.; Nylund, L.; Candela, M.; Ostan, R.; Bucci, L.; Pini, E.; Nikkïla, J.; Monti, D.; Satokari, R.; Franceschi, C.; et al. Through Ageing, and Beyond: Gut Microbiota and Inflammatory Status in Seniors and Centenarians. PLoS ONE 2010, 5, e10667. [Google Scholar] [CrossRef]
- Vázquez-Baeza, Y.; Hyde, E.R.; Suchodolski, J.S.; Knight, R. Dog and human inflammatory bowel disease rely on overlapping yet distinct dysbiosis networks. Nat. Microbiol. 2016, 1, 16177. [Google Scholar] [CrossRef]
- Gevers, D.; Kugathasan, S.; Denson, L.A.; Vázquez-Baeza, Y.; Van Treuren, W.; Ren, B.; Schwager, E.; Knights, D.; Song, S.J.; Yassour, M.; et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 2014, 15, 382–392. [Google Scholar] [CrossRef] [PubMed]
- Swidsinski, A.; Dörffel, Y.; Loening-Baucke, V.; Theissig, F.; Rückert, J.C.; Ismail, M.; Rau, W.A.; Gaschler, D.; Weizenegger, M.; Kühn, S.; et al. Acute appendicitis is characterised by local invasion with Fusobacterium nucleatum/necrophorum. Gut 2011, 60, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Hamady, M.; Lozupone, C.; Turnbaugh, P.J.; Ramey, R.R.; Bircher, J.S.; Schlegel, M.L.; Tucker, T.A.; Schrenzel, M.D.; Knight, R.; et al. Evolution of Mammals and Their Gut Microbes. Science 2008, 320, 1647–1651. [Google Scholar] [CrossRef] [PubMed]
- Muegge, B.D.; Kuczynski, J.; Knights, D.; Clemente, J.C.; Gonzalez, A.; Fontana, L.; Henrissat, B.; Knight, R.; Gordon, J.I. Diet Drives Convergence in Gut Microbiome Functions across Mammalian Phylogeny and Within Humans. Science 2011, 332, 970–974. [Google Scholar] [CrossRef] [PubMed]
- Nelson, T.M.; Rogers, T.L.; Brown, M.V. The Gut Bacterial Community of Mammals from Marine and Terrestrial Habitats. PLoS ONE 2013, 8, e83655. [Google Scholar] [CrossRef]
- Kirchoff, N.S.; Udell, M.A.R.; Sharpton, T.J. The gut microbiome correlates with conspecific aggression in a small population of rescued dogs (Canis familiaris). PeerJ 2019, 7, e6103. [Google Scholar] [CrossRef]
- Song, S.J.; Lauber, C.; Costello, E.K.; Lozupone, C.A.; Humphrey, G.; Berg-Lyons, D.; Caporaso, J.G.; Knights, D.; Clemente, J.C.; Nakielny, S.; et al. Cohabiting family members share microbiota with one another and with their dogs. Elife 2013, 2, 1–22. [Google Scholar] [CrossRef]
- Emery, D.C.; Shoemark, D.K.; Batstone, T.E.; Waterfall, C.M.; Coghill, J.A.; Cerajewska, T.L.; Davies, M.; West, N.X.; Allen, S.J. 16S rRNA next generation sequencing analysis shows bacteria in Alzheimer’s Post-Mortem Brain. Front. Aging Neurosci. 2017, 9, 1–13. [Google Scholar] [CrossRef]
- Topál, J.; Román, V.; Turcsán, B. The dog (Canis familiaris) as a translational model of autism: It is high time we move from promise to reality. Wiley Interdiscip. Rev. Cogn. Sci. 2019, 10, e1495. [Google Scholar] [CrossRef]
- Hsiao, E.Y.; McBride, S.W.; Hsien, S.; Sharon, G.; Hyde, E.R.; McCue, T.; Codelli, J.A.; Chow, J.; Reisman, S.E.; Petrosino, J.F.; et al. Microbiota Modulate Behavioral and Physiological Abnormalities Associated with Neurodevelopmental Disorders. Cell 2013, 155, 1451–1463. [Google Scholar] [CrossRef]
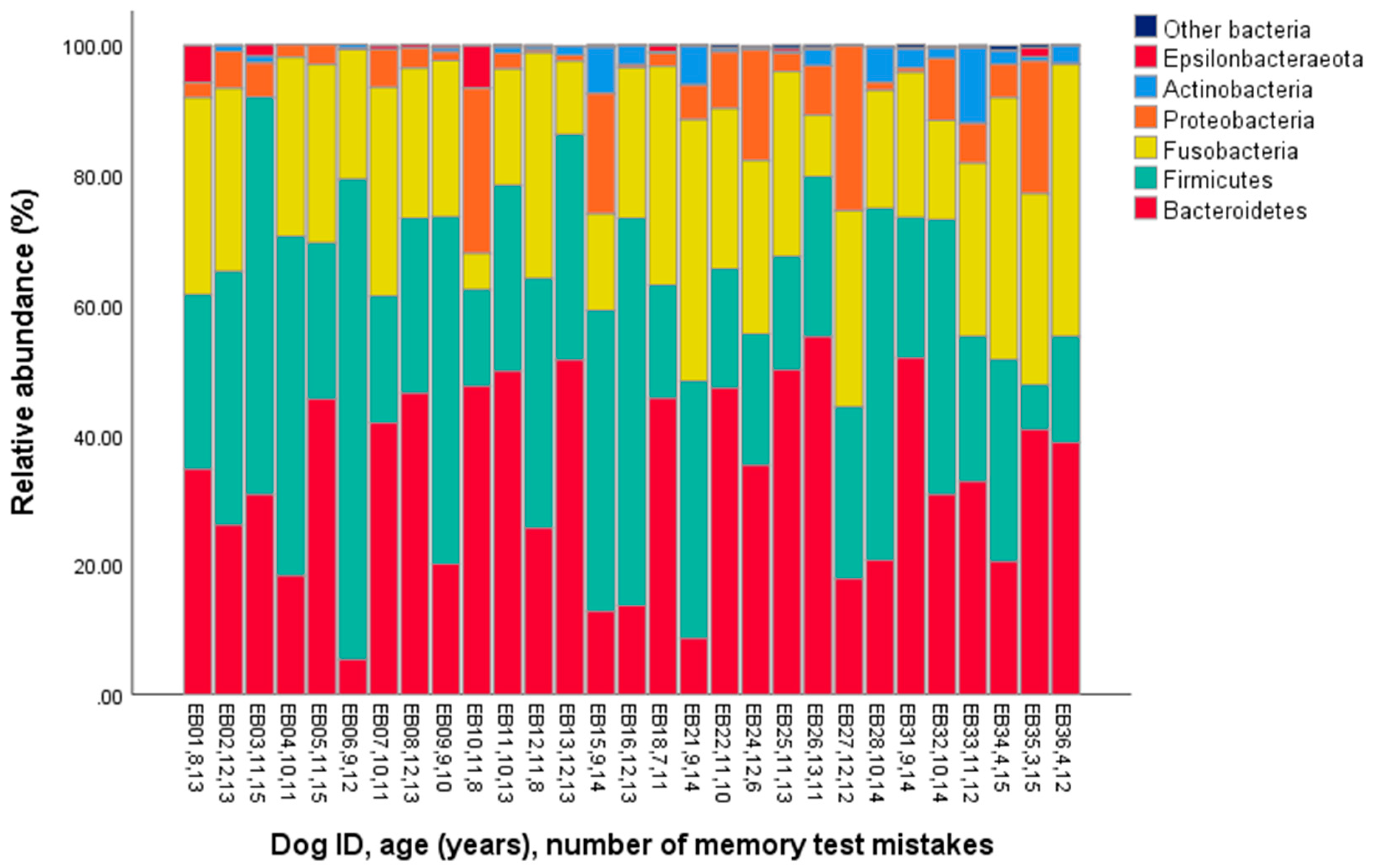

| Phylum | Mean | Standard Deviation | Minimum | Maximum |
| Actinobacteria | 1.9 | 2.7 | 0.0 | 11.7 |
| Bacteroidetes | 33.3 | 14.7 | 5.4 | 55.1 |
| Epsilonbacteraeota | 0.7 | 1.5 | 0.0 | 6.4 |
| Firmicutes | 33.0 | 16.5 | 6.8 | 73.9 |
| Fusobacteria | 24.4 | 10.1 | 0.0 | 41.9 |
| Proteobacteria | 6.5 | 7.5 | 0.1 | 25.4 |
| Other bacteria | 0.2 | 0.2 | 0.0 | 0.7 |
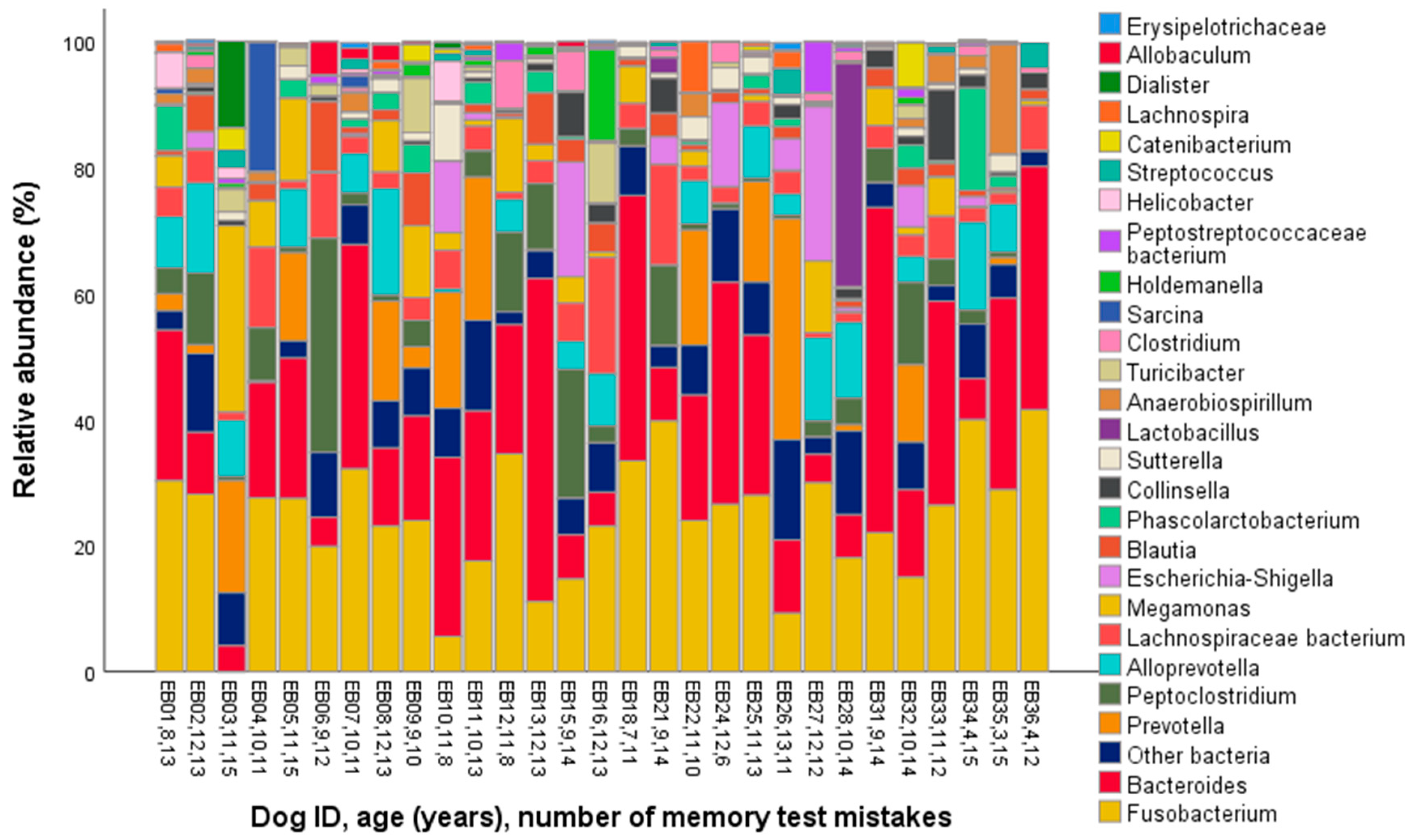
| Genus | Mean | Standard Deviation | Minimum | Maximum |
| Allobaculum | 0.5 | 1.1 | 0.0 | 5.4 |
| Alloprevotella | 5.2 | 5.3 | 0.0 | 16.9 |
| Anaerobiospirillum | 1.3 | 3.4 | 0.0 | 17.5 |
| Bacteroides | 21.1 | 14.1 | 4.1 | 51.6 |
| Blautia | 2.5 | 2.8 | 0.0 | 11.2 |
| Catenibacterium | 0.6 | 1.5 | 0.0 | 7.1 |
| Clostridium | 1.0 | 1.8 | 0.0 | 7.6 |
| Collinsella | 1.6 | 2.5 | 0.0 | 11.3 |
| Dialister | 0.5 | 2.6 | 0.0 | 13.8 |
| Erysipelotrichaceae bacterium | 0.1 | 0.3 | 0.0 | 1.2 |
| Escherichia-Shigella | 3.1 | 6.1 | 0.0 | 24.5 |
| Fusobacterium | 24.2 | 10.0 | 0.0 | 41.6 |
| Helicobacter | 0.6 | 1.6 | 0.0 | 6.4 |
| Holdemanella | 0.8 | 2.7 | 0.0 | 14.5 |
| Lachnospira | 0.5 | 1.5 | 0.0 | 8.0 |
| Lachnospiraceae bacterium | 5 | 4.4 | 0.8 | 18.5 |
| Lactobacillus | 1.3 | 6.6 | 0.0 | 35.5 |
| Megamonas | 4.6 | 6.3 | 0.0 | 29.6 |
| Peptoclostridium | 5.8 | 7.5 | 0.0 | 34.0 |
| Peptostreptococcaceae bacterium | 0.7 | 1.6 | 0.0 | 8.2 |
| Phascolarctobacterium | 1.8 | 3.3 | 0.0 | 16.3 |
| Prevotella | 6.2 | 9.4 | 0.0 | 35.1 |
| Sarcina | 0.8 | 3.8 | 0.0 | 20.5 |
| Streptococcus | 0.6 | 1.1 | 0.0 | 3.9 |
| Sutterella | 1.4 | 1.8 | 0.0 | 9.0 |
| Turicibacter | 1.1 | 2.4 | 0.0 | 9.5 |
| Other bacteria | 7.0 | 4.0 | 0.2 | 15.9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kubinyi, E.; Bel Rhali, S.; Sándor, S.; Szabó, A.; Felföldi, T. Gut Microbiome Composition is Associated with Age and Memory Performance in Pet Dogs. Animals 2020, 10, 1488. https://doi.org/10.3390/ani10091488
Kubinyi E, Bel Rhali S, Sándor S, Szabó A, Felföldi T. Gut Microbiome Composition is Associated with Age and Memory Performance in Pet Dogs. Animals. 2020; 10(9):1488. https://doi.org/10.3390/ani10091488
Chicago/Turabian StyleKubinyi, Eniko, Soufiane Bel Rhali, Sára Sándor, Attila Szabó, and Tamás Felföldi. 2020. "Gut Microbiome Composition is Associated with Age and Memory Performance in Pet Dogs" Animals 10, no. 9: 1488. https://doi.org/10.3390/ani10091488
APA StyleKubinyi, E., Bel Rhali, S., Sándor, S., Szabó, A., & Felföldi, T. (2020). Gut Microbiome Composition is Associated with Age and Memory Performance in Pet Dogs. Animals, 10(9), 1488. https://doi.org/10.3390/ani10091488






