Current and Future Antiviral Strategies to Tackle Gastrointestinal Viral Infections
Abstract
:1. Introduction
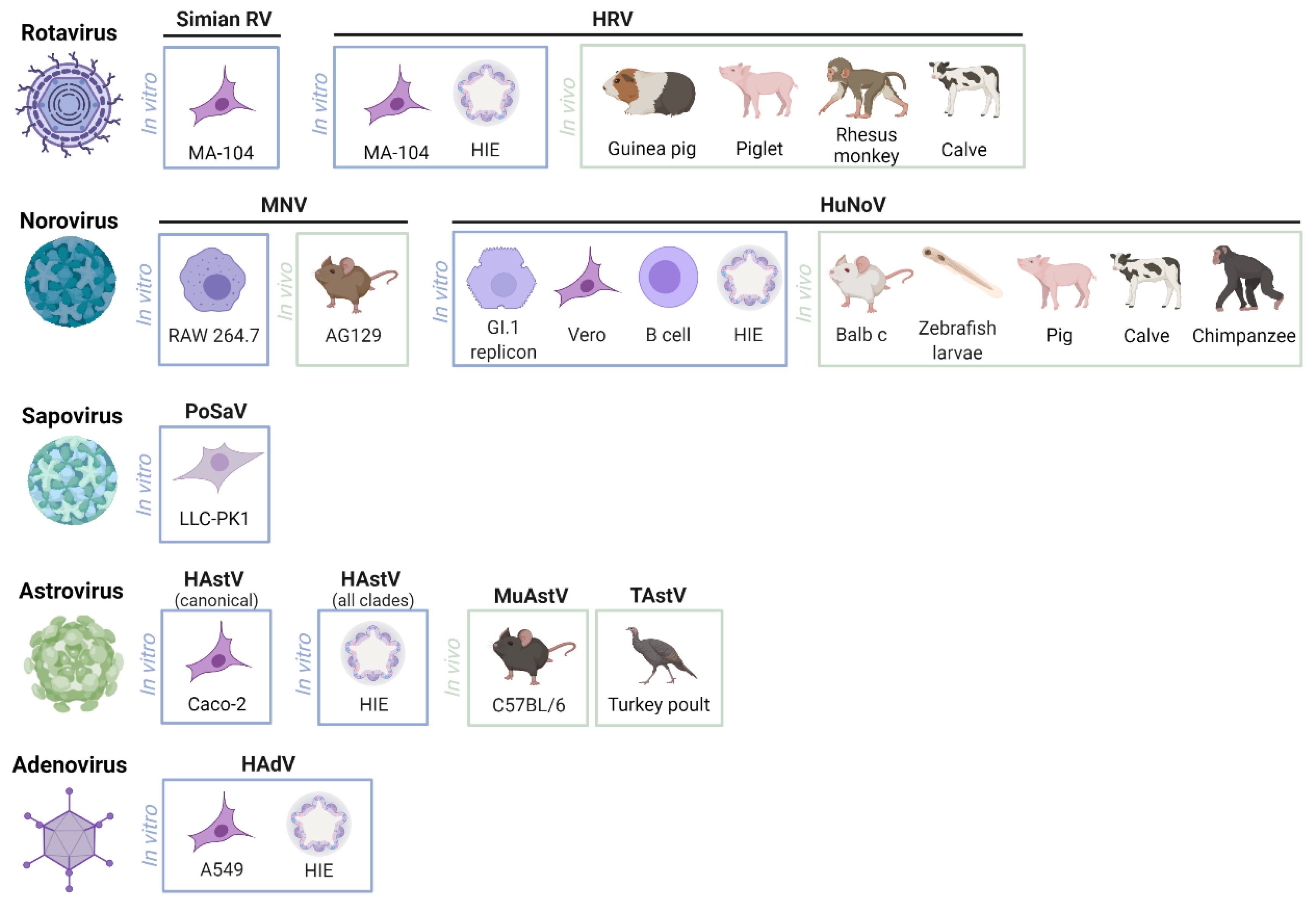
2. Model Systems to Study Viral Replication and Disease
2.1. In Vitro Systems
2.1.1. Rotavirus
2.1.2. Norovirus
2.1.3. Sapovirus
2.1.4. Astrovirus
2.1.5. Adenovirus
2.2. In Vivo Systems
2.2.1. Rotavirus
2.2.2. Norovirus
2.2.3. Astrovirus
3. Antiviral Targets and Known Antivirals
3.1. Rotavirus
3.1.1. Suppression of Virus Replication
3.1.2. Inhibition of Viroplasm Formation
3.1.3. Targeting Host Cell Factors Essential for Viral Replication and Others
3.2. Norovirus
3.2.1. Targeting Virus Binding to Host Cell Surface
3.2.2. Targeting the Viral Protease
3.2.3. Targeting the Viral RdRp
3.2.4. Targeting Host Cell Factors Essential for Viral Replication
3.2.5. Others
3.3. Sapovirus
3.4. Astrovirus
3.4.1. Targeting Virus RdRp
3.4.2. Others or Unknown Target
3.5. Adenovirus
3.5.1. Targeting Viral DNA Polymerase
3.5.2. Others
3.6. Conclusions
4. Future Perspectives and Challenges
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Klein, E.J.; Boster, D.R.; Stapp, J.R.; Wells, J.G.; Qin, X.; Clausen, C.R.; Swerdlow, D.L.; Braden, C.R.; Tarr, P.I. Diarrhea etiology in a children’s hospital emergency department: A prospective cohort study. Clin. Infect. Dis. 2006, 43, 807–813. [Google Scholar] [CrossRef]
- O’Ryan, M.; Prado, V.; Pickering, L.K. A millennium update on pediatric diarrheal illness in the developing world. Semin. Pediatr. Infect. Dis. 2005, 16, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Simpson, R.; Aliyu, S.; Iturriza-Gómara, M.; Desselberger, U.; Gray, J. Infantile viral gastroenteritis: On the way to closing the diagnostic gap. J. Med. Virol. 2003, 70, 258–262. [Google Scholar] [CrossRef]
- Quintero-Ochoa, G.; Romero-Argüelles, R.; Aviles-Hernández, A.; Cejudo-Flores, M.; Calleja-García, P.; Domínguez-Gámez, M.; Cantú-Bernal, S.; Icedo-García, R.; Soñanez-Organis, J.; Rosas-Rodríguez, J.; et al. Viral agents of gastroenteritis and their correlation with clinical symptoms in rotavirus-vaccinated children. Infect. Genet. Evol. 2019, 73, 190–196. [Google Scholar] [CrossRef] [PubMed]
- Wilhelmi, I.; Roman, E.; Sanchez-Fauquier, A. Viruses causing gastroenteritis. Clin. Microbiol. Infect. 2003, 9, 247–262. [Google Scholar] [CrossRef] [Green Version]
- Eckardt, A.J. Viral gastroenteritis in adults. Recent Pat. Anti Infect. Drug Discov. 2011, 6, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Bányai, K.; Estes, M.K.; Martella, V.; Parashar, U.D. Viral gastroenteritis. Lancet 2018, 392, 175–186. [Google Scholar] [CrossRef]
- Corcoran, M.S.; Van Well, G.T.J.; Van Loo, I.H.M. Diagnosis of viral gastroenteritis in children: Interpretation of real-time PCR results and relation to clinical symptoms. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 1663–1673. [Google Scholar] [CrossRef]
- Sidoti, F.; Ritta, M.; Costa, C.; Cavallo, R. Diagnosis of viral gastroenteritis: Limits and potential of currently available procedures. J. Infect. Dev. Ctries. 2015, 9, 551–561. [Google Scholar] [CrossRef] [Green Version]
- Troeger, C.; Forouzanfar, M.; Rao, P.C.; Khalil, I.; Brown, A.; Reiner, R.C., Jr.; Fullman, N.; Thompson, R.L.; Abajobir, A.; Ahmed, M.; et al. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 2017, 17, 909–948. [Google Scholar] [CrossRef] [Green Version]
- Tate, J.E.; Burton, A.H.; Boschi-Pinto, C.; Parashar, U.D. Global, regional and national estimates of rotavirus mortality in children <5 years of age, 2000–2013. Clin. Infect. Dis. 2016, 62, 96–105. [Google Scholar] [CrossRef] [Green Version]
- Aliabadi, N.; Antoni, S.; Mwenda, J.M.; Weldegebriel, G.; Biey, J.N.M.; Cheikh, D.; Fahmy, K.; Teleb, N.; Ashmony, H.A.; Ahmed, H.; et al. Global impact of rotavirus vaccine introduction on rotavirus hospitalisations among children under 5 years of age, 2008–2016: Findings from the Global Rotavirus Surveillance Network. Lancet Glob. Health 2019, 7, 893–903. [Google Scholar] [CrossRef] [Green Version]
- Burnett, E.; Parashar, U.D.; Tate, J.E. Global impact of rotavirus vaccination on diarrhea hospitalizations and deaths among children. J. Infect. Dis. 2020, 222, 1731–1739. [Google Scholar] [CrossRef]
- Johns Hopkins Bloomberg School of Public Health VIEW-Hub. International Vaccine Access Center (IVAC). Available online: https://view-hub.org (accessed on 29 December 2020).
- Vecchio, A.L.; Liguoro, I.; Dias, J.A.; Berkley, J.A.; Boey, C.; Cohen, M.B.; Cruchet, S.; Salazar-Lindo, E.; Podder, S.; Sandhu, B.; et al. Rotavirus immunization: Global coverage and local barriers for implementation. Vaccine 2017, 35, 1637–1644. [Google Scholar] [CrossRef] [PubMed]
- Desselberger, U. Differences of rotavirus vaccine effectiveness by country: Likely causes and contributing factors. Pathogens 2017, 6, 65. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, S.M.; Hall, A.J.; Robinson, A.E.; Verhoef, L.; Premkumar, P.; Parashar, U.D.; Koopmans, M.; Lopman, B.A. Global prevalence of norovirus in cases of gastroenteritis: A systematic review and meta-analysis. Lancet Infect. Dis. 2014, 14, 725–730. [Google Scholar] [CrossRef] [Green Version]
- Bartsch, S.M.; Lopman, B.A.; Ozawa, S.; Hall, A.J.; Lee, B.Y. Global economic burden of norovirus gastroenteritis. PLoS ONE 2016, 11, 0151219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Angarone, M.P.; Sheahan, A.; Kamboj, M. Norovirus in transplantation. Curr. Infect. Dis. Rep. 2016, 18. [Google Scholar] [CrossRef]
- Green, K.Y. Norovirus infection in immunocompromised hosts. Clin. Microbiol. Infect. 2014, 20, 717–723. [Google Scholar] [CrossRef] [Green Version]
- Desai, R.; Hembree, C.D.; Handel, A.; Matthews, J.E.; Dickey, B.W.; McDonald, S.; Hall, A.J.; Parashar, U.D.; Leon, J.S.; Lopman, B. Severe outcomes are associated with genogroup 2 genotype 4 norovirus outbreaks: A systematic literature review. Clin. Infect. Dis. 2012, 55, 189–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sherwood, J.; Mendelman, P.M.; Lloyd, E.; Liu, M.; Boslego, J.; Borkowski, A.; Jackson, A.; Faix, D. Efficacy of an intramuscular bivalent norovirus GI.1/GII.4 virus-like particle vaccine candidate in healthy US adults. Vaccine 2020, 38, 6442–6449. [Google Scholar] [CrossRef] [PubMed]
- Kim, L.; Liebowitz, D.; Lin, K.; Kasparek, K.; Pasetti, M.F.; Garg, S.J.; Gottlieb, K.; Trager, G.; Tucker, S.N. Safety and immunogenicity of an oral tablet norovirus vaccine, a phase I randomized, placebo-controlled trial. JCI Insight 2018, 3, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Becker-Dreps, S.; González, F.; Bucardo, F. Sapovirus. Curr. Opin. Infect. Dis. 2020, 33, 388–397. [Google Scholar] [CrossRef] [PubMed]
- Quan, P.-L.; Wagner, T.A.; Briese, T.; Torgerson, T.R.; Hornig, M.; Tashmukhamedova, A.; Firth, C.; Palacios, G.; de Leon, A.B.; Paddock, C.D.; et al. Astrovirus encephalitis in boy with X-linked agammaglobulinemia. Emerg. Infect. Dis. 2010, 16, 918–925. [Google Scholar] [CrossRef]
- Wunderli, W.; Meerbach, A.; Güngör, T.; Berger, C.; Greiner, O.; Caduff, R.; Trkola, A.; Bossart, W.; Gerlach, D.; Schibler, M.; et al. Astrovirus infection in hospitalized infants with severe combined immunodeficiency after allogeneic hematopoietic stem cell transplantation. PLoS ONE 2011, 6, 27483. [Google Scholar] [CrossRef] [Green Version]
- Bosch, A.; Pintó, R.M.; Guix, S. Human astroviruses. Clin. Microbiol. Rev. 2014, 27, 1048–1074. [Google Scholar] [CrossRef] [Green Version]
- Radke, J.R.; Cook, J.L. Human adenovirus infections. Curr. Opin. Infect. Dis. 2018, 31, 251–256. [Google Scholar] [CrossRef]
- Koci, M.D.; Moser, L.A.; Kelley, L.A.; Larsen, D.; Brown, C.C.; Schultz-Cherry, S. Astrovirus induces diarrhea in the absence of inflammation and cell death. J. Virol. 2003, 77, 11798–11808. [Google Scholar] [CrossRef] [Green Version]
- Cortez, V.; Meliopoulos, V.A.; Karlsson, E.A.; Hargest, V.; Johnson, C.; Schultz-Cherry, S. Astrovirus biology and pathogenesis. Annu. Rev. Virol. 2017, 4, 327–348. [Google Scholar] [CrossRef]
- Saxena, K.; Blutt, S.E.; Ettayebi, K.; Zeng, X.-L.; Broughman, J.R.; Crawford, S.E.; Karandikar, U.C.; Sastri, N.P.; Conner, M.E.; Opekun, A.R.; et al. Human intestinal enteroids: A new model to study human rotavirus infection, host restriction, and pathophysiology. J. Virol. 2016, 90, 43–56. [Google Scholar] [CrossRef] [Green Version]
- Holly, M.K.; Smith, J.G. Adenovirus infection of human enteroids reveals interferon sensitivity and preferential infection of goblet cells. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [Green Version]
- Kolawole, A.; Mirabelli, C.; Hill, D.R.; Svoboda, S.A.; Janowski, A.B.; Passalacqua, K.; Rodriguez, B.N.; Dame, M.K.; Freiden, P.; Berger, R.P.; et al. Astrovirus replication in human intestinal enteroids reveals multi-cellular tropism and an intricate host innate immune landscape. PLOS Pathog. 2019, 15, e1008057. [Google Scholar] [CrossRef] [Green Version]
- Ettayebi, K.; Crawford, S.E.; Murakami, K.; Broughman, J.R.; Karandikar, U.; Tenge, V.; Neill, F.H.; Blutt, S.E.; Zeng, X.-L.; Qu, L.; et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 2016, 353, 1387–1393. [Google Scholar] [CrossRef]
- Kolawole, A.O.; Wobus, C.E. Gastrointestinal organoid technology advances studies of enteric virus biology. PLoS Pathog. 2020, 16, e1008212. [Google Scholar] [CrossRef] [Green Version]
- Zachos, N.C.; Kovbasnjuk, O.; Foulke-Abel, J.; In, J.; Blutt, S.E.; de Jonge, H.R.; Estes, M.K.; Donowitz, M. Human enteroids/colonoids and intestinal organoids functionally recapitulate normal intestinal physiology and pathophysiology. J. Biol. Chem. 2016, 291, 3759–3766. [Google Scholar] [CrossRef] [Green Version]
- Kitamoto, N.; Ramig, R.F.; Matson, D.O.; Estes, M.K. Comparative growth of different rotavirus strains in differentiated cells (MA104, HepG2, and CaCo-2). Virology 1991, 184, 729–737. [Google Scholar] [CrossRef]
- Superti, F.; Tinari, A.; Baldassarri, L.; Donelli, G. HT-29 cells: A new substrate for rotavirus growth. Arch. Virol. 1991, 116, 159–173. [Google Scholar] [CrossRef] [PubMed]
- Papa, G.; Venditti, L.; Arnoldi, F.; Schraner, E.M.; Potgieter, C.; Borodavka, A.; Eichwald, C.; Burrone, O.R. Recombinant rotaviruses rescued by reverse genetics reveal the role of NSP5 hyperphosphorylation in the assembly of viral factories. J. Virol. 2019, 94. [Google Scholar] [CrossRef] [Green Version]
- Van Dycke, J.; Arnoldi, F.; Papa, G.; Vandepoele, J.; Burrone, O.R.; Mastrangelo, E.; Tarantino, D.; Heylen, E.; Neyts, J.; Rocha-Pereira, J. A single nucleoside viral polymerase inhibitor against norovirus, rotavirus, and sapovirus-induced diarrhea. J. Infect. Dis. 2018, 218, 1753–1758. [Google Scholar] [CrossRef] [PubMed]
- Eichwald, C.; Rodriguez, J.F.; Burrone, O.R. Characterization of rotavirus NSP2/NSP5 interactions and the dynamics of viroplasm formation. J. Gen. Virol. 2004, 85, 625–634. [Google Scholar] [CrossRef]
- Finkbeiner, S.R.; Zeng, X.-L.; Utama, B.; Atmar, R.L.; Shroyer, N.; Estes, M.K. Stem cell-derived human intestinal organoids as an infection model for rotaviruses. mBio 2012, 3, e00159-12. [Google Scholar] [CrossRef] [Green Version]
- Zou, W.Y.; Blutt, S.E.; Crawford, S.E.; Ettayebi, K.; Zeng, X.-L.; Saxena, K.; Ramani, S.; Karandikar, U.C.; Zachos, N.C.; Estes, M.K. Human intestinal enteroids: New models to study gastrointestinal virus infections. In Methods in Molecular Biology; Walker, J.M., Ed.; Springer: Berlin, Germany, 2017; Volume 1576, pp. 229–247. [Google Scholar]
- Wobus, C.; Karst, S.M.; Thackray, L.B.; Chang, K.-O.; Sosnovtsev, S.V.; Belliot, G.; Krug, A.; Mackenzie, J.; Green, K.Y.; Iv, H.W.V. Replication of norovirus in cell culture reveals a tropism for dendritic cells and macrophages. PLoS Biol. 2004, 2, e432. [Google Scholar] [CrossRef] [PubMed]
- Hirneisen, K.A.; Kniel, K.E. Comparing human norovirus surrogates: Murine norovirus and Tulane virus. J. Food Prot. 2013, 76, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Bae, J.; Schwab, K.J. Evaluation of murine norovirus, feline calicivirus, poliovirus and MS2 as surrogates for human norovirus in a model of viral persistence in surface water and groundwater. Appl. Environ. Microbiol. 2008, 74, 477–484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, K.-O.; George, D.W. Interferons and ribavirin effectively inhibit Norwalk virus replication in replicon-bearing cells. J. Virol. 2007, 81, 12111–12118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rocha-Pereira, J.; Nascimento, M.S.J.; Ma, Q.; Hilgenfeld, R.; Neyts, J.; Jochmans, D. The enterovirus protease inhibitor rupintrivir exerts cross-genotypic anti-norovirus activity and clears cells from the norovirus replicon. Antimicrob. Agents Chemother. 2014, 58, 4675–4681. [Google Scholar] [CrossRef] [Green Version]
- Rocha-Pereira, J.; Jochmans, D.; Debing, Y.; Verbeken, E.; Nascimento, M.S.J.; Neyts, J. The viral polymerase inhibitor 2′-C-methylcytidine inhibits Norwalk virus replication and protects against norovirus-induced diarrhea and mortality in a mouse model. J. Virol. 2013, 87, 11798–11805. [Google Scholar] [CrossRef] [Green Version]
- Lohmann, V. HCV replicons: Overview and basic protocols. Methods Mol. Biol. 2009, 510, 145–163. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Stoltzfus, G.T.; Zhu, C.; Jung, K.; Wang, Q.; Saif, L.J. Attempts to grow human noroviruses, a sapovirus and a bovine norovirus in vitro. PLoS ONE 2018, 13, e0178157. [Google Scholar] [CrossRef]
- Todd, K.V.; Tripp, R.A. Vero cells as a mammalian cell substrate for human norovirus. Viruses 2020, 12, 439. [Google Scholar] [CrossRef] [Green Version]
- Jones, M.K.; Watanabe, M.; Zhu, S.; Graves, C.; Keyes, L.R.; Grau, K.R.; Gonzalez-Hernandez, M.B.; Iovine, N.M.; Wobus, C.; Vinje, J.; et al. Enteric bacteria promote human and mouse norovirus infection of B cells. Science 2014, 346, 755–759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Straub, T.M.; Zu Bentrup, K.H.; Coghlan, P.O.; Dohnalkova, A.; Mayer, B.K.; Bartholomew, R.A.; Valdez, C.O.; Bruckner-Lea, C.J.; Gerba, C.P.; Abbaszadegan, M.A.; et al. In vitro cell culture infectivity assay for human noroviruses. Emerg. Infect. Dis. 2007, 13, 396–403. [Google Scholar] [CrossRef]
- Herbst-Kralovetz, M.M.; Radtke, A.L.; Lay, M.K.; Hjelm, B.E.; Bolick, A.N.; Sarker, S.S.; Atmar, R.L.; Kingsley, D.H.; Arntzen, C.J.; Estes, M.K.; et al. Lack of norovirus replication and histo-blood group antigen expression in 3-dimensional intestinal epithelial cells. Emerg. Infect. Dis. 2013, 19, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Hosmillo, M.; Chaudhry, Y.; Nayak, K.; Sorgeloos, F.; Koo, B.-K.; Merenda, A.; Lillestol, R.; Drumright, L.; Zilbauer, M.; Goodfellow, I. Norovirus replication in human intestinal epithelial cells is restricted by the interferon-induced JAK/STAT signaling pathway and RNA polymerase II-mediated transcriptional responses. mBio 2020, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Costantini, V.; Morantz, E.K.; Browne, H.; Ettayebi, K.; Zeng, X.-L.; Atmar, R.L.; Estes, M.K.; Vinje, J. Human norovirus replication in human intestinal enteroids as model to evaluate virus inactivation. Emerg. Infect. Dis. 2018, 24, 1453–1464. [Google Scholar] [CrossRef] [Green Version]
- Estes, M.K.; Ettayebi, K.; Tenge, V.R.; Murakami, K.; Karandikar, U.; Lin, S.-C.; Ayyar, B.V.; Cortes-Penfield, N.W.; Haga, K.; Neill, F.H.; et al. Human norovirus cultivation in nontransformed stem cell-derived human intestinal enteroid cultures: Success and challenges. Viruses 2019, 11, 638. [Google Scholar] [CrossRef] [Green Version]
- Zhang, N.; Tan, M.; Zhong, W.; Xia, M.; Huang, P.; Jiang, X. Human intestinal organoids express histo-blood group antigens, bind norovirus VLPs, and support limited norovirus replication. Sci. Rep. 2017, 7, 12621. [Google Scholar] [CrossRef] [Green Version]
- Flynn, W.T.; Saif, L.J.; Moorhead, P.D. Pathogenesis of porcine enteric calicivirus-like virus in four-day-old gnotobiotic pigs. Am. J. Vet. Res. 1988, 49, 819–825. [Google Scholar]
- Chang, K.-O.; Sosnovtsev, S.V.; Belliot, G.; Kim, Y.; Saif, L.J.; Green, K.Y. Bile acids are essential for porcine enteric calicivirus replication in association with down-regulation of signal transducer and activator of transcription 1. Proc. Natl. Acad. Sci. USA 2004, 101, 8733–8738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, Z.; Yokoyama, M.; Chen, N.; Oka, T.; Jung, K.; Chang, K.-O.; Annamalai, T.; Wang, Q.; Saif, L.J. Mechanism of cell culture adaptation of an enteric calicivirus, the porcine sapovirus Cowden strain. J. Virol. 2016, 90, 1345–1358. [Google Scholar] [CrossRef] [Green Version]
- Takagi, H.; Oka, T.; Shimoike, T.; Saito, H.; Kobayashi, T.; Takahashi, T.; Tatsumi, C.; Kataoka, M.; Wang, Q.; Saif, L.J.; et al. Human sapovirus propagation in human cell lines supplemented with bile acids. Proc. Natl. Acad. Sci. USA 2020, 117, 32078–32085. [Google Scholar] [CrossRef]
- Brinker, J.P.; Blacklow, N.R.; Herrmann, J.E. Human astrovirus isolation and propagation in multiple cell lines. Arch. Virol. 2000, 145, 1847–1856. [Google Scholar] [CrossRef] [PubMed]
- Jogler, C.; Hoffmann, D.; Theegarten, D.; Grunwald, T.; Überla, K.; Wildner, O. Replication properties of human adenovirus in vivo and in cultures of primary cells from different animal species. J. Virol. 2006, 80, 3549–3558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cromeans, T.L.; Lu, X.; Erdman, D.D.; Humphrey, C.D.; Hill, V. Development of plaque assays for adenoviruses 40 and 41. J. Virol. Methods 2008, 151, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Tiemessen, C.; Kidd, A.H. Adenovirus type 40 and 41 growth in vitro: Host range diversity reflected by differences in patterns of DNA replication. J. Virol. 1994, 68, 1239–1244. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siqueira-Silva, J.; Yeda, F.P.; Favier, A.-L.; Mezin, P.; Silva, M.L.; Barrella, K.M.; Mehnert, D.U.; Fender, P.; Hársi, C.M. Infection kinetics of human adenovirus serotype 41 in HEK 293 cells. Memórias Inst. Oswaldo Cruz 2009, 104, 736–744. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolf, J.L.; Cukor, G.; Blacklow, N.R.; Dambrauskas, R.; Trier, J.S. Susceptibility of mice to rotavirus infection: Effects of age and administration of corticosteroids. Infect. Immun. 1981, 33, 565–574. [Google Scholar] [CrossRef] [Green Version]
- Bridger, J.C. A definition of bovine rotavirus virulence. J. Gen. Virol. 1994, 75, 2807–2812. [Google Scholar] [CrossRef] [PubMed]
- Franco, M.A.; Angel, J.; Greenberg, H. Immunity and correlates of protection for rotavirus vaccines. Vaccine 2006, 24, 2718–2731. [Google Scholar] [CrossRef]
- Souza, M.; Azevedo, M.S.P.; Jung, K.; Cheetham, S.; Saif, L.J. Pathogenesis and immune responses in gnotobiotic calves after infection with the genogroup II.4-HS66 strain of human norovirus. J. Virol. 2008, 82, 1777–1786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheetham, S.; Souza, M.; Meulia, T.; Grimes, S.; Han, M.G.; Saif, L.J. Pathogenesis of a genogroup II human norovirus in gnotobiotic pigs. J. Virol. 2006, 80, 10372–10381. [Google Scholar] [CrossRef] [Green Version]
- Ciarlet, M.; Conner, M.E.; Finegold, M.J.; Estes, M.K. Group a rotavirus infection and age-dependent diarrheal disease in rats: A new animal model to study the pathophysiology of rotavirus infection. J. Virol. 2002, 76, 41–57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- VanCott, J.L.; McNeal, M.M.; Choi, A.H.; Ward, R.L. The role of interferons in rotavirus infections and protection. J. Interf. Cytokine Res. 2003, 23, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Gozalbo-Rovira, R.; Santiso-Bellón, C.; Buesa, J.; Rubio-Del-Campo, A.; Vila-Vicent, S.; Muñoz, C.; Yebra, M.; Monedero, V.; Rodríguez-Díaz, J. Microbiota depletion promotes human rotavirus replication in an adult mouse model. bioRxiv 2021, 9, 846. [Google Scholar] [CrossRef]
- Bok, K.; Parra, G.; Mitra, T.; Abente, E.; Shaver, C.K.; Boon, D.; Engle, R.; Yu, C.; Kapikian, A.Z.; Sosnovtsev, S.V.; et al. Chimpanzees as an animal model for human norovirus infection and vaccine development. Proc. Natl. Acad. Sci. USA 2011, 108, 325–330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taube, S.; Kolawole, A.; Höhne, M.; Wilkinson, J.E.; Handley, S.A.; Perry, J.W.; Thackray, L.B.; Akkina, R.; Wobus, C.E. A mouse model for human norovirus. mBio 2013, 4, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Kolawole, A.; Gonzalez-Hernandez, M.B.; Turula, H.; Yu, C.; Elftman, M.D.; Wobus, C.E. Oral norovirus infection is blocked in mice lacking Peyer’s patches and mature M cells. J. Virol. 2016, 90, 1499–1506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Dycke, J.; Ny, A.; Conceição-Neto, N.; Maes, J.; Hosmillo, M.; Cuvry, A.; Goodfellow, I.; Nogueira, T.C.; Verbeken, E.; Matthijnssens, J.; et al. A robust human norovirus replication model in zebrafish larvae. PLoS Pathog. 2019, 15, e1008009. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Dycke, J.; Cuvry, A.; Knickmann, J.; Ny, A.; Rakers, S.; Taube, S.; de Witte, P.; Neyts, J.; Rocha-Pereira, J. Infection of zebrafish larvae with human norovirus and evaluation of the in vivo efficacy of small-molecule inhibitors. Nat. Protoc. 2021, 16, 1830–1849. [Google Scholar] [CrossRef]
- Guarin, M.; Faelens, R.; Giusti, A.; De Croze, N.; Léonard, M.; Cabooter, D.; Annaert, P.; de Witte, P.; Ny, A. Spatiotemporal imaging and pharmacokinetics of fluorescent compounds in zebrafish eleuthero-embryos after different routes of administration. Sci. Rep. 2021, 11, 1–15. [Google Scholar] [CrossRef]
- MacRae, C.A.; Peterson, R.T. Zebrafish as tools for drug discovery. Nat. Rev. Drug Discov. 2015, 14, 721–731. [Google Scholar] [CrossRef]
- Marvin, S.A.; Huerta, C.T.; Sharp, B.; Freiden, P.; Cline, T.D.; Schultz-Cherry, S. Type I interferon response limits astrovirus replication and protects against increased barrier permeability in vitro and in vivo. J. Virol. 2016, 90, 1988–1996. [Google Scholar] [CrossRef] [Green Version]
- Desselberger, U. Rotaviruses. Virus Res. 2014, 190, 75–96. [Google Scholar] [CrossRef] [Green Version]
- Mayor, S.; Parton, R.; Donaldson, J.G. Clathrin-independent pathways of endocytosis. Cold Spring Harb. Perspect. Biol. 2014, 6, a016758. [Google Scholar] [CrossRef] [Green Version]
- Martín, C.S.-S.; Lüpez, T.; Arias, C.F.; Lüpez, S. Characterization of rotavirus cell entry. J. Virol. 2004, 78, 2310–2318. [Google Scholar] [CrossRef] [Green Version]
- Jayaram, H.; Estes, M.; Prasad, B. Emerging themes in rotavirus cell entry, genome organization, transcription and replication. Virus Res. 2004, 101, 67–81. [Google Scholar] [CrossRef]
- Chen, S.; Ding, S.; Yin, Y.; Xu, L.; Li, P.; Peppelenbosch, M.P.; Pan, Q.; Wang, W. Suppression of pyrimidine biosynthesis by targeting DHODH enzyme robustly inhibits rotavirus replication. Antivir. Res. 2019, 167, 35–44. [Google Scholar] [CrossRef]
- Eichwald, C.; De Lorenzo, G.; Schraner, E.M.; Papa, G.; Bollati, M.; Swuec, P.; de Rosa, M.; Milani, M.; Mastrangelo, E.; Ackermann, M.; et al. Identification of a small molecule that compromises the structural integrity of viroplasms and rotavirus double-layered particles. J. Virol. 2018, 92, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Rossignol, J.-F. Nitazoxanide: A first-in-class broad-spectrum antiviral agent. Antivir. Res. 2014, 110, 94–103. [Google Scholar] [CrossRef] [Green Version]
- La Frazia, S.; Ciucci, A.; Arnoldi, F.; Coira, M.; Gianferretti, P.; Angelini, M.; Belardo, G.; Burrone, O.R.; Rossignol, J.-F.; Santoro, M.G. Thiazolides, a new class of antiviral agents effective against rotavirus infection, target viral morphogenesis, inhibiting viroplasm formation. J. Virol. 2013, 87, 11096–11106. [Google Scholar] [CrossRef] [Green Version]
- Rossignol, J.-F.; El-Gohary, Y.M. Nitazoxanide in the treatment of viral gastroenteritis: A randomized double-blind placebo-controlled clinical trial. Aliment. Pharmacol. Ther. 2006, 24, 1423–1430. [Google Scholar] [CrossRef]
- Teran, C.G.; Teran-Escalera, C.N.; Villarroel, P. Nitazoxanide vs. probiotics for the treatment of acute rotavirus diarrhea in children: A randomized, single-blind, controlled trial in Bolivian children. Int. J. Infect. Dis. 2009, 13, 518–523. [Google Scholar] [CrossRef] [Green Version]
- Tohmé, M.J.; Giménez, M.; Peralta, A.; Colombo, M.; Delgui, L. Ursolic acid: A novel antiviral compound inhibiting rotavirus infection in vitro. Int. J. Antimicrob. Agents 2019, 54, 601–609. [Google Scholar] [CrossRef]
- Holloway, G.; Truong, T.T.; Coulson, B.S. Rotavirus antagonizes cellular antiviral responses by inhibiting the nuclear accumulation of STAT1, STAT2 and NF-κB. J. Virol. 2009, 83, 4942–4951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Malik, Y.S.; Kobayashi, N. Therapeutics and immunoprophylaxis against noroviruses and rotaviruses: The past, present and future. Curr. Drug Metab. 2018, 19, 170–191. [Google Scholar] [CrossRef]
- Shen, Z.; He, H.; Wu, Y.; Li, J. Cyclosporin A inhibits rotavirus replication and restores interferon-beta signaling pathway in vitro and in vivo. PLoS ONE 2013, 8, e71815. [Google Scholar] [CrossRef] [Green Version]
- Chanda, S.D.; Banerjee, A.; Chawla, S.M. Cordycepin an adenosine analogue executes anti rotaviral effect by stimulating induction of type I interferon. J. Virol. Antivir. Res. 2015, 4. [Google Scholar] [CrossRef]
- Hardy, M.E.; Hendricks, J.M.; Paulson, J.M.; Faunce, N.R. 18 β-glycyrrhetinic acid inhibits rotavirus replication in culture. Virol. J. 2012, 9, 96. [Google Scholar] [CrossRef] [Green Version]
- Hendricks, J.M.; Hoffman, C.; Pascual, D.W.; Hardy, M.E. 18β-glycyrrhetinic acid delivered orally induces isolated lymphoid follicle maturation at the intestinal mucosa and attenuates rotavirus shedding. PLoS ONE 2012, 7, 49491. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.; Wang, Y.; Li, P.; Yin, Y.; Bijvelds, M.J.; de Jonge, H.R.; Peppelenbosch, M.P.; Kainov, D.E.; Pan, Q. Drug screening identifies gemcitabine inhibiting rotavirus through alteration of pyrimidine nucleotide synthesis pathway. Antivir. Res. 2020, 180, 104823. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Chen, S.; Hakim, M.S.; Wang, W.; Xu, L.; Dang, W.; Qu, C.; Verhaar, A.P.; Su, J.; Fuhler, G.M.; et al. 6-Thioguanine inhibits rotavirus replication through suppression of Rac1 GDP/GTP cycling. Antivir. Res. 2018, 156, 92–101. [Google Scholar] [CrossRef]
- Tohmé, M.J.; Delgui, L.R. Advances in the development of antiviral compounds for rotavirus infections. mBio 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Cunha, J.B.; Wobus, C.E. Select membrane proteins modulate MNV-1 infection of macrophages and dendritic cells in a cell type-specific manner. Virus Res. 2016, 222, 64–70. [Google Scholar] [CrossRef]
- Haga, K.; Fujimoto, A.; Takai-Todaka, R.; Miki, M.; Doan, Y.H.; Murakami, K.; Yokoyama, M.; Murata, K.; Nakanishi, A.; Katayama, K. Functional receptor molecules CD300lf and CD300ld within the CD300 family enable murine noroviruses to infect cells. Proc. Natl. Acad. Sci. USA 2016, 113, 6248–6255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graziano, V.R.; Wei, J.; Wilen, C.B. Norovirus attachment and entry. Viruses 2019, 11, 495. [Google Scholar] [CrossRef] [Green Version]
- Conley, M.; McElwee, M.; Azmi, L.; Gabrielsen, M.; Byron, O.; Goodfellow, I.G.; Bhella, D. Calicivirus VP2 forms a portal-like assembly following receptor engagement. Nat. Cell Biol. 2019, 565, 377–381. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.-H.; Chung, M.S.; Kim, K.H. Structure and function of caliciviral RNA polymerases. Viruses 2017, 9, 329. [Google Scholar] [CrossRef] [Green Version]
- Denison, M.R. Seeking membranes: Positive-strand RNA virus replication complexes. PLoS Biol. 2008, 6, e270. [Google Scholar] [CrossRef] [Green Version]
- Hyde, J.L.; MacKenzie, J.M. Subcellular localization of the MNV-1 ORF1 proteins and their potential roles in the formation of the MNV-1 replication complex. Virology 2010, 406, 138–148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharp, T.M.; Guix, S.; Katayama, K.; Crawford, S.E.; Estes, M.K. Inhibition of cellular protein secretion by Norwalk virus nonstructural protein p22 requires a mimic of an endoplasmic reticulum export signal. PLoS ONE 2010, 5, e13130. [Google Scholar] [CrossRef] [Green Version]
- Thorne, L.G.; Goodfellow, I. Norovirus gene expression and replication. J. Gen. Virol. 2014, 95, 278–291. [Google Scholar] [CrossRef]
- Rocha-Pereira, J.; Neyts, J.; Jochmans, D. Norovirus: Targets and tools in antiviral drug discovery. Biochem. Pharmacol. 2014, 91, 1–11. [Google Scholar] [CrossRef]
- Picarazzi, F.; Vicenti, I.; Saladini, F.; Zazzi, M.; Mori, M. Targeting the RdRp of emerging RNA viruses: The structure-based drug design challenge. Molecules 2020, 25, 5695. [Google Scholar] [CrossRef]
- Hennessy, E.P.; Green, A.D.; Connor, M.P.; Darby, R.; Macdonald, P. Norwalk virus infection and disease is associated with ABO histo–blood group type. J. Infect. Dis. 2003, 188, 176–177. [Google Scholar] [CrossRef] [Green Version]
- Rydell, G.E.; Nilsson, J.; Rodriguez-Diaz, J.; Ruvoën-Clouet, N.; Svensson, L.; Le Pendu, J.; Larson, G. Human noroviruses recognize sialyl Lewis x neoglycoprotein. Glycobiology 2008, 19, 309–320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamura, M.; Natori, K.; Kobayashi, M.; Miyamura, T.; Takeda, N. Genogroup II noroviruses efficiently bind to heparan sulfate proteoglycan associated with the cellular membrane. J. Virol. 2004, 78, 3817–3826. [Google Scholar] [CrossRef] [Green Version]
- Taube, S.; Perry, J.W.; Yetming, K.; Patel, S.P.; Auble, H.; Shu, L.; Nawar, H.F.; Lee, C.H.; Connell, T.D.; Shayman, J.A.; et al. Ganglioside-linked terminal sialic acid moieties on murine macrophages function as attachment receptors for murine noroviruses. J. Virol. 2009, 83, 4092–4101. [Google Scholar] [CrossRef] [Green Version]
- Huang, P.; Farkas, T.; Zhong, W.; Tan, M.; Thornton, S.; Morrow, A.L.; Jiang, X. Norovirus and histo-blood group antigens: Demonstration of a wide spectrum of strain specificities and classification of two major binding groups among multiple binding patterns. J. Virol. 2005, 79, 6714–6722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shirato, H.; Ogawa, S.; Ito, H.; Sato, T.; Kameyama, A.; Narimatsu, H.; Xiaofan, Z.; Miyamura, T.; Wakita, T.; Ishii, K.; et al. Noroviruses distinguish between type 1 and type 2 histo-blood group antigens for binding. J. Virol. 2008, 82, 10756–10767. [Google Scholar] [CrossRef] [Green Version]
- Almand, E.A.; Moore, M.D.; Jaykus, L.-A. Norovirus binding to ligands beyond histo-blood group antigens. Front. Microbiol. 2017, 8, 2549. [Google Scholar] [CrossRef] [Green Version]
- Farkas, T.; Yang, K.; Le Pendu, J.; Baines, J.D.; Cardin, R.D. The coxsackievirus and adenovirus receptor, a required host factor for recovirus infection, is a putative enteric calicivirus receptor. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Makino, A.; Shimojima, M.; Miyazawa, T.; Kato, K.; Tohya, Y.; Akashi, H. Junctional adhesion molecule 1 is a functional receptor for feline calicivirus. J. Virol. 2006, 80, 4482–4490. [Google Scholar] [CrossRef] [Green Version]
- Shirato, H. Norovirus and histo-blood group antigens. Jpn. J. Infect. Dis. 2011, 64, 95–103. [Google Scholar]
- Ali, E.S.; Rajapaksha, H.; Carr, J.; Petrovsky, N. Norovirus drug candidates that inhibit viral capsid attachment to human histo-blood group antigens. Antivir. Res. 2016, 133, 14–22. [Google Scholar] [CrossRef] [Green Version]
- Cao, S.; Lou, Z.; Tan, M.; Chen, Y.; Liu, Y.; Zhang, Z.; Zhang, X.C.; Jiang, X.; Li, X.; Rao, Z. Structural basis for the recognition of blood group trisaccharides by norovirus. J. Virol. 2007, 81, 5949–5957. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kitano, M.; Hosmillo, M.; Emmott, E.; Lu, J.; Goodfellow, I. Selection and characterization of rupintrivir-resistant Norwalk virus replicon cells in vitro. Antimicrob. Agents Chemother. 2018, 62, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.; Lovell, S.; Tiew, K.-C.; Mandadapu, S.R.; Alliston, K.R.; Battaile, K.; Groutas, W.C.; Chang, K.-O. Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses and coronaviruses. J. Virol. 2012, 86, 11754–11762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, K.-O.; Kim, Y.; Lovell, S.; Rathnayake, A.D.; Groutas, W.C. Antiviral drug discovery: Norovirus proteases and development of inhibitors. Viruses 2019, 11, 197. [Google Scholar] [CrossRef] [Green Version]
- Rathnayake, A.D.; Kim, Y.; Dampalla, C.S.; Nguyen, H.N.; Jesri, A.-R.M.; Kashipathy, M.M.; Lushington, G.H.; Battaile, K.P.; Lovell, S.; Chang, K.-O.; et al. Structure-guided optimization of dipeptidyl inhibitors of norovirus 3CL protease. J. Med. Chem. 2020, 63, 11945–11963. [Google Scholar] [CrossRef] [PubMed]
- Bassetto, M.; Van Dycke, J.; Neyts, J.; Brancale, A.; Rocha-Pereira, J. Targeting the viral polymerase of diarrhea-causing viruses as a strategy to develop a single broad-spectrum antiviral therapy. Viruses 2019, 11, 173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lanier, R.S.D.; Kolawole, A.; Hosmillo, M. CMX521: A nucleoside with pan-genotypic activity against norovirus. In Proceedings of the 31st International Conference on Antiviral Research, Porto, Portugal, 11–15 June 2018. [Google Scholar]
- Gardelli, C.; Attenni, B.; Donghi, M.; Meppen, M.; Pacini, B.; Harper, S.; Di Marco, A.; Fiore, F.; Giuliano, C.; Pucci, V.; et al. Phosphoramidate prodrugs of 2′-C-methylcytidine for therapy of hepatitis C virus infection. J. Med. Chem. 2009, 52, 5394–5407. [Google Scholar] [CrossRef]
- Rocha-Pereira, J.; Jochmans, D.; Neyts, J. Prophylactic treatment with the nucleoside analogue 2′-C-methylcytidine completely prevents transmission of norovirus. J. Antimicrob. Chemother. 2014, 70, 190–197. [Google Scholar] [CrossRef] [Green Version]
- Rocha-Pereira, J.; Kolawole, A.; Verbeken, E.; Wobus, C.E.; Neyts, J. Post-exposure antiviral treatment of norovirus infections effectively protects against diarrhea and reduces virus shedding in the stool in a mortality mouse model. Antivir. Res. 2016, 132, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Rocha-Pereira, J.; Van Dycke, J.; Neyts, J. Treatment with a nucleoside polymerase inhibitor reduces shedding of murine norovirus in stool to undetectable levels without emergence of drug-resistant variants. Antimicrob. Agents Chemother. 2016, 60, 1907–1911. [Google Scholar] [CrossRef] [Green Version]
- Kolawole, A.O.; Rocha-Pereira, J.; Elftman, M.D.; Neyts, J.; Wobus, C.E. Inhibition of human norovirus by a viral polymerase inhibitor in the B cell culture system and in the mouse model. Antivir. Res. 2016, 132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, Z.; Tucker, K.; Lin, X.; Kao, C.C.; Shaw, K.; Tan, H.; Symons, J.; Behera, I.; Rajwanshi, V.K.; Dyatkina, N.; et al. Biochemical evaluation of the inhibition properties of favipiravir and 2′-C-methyl-cytidine triphosphates against human and mouse norovirus RNA polymerases. Antimicrob. Agents Chemother. 2015, 59, 7504–7516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tuipulotu, D.E.; Fumian, T.M.; Netzler, N.E.; MacKenzie, J.M.; White, P.A. The adenosine analogue NITD008 has potent antiviral activity against human and animal caliciviruses. Viruses 2019, 11, 496. [Google Scholar] [CrossRef] [Green Version]
- Brown, L.-A.K.; Freemantle, N.; Breuer, J.; Dehbi, H.-M.; Chowdhury, K.; Jones, G.; Ikeji, F.; Ndoutoumou, A.; Santhirakumar, K.; Longley, N.; et al. Early antiviral treatment in outpatients with COVID-19 (FLARE): A structured summary of a study protocol for a randomised controlled trial. Trials 2021, 22, 1–4. [Google Scholar] [CrossRef]
- Ruis, C.; Brown, L.-A.; Roy, S.; Atkinson, C.; Williams, R.; Burns, S.; Yara-Romero, E.; Jacobs, M.; Goldstein, R.; Breuer, J.; et al. Mutagenesis in norovirus in response to favipiravir treatment. N. Engl. J. Med. 2018, 379, 2173–2176. [Google Scholar] [CrossRef] [Green Version]
- Arias, A.; Thorne, L.; Goodfellow, I. Favipiravir elicits antiviral mutagenesis during virus replication in vivo. eLife 2014, 3, e03679. [Google Scholar] [CrossRef]
- Rocha-Pereira, J.; Jochmans, D.; Dallmeier, K.; Leyssen, P.; Nascimento, M.; Neyts, J. Favipiravir (T-705) inhibits in vitro norovirus replication. Biochem. Biophys. Res. Commun. 2012, 424, 777–780. [Google Scholar] [CrossRef]
- Leyssen, P.; Balzarini, J.; De Clercq, E.; Neyts, J. The predominant mechanism by which ribavirin exerts its antiviral activity in vitro against flaviviruses and paramyxoviruses is mediated by inhibition of IMP dehydrogenase. J. Virol. 2005, 79, 1943–1947. [Google Scholar] [CrossRef] [Green Version]
- Eltahla, A.A.; Lim, K.L.; Eden, J.-S.; Kelly, A.G.; MacKenzie, J.M.; White, P.A. Nonnucleoside inhibitors of norovirus RNA polymerase: Scaffolds for rational drug design. Antimicrob. Agents Chemother. 2014, 58, 3115–3123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferla, S.; Netzler, N.E.; Ferla, S.; Veronese, S.; Tuipulotu, D.E.; Guccione, S.; Brancale, A.; White, P.; Bassetto, M. In silico screening for human norovirus antivirals reveals a novel non-nucleoside inhibitor of the viral polymerase. Sci. Rep. 2018, 8, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Mastrangelo, E.; Pezzullo, M.; Tarantino, D.; Petazzi, R.; Germani, F.; Kramer, D.; Robel, I.; Rohayem, J.; Bolognesi, M.; Milani, M. Structure-based inhibition of norovirus RNA-dependent RNA polymerases. J. Mol. Biol. 2012, 419, 198–210. [Google Scholar] [CrossRef]
- Tarantino, D.; Pezzullo, M.; Mastrangelo, E.; Croci, R.; Rohayem, J.; Robel, I.; Bolognesi, M.; Milani, M. Naphthalene-sulfonate inhibitors of human norovirus RNA-dependent RNA-polymerase. Antivir. Res. 2014, 102, 23–28. [Google Scholar] [CrossRef] [Green Version]
- Netzler, N.E.; Tuipulotu, D.E.; Eltahla, A.A.; Lun, J.H.; Ferla, S.; Brancale, A.; Urakova, N.; Frese, M.; Strive, T.; Mackenzie, J.M.; et al. Broad-spectrum non-nucleoside inhibitors for caliciviruses. Antivir. Res. 2017, 146, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Croci, R.; Tarantino, D.; Milani, M.; Pezzullo, M.; Rohayem, J.; Bolognesi, M.; Mastrangelo, E. PPNDS inhibits murine norovirus RNA-dependent RNA-polymerase mimicking two RNA stacking bases. FEBS Lett. 2014, 588, 1720–1725. [Google Scholar] [CrossRef]
- Croci, R.; Pezzullo, M.; Tarantino, D.; Milani, M.; Tsay, S.-C.; Sureshbabu, R.; Tsai, Y.-J.; Mastrangelo, E.; Rohayem, J.; Bolognesi, M.; et al. Structural bases of norovirus RNA dependent RNA polymerase inhibition by novel suramin-related compounds. PLoS ONE 2014, 9, 91765. [Google Scholar] [CrossRef] [PubMed]
- Maloney, N.S.; Thackray, L.B.; Goel, G.; Hwang, S.; Duan, E.; Vachharajani, P.; Xavier, R.; Virgin, H.W. Essential cell-autonomous role for interferon (IFN) regulatory factor 1 in IFN—Mediated inhibition of norovirus replication in macrophages. J. Virol. 2012, 86, 12655–12664. [Google Scholar] [CrossRef] [Green Version]
- Thackray, L.B.; Duan, E.; Lazear, H.; Kambal, A.; Schreiber, R.D.; Diamond, M.S.; Virgin, H.W. Critical role for interferon regulatory factor 3 (IRF-3) and IRF-7 in type I interferon-mediated control of murine norovirus replication. J. Virol. 2012, 86, 13515–13523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Changotra, H.; Jia, Y.; Moore, T.N.; Liu, G.; Kahan, S.M.; Sosnovtsev, S.V.; Karst, S.M. Type I and type II Interferons inhibit the translation of murine norovirus proteins. J. Virol. 2009, 83, 5683–5692. [Google Scholar] [CrossRef] [Green Version]
- Rocha-Pereira, J.; Jacobs, S.; Noppen, S.; Verbeken, E.; Michiels, T.; Neyts, J. Interferon lambda (IFN-λ) efficiently blocks norovirus transmission in a mouse model. Antivir. Res. 2018, 149, 7–15. [Google Scholar] [CrossRef]
- Arthur, S.E.; Sorgeloos, F.; Hosmillo, M.; Goodfellow, I.G. Epigenetic suppression of interferon lambda receptor expression leads to enhanced human norovirus replication in vitro. mBio 2019, 10, e02155-19. [Google Scholar] [CrossRef] [Green Version]
- Tuipulotu, D.E.; Netzler, N.E.; Lun, J.H.; Mackenzie, J.M.; White, P.A. TLR7 agonists display potent antiviral effects against norovirus infection via innate stimulation. Antimicrob. Agents Chemother. 2018, 62, e02417-17. [Google Scholar] [CrossRef] [Green Version]
- Lee, W.; Kim, M.; Lee, S.-H.; Jung, H.-G.; Oh, J.-W. Prophylactic efficacy of orally administered Bacillus poly-γ-glutamic acid, a non-LPS TLR4 ligand, against norovirus infection in mice. Sci. Rep. 2018, 8, 8667. [Google Scholar] [CrossRef]
- Vashist, S.; Urena, L.; Gonzalez-Hernandez, M.B.; Choi, J.; De Rougemont, A.; Rocha-Pereira, J.; Neyts, J.; Hwang, S.; Wobus, C.; Goodfellow, I. Molecular chaperone Hsp90 is a therapeutic target for noroviruses. J. Virol. 2015, 89, 6352–6363. [Google Scholar] [CrossRef] [Green Version]
- Walker, N.F.; Youkee, D.; Brown, C.S.; Lado, M.; Johnson, O. Management of Ebola virus disease: Is environmental decontamination effective? J. Infect. Dis. 2016, 215, 485–486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graci, J.D.; Cameron, C.E. Mechanisms of action of ribavirin against distinct viruses. Rev. Med. Virol. 2005, 16, 37–48. [Google Scholar] [CrossRef]
- Furuta, Y.; Komeno, T.; Nakamura, T. Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proc. Jpn. Acad. Ser. B 2017, 93, 449–463. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Janowski, A.B.; Dudley, H.; Wang, D. Antiviral activity of ribavirin and favipiravir against human astroviruses. J. Clin. Virol. 2020, 123, 104247. [Google Scholar] [CrossRef]
- Hargest, V.; Sharp, B.; Livingston, B.; Cortez, V.; Schultz-Cherry, S. Astrovirus replication is inhibited by nitazoxanide in vitro and in vivo. J. Virol. 2020, 94, 1–8. [Google Scholar] [CrossRef]
- De Clercq, E. Antivirals and antiviral strategies. Nat. Rev. Genet. 2004, 2, 704–720. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M.S.; El-Hameed, R.H.A.; Sayed, A.I.; Soror, S.H. Novel antiviral compounds against gastroenteric viral infections. Arch. Pharm. 2015, 348, 194–205. [Google Scholar] [CrossRef]
- Bordigoni, P.; Carret, A.-S.; Venard, V.; Witz, F.; Le Faou, A. Treatment of adenovirus infections in patients undergoing allogeneic hematopoietic stem cell transplantation. Clin. Infect. Dis. 2001, 32, 1290–1297. [Google Scholar] [CrossRef] [PubMed]
- Ljungman, P.; Ribaud, P.; Eyrich, M.; Matthes-Martin, S.; Einsele, H.; Bleakley, M.; Machaczka, M.; Bierings, M.B.; Bosi, A.; Gratecos, N.; et al. Cidofovir for adenovirus infections after allogeneic hematopoietic stem cell transplantation: A survey by the infectious diseases working party of the European Group for blood and marrow transplantation. Bone Marrow Transplant. 2003, 31, 481–486. [Google Scholar] [CrossRef] [Green Version]
- Hoffman, J.A.; Shah, A.J.; Ross, L.A.; Kapoor, N. Adenoviral infections and a prospective trial of cidofovir in pediatric hematopoietic stem cell transplantation. Biol. Blood Marrow Transplant. 2001, 7, 388–394. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yusuf, U.; Hale, G.A.; Carr, J.; Gu, Z.; Benaim, E.; Woodard, P.; Kasow, K.A.; Horwitz, E.M.; Leung, W.; Srivastava, D.K.; et al. Cidofovir for the treatment of adenoviral infection in pediatric hematopoietic stem cell transplant patients. Transplantation 2006, 81, 1398–1404. [Google Scholar] [CrossRef]
- Alcamo, A.M.; Wolf, M.S.; Alessi, L.J.; Chong, H.J.; Green, M.; Williams, J.V.; Simon, D.W. Successful use of cidofovir in an immunocompetent child with severe adenoviral sepsis. Pediatrics 2020, 145, e20191632. [Google Scholar] [CrossRef]
- Kajon, A.E.; Lynch, J.P. Adenovirus: Epidemiology, global spread of novel serotypes, and advances in treatment and prevention. Semin. Respir. Crit. Care Med. 2016, 37, 586–602. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lopez, S.M.; Michaels, M.G.; Green, M. Adenovirus infection in pediatric transplant recipients: Are effective antiviral agents coming our way? Curr. Opin. Organ. Transplant. 2018, 23, 395–399. [Google Scholar] [CrossRef]
- Hartline, C.B.; Gustin, K.M.; Wan, W.B.; Ciesla, S.L.; Beadle, J.R.; Hostetler, K.; Kern, E.R. Ether lipid-ester prodrugs of acyclic nucleoside phosphonates: Activity against adenovirus replication in vitro. J. Infect. Dis. 2005, 191, 396–399. [Google Scholar] [CrossRef] [Green Version]
- Grimley, M.S.; Chemaly, R.F.; Englund, J.A.; Kurtzberg, J.; Chittick, G.; Brundage, T.M.; Bae, A.; Morrison, M.E.; Prasad, V.K. Brincidofovir for asymptomatic adenovirus viremia in pediatric and adult allogeneic hematopoietic cell transplant recipients: A randomized placebo-controlled phase II trial. Biol. Blood Marrow Transplant. 2017, 23, 512–521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naesens, L.; Lenaerts, L.; Andrei, G.; Snoeck, R.; Van Beers, D.; Holý, A.; Balzarini, J.; De Clercq, E. Antiadenovirus activities of several classes of nucleoside and nucleotide analogues. Antimicrob. Agents Chemother. 2005, 49, 1010–1016. [Google Scholar] [CrossRef] [Green Version]
- Toth, K.; Hussein, I.T.M.; Tollefson, A.E.; Ying, B.; Spencer, J.F.; Eagar, J.; James, S.H.; Prichard, M.N.; Wold, W.S.M.; Bowlin, T.L. Filociclovir is a potent in vitro and in vivo inhibitor of human adenoviruses. Antimicrob. Agents Chemother. 2020, 64. [Google Scholar] [CrossRef] [PubMed]
- Dodge, M.J.; MacNeil, K.M.; Tessier, T.M.; Weinberg, J.B.; Mymryk, J.S. Emerging antiviral therapeutics for human adenovirus infection: Recent developments and novel strategies. Antivir. Res. 2021, 188, 105034. [Google Scholar] [CrossRef] [PubMed]
| Antiviral Compound | Class of Inhibitor | Stage of Viral Life Cycle | Molecular Target | Mechanism of Action |
|---|---|---|---|---|
| 2CMC | Nucleoside analogue (cytidine) | Genome replication | RdRp | Direct inhibition of viral RdRp acting as final chain terminator |
| 7DMA | Nucleoside analogue (adenosine) | |||
| Brequinar | Quinolinecarboxylic acid | DHODH | Blocking de novo pyrimidine biosynthesis by inhibition of DHODH | |
| Leflunomide | Isoxazole derivative | |||
| ML-60218 | Indazole sulfonamide | Viroplasm formation | VPs; DLPs | Disruption of VPs and hampering formation of new VPs; Induction of structural damage into DLPs hampering VP6 formation |
| Nitazoxanide | Thiazolide | VPs | Inhibition of VP7 maturation, hampering VP formation; Interference in viral morphogenesis | |
| Ursolic acid | Triterpenoid | Lipid droplets | Decreases lipid droplets availability required for VP formation | |
| Cyclosporine | Cyclic peptide | Host factor | IFN signalling pathway | Increase expression of type I IFN |
| Cordycepin | Adenosine analogue | |||
| 18βGRA | Aglycone | PI3K/Akt pathway | Modulation of PI3K/Akt pathway, increasing cell apoptosis and preventing virus replication |
| Antiviral Compound | Class of Inhibitor | Stage of Viral Life Cycle | Molecular Target | Mechanism of Action |
|---|---|---|---|---|
| Citrate | Carbohydrate analogue | Viral entry | Viral capsid | Blocks binding of P domain of viral capsid to HBGAs |
| Rupintrivir | Peptidomimetic inhibitor | Translation | Viral protease | Inhibition of NoV 3CLpro blocking the cleavage of NS polyprotein, essential for production of viral progeny |
| CMX521 | Purine nucleoside | Genome replication | RdRp | Direct inhibition of viral RdRp acting as final chain terminator |
| 2CMC | Nucleoside analogue (cytidine) | |||
| 7DMA | Nucleoside analogue (adenosine) | |||
| NITD008 | Nucleoside analogue (adenosine) | |||
| Favipiravir | Nucleoside analogue (pyrazine) | Direct inhibition of viral RdRp by competition with ATP and GTP at the initiation and elongation steps; Lethal mutagenesis | ||
| Ribavirin | Nucleoside analogue (guanosine) | Inhibition of viral RdRp by depletion of intracellular GTP pools | ||
| NAF2 | Non-nucleoside analogue | Allosteric inhibition of RdRp | ||
| Suramin | ||||
| PPDS | ||||
| NF023 | ||||
| Resiquimod | TLR agonist | Host factor | TLR7 | Stimulation of IFN production by TLR7 agonism |
| γ-PGA | TLR4 | |||
| 17-DMAG | - | Hsp90 | Inhibition of Hsp90 activity | |
| Nitazoxanide | Thiazolide | Other | Not known | Not known |
| Virus | Antiviral Compound | Class of Inhibitor | Stage of Viral Life Cycle | Molecular Target | Mechanism of Action |
|---|---|---|---|---|---|
| SaV | 2CMC | Nucleoside analogue (cytidine) | Genome replication | RdRp | Direct inhibition of viral RdRp acting as final chain terminator |
| 7DMA | Nucleoside analogue (adenosine) | ||||
| AstV | Ribavirin | Nucleoside analogue (guanosine) | Genome replication | RdRp | Inhibition of viral RdRp |
| Favipiravir | Nucleoside analogue (pyrazine) | ||||
| Nitazoxanide | Thiazolide | Other | Not known | Possible induction of IFN response by activation of protein kinase R | |
| AdV | Cidofovir | Nucleoside analogue (cytosine) | Genome replication | Viral DNA polymerase | Direct inhibition of viral DNA polymerase acting as final chain terminator |
| Brincidofovir | Nucleoside analogue (cytosine) | ||||
| Compound 7f and 12a | Pyrrolopyrimidine derivatives |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos-Ferreira, N.; Van Dycke, J.; Neyts, J.; Rocha-Pereira, J. Current and Future Antiviral Strategies to Tackle Gastrointestinal Viral Infections. Microorganisms 2021, 9, 1599. https://doi.org/10.3390/microorganisms9081599
Santos-Ferreira N, Van Dycke J, Neyts J, Rocha-Pereira J. Current and Future Antiviral Strategies to Tackle Gastrointestinal Viral Infections. Microorganisms. 2021; 9(8):1599. https://doi.org/10.3390/microorganisms9081599
Chicago/Turabian StyleSantos-Ferreira, Nanci, Jana Van Dycke, Johan Neyts, and Joana Rocha-Pereira. 2021. "Current and Future Antiviral Strategies to Tackle Gastrointestinal Viral Infections" Microorganisms 9, no. 8: 1599. https://doi.org/10.3390/microorganisms9081599
APA StyleSantos-Ferreira, N., Van Dycke, J., Neyts, J., & Rocha-Pereira, J. (2021). Current and Future Antiviral Strategies to Tackle Gastrointestinal Viral Infections. Microorganisms, 9(8), 1599. https://doi.org/10.3390/microorganisms9081599







