A Newly Identified LncRNA LncIMF4 Controls Adipogenesis of Porcine Intramuscular Preadipocyte through Attenuating Autophagy to Inhibit Lipolysis
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal
2.2. Cell Culture
2.3. Cell Transfection
2.4. Hematoxylin–Eosin Staining
2.5. Fluorescence in Situ Hybridization
2.6. Cytoplasmic and Nuclear RNA Extraction
2.7. RNA Extraction and Real-Time PCR
2.8. Western Blotting
2.9. Cell Counting Kit (CCK8) and EdU Assays
2.10. Flow Cytometry
2.11. Oil Red O Staining
2.12. Monodansylcadaverine (MDC) Staining
2.13. Statistical Analysis
3. Results
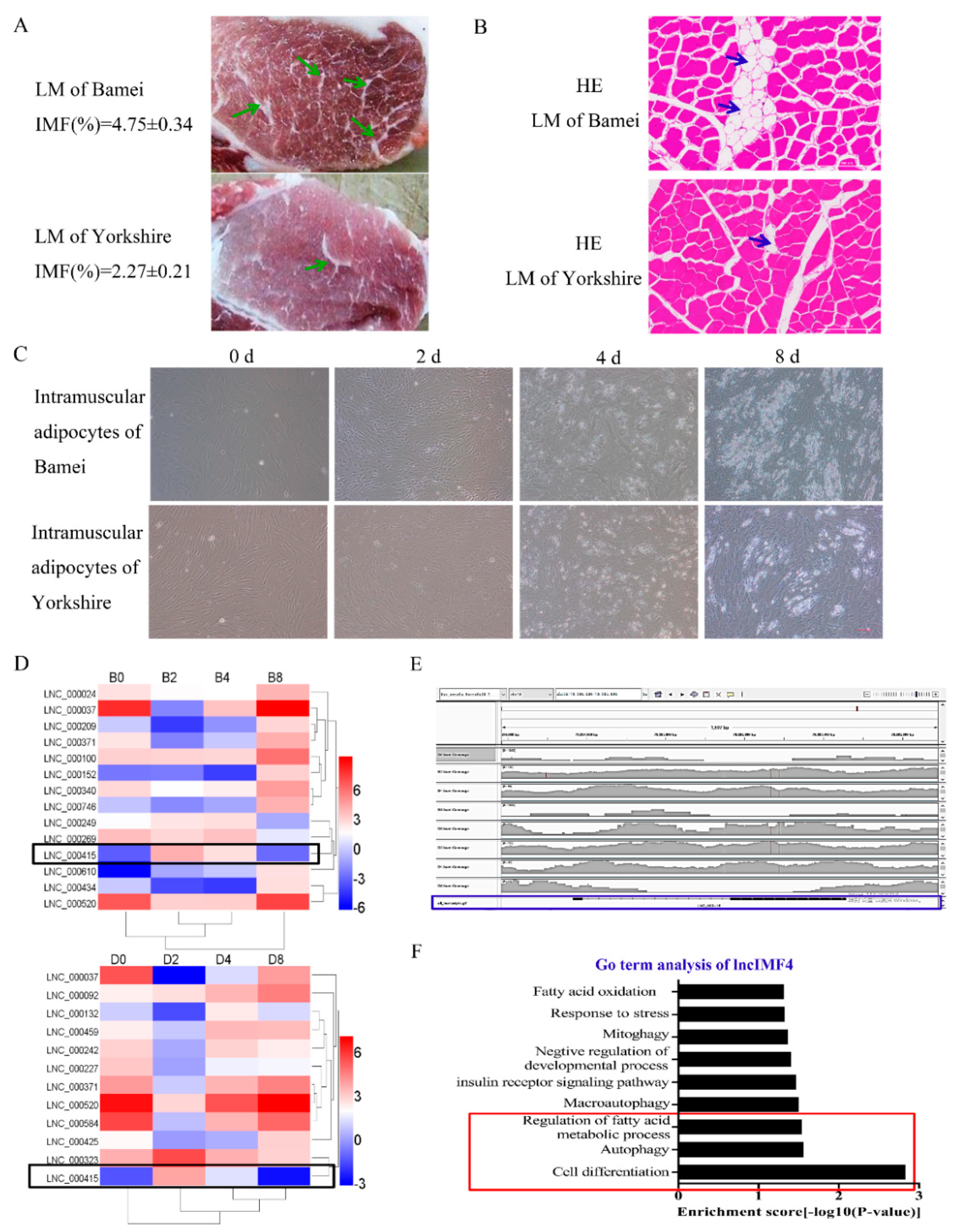
3.1. LncIMF4 May be A Novel LncRNA Implicated in Intramuscular Fat Deposition
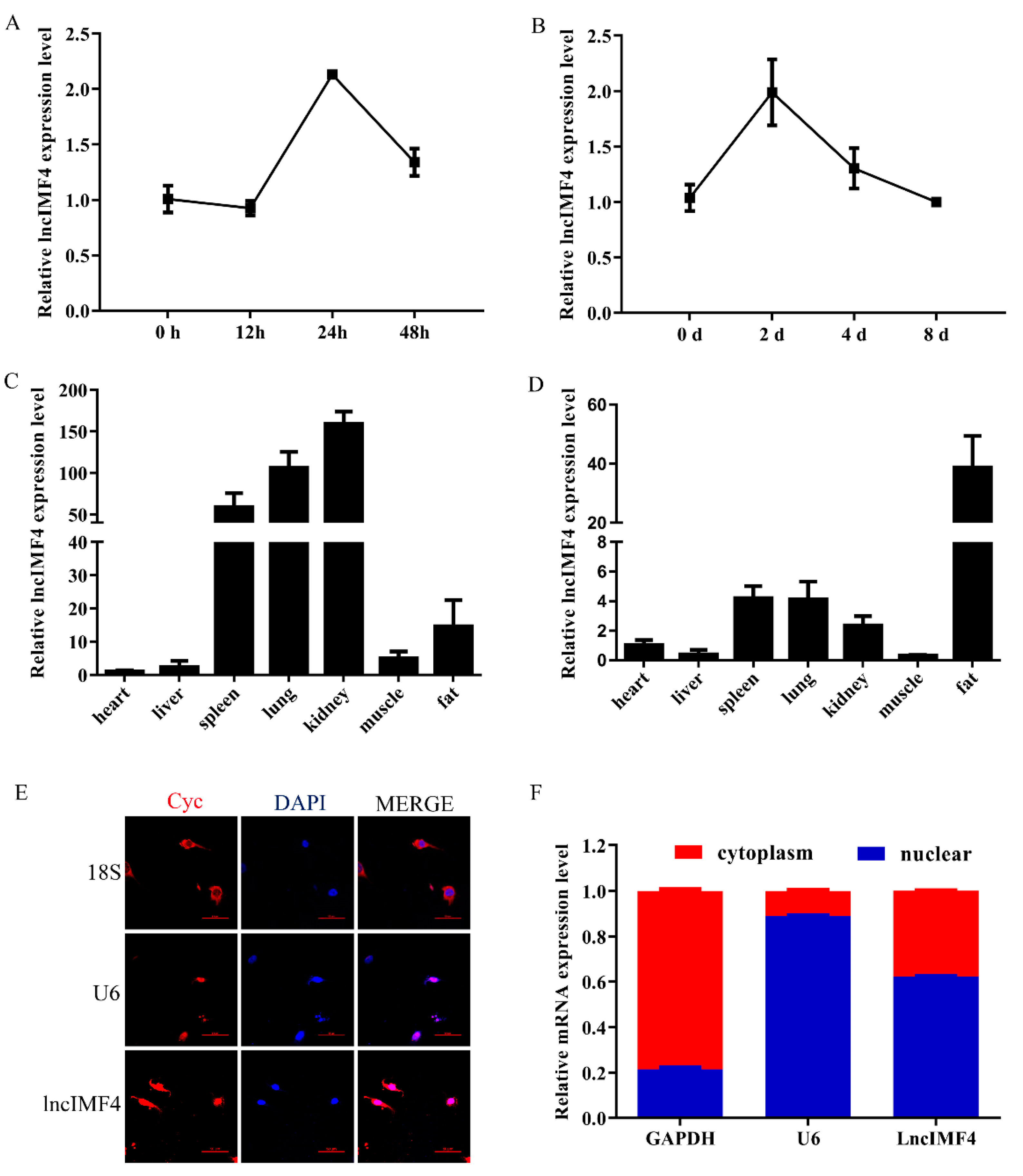
3.2. The Expression Pattern of LncIMF4 in Pig and Its Subcellular Location in Adipocyte
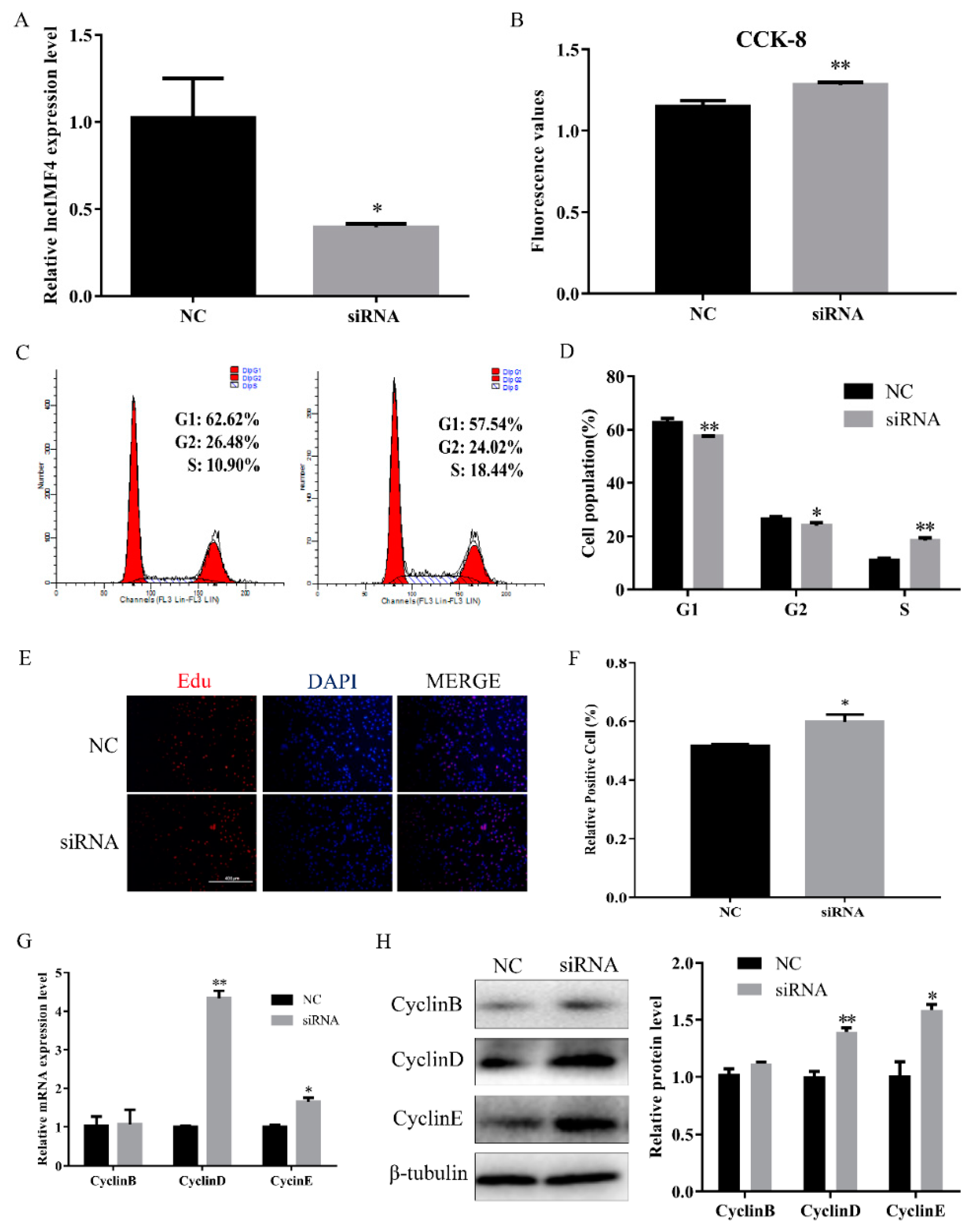
3.3. LncIMF4 Knockdown Promoted the Proliferation of Porcine Intramuscular Preadipocytes
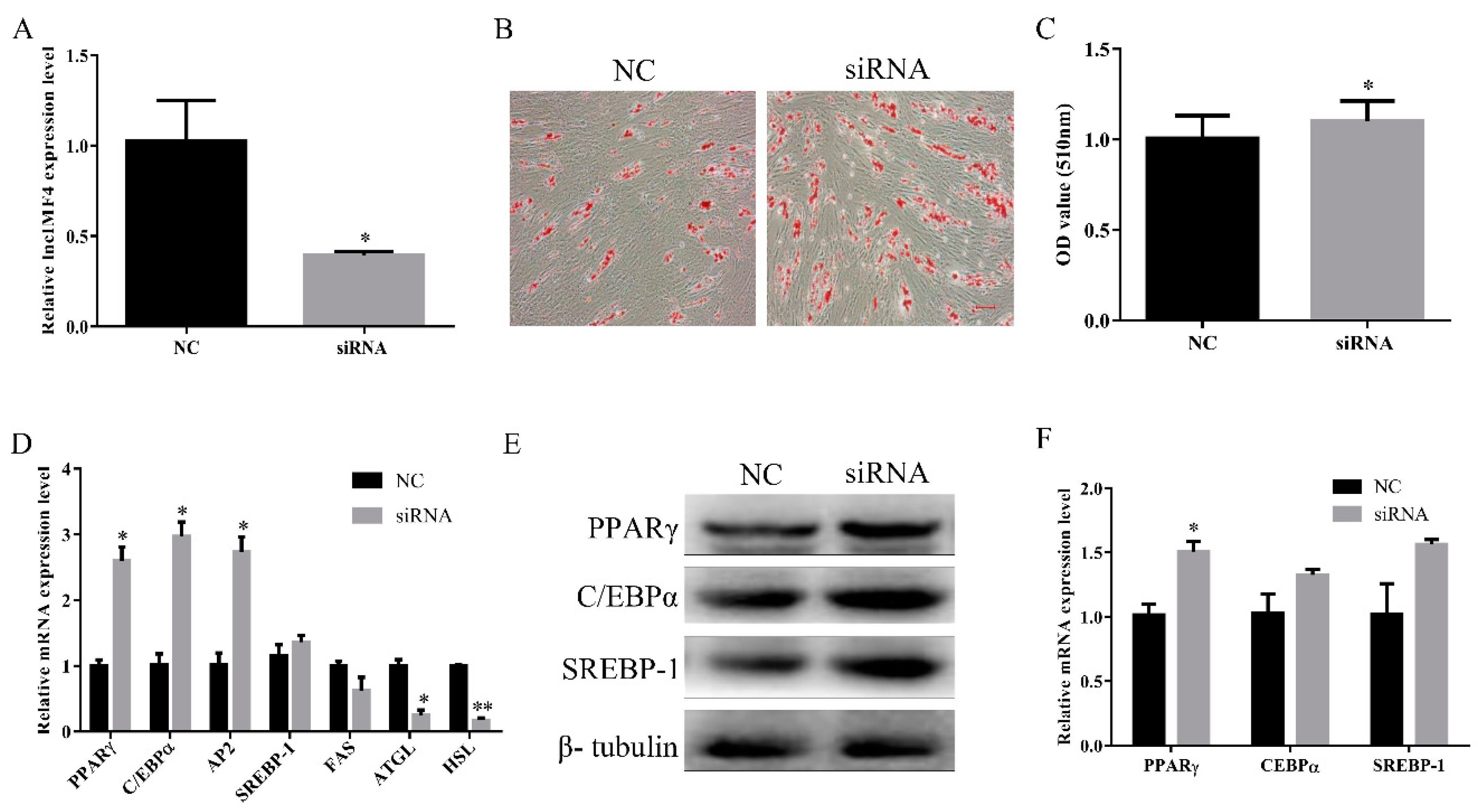
3.4. LncIMF4 Knockdown Promoted Intramuscular Adipogenic Differentiation
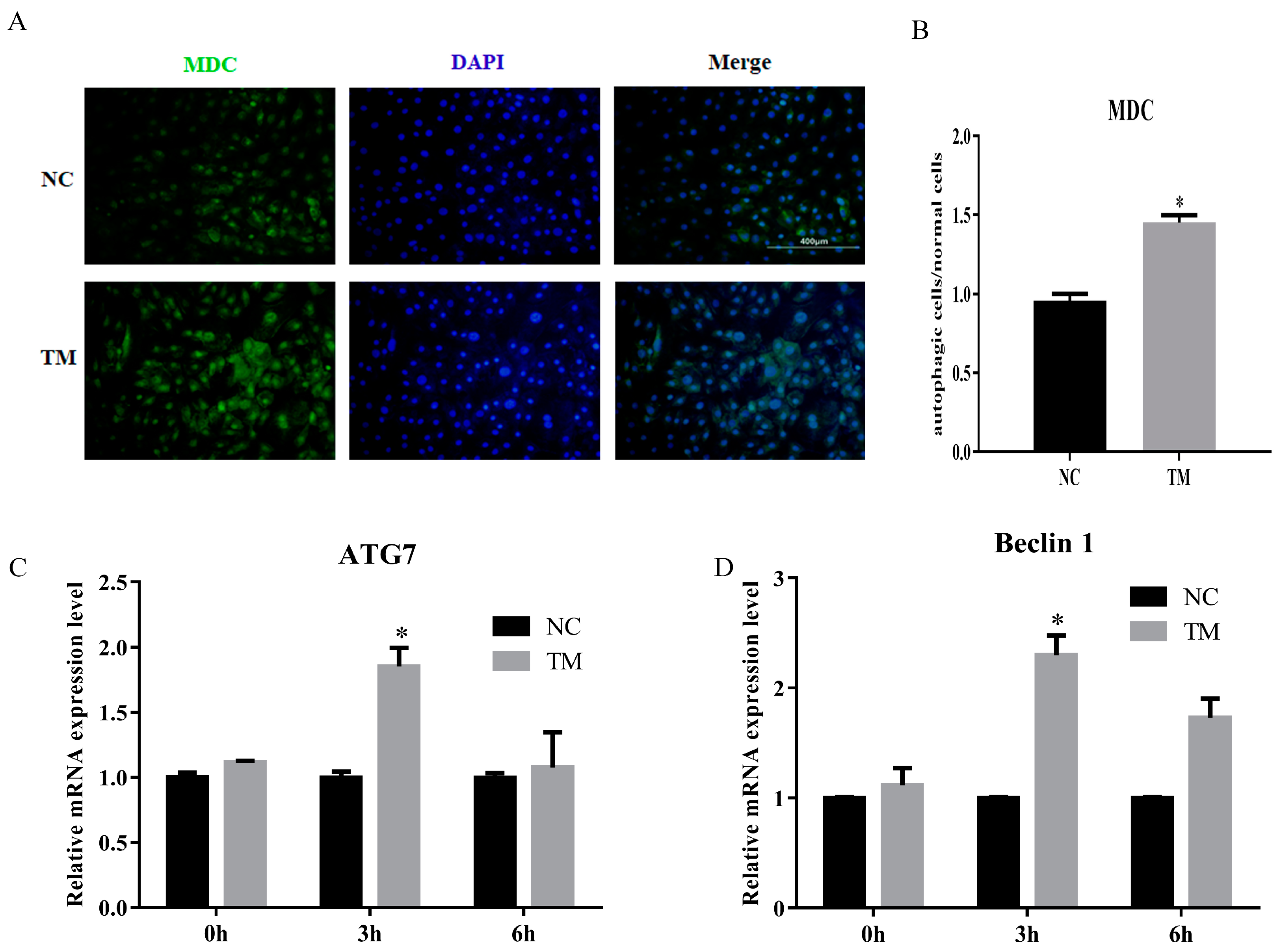
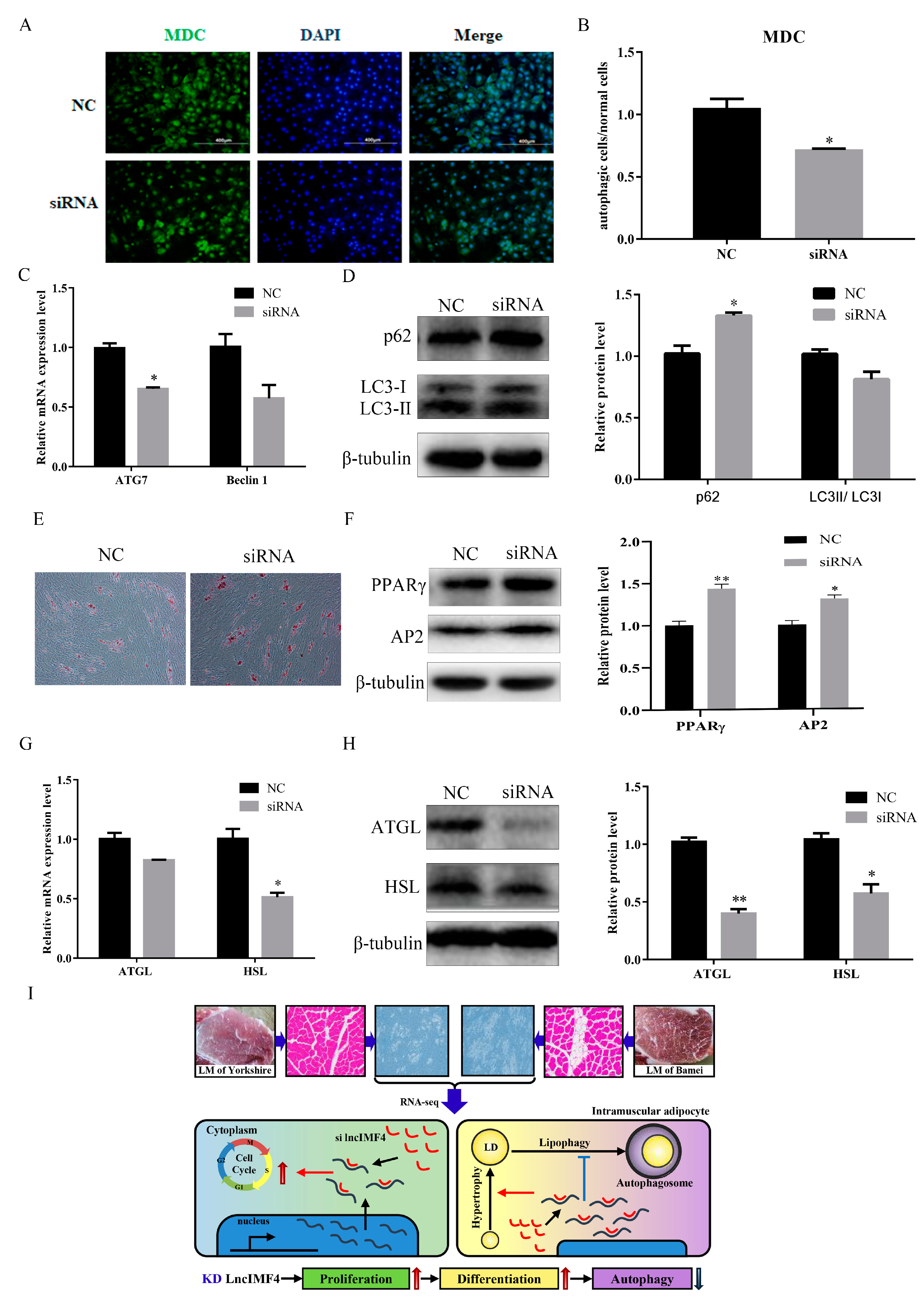
3.5. LncIMF4 Knockdown Inhibited Lipolysis by Attenuating Autophagy in Porcine Intramuscular Adipocytes
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Hocquette, J.F.; Gondret, F.; Baéza, E.; Médale, F.; Jurie, C.; Pethick, D.W. Intramuscular fat content in meat-producing animals: Development, genetic and nutritional control, and identification of putative markers. Animal 2010, 4, 303–319. [Google Scholar] [CrossRef]
- Gao, S.Z.; Zhao, S.M. Physiology, affecting factors and strategies for control of pig meat intramuscular fat. Recent. Pat. Food Nutr. Agric. 2009, 1, 59–74. [Google Scholar]
- Watkins, P.J.; Frank, D.; Singh, T.K.; Young, O.A.; Warner, R.D. Sheep meat flavor and the effect of different feeding systems: A review. J. Agric. Food Chem. 2013, 61, 3561–3579. [Google Scholar] [CrossRef]
- Mateescu, R.G.; Garrick, D.J.; Garmyn, A.J.; VanOverbeke, D.L.; Mafi, G.G.; Reecy, J.M. Genetic parameters for sensory traits in longissimus muscle and their associations with tenderness, marbling score, and intramuscular fat in Angus cattle. J. Anim. Sci. 2015, 93, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Lopes, P.A.; Costa, A.S.; Costa, P.; Pires, V.M.; Madeira, M.S.; Achega, F.; Pinto, R.M.; Prates, J.A. Contrasting cellularity on fat deposition in the subcutaneous adipose tissue and longissimus lumborum muscle from lean and fat pigs under dietary protein reduction. Animal 2014, 8, 629–637. [Google Scholar] [CrossRef]
- Cai, R.; Tang, G.R.; Zhang, Q.; Yong, W.L.; Zhang, W.R.; Xiao, J.H.; Wei, C.S.; He, C.; Yang, G.S.; Pang, W.J. A novel lnc-RNA, named lnc-ORA, is identified by RNA-Seq analysis, and its knockdown inhibits adipogenesis by regulating the PI3K/AKT/mTOR signaling pathway. Cells 2019, 8, 477. [Google Scholar] [CrossRef]
- Wei, G.H.; Wang, X. lncRNA MEG3 inhibit proliferation and metastasis of gastric cancer via p53 signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3850–3856. [Google Scholar]
- Yi, F.; Zhang, P.; Wang, Y.; Xu, Y.; Zhang, Z.X.; Ma, W.Z.; Xu, B.; Xia, Q.; Du, Q. Long non-coding RNA slincRAD functions in methylation regulation during the early stage of mouse adipogenesis. RNA Biol. 2019, 19, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Yang, W.J.; Guo, Y.J.; Chen, W.; Zheng, P.; Zeng, J.S.; Tong, W.S. Exosomal lncRNA GAS5 regulates the apoptosis of macrophages and vascular endothelial cells in atherosclerosis. PLoS ONE 2017, 12, e0185406. [Google Scholar] [CrossRef]
- Pang, W.J.; Lin, L.G.; Xiong, Y.; Wei, N.; Wang, Y.; Shen, Q.W.; Yang, G.S. Knockdown of PU.1 AS lncRNA inhibits adipogenesis through enhancing PU.1 mRNA translation. J. Cell Biochem. 2013, 114, 2500–2512. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hua, L.S.; Chen, J.F.; Zhang, J.Q.; Bai, X.X.; Gao, B.W.; Li, C.J.; Shi, Z.H.; Sheng, W.D.; Gao, Y.; et al. Identification and characterization of long non-coding RNAs in subcutaneous adipose tissue from castrated and intact full-sib pair Huainan male pigs. BMC Genom. 2017, 18, 542. [Google Scholar] [CrossRef] [PubMed]
- Wei, N.; Wang, Y.; Xu, R.X.; Wang, G.Q.; Xiong, Y.; Yu, T.Y.; Yang, G.S.; Pang, W.J. PU.1 antisense lncRNA against its mRNA translation promotes adipogenesis in porcine preadipocytes. Anim. Genet. 2015, 46, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Miao, Z.G.; Wang, S.; Zhang, J.Z.; Wei, P.P.; Guo, L.P.; Liu, D.Y.; Wang, Y.M.; Shi, M.Y. Identification and comparison of long non-conding RNA in Jinhua and Landrace pigs. Biochem. Biophys. Res. Commun. 2018, 506, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Panigrahi, D.P.; Patil, S.; Bhutia, S.K. Autophagy in health and disease: A comprehensive review. Biomed. Pharmacother 2018, 104, 485–495. [Google Scholar] [CrossRef]
- Kaur, J.; Debnath, J. Autophagy at the crossroads of catabolism and anabolism. Nat. Rev. Mol. Cell Biol. 2015, 16, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Carroll, B.; Korolchuk, V.I.; Sarkar, S. Amino acids and autophagy: Cross-talk and co-operation to control cellular homeostasis. Amino Acids 2015, 47, 2065–2088. [Google Scholar] [CrossRef]
- Ha, J.; Guan, K.L.; Kim, J. AMPK and autophagy in glucose/glycogen metabolism. Mol. Aspects Med. 2015, 46, 46–62. [Google Scholar] [CrossRef]
- Karsli-Uzunbas, G.; Guo, J.Y.; Price, S.; Teng, X.; Laddha, S.V.; Khor, S.; Kalaany, N.Y.; Jacks, T.; Chan, C.S.; Rabinowitz, J.D.; et al. Autophagy is required for glucose homeostasis and lung tumor maintenance. Cancer Discov. 2014, 4, 914–927. [Google Scholar] [CrossRef]
- Sun, Y.M.; Chen, X.C.; Qin, J.; Liu, S.G.; Zhao, R.; Yu, T.Y.; Chu, G.Y.; Yang, G.S.; Pang, W.J. Comparative analysis of long noncoding RNAs expressed during intramuscular adipocytes adipogenesis in fat-type and lean-type pigs. J. Agric. Food Chem. 2018, 66, 12122–12130. [Google Scholar] [CrossRef]
- Xiong, Y.; Yue, F.; Jia, Z.H.; Gao, Y.; Jin, W.; Hu, K.P.; Zhang, Y.; Zhu, D.H.; Yang, G.S.; Kuang, S.H. A novel brown adipocyte-enriched long non-coding RNA that is required for brown adipocyte differentiation and sufficient to drive thermogenic gene program in white adipocytes. Biochim. Biophys. Acta. Mol. Cell Biol. Lipids 2018, 1863, 409–419. [Google Scholar] [CrossRef]
- Mo, D.L.; Yu, K.F.; Chen, H.; Chen, L.X.; Liu, X.H.; He, Z.Y.; Cong, P.Q.; Chen, Y.S. Transcriptome landscape of porcine intramuscular adipocytes during differentiation. J. Agric. Food Chem. 2017, 65, 6317–6328. [Google Scholar] [CrossRef] [PubMed]
- Sarantopoulos, C.N.; Banyard, D.A.; Ziegler, M.E.; Sun, B.; Shaterian, A.; Widgerow, A.D. Elucidating the preadipocyte and its role in adipocyte formation: A Comprehensive Review. Stem Cell Rev. Rep. 2018, 14, 27–42. [Google Scholar] [CrossRef] [PubMed]
- Green, C.R.; Wallace, M.; Divakaruni, A.S.; Phillips, S.A.; Murphy, A.N.; Ciaraldi, T.P.; Metallo, C.M. Branched-chain amino acid catabolism fuels adipocyte differentiation and lipogenesis. Nat. Chem. Biol. 2016, 12, 15–21. [Google Scholar] [CrossRef]
- Khaldoun, S.A.; Emond-Boisjoly, M.A.; Chateau, D.; Carrière, V.; Lacasa, M.; Rousset, M.; Demignot, S.; Morel, E. Autophagosomes contribute to intracellular lipid distribution in enterocytes. Mol. Biol. Cell 2014, 25, 118–132. [Google Scholar] [CrossRef]
- Sun, L.; Lin, J.D. Function and mechanism of long noncoding RNAs in adipocyte biology. Diabetes 2019, 68, 887–896. [Google Scholar] [CrossRef]
- Galic, S.; Oakhill, J.S.; Steinberg, G.R. Adipose tissue as an endocrine organ. Mol. Cell Endocrinol. 2010, 316, 129–139. [Google Scholar] [CrossRef]
- Chen, S.Z.; Chen, Y.; Qian, Q.F.; Wang, X.W.; Chang, Y.T.; Ju, S.H.; Xu, Y.D.; Zhang, C.; Qin, N.; Ding, H.; et al. Gene amplification derived a cancer-testis long noncoding RNA PCAT6 regulates cell proliferation and migration in hepatocellular carcinoma. Cancer Med. 2019, 8, 3017–3025. [Google Scholar] [CrossRef]
- Tong, Y.X.; Wang, M.S.; Dai, Y.N.; Bao, D.J.; Zhang, J.J.; Pan, H.Y. LncRNA HOXA-AS3 sponges miR-29c to facilitate cell proliferation, metastasis, and EMT Process and activate the MEK/ERK signaling pathway in hepatocellular carcinoma. Hum. Gene. Ther. Clin. Dev. 2019, 30, 129–141. [Google Scholar] [CrossRef]
- Nuermaimaiti, N.; Liu, J.; Liang, X.D.; Jiao, Y.; Zhang, D.; Liu, L.; Meng, X.Y.; Guan, Y.Q. Effect of lncRNA HOXA11-AS1 on adipocyte differentiation in human adipose-derived stem cells. Biochem. Biophys. Res. Commun. 2018, 495, 1878–1884. [Google Scholar] [CrossRef]
- Xiao, T.F.; Liu, L.H.; Li, H.L.; Sun, Y.; Luo, H.X.; Li, T.P.; Wang, S.H.; Dalton, S.; Zhao, R.C.; Chen, R.S. Long noncoding RNA ADINR regulates adipogenesis by transcriptionally activating C/EBPalpha. Stem Cell Reports 2015, 5, 856–865. [Google Scholar] [CrossRef]
- Zhu, E.D.; Zhang, J.J.; Li, Y.C.; Yuan, H.R.; Zhou, J.; Wang, B.L. Long noncoding RNA Plnc1 controls adipocyte differentiation by regulating peroxisome proliferator-activated receptor gamma. FASEB J. 2019, 33, 2396–2408. [Google Scholar] [CrossRef] [PubMed]
- Levine, B.; Kroemer, G. Autophagy in the pathogenesis of disease. Cell 2008, 132, 27–42. [Google Scholar] [CrossRef] [PubMed]
- Koga, H.; Kaushik, S.; Cuervo, A.M. Altered lipid content inhibits autophagic vesicular fusion. FASEB J. 2010, 24, 3052–3065. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Li, Z.; Zhang, Q.; Wei, S.; Li, B.W.; Zhang, X.; Zhang, L.; Li, Q.; Xu, H.; Xu, Z.K. Silencing of AQP3 induces apoptosis of gastric cancer cells via downregulation of glycerol intake and downstream inhibition of lipogenesis and autophagy. Onco. Targets Ther. 2017, 10, 2791–2804. [Google Scholar] [CrossRef] [PubMed]
- Juarez-Rojas, J.G.; Reyes-Soffer, G.; Conlon, D.; Ginsberg, H.N. Autophagy and cardiometabolic risk factors. Rev. Endocr. Metab. Disord. 2014, 15, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Desjardins, E.M.; Steinberg, G.R. Emerging role of AMPK in brown and beige adipose tissue (BAT): Implications for obesity, insulin resistance, and type 2 diabetes. Curr. Diab. Rep. 2018, 18, 80. [Google Scholar] [CrossRef] [PubMed]
| Name | Forward (5′→3′) | Reverse (5′→3′) |
|---|---|---|
| PPARγ | AGGACTACCAAAGTGCCATCAAA | GAGGCTTTATCCCCACAGACAC |
| AP2 | GAGCACCATAACCTTAGATGGA | AAATTCTGGTAGCCGTGACA |
| C/EBPα | CGATGCTCTTAGCTGAGTGT | GGTCCAAGAATTTCACCTCT |
| SREBP1 | GGAGCCATGGATTGCACATT | GGCCCGGGAAGTCACTGT |
| Cyclin B | AATCCCTTCTTGTGGTTA | CTTAGATGTGGCATACTTG |
| MyoD | TACACCGACAACTCCATCCG | GAGGGCGGGTTGGAAATGAA |
| Cyclin E | CAGAGCAGCGAGCAGGAGC | GCAAGCTGCTTCCACACCACAT |
| FAS | CCCCGAATCTGCACTACCAC | AGTTGGGCTGAAGGATGACG |
| ATGL | TCACCAACACCAGCATCCA | GCACATCTCTCGAAGCACCA |
| HSL | CACTGACTGCTGACCCCAAG | TCCTCACTGTCCTGTCCTTCAC |
| ATG7 | GATTGCCTGGTGGGTGGTAA | CATGGCTTTCGATGAGCTGC |
| Beclin1 | AGTAGGTGAAGGCTAGGCGA | AGCTCGTGTCCAGTTTCAGG |
| GAPDH LncIMF4 | AGGTCGGAGTGAACGGATTTG GTGGATTGGGAGCCTGCTAT | ACCATGTAGTGGAGGTCAATGAAG ACACTCCATGGCCTGGTAAAA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, Y.; Cai, R.; Wang, Y.; Zhao, R.; Qin, J.; Pang, W. A Newly Identified LncRNA LncIMF4 Controls Adipogenesis of Porcine Intramuscular Preadipocyte through Attenuating Autophagy to Inhibit Lipolysis. Animals 2020, 10, 926. https://doi.org/10.3390/ani10060926
Sun Y, Cai R, Wang Y, Zhao R, Qin J, Pang W. A Newly Identified LncRNA LncIMF4 Controls Adipogenesis of Porcine Intramuscular Preadipocyte through Attenuating Autophagy to Inhibit Lipolysis. Animals. 2020; 10(6):926. https://doi.org/10.3390/ani10060926
Chicago/Turabian StyleSun, Yunmei, Rui Cai, Yingqian Wang, Rui Zhao, Jin Qin, and Weijun Pang. 2020. "A Newly Identified LncRNA LncIMF4 Controls Adipogenesis of Porcine Intramuscular Preadipocyte through Attenuating Autophagy to Inhibit Lipolysis" Animals 10, no. 6: 926. https://doi.org/10.3390/ani10060926
APA StyleSun, Y., Cai, R., Wang, Y., Zhao, R., Qin, J., & Pang, W. (2020). A Newly Identified LncRNA LncIMF4 Controls Adipogenesis of Porcine Intramuscular Preadipocyte through Attenuating Autophagy to Inhibit Lipolysis. Animals, 10(6), 926. https://doi.org/10.3390/ani10060926






