Detection of Pseudorabies Virus in Wild Boar Foetus
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Serologic Analysis
2.3. Molecular Analysis
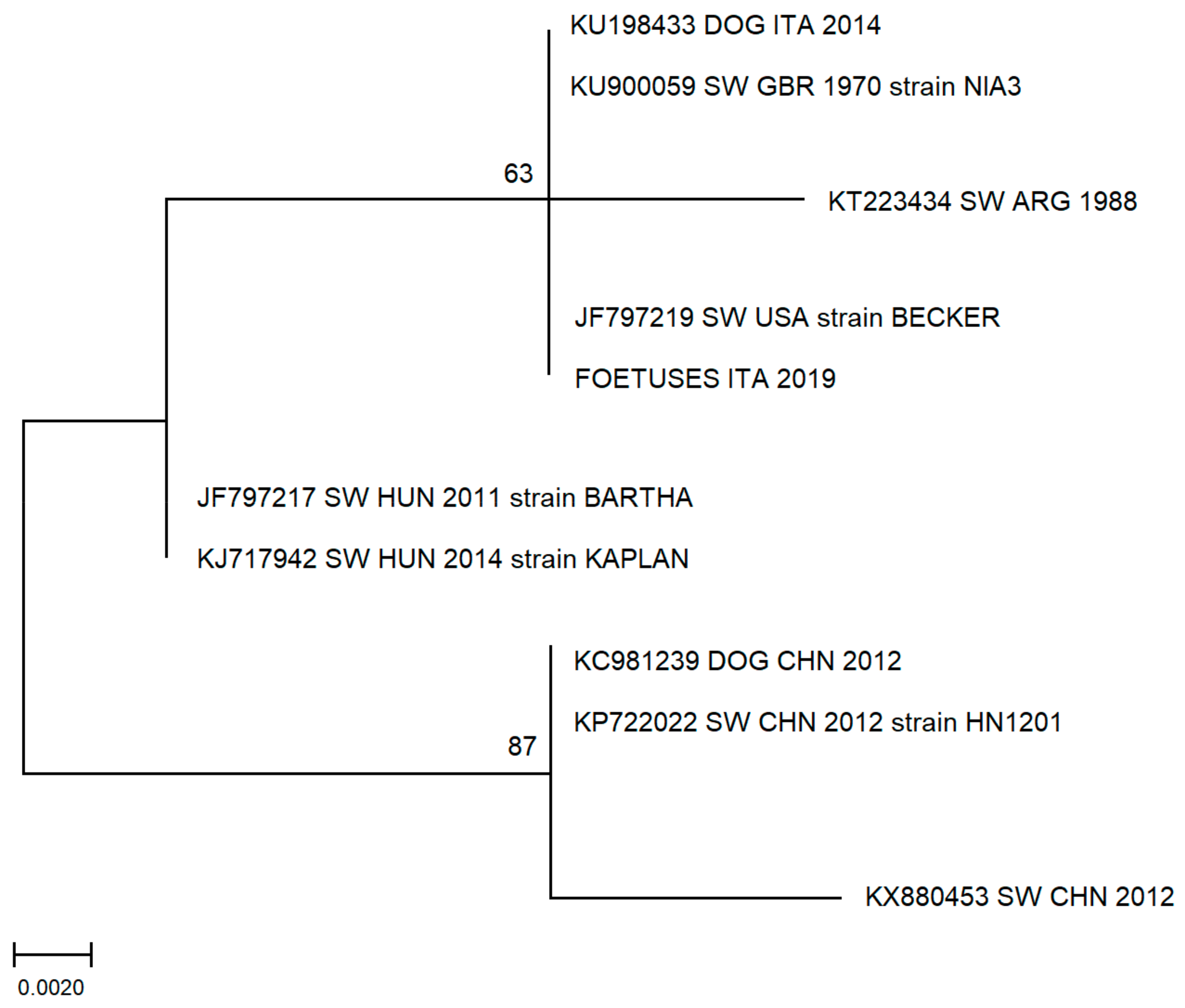
2.4. Phylogenetic Analysis
2.5. Statistical Analyses
3. Results
3.1. Serologic Analysis
3.2. Molecular Analysis
3.3. Phylogenetic Analysis
3.4. Statistical Analyses
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Aujeszky, A. Ueber eine neue Infektionskrankheit bei Haustieren. Cent. für Bakteriol. Parasitenkd. und Infekt. 1902, 32, 353–357. [Google Scholar]
- Müller, T.; Hahn, E.C.; Tottewitz, F.; Kramer, M.; Klupp, B.G.; Mettenleiter, T.C.; Freuling, C. Pseudorabies virus in wild swine: A global perspective. Arch. Virol. 2011, 156, 1691–1705. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Fons, F.; Vidal, D.; Höfle, U.; Vicente, J.; Gortázar, C. Aujeszky’s disease virus infection patterns in European wild boar. Vet. Microbiol. 2007, 120, 241–250. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.Q.; Adams, M.J.; Carstens, E.B.; Lefkowitz, E.J. Herpesvirales. In Virus Taxonomy; Elsevier: Amsterdam, The Netherlands, 2012; pp. 99–107. [Google Scholar]
- Romero, C.H.; Meade, P.N.; Shultz, J.E.; Chung, H.Y.; Gibbs, E.P.; Hahn, E.C.; Lollis, G. Venereal transmission of pseudorabies viruses indigenous to feral swine. J. Wildl. Dis. 2001, 37, 289–296. [Google Scholar] [CrossRef]
- Verin, R.; Varuzza, P.; Mazzei, M.; Poli, A. Serologic, molecular, and pathologic survey of pseudorabies virus infection in hunted wild boars (Sus scrofa) in Italy. J. Wildl. Dis. 2014, 50, 559–565. [Google Scholar] [CrossRef]
- González-Barrio, D.; Martín-Hernando, M.P.; Ruiz-Fons, F. Shedding patterns of endemic Eurasian wild boar (Sus scrofa) pathogens. Res. Vet. Sci. 2015, 102, 206–211. [Google Scholar] [CrossRef]
- Hahn, E.C.; Page, G.R.; Hahn, P.S.; Gillis, K.D.; Romero, C.; Annelli, J.A.; Gibbs, E.P.J. Mechanisms of transmission of Aujeszky’s disease virus originating from feral swine in the United States. Vet. Microbiol. 1997, 55, 123–130. [Google Scholar] [CrossRef]
- Hernández, F.A.; Sayler, K.A.; Bounds, C.; Milleson, M.P.; Carr, A.N.; Wisely, S.M. Evidence of pseudorabies virus shedding in feral swine (Sus scrofa) populations of Florida, USA. J. Wildl. Dis. 2018, 54, 45–53. [Google Scholar] [CrossRef]
- Moreno, A.; Sozzi, E.; Grilli, G.; Gibelli, L.R.; Gelmetti, D.; Lelli, D.; Chiari, M.; Prati, P.; Alborali, G.L.; Boniotti, M.B.; et al. Detection and molecular analysis of Pseudorabies virus strains isolated from dogs and a wild boar in Italy. Vet. Microbiol. 2015, 177, 359–365. [Google Scholar] [CrossRef]
- Pomeranz, L.E.; Reynolds, A.E.; Hengartner, C.J. Molecular Biology of Pseudorabies Virus: Impact on Neurovirology and Veterinary Medicine. Microbiol. Mol. Biol. Rev. 2005, 69, 462–500. [Google Scholar] [CrossRef]
- Macdiarmid, S.C. Aujeszky’s disease eradication in New Zealand. Aust. Vet. J. 2000, 78, 470–471. [Google Scholar] [CrossRef] [PubMed]
- Hahn, E.C.; Fadl-Alla, B.; Lichtensteiger, C.A. Variation of Aujeszky’s disease viruses in wild swine in USA. Vet. Microbiol. 2010, 143, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Adriana, A.; Porea, D.; Aniţă, D.; Savuţa, G. SEROLOGICAL STUDY OF SELECTED VIRAL PATHOGENS IN WILD BOAR FROM EASTERN ROMANIA. Development of Translational Research Capacity: Development of Vaccines from Concept to Preclinical Evaluation View Project REGIONAL CENTER OF ADVANCED RESEARCH FOR EMERGING D; Lucrări ştiinţifice: Cluj, Romania, 2015. [Google Scholar]
- Caruso, C.; Vitale, N.; Prato, R.; Radaelli, M.C.; Zoppi, S.; Possidente, R.; Dondo, A.; Chiavacci, L.; Maria, A.; Martin, M.; et al. Pseudorabies virus in North-West Italian wild boar (Sus scrofa) populations: Prevalence and risk factors to support a territorial risk-based surveillance. Vet. Ital. 2018, 54, 337–341. [Google Scholar] [PubMed]
- Vicente, J.; León-Vizcaíno, L.; Gortázar, C.; Cubero, M.J.; González, M.; Martín-Atance, P. Antibodies to selected viral and bacterial pathogens in European wild boars from southcentral Spain. J. Wildl. Dis. 2002, 38, 649–652. [Google Scholar] [CrossRef] [PubMed]
- Vicente, J.; Ruiz-Fons, F.; Vidal, D.; Höfle, U.; Acevedo, P.; Villanúa, D.; Fernández-de-Mera, I.G.; Martín, M.P.; Gortázar, C. Serosurvey of Aujeszky’s disease virus infection in European wild boar in Spain. Vet. Rec. 2005, 156, 408–412. [Google Scholar] [CrossRef]
- Vengust, G.; Valencak, Z.; Bidovec, A. Presence of antibodies against Aujeszky’s disease virus in wild boar (Sus scrofa) in Slovenia. J. Wildl. Dis. 2005, 41, 800–802. [Google Scholar] [CrossRef]
- Sedlak, K.; Bartova, E.; Machova, J. Antibodies to Selected Viral Disease Agents in Wild Boars from the Czech Republic. J. Wildl. Dis. 2008, 44, 777–780. [Google Scholar] [CrossRef]
- Müller, T.; Klupp, B.G.; Freuling, C.; Hoffmann, B.; Mojcicz, M.; Capua, I.; Palfi, V.; Toma, B.; Lutz, W.; Ruiz-Fon, F.; et al. Characterization of pseudorabies virus of wild boar origin from Europe. Epidemiol. Infect. 2010, 138, 1590–1600. [Google Scholar] [CrossRef]
- Pannwitz, G.; Freuling, C.; Denzin, N.; Schaarschmidt, U.; Nieper, H.; Hlinak, A.; Burkhardt, S.; Klopries, M.; Dedek, J.; Hoffmann, L.; et al. A long-term serological survey on Aujeszky’s disease virus infections in wild boar in East Germany. Epidemiol. Infect. 2012, 140, 348–353. [Google Scholar] [CrossRef]
- Cano-Manuel, F.J.; López-Olvera, J.; Fandos, P.; Soriguer, R.C.; Pérez, J.M.; Granados, J.E. Long-term monitoring of 10 selected pathogens in wild boar (Sus scrofa) in Sierra Nevada National Park, southern Spain. Vet. Microbiol. 2014, 174, 148–154. [Google Scholar] [CrossRef]
- Herrmann, S.-C.; Heppner, B.; Ludwig, H. Pseudorabies Viruses from Clinical Outbreaks and Latent Infections Grouped into Four Major Genome Types. In Latent Herpes Virus Infections in Veterinary Medicine; Springer: Amsterdam, The Netherlands, 1984; pp. 387–401. [Google Scholar]
- Lari, A.; Lorenzi, D.; Nigrelli, D.; Brocchi, E.; Faccini, S.; Poli, A. Pseudorabies virus in European wild boar from Central Italy. J. Wildl. Dis. 2006, 42, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Spradbrow, P.B. Veterinary virology. Vet. Microbiol. 1990, 21, 380–381. [Google Scholar] [CrossRef]
- Craig, R.A. Diseases of Swine; Wiley-Blackwell: Hoboken, NJ, USA, 2011. [Google Scholar]
- Wittmann, G. Aujeszky’s Disease (Pseudorabies) in Pigs; Springer: Berlin, Germany, 1989. [Google Scholar]
- Yu, X.; Zhou, Z.; Hu, D.; Zhang, Q.; Han, T.; Li, X.; Gu, X.; Yuan, L.; Zhang, S.; Wang, B.; et al. Pathogenic pseudorabies virus, China, 2012. Emerg. Infect. Dis. 2014, 20, 102–104. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, K.V.; Burriel, A.R.; Filioussis, G.; Psychas, V.; Nauwynck, H.; Kritas, S. Aujeszky’s Disease (Pseudorabies). An old threat in current pig industry? Part I. Pathogenetic information and implications. J. Hell. Vet. Med. Soc. 2017, 62, 29. [Google Scholar] [CrossRef]
- Hsu, F.S.; Chu, R.M.; Lee, R.C.T.; Chu, S.H.J. Placental lesions caused by pseudorabies virus in pregnant sows. J. Am. Vet. Med. Assoc. 1980, 177, 636–641. [Google Scholar]
- Carnevali, L.; Pedrotti, L.; Riga, F.; Toso, S. Banca Dati Ungulati: Status, distribuzione, consistenza, gestione e prelievo venatorio delle popolazioni di Ungulati in Italia. Rapporto 2001–2005. Biol. Cons. Fauna 2009, 117, 77–92. [Google Scholar]
- Arenas-Montes, A.; García-Bocanegra, I.; Paniagua, J.; Franco, J.J.; Miró, F.; Fernández-Morente, M.; Carbonero, A.; Arenas, A. Blood sampling by puncture in the cavernous sinus from hunted wild boar. Eur. J. Wildl. Res. 2013, 59, 299–303. [Google Scholar] [CrossRef]
- Yoon, H.A.; Seong, K.E.; Aleyas, A.G.; Seong, O.P.; Lee, J.H.; Joon, S.C.; Jeong, G.C.; Song, H.J. Molecular survey of latent pseudorabies virus infection in nervous tissues of slaughtered pigs by nested and real-time PCR. J. Microbiol. 2005, 43, 430–436. [Google Scholar]
- Mettenleiter, T.C.; Spear, P.G. Glycoprotein gB (gII) of pseudorabies virus can functionally substitute for glycoprotein gB in herpes simplex virus type 1. J. Virol. 1994, 68, 500–504. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Pizzurro, F.; Mangone, I.; Zaccaria, G.; De Luca, E.; Malatesta, D.; Innocenti, M.; Carmine, I.; Cito, F.; Marcacci, M.; Di Sabatino, D.; et al. Whole-genome sequence of a suid herpesvirus-1 strain isolated from the brain of a hunting dog in Italy. Genome Announc. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Montagnaro, S.; Sasso, S.; De Martino, L.; Longo, M.; Lovane, V.; Ghlurmino, G.; Plsanelli, G.; Nava, D.; Baldl, L.; Pagninl, U. Prevalence of antibodies to selected viral and bacterial pathogens in wild boar (Sus scrofa) in Campania Region, Italy. J. Wildl. Dis. 2010, 46, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Ercolini, C.; Ferrari, A.; Fisichella, S.; Guerci Lena, P.; Mandola, M.L.; Masoero, L.; Mignone, W.; Perruchon, M.; Poggi, M. Serological survey of Wild boar (Sus scrofa) in Liguria, Italy. J. Mt. Ecol. 2014, 3. [Google Scholar]
- Chiari, M.; Ferrari, N.; Bertoletti, M.; Avisani, D.; Cerioli, M.; Zanoni, M.; Alborali, L.G.; Lanfranchi, P.; Lelli, D.; Martin, A.M.; et al. Long-Term Surveillance of Aujeszky’s Disease in the Alpine Wild Boar (Sus scrofa). Ecohealth 2015, 12, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Capua, I.; Fico, R.; Banks, M.; Tamba, M.; Calzetta, G. Isolation and characterisation of an Aujeszky’s disease virus naturally infecting a wild boar (Sus scrofa). In Proceedings of the Veterinary Microbiology; Elsevier: Amsterdam, The Netherlands, 1997; Volume 55, pp. 141–146. [Google Scholar]
| SEROPOSITIVES | SERONEGATIVES | |||
|---|---|---|---|---|
| MALES | 14/20 (70%) | 6/20 (30%) | ||
| FEMALES | 26/34 (77%) | 8/34 (23%) | ||
| PREGNANT 19/24 (79%) | NO PREGNANT 7/10 (70%) | PREGNANT 5/24 (21%) | NO PREGNANT 3/10 (30%) | |
| TOT | 40/54 (74%) | 14/54 (26%) | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pacini, M.I.; Forzan, M.; Cilia, G.; Bernardini, L.; Marzoli, F.; Pedonese, F.; Bandecchi, P.; Fratini, F.; Mazzei, M. Detection of Pseudorabies Virus in Wild Boar Foetus. Animals 2020, 10, 366. https://doi.org/10.3390/ani10020366
Pacini MI, Forzan M, Cilia G, Bernardini L, Marzoli F, Pedonese F, Bandecchi P, Fratini F, Mazzei M. Detection of Pseudorabies Virus in Wild Boar Foetus. Animals. 2020; 10(2):366. https://doi.org/10.3390/ani10020366
Chicago/Turabian StylePacini, Maria Irene, Mario Forzan, Giovanni Cilia, Lucrezia Bernardini, Filippo Marzoli, Francesca Pedonese, Patrizia Bandecchi, Filippo Fratini, and Maurizio Mazzei. 2020. "Detection of Pseudorabies Virus in Wild Boar Foetus" Animals 10, no. 2: 366. https://doi.org/10.3390/ani10020366
APA StylePacini, M. I., Forzan, M., Cilia, G., Bernardini, L., Marzoli, F., Pedonese, F., Bandecchi, P., Fratini, F., & Mazzei, M. (2020). Detection of Pseudorabies Virus in Wild Boar Foetus. Animals, 10(2), 366. https://doi.org/10.3390/ani10020366







