Altered Fecal Microbiotas and Organic Acid Concentrations Indicate Possible Gut Dysbiosis in University Rugby Players: An Observational Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics Statements and Study Participants
2.2. Analysis of the Fecal Microbiota
2.3. Analysis of the Concentrations of Fecal Organic Acids
2.4. Statistical Analysis
3. Results
3.1. Corrections of the Experimental Groups
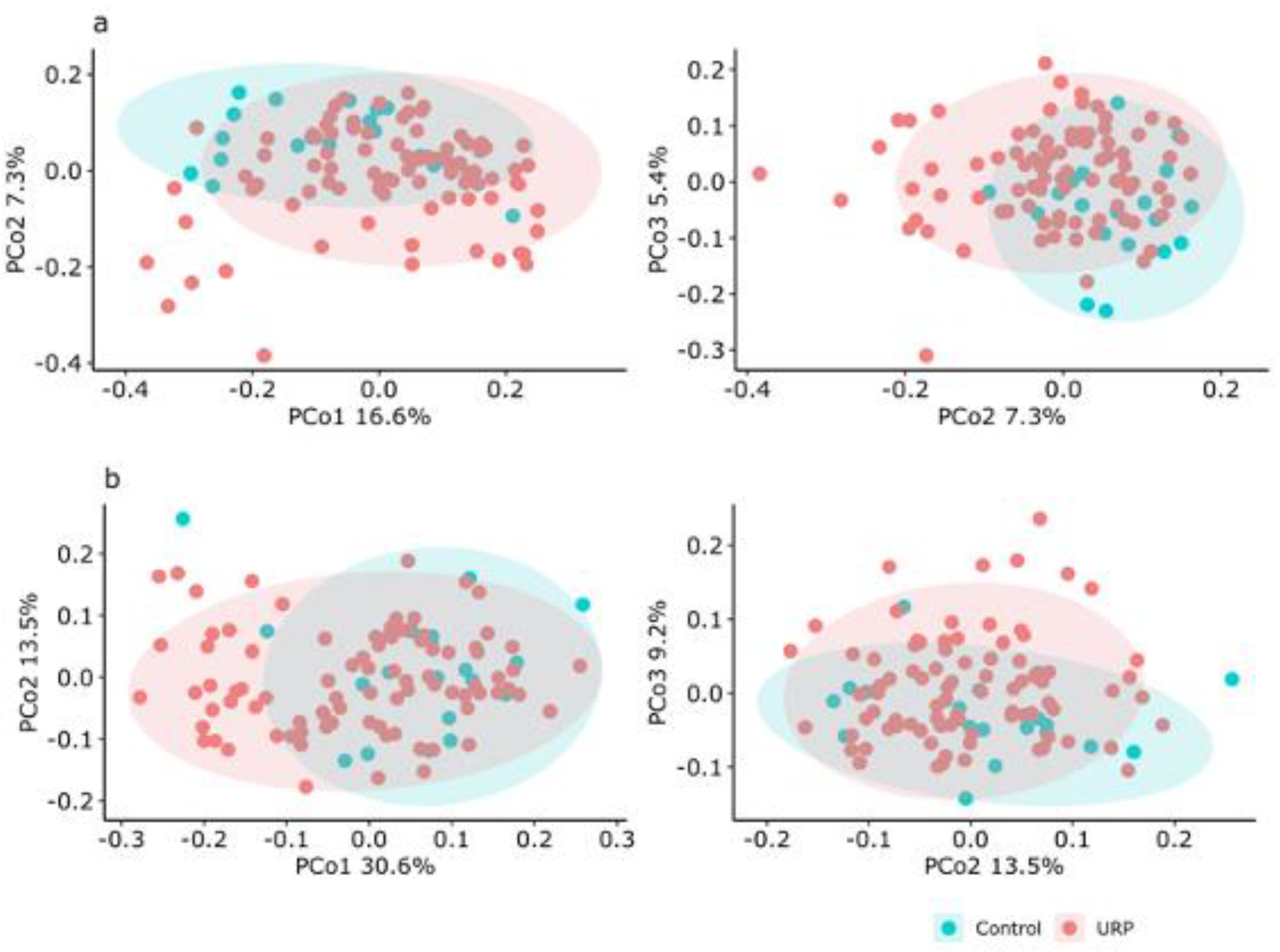
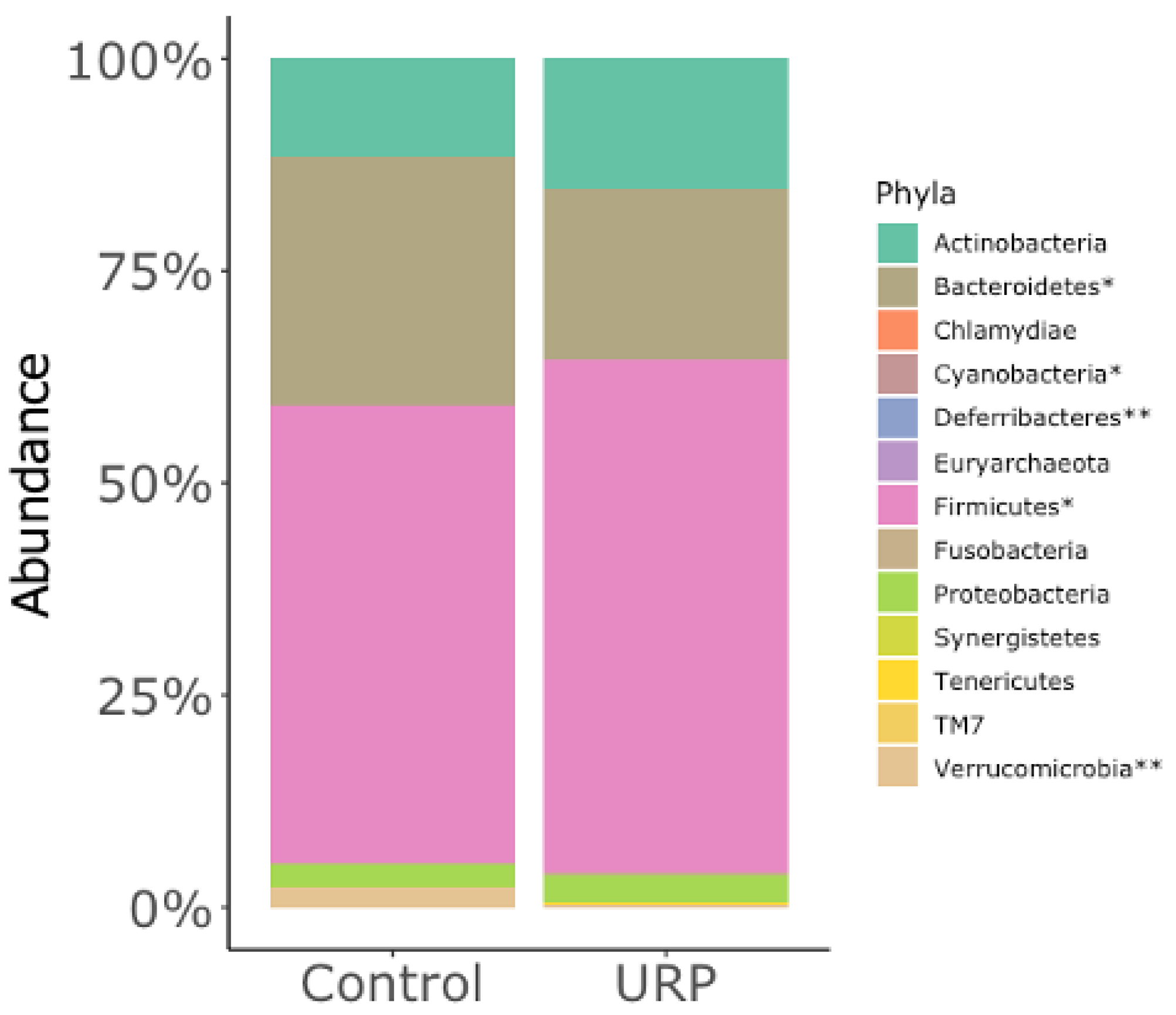
3.2. Analysis of the Fecal Microbiota
3.3. Concentrations of Fecal Organic Acids
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cryan, J.F.; O’Riordan, K.J.; Sandhu, K.; Peterson, V.; Dinan, T.G. The gut microbiome in neurological disorders. Lancet Neurol 2020, 19, 179–194. [Google Scholar] [CrossRef]
- Milosevic, I.; Vujovic, A.; Barac, A.; Djelic, M.; Korac, M.; Radovanovic Spurnic, A.; Gmizic, I.; Stevanovic, O.; Djordjevic, V.; Lekic, N.; et al. Gut-Liver Axis, Gut Microbiota, and Its Modulation in the Management of Liver Diseases: A Review of the Literature. Int. J. Mol. Sci. 2019, 20, 395. [Google Scholar] [CrossRef] [Green Version]
- Sochocka, M.; Donskow-Lysoniewska, K.; Diniz, B.S.; Kurpas, D.; Brzozowska, E.; Leszek, J. The Gut Microbiome Alterations and Inflammation-Driven Pathogenesis of Alzheimer’s Disease-a Critical Review. Mol. Neurobiol. 2019, 56, 1841–1851. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Alammar, N.; Singh, R.; Nanavati, J.; Song, Y.; Chaudhary, R.; Mullin, G.E. Gut Microbial Dysbiosis in the Irritable Bowel Syndrome: A Systematic Review and Meta-Analysis of Case-Control Studies. J. Acad. Nutr. Diet. 2020, 120, 565–586. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ortiz-Alvarez, L.; Xu, H.; Martinez-Tellez, B. Influence of Exercise on the Human Gut Microbiota of Healthy Adults: A Systematic Review. Clin. Transl. Gastroenterol. 2020, 11, e00126. [Google Scholar] [CrossRef]
- McNabney, S.M.; Henagan, T.M. Short Chain Fatty Acids in the Colon and Peripheral Tissues: A Focus on Butyrate, Colon Cancer, Obesity and Insulin Resistance. Nutrients 2017, 9, 1348. [Google Scholar] [CrossRef] [Green Version]
- Allen, J.M.; Mailing, L.J.; Niemiro, G.M.; Moore, R.; Cook, M.D.; White, B.A.; Holscher, H.D.; Woods, J.A. Exercise Alters Gut Microbiota Composition and Function in Lean and Obese Humans. Med. Sci. Sports Exerc. 2018, 50, 747–757. [Google Scholar] [CrossRef] [PubMed]
- Bressa, C.; Bailen-Andrino, M.; Perez-Santiago, J.; Gonzalez-Soltero, R.; Perez, M.; Montalvo-Lominchar, M.G.; Mate-Munoz, J.L.; Dominguez, R.; Moreno, D.; Larrosa, M. Differences in gut microbiota profile between women with active lifestyle and sedentary women. PLoS ONE 2017, 12, e0171352. [Google Scholar] [CrossRef] [Green Version]
- Allen, J.M.; Berg Miller, M.E.; Pence, B.D.; Whitlock, K.; Nehra, V.; Gaskins, H.R.; White, B.A.; Fryer, J.D.; Woods, J.A. Voluntary and forced exercise differentially alters the gut microbiome in C57BL/6J mice. J. Appl. Physiol. (1985) 2015, 118, 1059–1066. [Google Scholar] [CrossRef] [Green Version]
- Jang, L.G.; Choi, G.; Kim, S.W.; Kim, B.Y.; Lee, S.; Park, H. The combination of sport and sport-specific diet is associated with characteristics of gut microbiota: An observational study. J. Int. Soc. Sports Nutr. 2019, 16, 21. [Google Scholar] [CrossRef] [Green Version]
- Morishima, S.; Aoi, W.; Kawamura, A.; Kawase, T.; Takagi, T.; Naito, Y.; Tsukahara, T.; Inoue, R. Intensive, prolonged exercise seemingly causes gut dysbiosis in female endurance runners. J. Clin. Biochem. Nutr. 2021, 68, 253–258. [Google Scholar] [CrossRef]
- Leiper, J.B.; Nicholas, C.W.; Ali, A.; Williams, C.; Maughan, R.J. The effect of intermittent high-intensity running on gastric emptying of fluids in man. Med. Sci. Sports Exerc. 2005, 37, 240–247. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, E.P.; Burini, R.C. The impact of physical exercise on the gastrointestinal tract. Curr. Opin. Clin. Nutr. Metab. Care 2009, 12, 533–538. [Google Scholar] [CrossRef] [PubMed]
- ter Steege, R.W.; Kolkman, J.J. Review article: The pathophysiology and management of gastrointestinal symptoms during physical exercise, and the role of splanchnic blood flow. Aliment. Pharmacol. Ther. 2012, 35, 516–528. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, N.R.; DiMarco, N.M.; Langley, S. American Dietetic Association, Dietitians of Canada, American College of Sports Medicine: Nutrition and Athletic Performance. Position of the American Dietetic Association, Dietitians of Canada, and the American College of Sports Medicine: Nutrition and athletic performance. J. Am. Diet. Assoc. 2009, 109, 509–527. [Google Scholar] [CrossRef] [PubMed]
- Clarke, S.F.; Murphy, E.F.; O’Sullivan, O.; Lucey, A.J.; Humphreys, M.; Hogan, A.; Hayes, P.; O’Reilly, M.; Jeffery, I.B.; Wood-Martin, R.; et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut 2014, 63, 1913–1920. [Google Scholar] [CrossRef] [Green Version]
- Egan, B. Protein intake for athletes and active adults: Current concepts and controversies. Nutr. Bull. 2016, 41, 202–213. [Google Scholar] [CrossRef] [Green Version]
- Clark, A.; Mach, N. Exercise-induced stress behavior, gut-microbiota-brain axis and diet: A systematic review for athletes. J. Int. Soc. Sports Nutr. 2016, 13, 43. [Google Scholar] [CrossRef] [Green Version]
- Nirengi, S.; Fujibayashi, M.; Furuno, S.; Uchibe, A.; Kawase, Y.; Sukino, S.; Kawaguchi, Y.; Minato, S.; Kotani, K.; Sakane, N. Nonalcoholic Fatty Liver Disease in University Rugby Football Players. Front. Endocrinol. (Lausanne) 2018, 9, 341. [Google Scholar] [CrossRef] [Green Version]
- Leung, C.; Rivera, L.; Furness, J.B.; Angus, P.W. The role of the gut microbiota in NAFLD. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 412–425. [Google Scholar] [CrossRef]
- World Health Orgnization. Obesity: Preventing and Managing the Global Epidemic; Report of a WHO Consultation; World Health Orgnization: Geneva, Switzerland, 2000. [Google Scholar]
- Inoue, R.; Sakaue, Y.; Sawai, C.; Sawai, T.; Ozeki, M.; Romero-Perez, G.A.; Tsukahara, T. A preliminary investigation on the relationship between gut microbiota and gene expressions in peripheral mononuclear cells of infants with autism spectrum disorders. Biosci. Biotechnol. Biochem. 2016, 80, 2450–2458. [Google Scholar] [CrossRef] [Green Version]
- Andoh, A.; Inoue, R.; Kawada, Y.; Morishima, S.; Inatomi, O.; Ohno, M.; Bamba, S.; Nishida, A.; Kawahara, M.; Naito, Y. Elemental diet induces alterations of the gut microbial community in mice. J. Clin. Biochem. Nutr. 2019, 65, 118–124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahe, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, R.C. UCHIME2: Improved chimera prediction for amplicon sequencing. bioRxiv 2016. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [Green Version]
- Tsukahara, T.; Matsukawa, N.; Tomonaga, S.; Inoue, R.; Ushida, K.; Ochiai, K. High-sensitivity detection of short-chain fatty acids in porcine ileal, cecal, portal and abdominal blood by gas chromatography-mass spectrometry. Anim. Sci. J. 2014, 85, 494–498. [Google Scholar] [CrossRef]
- Shin, J.H.; Park, Y.H.; Sim, M.; Kim, S.A.; Joung, H.; Shin, D.M. Serum level of sex steroid hormone is associated with diversity and profiles of human gut microbiome. Res. Microbiol. 2019, 170, 192–201. [Google Scholar] [CrossRef]
- Gotshalk, L.A.; Loebel, C.C.; Nindl, B.C.; Putukian, M.; Sebastianelli, W.J.; Newton, R.U.; Hakkinen, K.; Kraemer, W.J. Hormonal responses of multiset versus single-set heavy-resistance exercise protocols. Can. J. Appl. Physiol. 1997, 22, 244–255. [Google Scholar] [CrossRef]
- Kraemer, W.J.; Gordon, S.E.; Fleck, S.J.; Marchitelli, L.J.; Mello, R.; Dziados, J.E.; Friedl, K.; Harman, E.; Maresh, C.; Fry, A.C. Endogenous anabolic hormonal and growth factor responses to heavy resistance exercise in males and females. Int. J. Sports Med. 1991, 12, 228–235. [Google Scholar] [CrossRef] [PubMed]
- Medawar, E.; Haange, S.-B.; Rolle-Kampczyk, U.; Engelmann, B.; Dietrich, A.; Thieleking, R.; Wiegank, C.; Fries, C.; Horstmann, A.; Villringer, A.; et al. Gut microbiota link dietary fiber intake and short-chain fatty acid metabolism with eating behaviour. medRxiv 2021. [Google Scholar] [CrossRef]
- Gomez-Arango, L.F.; Barrett, H.L.; Wilkinson, S.A.; Callaway, L.K.; McIntyre, H.D.; Morrison, M.; Dekker Nitert, M. Low dietary fiber intake increases Collinsella abundance in the gut microbiota of overweight and obese pregnant women. Gut Microbes 2018, 9, 189–201. [Google Scholar] [CrossRef] [Green Version]
- Tomsett, K.I.; Barrett, H.L.; Dekker, E.E.; Callaway, L.K.; McIntyre, D.H.; Dekker Nitert, M. Dietary Fiber Intake Alters Gut Microbiota Composition but Does Not Improve Gut Wall Barrier Function in Women with Future Hypertensive Disorders of Pregnancy. Nutrients 2020, 12, 3862. [Google Scholar] [CrossRef]
- Gevers, D.; Kugathasan, S.; Denson, L.A.; Vazquez-Baeza, Y.; Van Treuren, W.; Ren, B.; Schwager, E.; Knights, D.; Song, S.J.; Yassour, M.; et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 2014, 15, 382–392. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cani, P.D.; de Vos, W.M. Next-Generation Beneficial Microbes: The Case of Akkermansia muciniphila. Front. Microbiol. 2017, 8, 1765. [Google Scholar] [CrossRef] [PubMed]
- Connors, J.; Dawe, N.; Van Limbergen, J. The Role of Succinate in the Regulation of Intestinal Inflammation. Nutrients 2018, 11, 25. [Google Scholar] [CrossRef] [Green Version]
- Tsukahara, T.; Ushida, K. Succinate accumulation in pig large intestine during antibiotic-associated diarrhea and the constitution of succinate-producing flora. J. Gen. Appl. Microbiol. 2002, 48, 143–154. [Google Scholar] [CrossRef] [Green Version]
- Vernia, P.; Caprilli, R.; Latella, G.; Barbetti, F.; Magliocca, F.M.; Cittadini, M. Fecal lactate and ulcerative colitis. Gastroenterology 1988, 95, 1564–1568. [Google Scholar] [CrossRef]
- Siigur, U.; Norin, K.E.; Allgood, G.; Schlagheck, T.; Midtvedt, T. Concentrations and Correlations of Faecal Short-chain Fatty Acids and Faecal Water Content in Man. Microb. Ecol. Health Dis. 1994, 7, 287–294. [Google Scholar] [CrossRef]
- Barton, W.; Penney, N.C.; Cronin, O.; Garcia-Perez, I.; Molloy, M.G.; Holmes, E.; Shanahan, F.; Cotter, P.D.; O’Sullivan, O. The microbiome of professional athletes differs from that of more sedentary subjects in composition and particularly at the functional metabolic level. Gut 2018, 67, 625–633. [Google Scholar] [CrossRef] [PubMed]
| URPs † | |||
|---|---|---|---|
| Controls | FWs | BKs | |
| Age | 20.1 ± 1.8 | 20.5 ± 1.2 | 20.8 ± 1.3 |
| BMI | 20.8 ± 2.6 *a | 30.7 ± 3.6 b | 26.1 ± 1.8 c |
| Index | Controls | URP | p-Value |
|---|---|---|---|
| Chao1 | 206.74 ± 71.07 | 164.85 ± 47.84 | 0.02 * |
| Shannon | 4.75 ± 0.55 | 4.53 ± 0.52 | 0.13 |
| Taxonomy | Control | URP | p-Value | ||
|---|---|---|---|---|---|
| Order | Family | Genus | |||
| Higher in URP | |||||
| Clostridiales | Veillonellaceae | Megamonas | 0.299 ± 1.303 | 4.698 ± 8.953 | <0.001 ** |
| Coriobacteriales | Coriobacteriaceae | Collinsella | 2.512 ± 3.064 | 5.404 ± 5.989 | 0.025 * |
| Pasteurellales | Pasteurellaceae | Haemophilus | 0.157 ± 0.475 | 0.795 ± 2.305 | 0.004 ** |
| Clostridiales | Clostridiaceae | SMB53 | 0.125 ± 0.124 | 0.730 ± 1.934 | 0.010 * |
| Enterobacteriales | Enterobacteriaceae | Enterobacter | 0.020 ± 0.045 | 0.433 ± 1.193 | 0.002 ** |
| Fusobacteriales | Fusobacteriaceae | Unclassified | 0.001 ± 0.005 | 0.287 ± 0.924 | 0.028 * |
| Actinomycetales | Actinomycetaceae | Actinomyces | 0.076 ± 0.108 | 0.164 ± 0.198 | 0.019 * |
| Lower in URP | |||||
| Clostridiales | Lachnospiraceae | Clostridium | 0.587 ± 0.645 | 0.521 ± 1.547 | 0.013 * |
| Clostridiales | Ruminococcaceae | Unclassified | 0.178 ± 0.275 | 0.107 ± 0.304 | 0.012 * |
| Coriobacteriales | Coriobacteriaceae | Eggerthella | 0.150 ± 0.153 | 0.075 ± 0.090 | 0.027 * |
| Bacteroidales | [Odoribacteraceae] | Odoribacter | 0.200 ± 0.274 | 0.053 ± 0.140 | 0.001 ** |
| Desulfovibrionales | Desulfovibrionaceae | Bilophila | 0.241 ± 0.262 | 0.053 ± 0.093 | 0.001 ** |
| Clostridiales | Unclassified | Unclassified | 0.593 ± 1.249 | 0.318 ± 0.924 | 0.008 ** |
| Bacteroidales | Rikenellaceae | Unclassified | 1.283 ± 1.328 | 0.406 ± 1.128 | <0.001 ** |
| Clostridiales | Ruminococcaceae | Oscillospira | 1.978 ± 1.478 | 0.938 ± 1.277 | <0.001 ** |
| Verrucomicrobiales | Verrucomicrobiaceae | Akkermansia | 2.322 ± 5.611 | 0.436 ± 2.347 | 0.010 ** |
| Bacteroidales | Bacteroidaceae | Bacteroides | 23.771 ± 11.922 | 14.765 ± 11.455 | 0.004 ** |
| Organic Acids | Controls | URP | p-Value |
|---|---|---|---|
| (mmol/kg Wet Feces) | |||
| Succinate | 0.3 ± 0.5 | 1.6 ± 3.1 | <0.001 ** |
| Lactate | 0.2 ± 0.6 | 0.0 ± 0.1 | 0.27 |
| Formiate | 0.2 ± 0.4 | 0.3 ± 1.2 | 0.58 |
| Acetate | 35.3 ± 18.2 | 38.5 ± 20.2 | 0.51 |
| Propionate | 11.6 ± 5.2 | 10.4 ± 8.5 | 0.44 |
| isoButyrate | 3.0 ± 4.6 | 0.5 ± 1.5 | 0.03 * |
| nButyrate | 7.4 ± 6.7 | 5.2 ± 4.6 | 0.19 |
| isoValerate | 1.3 ± 1.0 | 0.0 ± 0.2 | <0.001 ** |
| nValerate | <0.1 | 0.0 ± 0.2 | 0.32 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morishima, S.; Oda, N.; Ikeda, H.; Segawa, T.; Oda, M.; Tsukahara, T.; Kawase, Y.; Takagi, T.; Naito, Y.; Fujibayashi, M.; et al. Altered Fecal Microbiotas and Organic Acid Concentrations Indicate Possible Gut Dysbiosis in University Rugby Players: An Observational Study. Microorganisms 2021, 9, 1687. https://doi.org/10.3390/microorganisms9081687
Morishima S, Oda N, Ikeda H, Segawa T, Oda M, Tsukahara T, Kawase Y, Takagi T, Naito Y, Fujibayashi M, et al. Altered Fecal Microbiotas and Organic Acid Concentrations Indicate Possible Gut Dysbiosis in University Rugby Players: An Observational Study. Microorganisms. 2021; 9(8):1687. https://doi.org/10.3390/microorganisms9081687
Chicago/Turabian StyleMorishima, So, Naoko Oda, Hiromi Ikeda, Tomohiro Segawa, Machi Oda, Takamitsu Tsukahara, Yasuharu Kawase, Tomohisa Takagi, Yuji Naito, Mami Fujibayashi, and et al. 2021. "Altered Fecal Microbiotas and Organic Acid Concentrations Indicate Possible Gut Dysbiosis in University Rugby Players: An Observational Study" Microorganisms 9, no. 8: 1687. https://doi.org/10.3390/microorganisms9081687
APA StyleMorishima, S., Oda, N., Ikeda, H., Segawa, T., Oda, M., Tsukahara, T., Kawase, Y., Takagi, T., Naito, Y., Fujibayashi, M., & Inoue, R. (2021). Altered Fecal Microbiotas and Organic Acid Concentrations Indicate Possible Gut Dysbiosis in University Rugby Players: An Observational Study. Microorganisms, 9(8), 1687. https://doi.org/10.3390/microorganisms9081687







