The Only Chemoreceptor Encoded by che Operon Affects the Chemotactic Response of Agrobacterium to Various Chemoeffectors
Abstract
1. Introduction
2. Materials and Methods
2.1. Primers, Plasmids, Bacterial Strains and Growth Conditions
2.2. DNA Manipulations
2.3. Mutagenesis and Complementation of Atu0514
2.4. Chemotaxis Assays
2.5. Bacterial Two-Hybrid Analyses
2.6. Fluorescence Microscopy
2.7. Statistical Analysis
3. Results
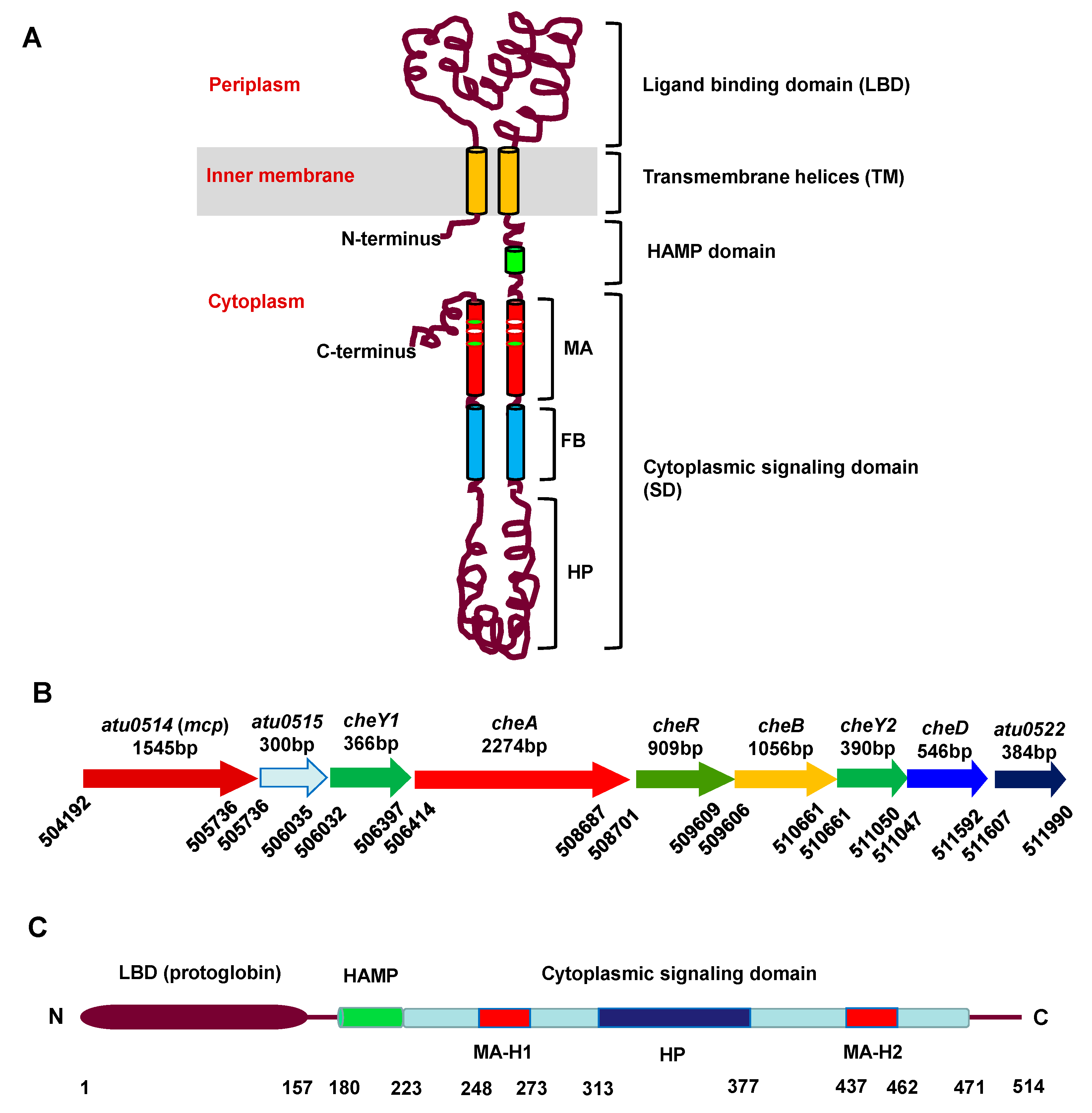
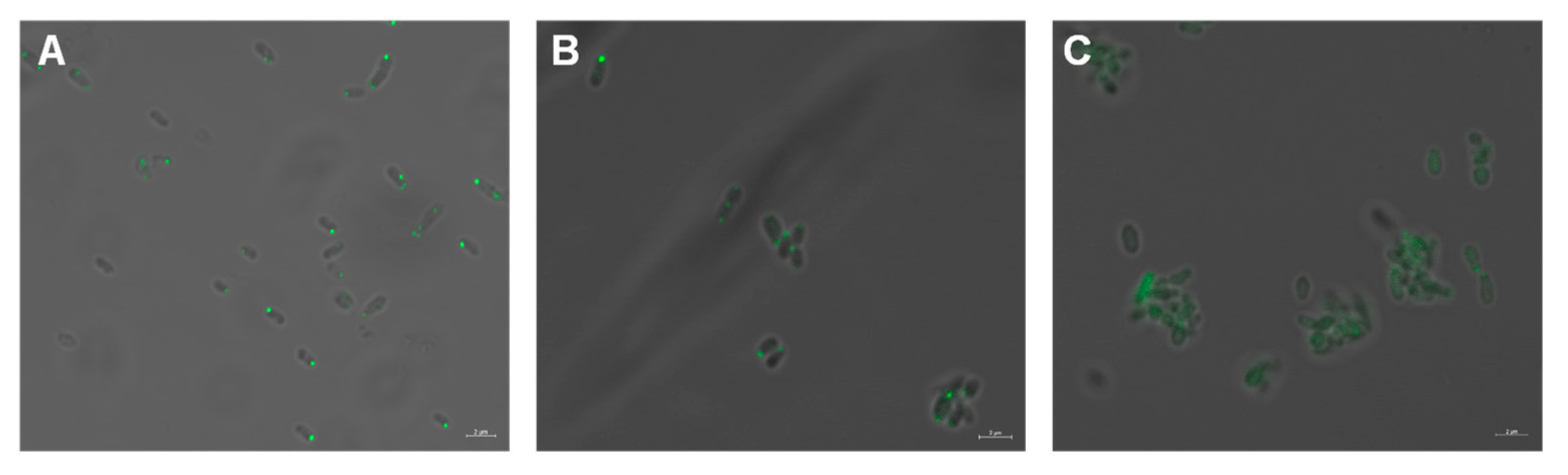
3.1. MCP514 Is a Cytoplasmic Chemoreceptor, but Localized at Cell Poles
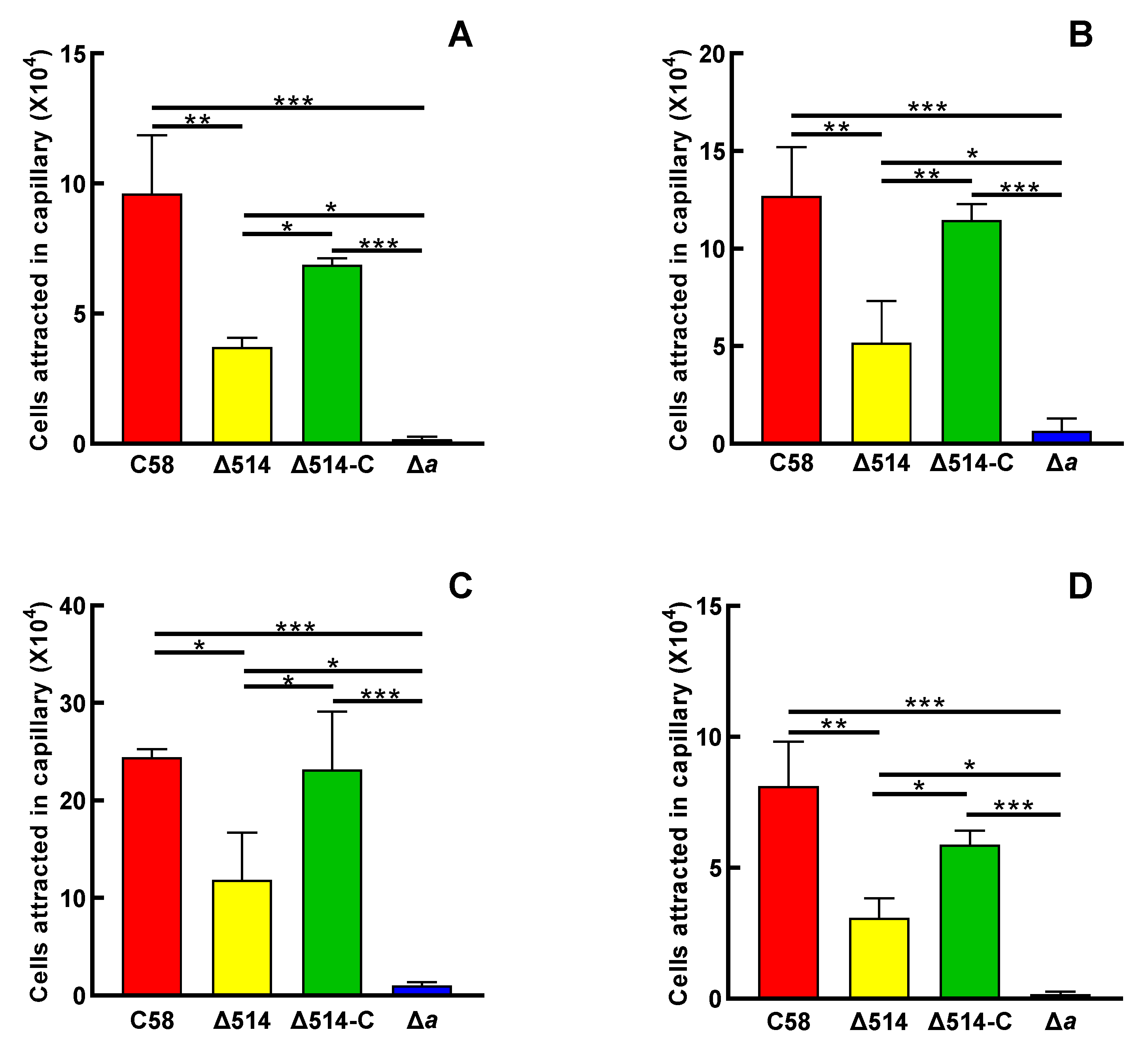
3.2. MCP514 Significantly Affects the Chemotactic Response of Agrobacterium Fabrum
3.3. Both CheW1 and CheW2 Interact with MCP514 but Do Not Affect the Cellular Localization of MCP514
3.4. Helical Structure of Hairpin Subdomain Is Required for the Cellular Localization of MCP514
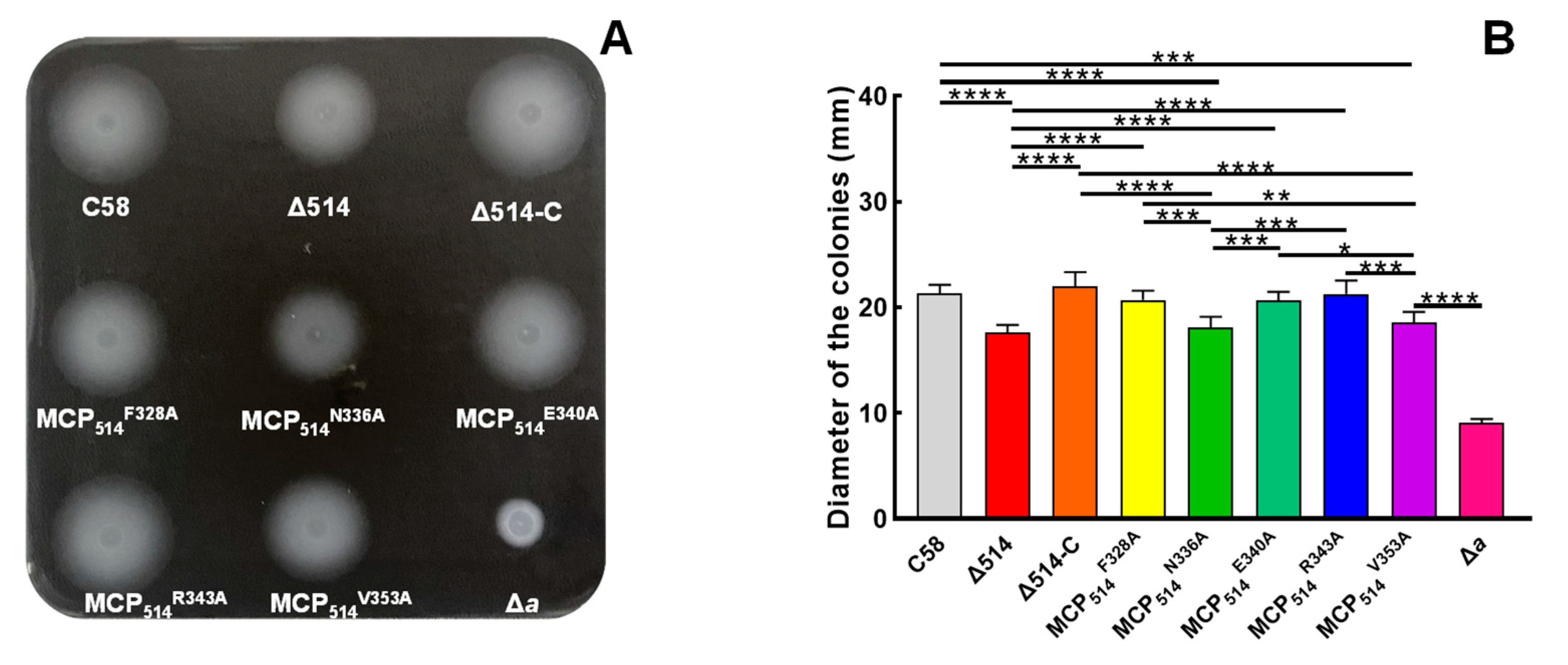
3.5. Two Key Residues of Hairpin Subdomain Play a Key Role in Maintaining the Chemotactic Function of MCP514
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dogra, G.; Purschke, F.G.; Wagner, V.; Haslbeck, M.; Kriehuber, T.; Hughes, J.G.; Van Tassell, M.L.; Gilbert, C.; Niemeyer, M.; Ray, W.K.; et al. Sinorhizobium meliloti CheA complexed with CheS exhibits enhanced binding to CheY1, resulting in accelerated CheY1 dephosphorylation. J. Bacteriol. 2012, 194, 1075–1087. [Google Scholar] [CrossRef]
- Sourjik, V.; Wingreen, N.S. Responding to chemical gradients: Bacterial chemotaxis. Curr. Opin. Cell Biol. 2012, 24, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Parkinson, J.S.; Hazelbauer, G.L.; Falke, J.J. Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol. 2015, 23, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Hazelbauer, G.L.; Falke, J.J.; Parkinson, J.S. Bacterial chemoreceptors: High-performance signaling in networked arrays. Trends Biochem. Sci. 2008, 33, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Sourjik, V. Receptor clustering and signal processing in E. coli chemotaxis. Trends Microbiol. 2004, 12, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Wadhams, G.; Martin, A.C.; Porter, S.; Maddock, J.; Mantotta, J.; King, H.; Armitage, J. TlpC, a novel chemotaxis protein in Rhodobacter sphaeroides, localizes to a discrete region in the cytoplasm. Mol. Microbiol. 2003, 46, 1211–1221. [Google Scholar] [CrossRef] [PubMed]
- Bi, S.; Lai, L. Bacterial chemoreceptors and chemoeffectors. Cell. Mol. Life Sci. 2015, 72, 691–708. [Google Scholar] [CrossRef]
- Karmakar, R. State of the art of bacterial chemotaxis. J. Basic Microbiol. 2021, 61, 366–379. [Google Scholar] [CrossRef]
- Ortega, A.; Zhulin, I.; Krell, T. Sensory repertoire of bacterial chemoreceptors. Microbiol. Mol. Biol. Rev. 2017, 81, e00033-17. [Google Scholar] [CrossRef]
- Sampedro, I.; Parales, R.E.; Krell, T.; Hill, J.E. Pseudomonas chemotaxis. FEMS Microbiol. Rev. 2015, 39, 17–46. [Google Scholar] [CrossRef]
- Gavira, J.A.; Gumerov, V.M.; Rico-Jimenez, M.; Petukh, M.; Upadhyay, A.A.; Ortega, A.; Matilla, M.A.; Zhulin, I.B.; Krell, T. How bacterial chemoreceptors evolve novel ligand specificities. mBio 2020, 11, e03066-19. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Huang, Z.; Yang, J. Is there any crosstalk between the chemotaxis and virulence induction signaling in Agrobacterium tumefaciens? Biotechnol. Adv. 2017, 35, 505–511. [Google Scholar] [CrossRef]
- Adadevoh, J.S.; Triolo, S.; Ramsburg, C.A.; Ford, R.M. Chemotaxis increases the residence time of bacteria in granular media containing distributed contaminant sources. Environ. Sci. Technol. 2016, 50, 181–187. [Google Scholar] [CrossRef] [PubMed]
- Ud-Din, A.; Khan, M.F.; Roujeinikova, A. Broad specificity of amino acid chemoreceptor CtaA of Pseudomonas fluorescens is afforded by plasticity of its amphipathic ligand-binding pocket. Mol. Plant Microbe Interact. 2020, 33, 612–623. [Google Scholar] [CrossRef]
- Yang, W.; Briegel, A. Diversity of bacterial chemosensory arrays. Trends Microbiol. 2020, 28, 68–80. [Google Scholar] [CrossRef] [PubMed]
- Zhulin, I. The superfamily of chemotaxis transducers: From physiology to genomics and back. Adv. Microb. Physiol. 2001, 45, 157–198. [Google Scholar] [CrossRef]
- Lacal, J.; Garcia-Fontana, C.; Munoz-Martinez, F.; Ramos, J.L.; Krell, T. Sensing of environmental signals: Classification of chemoreceptors according to the size of their ligand binding regions. Environ. Microbiol. 2010, 12, 2873–2884. [Google Scholar] [CrossRef]
- Schultz, J.; Milpetz, F.; Bork, P.; Ponting, C. SMART, a simple modular architecture research tool: Identification of signaling domains. Proc. Natl. Acad. Sci. USA 1998, 95, 5857–5864. [Google Scholar] [CrossRef]
- Xie, Z.; Ulrich, L.E.; Zhulin, I.B.; Alexandre, G. PAS domain containing chemoreceptor couples dynamic changes in metabolism with chemotaxis. Proc. Natl. Acad. Sci. USA 2010, 107, 2235–2240. [Google Scholar] [CrossRef]
- Schweinitzer, T.; Mizote, T.; Ishikawa, N.; Dudnik, A.; Inatsu, S.; Schreiber, S.; Suerbaum, S.; Aizawa, S.; Josenhans, C. Functional characterization and mutagenesis of the proposed behavioral sensor TlpD of Helicobacter pylori. J. Bacteriol. 2008, 190, 3244–3255. [Google Scholar] [CrossRef] [PubMed]
- Hou, S.; Larsen, R.; Boudko, D.; Riley, C.; Karatan, E.; Zimmer, M.; Ordal, G.; Alam, M. Myoglobin-like aerotaxis transducers in archaea and bacteria. Nature 2000, 403, 540–544. [Google Scholar] [CrossRef]
- Collins, K.D.; Lacal, J.; Ottemann, K.M. Internal sense of direction: Sensing and signaling from cytoplasmic chemoreceptors. Microbiol. Mol. Biol. Rev. 2014, 78, 672–684. [Google Scholar] [CrossRef]
- Escobar, M.A.; Dandekar, A.M. Agrobacterium tumefaciens as an agent of disease. Trends Plant Sci. 2003, 8, 380–386. [Google Scholar] [CrossRef]
- Pitzschke, A.; Hirt, H. New insights into an old story: Agrobacterium-induced tumour formation in plants by plant transformation. EMBO J. 2010, 29, 1021–1032. [Google Scholar] [CrossRef]
- Guo, M.; Bian, X.; Wu, X.; Wu, M. Agrobacterium-mediated genetic transformation: History and progress. In Genetic Transformation; Alvarez, M., Ed.; InTech: Rijeka, Croatia, 2011. [Google Scholar]
- Kado, C.I. Historical account on gaining insights on the mechanism of crown gall tumorigenesis induced by Agrobacterium tumefaciens. Front. Microbiol. 2014, 5, 340. [Google Scholar] [CrossRef]
- Yuan, Z.C.; Williams, M. A really useful pathogen, Agrobacterium tumefaciens. Plant Cell 2012, 24, tpc.112.tt1012. [Google Scholar] [CrossRef] [PubMed]
- Hawes, M.; Smith, L. Requirement for chemotaxis in pathogenicity of Agrobacterium tumefaciens on roots of soil-grown pea plants. J. Bacteriol. 1989, 171, 5668–5671. [Google Scholar] [CrossRef] [PubMed]
- Ashby, A.; Watson, M.; Loake, G.; Shaw, C. Ti plasmid-specified chemotaxis of Agrobacterium tumefaciens C58C1 toward vir-inducing phenolic compounds and soluble factors from monocotyledonous and dicotyledonous plants. J. Bacteriol. 1988, 170, 4181–4187. [Google Scholar] [CrossRef] [PubMed]
- Loake, G.; Ashby, A.; Shaw, C. Attraction of Agrobacterium tumefaciens C58C1 towards sugars involves a highly sensitive chemotaxis system. J. Gen. Microbiol. 1988, 134, 1427–1432. [Google Scholar] [CrossRef][Green Version]
- Parke, D.; Ornston, L.; Nester, E. Chemotaxis to plant phenolic inducers of virulence genes is constitutively expressed in the absence of the Ti plasmid in Agrobacterium tumefaciens. J. Bacteriol. 1987, 169, 5336–5338. [Google Scholar] [CrossRef]
- Winans, S. Two-way chemical signaling in Agrobacterium-plant interactions. Microbiol. Rev. 1992, 56, 12–31. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.W.; Setubal, J.C.; Kaul, R.; Monks, D.E.; Kitajima, J.P.; Okura, V.K.; Zhou, Y.; Chen, L.; Wood, G.E.; Almeida, N.F., Jr.; et al. The genome of the natural genetic engineer Agrobacterium tumefaciens C58. Science 2001, 294, 2317–2323. [Google Scholar] [CrossRef] [PubMed]
- Goodner, B.; Hinkle, G.; Gattung, S.; Miller, N.; Blanchard, M.; Qurollo, B.; Goldman, B.S.; Cao, Y.; Askenazi, M.; Halling, C.; et al. Genome sequence of the plant pathogen and biotechnology agent Agrobacterium tumefaciens C58. Science 2001, 294, 2323–2328. [Google Scholar] [CrossRef]
- Sambrock, J.; Russel, D. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Cangelosi, G.A.; Best, E.A.; Martinetti, G.; Nester, E.W. Genetic analysis of Agrobacterium. Methods Enzymol. 1991, 204, 384–397. [Google Scholar] [CrossRef] [PubMed]
- Gelvin, S.B. Agrobacterium virulence gene induction. Methods Mol. Biol. 2006, 343, 77–84. [Google Scholar] [CrossRef]
- Guo, M.; Hou, Q.; Hew, C.L.; Pan, S.Q. Agrobacterium VirD2-binding protein is involved in tumorigenesis and redundantly encoded in conjugative transfer gene clusters. Mol. Plant Microbe Interact. 2007, 20, 1201–1212. [Google Scholar] [CrossRef]
- Guo, M.L.; Zhu, Q.; Gao, D.K. Development and optimization of method for generating unmarked A. tumefaciens mutants. Prog. Biochem. Biophys. 2009, 36, 556–565. [Google Scholar] [CrossRef]
- Huang, Z.; Zhou, Q.; Sun, P.; Yang, J.; Guo, M. Two Agrobacterium tumefaciens CheW proteins are incorporated into one chemosensory pathway with different efficiencies. Mol. Plant Microbe Interact. 2018, 31, 460–470. [Google Scholar] [CrossRef]
- Higuchi, R. Using PCR to engineer DNA. In PCR Technology; Palgrave Macmillan: London, UK, 1989; pp. 61–70. [Google Scholar]
- Adler, J. A method for measuring chemotaxis and use of the method to determine optimum conditions for chemotaxis by Escherichia coli. Microbiology 1973, 74, 77–91. [Google Scholar] [CrossRef]
- Merritt, P.M.; Danhorn, T.; Fuqua, C. Motility and chemotaxis in Agrobacterium tumefaciens surface attachment and biofilm formation. J. Bacteriol. 2007, 189, 8005–8014. [Google Scholar] [CrossRef]
- Miller, J. Experiments in Molecular Genetics; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 1972; Volume 172. [Google Scholar]
- Alexander, R.; Zhulin, I. Evolutionary genomics reveals conserved structural determinants of signaling and adaptation in microbial chemoreceptors. Proc. Natl. Acad. Sci. USA 2007, 104, 2885–2890. [Google Scholar] [CrossRef]
- Hirokawa, T.; Boon-Chieng, S.; Mitaku, S. SOSUI: Classification and secondary structure prediction system for membrane proteins. Bioinformatics 1998, 14, 378–379. [Google Scholar] [CrossRef]
- Juretić, D.; Zoranić, L.; Zucić, D. Basic charge clusters and predictions of membrane protein topology. J. Chem. Inf. Comput. Sci. 2002, 42, 620–632. [Google Scholar] [CrossRef]
- Greenfield, D.; McEvoy, A.L.; Shroff, H.; Crooks, G.E.; Wingreen, N.S.; Betzig, E.; Liphardt, J. Self-organization of the Escherichia coli chemotaxis network imaged with super-resolution light microscopy. PLoS Biol. 2009, 7, e1000137. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q. Functional Identification of Methyl Accepting Chemotaxis Protein Encoded by Gene atu0514 Located in the Chemotaxis Operon of Agrobacterium tumefaciens. Master’s Thesis, Yangzhou University, Yangzhou, China, 2020. [Google Scholar]
- Kim, K.K.; Yokota, H.; Sangjin, K. Four-helical-bundle structure of the cytoplasmic domain of a serine chemotaxis receptor. Nature 1999, 400, 787–792. [Google Scholar] [CrossRef] [PubMed]
- Ames, P.; Studdert, C.; Reiser, R.; Parkinson, J. Collaborative signaling by mixed chemoreceptor teams in Escherichia coli. Proc. Natl. Acad. Sci. USA 2002, 99, 7060–7065. [Google Scholar] [CrossRef] [PubMed]
- Studdert, C.; Parkinson, J. Crosslinking snapshots of bacterial chemoreceptor squads. Proc. Natl. Acad. Sci. USA 2004, 101, 2117–2122. [Google Scholar] [CrossRef] [PubMed]
- Studdert, C.A.; Parkinson, J.S. Insights into the organization and dynamics of bacterial chemoreceptor clusters through in vivo crosslinking studies. Proc. Natl. Acad. Sci. USA 2005, 102, 15623–15628. [Google Scholar] [CrossRef]
- Li, M.; Khursigara, C.M.; Subramaniam, S.; Hazelbauer, G.L. Chemotaxis kinase CheA is activated by three neighbouring chemoreceptor dimers as effectively as by receptor clusters. Mol. Microbiol. 2011, 79, 677–685. [Google Scholar] [CrossRef]
- Gosink, K.K.; Zhao, Y.; Parkinson, J.S. Mutational analysis of N381, a key trimer contact residue in Tsr, the Escherichia coli serine chemoreceptor. J. Bacteriol. 2011, 193, 6452–6460. [Google Scholar] [CrossRef]
- Wright, E.; Deakin, W.; Shaw, C. A chemotaxis cluster from Agrobacterium tumefaciens. Gene 1998, 220, 83–89. [Google Scholar] [CrossRef]
- McCullen, C.A.; Binns, A.N. Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annu. Rev. Cell Dev. Biol. 2006, 22, 101–127. [Google Scholar] [CrossRef]
- Gelvin, S.B. Traversing the cell: Agrobacterium T-DNA’s journey to the host genome. Front. Plant Sci. 2012, 3, 52. [Google Scholar] [CrossRef] [PubMed]
- Pitzschke, A. Agrobacterium infection and plant defense—Transformation success hangs by a thread. Front. Plant Sci. 2013, 4, 519. [Google Scholar] [CrossRef] [PubMed]
- Szurmant, H.; Ordal, G.W. Diversity in chemotaxis mechanisms among the bacteria and archaea. Microbiol. Mol. Biol. Rev. 2004, 68, 301–319. [Google Scholar] [CrossRef] [PubMed]
- Meier, V.M.; Scharf, B.E. Cellular localization of predicted transmembrane and soluble chemoreceptors in Sinorhizobium meliloti. J. Bacteriol. 2009, 191, 5724–5733. [Google Scholar] [CrossRef]
- Briegel, A.; Ladinsky, M.S.; Oikonomou, C.; Jones, C.W.; Harris, M.J.; Fowler, D.J.; Chang, Y.W.; Thompson, L.K.; Armitage, J.P.; Jensen, G.J. Structure of bacterial cytoplasmic chemoreceptor arrays and implications for chemotactic signaling. Elife 2014, 3, e02151. [Google Scholar] [CrossRef]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ye, J.; Gao, M.; Zhou, Q.; Wang, H.; Xu, N.; Guo, M. The Only Chemoreceptor Encoded by che Operon Affects the Chemotactic Response of Agrobacterium to Various Chemoeffectors. Microorganisms 2021, 9, 1923. https://doi.org/10.3390/microorganisms9091923
Ye J, Gao M, Zhou Q, Wang H, Xu N, Guo M. The Only Chemoreceptor Encoded by che Operon Affects the Chemotactic Response of Agrobacterium to Various Chemoeffectors. Microorganisms. 2021; 9(9):1923. https://doi.org/10.3390/microorganisms9091923
Chicago/Turabian StyleYe, Jingyang, Miaomiao Gao, Qingxuan Zhou, Hao Wang, Nan Xu, and Minliang Guo. 2021. "The Only Chemoreceptor Encoded by che Operon Affects the Chemotactic Response of Agrobacterium to Various Chemoeffectors" Microorganisms 9, no. 9: 1923. https://doi.org/10.3390/microorganisms9091923
APA StyleYe, J., Gao, M., Zhou, Q., Wang, H., Xu, N., & Guo, M. (2021). The Only Chemoreceptor Encoded by che Operon Affects the Chemotactic Response of Agrobacterium to Various Chemoeffectors. Microorganisms, 9(9), 1923. https://doi.org/10.3390/microorganisms9091923





