Surface Waters and Urban Brown Rats as Potential Sources of Human-Infective Cryptosporidium and Giardia in Vienna, Austria
Abstract
1. Introduction
2. Materials and Methods
2.1. Trapping
2.2. Cryptosporidium and Giardia Quantification in Water Samples
2.3. DNA Extraction
2.4. Identification of Cryptosporidiumand Giardia Species and Genotypes
2.5. Statistical Analysis and Mapping
2.6. Ethical Statement
3. Results
3.1. Trapping
3.2. Prevalence and Identification of Protozoa in Rat Faeces
3.2.1. Cryptosporidium
3.2.2. Eimeria
3.2.3. Giardia
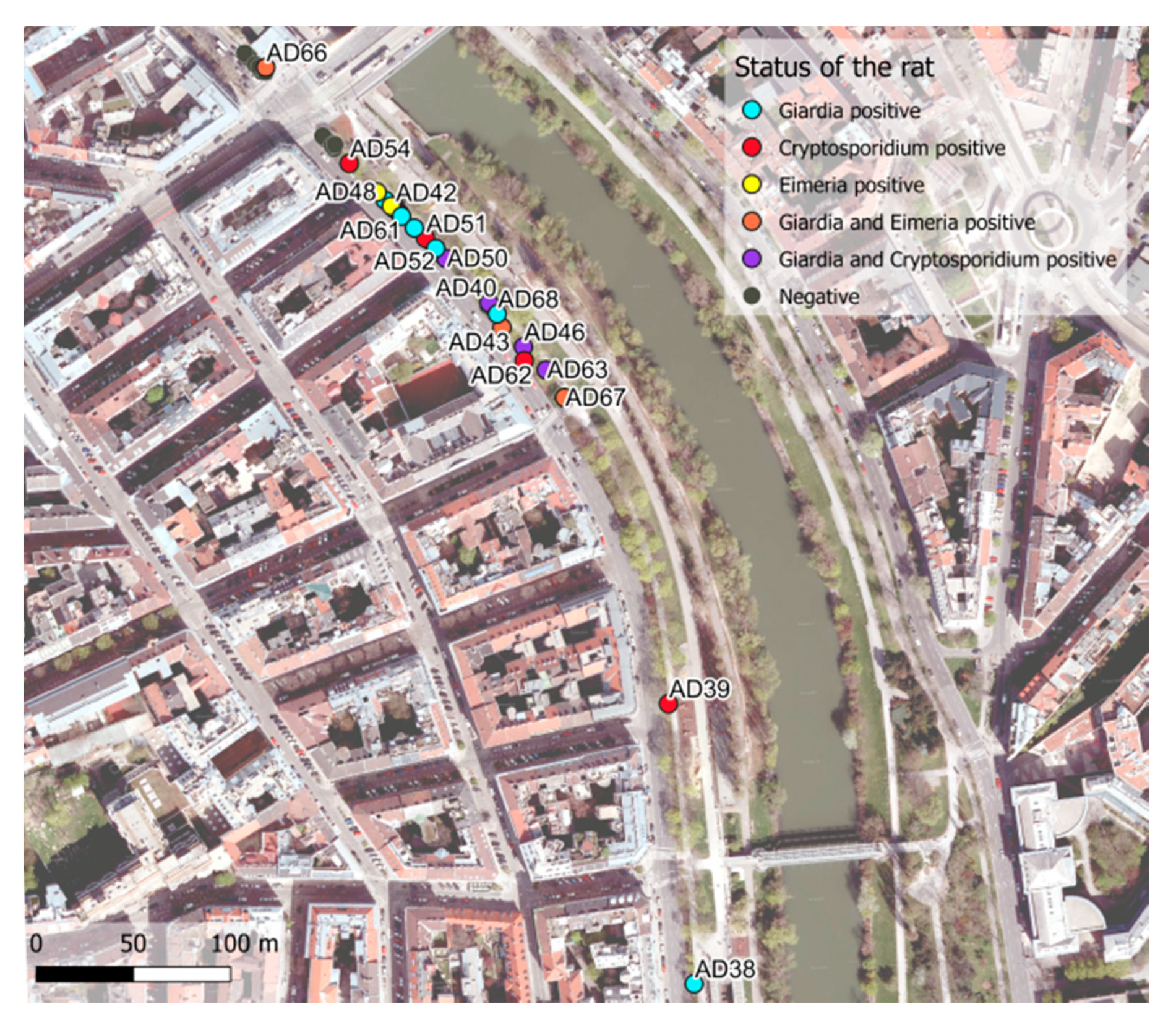
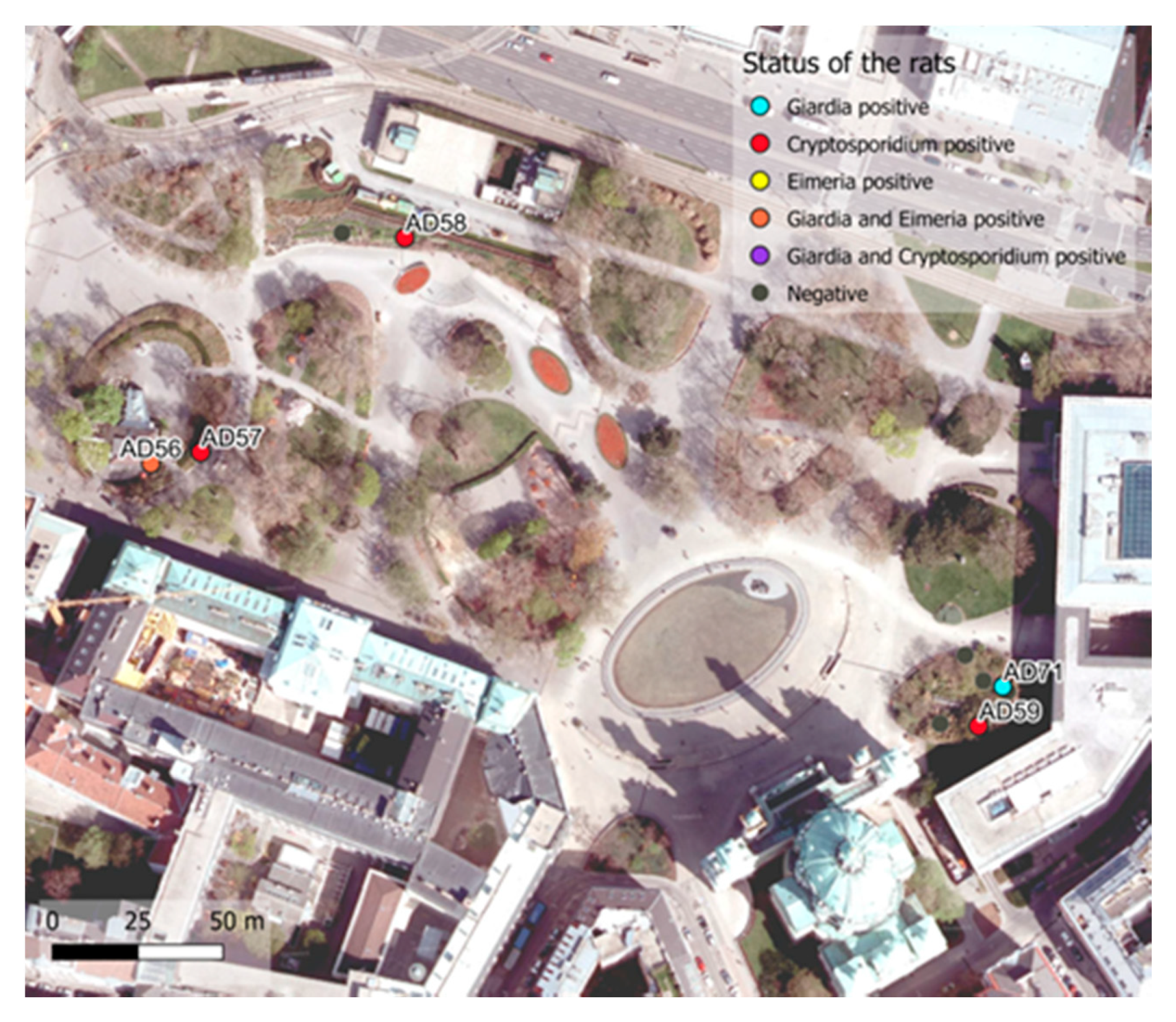
3.3. Land-Use Data
3.4. Predictors of Cryptosporidium, Eimeria and Giardia Shedding in Urban Rats
3.5. Prevalence and Identification of Cryptosporidium and Giardia in Surface Water Samples
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kirschner, A.K.T.; Reischer, G.H.; Jakwerth, S.; Savio, D.; Ixenmaier, S.; Toth, E.; Sommer, R.; Mach, R.L.; Linke, R.; Eiler, A.; et al. Multiparametric monitoring of microbial faecal pollution reveals the dominance of human contamination along the whole Danube River. Water Res. 2017, 124, 543–555. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Wang, X.; Yang, R.; Zhao, W.; Li, N.; Guo, Y.; Xiao, L.; Feng, Y. Molecular characterization of the waterborne pathogens Cryptosporidium spp., Giardia duodenalis, Enterocytozoon bieneusi, Cyclospora cayetanensis and Eimeria spp. in wastewater and sewage in Guangzhou, China. Parasites Vectors 2021, 14, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Efstratiou, A.; Ongerth, J.E.; Karanis, P. Waterborne transmission of protozoan parasites: Review of worldwide outbreaks—An update 2011–2016. Water Res. 2017, 114, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.C.A.; Ash, A. Molecular epidemiology of Giardia and Cryptosporidium infections. Infect. Genet. Evol. 2016, 40, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Capewell, P.; Krumrie, S.; Katzer, F.; Alexander, C.L.; Weir, W. Molecular Epidemiology of Giardia Infections in the Genomic Era. Trends Parasitol. 2021, 37, 142–153. [Google Scholar] [CrossRef]
- Plutzer, J.; Lassen, B.; Jokelainen, P.; Djurković-Djaković, O.; Kucsera, I.; Dorbek-Kolin, E.; Šoba, B.; Sréter, T.; Imre, K.; Omeragić, J.; et al. Review of Cryptosporidium and Giardia in the eastern part of Europe, 2016. Eurosurveillance 2018, 23, 2011–2015. [Google Scholar] [CrossRef]
- Innes, E.A.; Chalmers, R.M.; Wells, B.; Pawlowic, M.C. A One Health Approach to Tackle Cryptosporidiosis. Trends Parasitol. 2020, 36, 290–303. [Google Scholar] [CrossRef]
- Feng, Y.; Xiao, L. Zoonotic potential and molecular epidemiology of Giardia species and Giardiasis. Clin. Microbiol. Rev. 2011, 24, 110–140. [Google Scholar] [CrossRef]
- Sulaiman, I.M.; Fayer, R.; Bern, C.; Gilman, R.H.; Trout, J.M.; Schantz, P.M.; Das, P.; Lal, A.A.; Xiao, L. Triosephosphate Isomerase Gene Characterization and Potential Zoonotic Transmission of Giardia duodenalis. Emerg. Infect. Dis. 2003. [Google Scholar] [CrossRef]
- Baldursson, S.; Karanis, P. Waterborne transmission of protozoan parasites: Review of worldwide outbreaks—An update 2004–2010. Water Res. 2011, 45, 6603–6614. [Google Scholar] [CrossRef]
- Feng, Y.; Ryan, U.M.; Xiao, L. Genetic Diversity and Population Structure of Cryptosporidium. Trends Parasitol. 2018, 34, 997–1011. [Google Scholar] [CrossRef] [PubMed]
- Zahedi, A.; Ryan, U. Cryptosporidium—An update with an emphasis on foodborne and waterborne transmission. Res. Vet. Sci. 2020, 132, 500–512. [Google Scholar] [CrossRef] [PubMed]
- Heyworth, M.F. Giardia duodenalis genetic assemblages and hosts. Parasite 2016, 23. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.C.A.; Monis, P. Giardia-From Genome to Proteome, 1st ed.; Elsevier Ltd.: Amsterdam, The Netherlands, 2012; Volume 78, ISBN 9780123943033. [Google Scholar]
- Ryan, U.; Cacciò, S.M. Zoonotic potential of Giardia. Int. J. Parasitol. 2013, 43, 943–956. [Google Scholar] [CrossRef] [PubMed]
- The World’s Cities in 2018—Data Booklet; United Nations: New York, NY, USA, 2018.
- Feng, A.Y.T.; Himsworth, C.G. The secret life of the city rat: A review of the ecology of urban Norway and black rats (Rattus norvegicus and Rattus rattus). Urban Ecosyst. 2014, 17, 149–162. [Google Scholar] [CrossRef]
- Koehler, A.V.; Wang, T.; Haydon, S.R.; Gasser, R.B. Cryptosporidium viatorum from the native Australian swamp rat Rattus lutreolus—An emerging zoonotic pathogen? Int. J. Parasitol. Parasites Wildl. 2018, 7, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Hassell, J.M.; Begon, M.; Ward, M.J.; Fèvre, E.M. Urbanization and Disease Emergence: Dynamics at the Wildlife–Livestock–Human Interface. Trends Ecol. Evol. 2017, 32, 55–67. [Google Scholar] [CrossRef] [PubMed]
- Easterbrook, J.D.; Kaplan, J.B.; Vanasco, N.B.; Reeves, W.K.; Purcell, R.H.; Kosoy, M.Y.; Glass, G.E.; Watson, J.; Klein, S.L. A survey of zoonotic pathogens carried by Norway rats in Baltimore, Maryland, USA. Epidemiol. Infect. 2007, 135, 1192–1199. [Google Scholar] [CrossRef]
- Meerburg, B.G.; Singleton, G.R.; Kijlstra, A. Rodent-borne diseases and their risks for public health Rodent-borne diseases and their risks for public health. Crit. Rev. Microbiol. 2009, 35, 221–270. [Google Scholar] [CrossRef]
- Himsworth, C.G.; Parsons, K.L.; Jardine, C.; Patrick, D.M. Rats, cities, people, and pathogens: A systematic review and narrative synthesis of literature regarding the ecology of rat-associated zoonoses in urban centers. Vector-Borne Zoonotic Dis. 2013, 13, 349–359. [Google Scholar] [CrossRef]
- Ng-Hublin, J.S.Y.; Singleton, G.R.; Ryan, U. Molecular characterization of Cryptosporidium spp. from wild rats and mice from rural communities in the Philippines. Infect. Genet. Evol. 2013, 16, 5–12. [Google Scholar] [CrossRef] [PubMed]
- Foo, C.; Farrell, J.; Boxell, A.; Robertson, I.; Ryan, U.M. Novel Cryptosporidium genotype in wild Australian mice (Mus domesticus). Appl. Environ. Microbiol. 2007, 73, 7693–7696. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Helmy, Y.A.; Spierling, N.G.; Schmidt, S.; Rosenfeld, U.M.; Reil, D.; Imholt, C.; Jacob, J.; Ulrich, R.G.; Aebischer, T.; Klotz, C. Occurrence and distribution of Giardia species in wild rodents in Germany. Parasites Vectors 2018, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Wang, R.; Zhao, W.; Qi, M.; Zhao, J.; Zhang, L.; Li, J.; Liu, A. Genotyping and subtyping of Giardia and Cryptosporidium isolates from commensal rodents in China. Parasitology 2015, 142, 800–806. [Google Scholar] [CrossRef] [PubMed]
- Desvars-Larrive, A.; Smith, S.; Munimanda, G.; Bourhy, P.; Waigner, T.; Odom, M.; Gliga, D.S.; Walzer, C. Prevalence and risk factors of Leptospira infection in urban brown rats (Rattus norvegicus), Vienna, Austria. Urban Ecosyst. 2020, 23, 775–784. [Google Scholar] [CrossRef]
- Vadell, M.V.; Cavia, R.; Suárez, O.V. Abundance, age structure and reproductive patterns of Rattus norvegicus and Mus musculus in two areas of the city of Buenos Aires. Int. J. Pest Manag. 2010, 56, 327–336. [Google Scholar] [CrossRef]
- ISO 15553:2006, Water Quality—Isolation and Identification of Cryptosporidium Oocysts and Giardia Cysts from Water; International Organization for Standardization: Geneva, Switzerland, 2006.
- Stevenson, M.E.; Blaschke, A.P.; Toze, S.; Sidhu, J.P.S.; Ahmed, W.; van Driezum, I.H.; Sommer, R.; Kirschner, A.K.T.; Cervero-Aragó, S.; Farnleitner, A.H.; et al. Biotin- and glycoprotein-coated microspheres as surrogates for studying filtration removal of Cryptosporidium parvum in a granular limestone aquifer medium. Appl. Environ. Microbiol. 2015, 81. [Google Scholar] [CrossRef]
- Ryan, U.; Xiao, L.; Read, C.; Zhou, L.; Lal, A.A.; Pavlasek, I. Identification of novel Cryptosporidium genotypes from the Czech Republic. Appl. Environ. Microbiol. 2003, 69, 4302–4307. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Kváč, M.; Vlnatá, G.; Ježková, J.; Horčičková, M.; Konečný, R.; Hlásková, L.; McEvoy, J.; Sak, B. Cryptosporidium occultus sp. n. (Apicomplexa: Cryptosporidiidae) in rats. Eur. J. Protistol. 2018, 63, 96–104. [Google Scholar] [CrossRef]
- Ježková, J.; Prediger, J.; Holubová, N.; Sak, B.; Konečný, R.; Feng, Y.; Xiao, L.; Rost, M.; McEvoy, J.; Kváč, M. Cryptosporidium ratti n. sp. (Apicomplexa: Cryptosporidiidae) and genetic diversity of Cryptosporidium spp. In brown rats (Rattus norvegicus) in the Czech Republic. Parasitology 2021, 148, 84–97. [Google Scholar] [CrossRef]
- Angley, L.P.; Combs, M.; Firth, C.; Frye, M.J.; Lipkin, I.; Richardson, J.L.; Munshi-South, J. Spatial variation in the parasite communities and genomic structure of urban rats in New York City. Zoonoses Public Health 2018, 65, e113–e123. [Google Scholar] [CrossRef]
- Himsworth, C.G.; Bidulka, J.; Parsons, K.L.; Feng, A.Y.T.; Tang, P.; Jardine, C.M.; Kerr, T.; Mak, S.; Robinson, J.; Patrick, D.M. Ecology of Leptospira interrogans in Norway Rats (Rattus norvegicus) in an Inner-City Neighborhood of Vancouver, Canada. PLoS Negl. Trop. Dis. 2013, 7. [Google Scholar] [CrossRef]
- Himsworth, C.G.; Bai, Y.; Kosoy, M.Y.; Wood, H.; Dibernardo, A.; Lindsay, R.; Bidulka, J.; Tang, P.; Jardine, C.; Patrick, D. An investigation of Bartonella spp., Rickettsia typhi, and seoul hantavirus in rats (Rattus spp.) from an inner-city neighborhood of Vancouver, Canada: Is pathogen presence a reflection of global and local rat population structure? Vector-Borne Zoonotic Dis. 2015, 15, 21–26. [Google Scholar] [CrossRef]
- Jarquín-Díaz, V.H.; Balard, A.; Mácová, A.; Jost, J.; Roth von Szepesbéla, T.; Berktold, K.; Tank, S.; Kvičerová, J.; Heitlinger, E. Generalist Eimeria species in rodents: Multilocus analyses indicate inadequate resolution of established markers. Ecol. Evol. 2020, 10, 1378–1389. [Google Scholar] [CrossRef]
- Šlapeta, J.R.; Modrý, D.; Votýpka, J.; Jirku, M.; Oborník, M.; Lukeš, J.; Koudela, B. Eimeria telekii n.sp. (Apicomplexa: Coccidia) from Lemniscomys striatus (Rodentia: Muridae): Morphology, pathology and phylogeny. Parasitology 2001, 122, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Jarquín-Díaz, V.H.; Balard, A.; Jost, J.; Kraft, J.; Dikmen, M.N.; Kvičerová, J.; Heitlinger, E. Detection and quantification of house mouse Eimeria at the species level—Challenges and solutions for the assessment of coccidia in wildlife. Int. J. Parasitol. Parasites Wildl. 2019, 10, 29–40. [Google Scholar] [CrossRef]
- Lebbad, M.; Mattsson, J.G.; Christensson, B.; Ljungström, B.; Backhans, A.; Andersson, J.O.; Svärd, S.G. From mouse to moose: Multilocus genotyping of Giardia isolates from various animal species. Vet. Parasitol. 2010, 168, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lang, P.; Huang, M.; Jing, B.; Karim, M.R.; Chao, L.; Wang, Z.; Lv, Y.; Li, J.; Qi, M. Molecular characterization of Cryptosporidium spp. and Giardia duodenalis in experimental rats in China. Parasitol. Int. 2020, 77, 102127. [Google Scholar] [CrossRef] [PubMed]
- Cacciò, S.M.; Thompson, R.C.A.; McLauchlin, J.; Smith, H.V. Unravelling Cryptosporidium and Giardia epidemiology. Trends Parasitol. 2005, 21, 430–437. [Google Scholar] [CrossRef]
- Peng, J.J.; Zou, Y.; Li, Z.X.; Liang, Q.L.; Song, H.Y.; Li, T.S.; Ma, Y.Y.; Zhu, X.Q.; Zhou, D.H. Prevalence and multilocus genotyping of Giardia duodenalis in Tan sheep (Ovis aries) in northwestern China. Parasitol. Int. 2020, 77, 102126. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Ortega, Y.; Cama, V.; Terrell, J.; Xiao, L. High intragenotypic diversity of Giardia duodenalis in dairy cattle on three farms. Parasitol. Res. 2008, 103, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Bonhomme, J.; Le Goff, L.; Lemée, V.; Gargala, G.; Ballet, J.J.; Favennec, L. Limitations of tpi and bg genes sub-genotyping for characterization of human Giardia duodenalis isolates. Parasitol. Int. 2011, 60, 327–330. [Google Scholar] [CrossRef]
- Li, F.; Wang, R.; Guo, Y.; Li, N.; Feng, Y.; Xiao, L. Zoonotic potential of Enterocytozoon bieneusi and Giardia duodenalis in horses and donkeys in northern China. Parasitol. Res. 2020, 119, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Lebbad, M.; Petersson, I.; Karlsson, L.; Botero-Kleiven, S.; Andersson, J.O.; Svenungsson, B.; Svärd, S.G. Multilocus genotyping of human Giardia isolates suggests limited zoonotic transmission and association between assemblage B and flatulence in children. PLoS Negl. Trop. Dis. 2011, 5, e1262. [Google Scholar] [CrossRef] [PubMed]
- Roellig, D.M.; Salzer, J.S.; Carroll, D.S.; Ritter, J.M.; Drew, C.; Gallardo-Romero, N.; Keckler, M.S.; Langham, G.; Hutson, C.L.; Karem, K.L.; et al. Identification of Giardia duodenalis and Enterocytozoon bieneusi in an epizoological investigation of a laboratory colony of prairie dogs, Cynomys ludovicianus. Vet. Parasitol. 2015, 210, 91–97. [Google Scholar] [CrossRef]
- Wilkes, G.; Edge, T.; Gannon, V.; Jokinen, C.; Lyautey, E.; Medeiros, D.; Neumann, N.; Ruecker, N.; Topp, E.; Lapen, D.R. Seasonal relationships among indicator bacteria, pathogenic bacteria, Cryptosporidium oocysts, Giardia cysts, and Hydrological indices for surface waters within an agricultural landscape. Water Res. 2009, 43, 2209–2223. [Google Scholar] [CrossRef]
- Plutzer, J.; Takó, M.H.; Márialigeti, K.; Törökné, A.; Karanis, P. First investigations into the prevalence of Cryptosporidium and Giardia spp. in Hungarian drinking water. J. Water Health 2007, 5, 573–584. [Google Scholar] [CrossRef]
- Ćirković, V.; Klun, I.; Utaaker, K.S.; Uzelac, A.; Tysnes, K.R.; Robertson, L.J.; Djurković-Djaković, O. Surface waters as a potential source of Giardia and Cryptosporidium in Serbia. Exp. Parasitol. 2020, 209, 107824. [Google Scholar] [CrossRef]
- Ajeagah, G.A.; Maria, C.; Mirela, P.; Constantin, O.; Palela, M.; Bahrim, G. An Ecological Assessment of the Pollution Status of the Danube River Basin in the Galati Region—Romania. J. Water Resour. Prot. 2013, 05, 876–886. [Google Scholar] [CrossRef][Green Version]
- Ryan, U.; Zahedi, A.; Paparini, A. Cryptosporidium in humans and animals—a one health approach to prophylaxis. Parasite Immunol. 2016, 38, 535–547. [Google Scholar] [CrossRef] [PubMed]
- Robert Koch-Institut. Infektionsepidemiologisches Jahrbuch Meldepflichtiger Krankheiten für 2019; Robert Koch-Institut: Berlin, Germany, 2020. [Google Scholar]
- Lee, M.F.; Auer, H.; Lindo, J.F.; Walochnik, J. Multilocus sequence analysis of Giardia spp. isolated from patients with diarrhea in Austria. Parasitol. Res. 2017, 116, 477–481. [Google Scholar] [CrossRef] [PubMed]
| Prevalence (%) | ||||
|---|---|---|---|---|
| Variable | n° Rats | Cryptosporidium | Eimeria | Giardia |
| Sex | ||||
| Male | 28/50 (56%) | 6/28 (21.4%) | 4/28 (14.3%) | 10/28 (35.7%) |
| Female | 22/50 (44%) | 7/22 (31.8%) | 3/22 (13.6%) | 7/22 (31.8%) |
| Age | ||||
| Sexually immature | 26/50 (52%) | 5/26 (19.2%) | 5/26 (19.2%) | 8/26 (30.8%) |
| Sexually mature | 24/50 (48%) | 8/24 (33.3%) | 2/24 (8.3%) | 9/24 (37.5%) |
| Body mass | ||||
| 0–200 g | 29/50 (58%) | 5/29 (17.2%) | 6/29 (20.7%) | 8/29 (27.6%) |
| >200 g | 21/50 (42%) | 8/21 (38.1%) | 1/21 (4.8%) | 9/21 (42.8%) |
| Body length (nose to anus) | ||||
| 100–200 mm | 32/50 (64%) | 7/32 (21.9%) | 6/32 (18.7%) | 9/32 (21.9%) |
| >200 mm | 18/50 (36%) | 6/18 (33.3%) | 1/18 (5.5%) | 8/18 (44.4%) |
| Trapping month | ||||
| March | 23/50 (46%) | 8/23 (34.8%) | 4/23 (17.4%) | 9/23 (39.1%) |
| April | 9/50 (18%) | 5/9 (55.5%) | 1/9 (11.1%) | 3/9 (33.3%) |
| May | 8/50 (16%) | 0/8 (0%) | 2/8 (25%) | 5/8 (62.5%) |
| June | 10/50 (20%) | 0/10 (0%) | 0/10 (0%) | 0/10 (0%) |
| Trapping site | ||||
| Danube Canal | 39/50 (78%) | 10/39 (25.6%) | 6/39 (15.4%) | 15/39 (38.5%) |
| Karlsplatz | 11/50 (22%) | 3/11 (27.3%) | 1/11 (9.1%) | 2/11 (18.2%) |
| Trapping Site | Transport | Building | Green Infrastructure | Blue Infrastructure |
|---|---|---|---|---|
| Danube Canal | 29.3% | 30.6% | 25.7% | 14.4% |
| Karlsplatz | 32.9% | 41.4% | 25.7% | 0% |
| Sampling Date | Volume Filtered (L) | Turbidity (NTU) | Cryptosporidium Oocysts/L | Giardia Cysts/L |
|---|---|---|---|---|
| 7 May 2019 | 10 | 23 | 3 | 1 |
| 4 June 2019 | 5 | 67 | 4 | 0.8 |
| 9 July 2019 | 7 | 70 | <0.6 | <0.6 |
| 6 August 2019 | 7 | 29 | <0.6 | 0.6 |
| 3 September 2019 | 7 | 24 | <0.6 | 2.9 |
| 8 October 2019 | 10 | 16 | 1 | 1 |
| 5 November 2019 | 10 | 11 | 2.8 | 57.6 |
| 10 December 2019 | 10 | 14 | 1.6 | 1.2 |
| 14 January 2020 | 10 | 5.8 | 1.2 | 3.2 |
| 24 February 2020 | 10 | 14 | <2 | <2 |
| 5 May 2020 | 10 | 10.5 | 4 | 96 |
| 9 June 2020 | 10 | 10 | <0.4 | 1.2 |
| 7 July 2020 | 5 | 80 | <0.8 | 0.8 |
| 11 August 2020 | 7 | 35 | 2.9 | <0.6 |
| 22 September 2020 | 15 | 13 | 0.3 | <0.3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cervero-Aragó, S.; Desvars-Larrive, A.; Lindner, G.; Sommer, R.; Häfeli, I.; Walochnik, J. Surface Waters and Urban Brown Rats as Potential Sources of Human-Infective Cryptosporidium and Giardia in Vienna, Austria. Microorganisms 2021, 9, 1596. https://doi.org/10.3390/microorganisms9081596
Cervero-Aragó S, Desvars-Larrive A, Lindner G, Sommer R, Häfeli I, Walochnik J. Surface Waters and Urban Brown Rats as Potential Sources of Human-Infective Cryptosporidium and Giardia in Vienna, Austria. Microorganisms. 2021; 9(8):1596. https://doi.org/10.3390/microorganisms9081596
Chicago/Turabian StyleCervero-Aragó, Silvia, Amélie Desvars-Larrive, Gerhard Lindner, Regina Sommer, Iveta Häfeli, and Julia Walochnik. 2021. "Surface Waters and Urban Brown Rats as Potential Sources of Human-Infective Cryptosporidium and Giardia in Vienna, Austria" Microorganisms 9, no. 8: 1596. https://doi.org/10.3390/microorganisms9081596
APA StyleCervero-Aragó, S., Desvars-Larrive, A., Lindner, G., Sommer, R., Häfeli, I., & Walochnik, J. (2021). Surface Waters and Urban Brown Rats as Potential Sources of Human-Infective Cryptosporidium and Giardia in Vienna, Austria. Microorganisms, 9(8), 1596. https://doi.org/10.3390/microorganisms9081596