Metagenomics Analysis Reveals an Extraordinary Inner Bacterial Diversity in Anisakids (Nematoda: Anisakidae) L3 Larvae
Abstract
1. Introduction
2. Material and Methods
2.1. Anisakids Collection and Taxonomic Identification
2.2. 16 S rRNA-rRNA Amplicons, Preparation and Illumina Sequencing
2.3. Bioinformatics Analysis and Data Availability
2.4. Statistical Analysis
3. Results
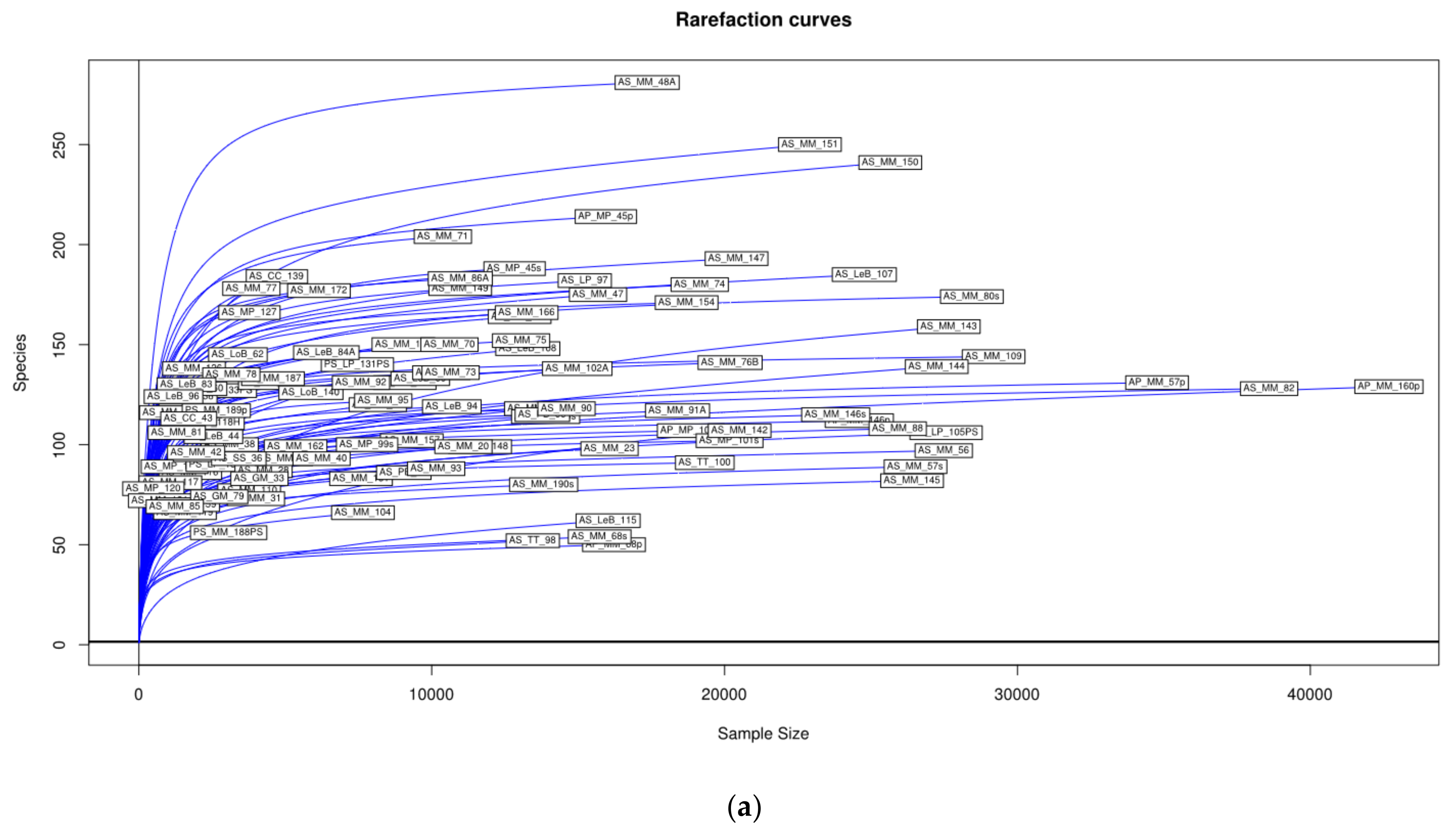
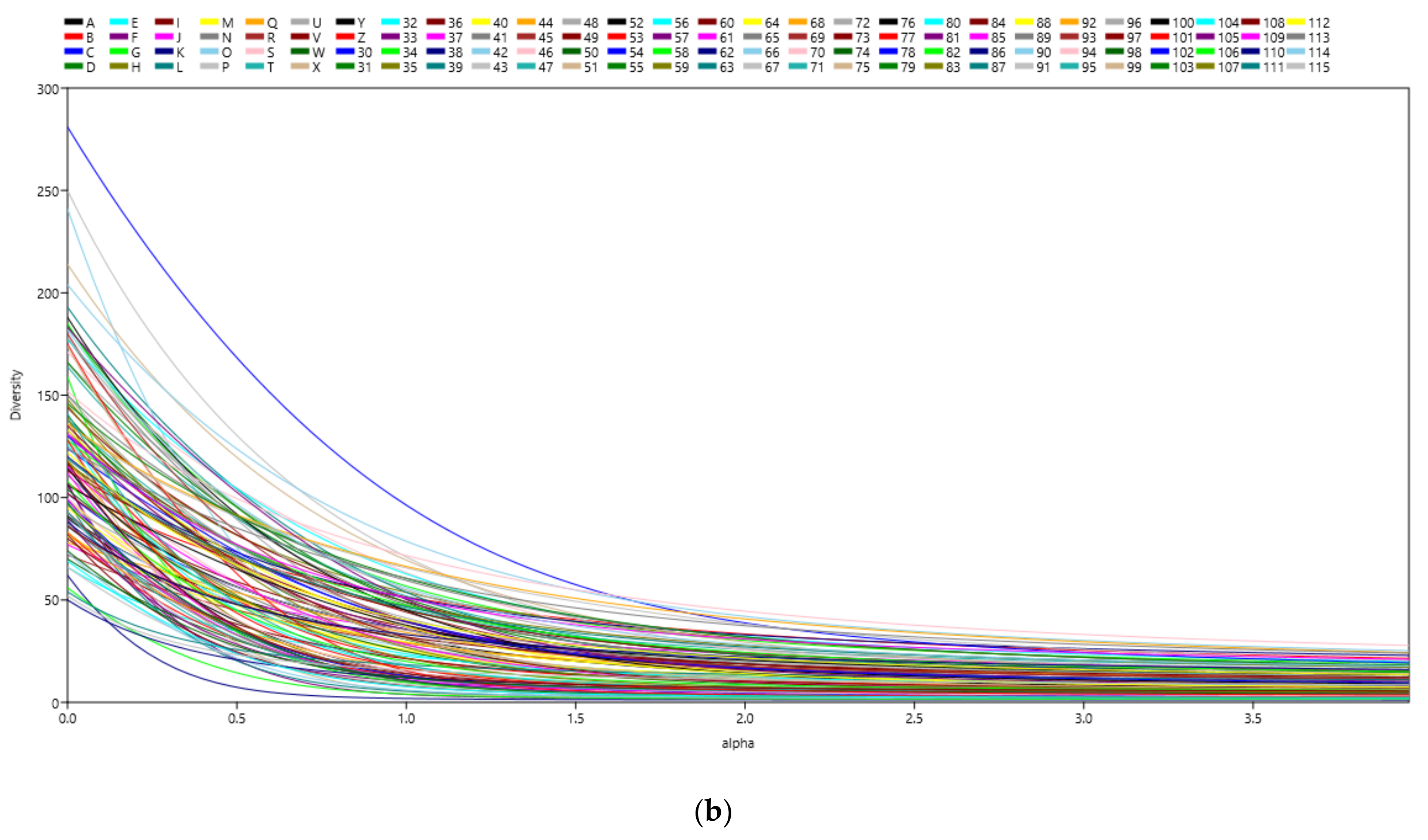
3.1. Microbial Richness and Diversity of Samples (Anisakids Individuals)
3.2. Quantitative Analysis (Correlation) Does Not Support Inference of Microbiota Structure
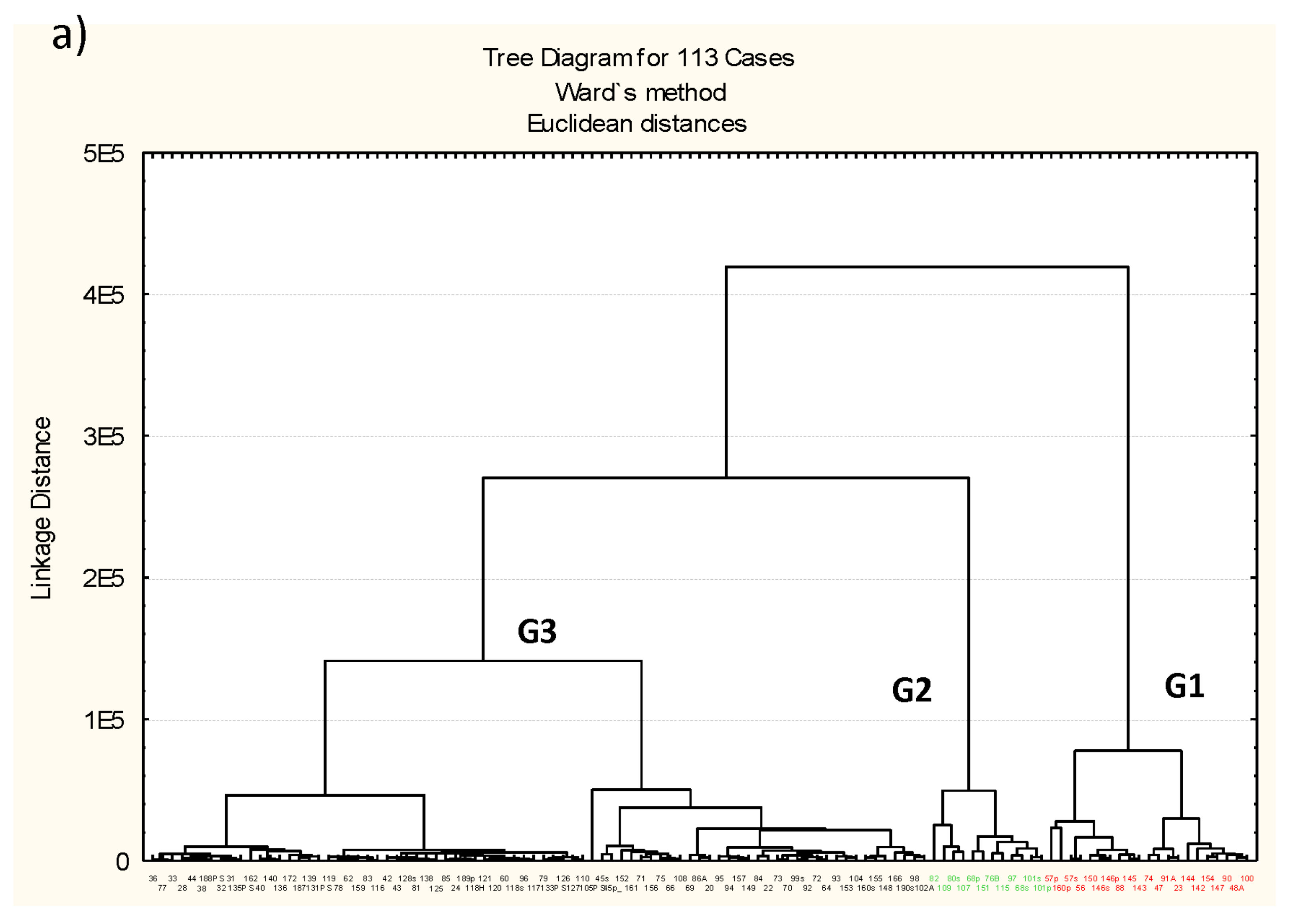
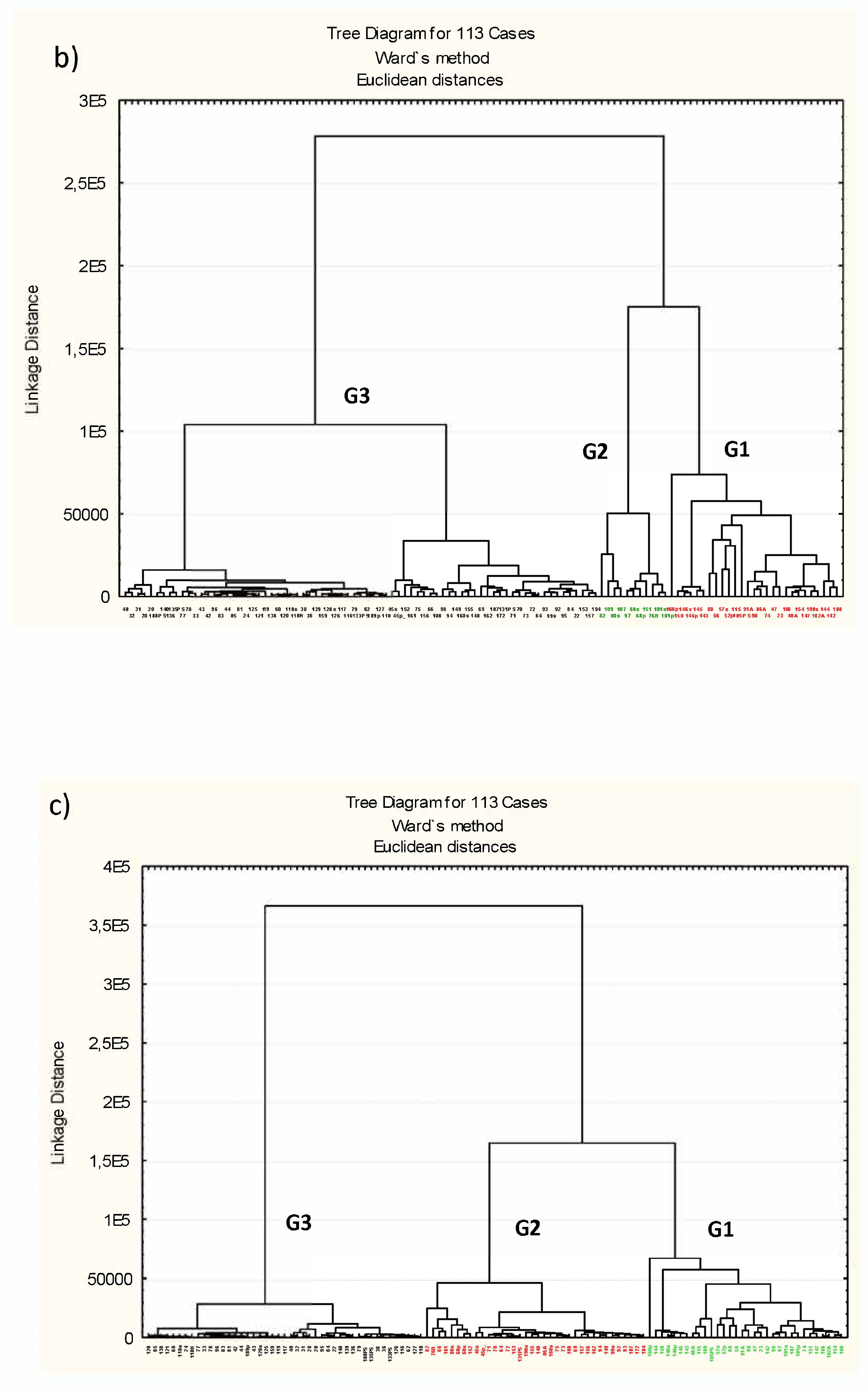
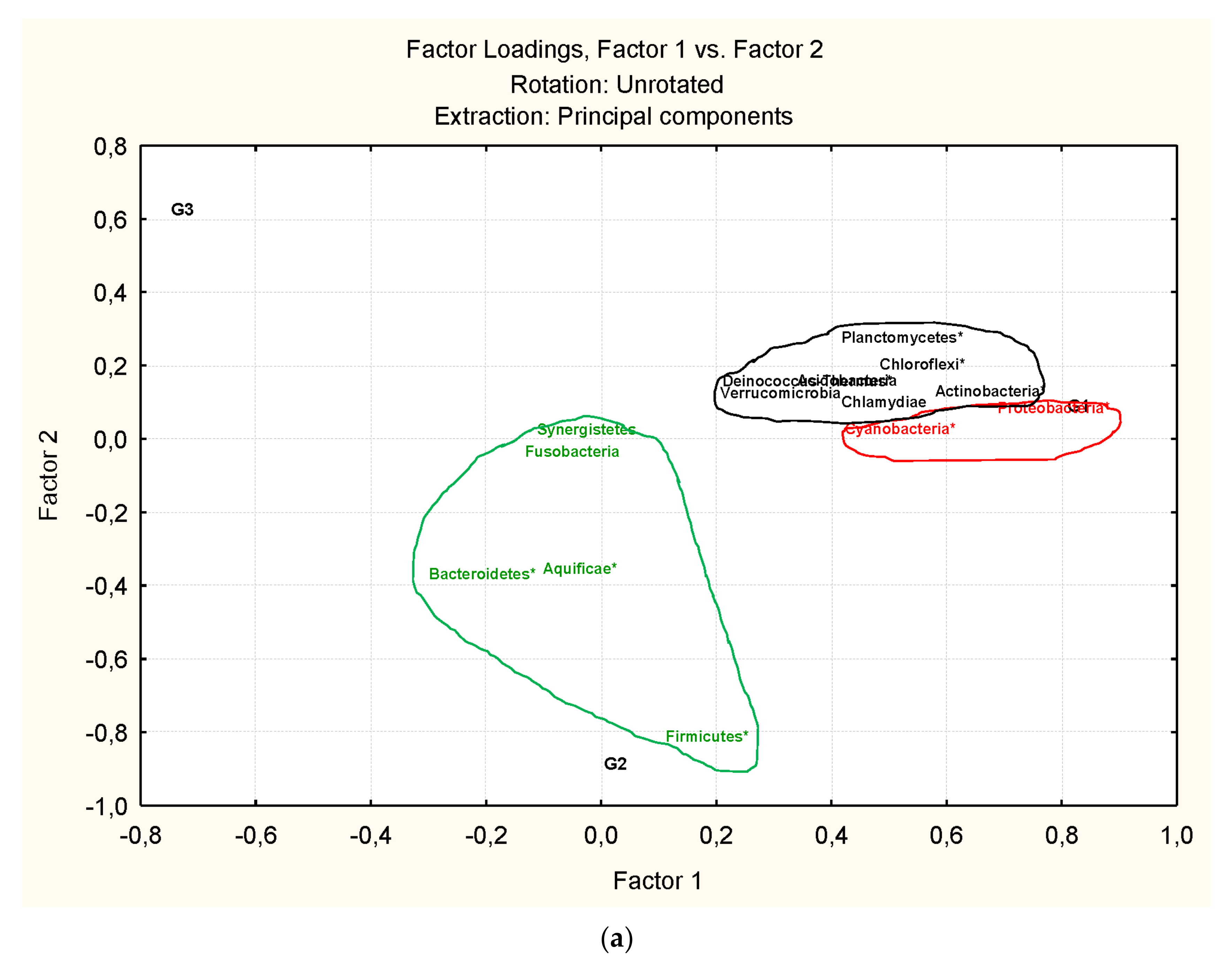
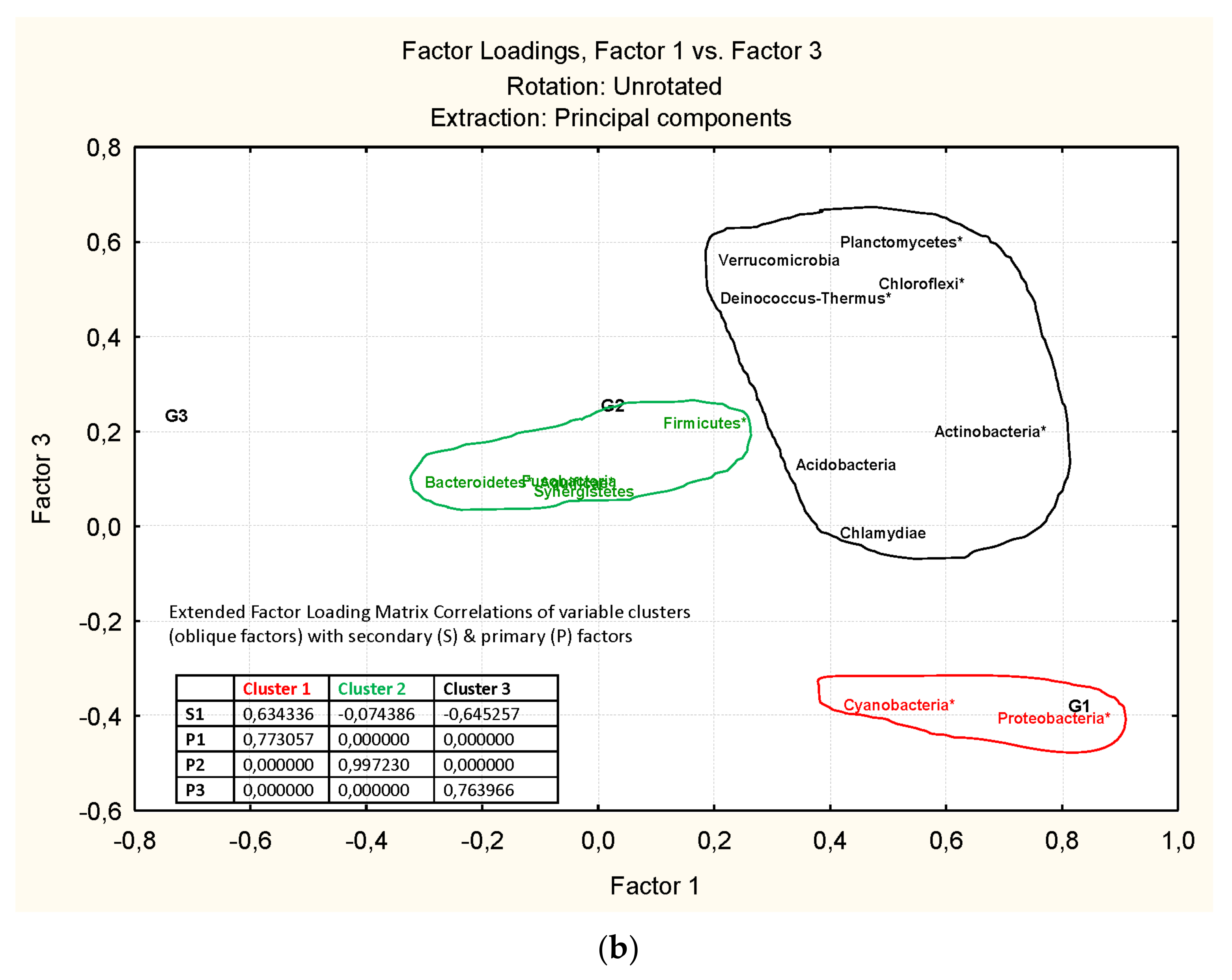
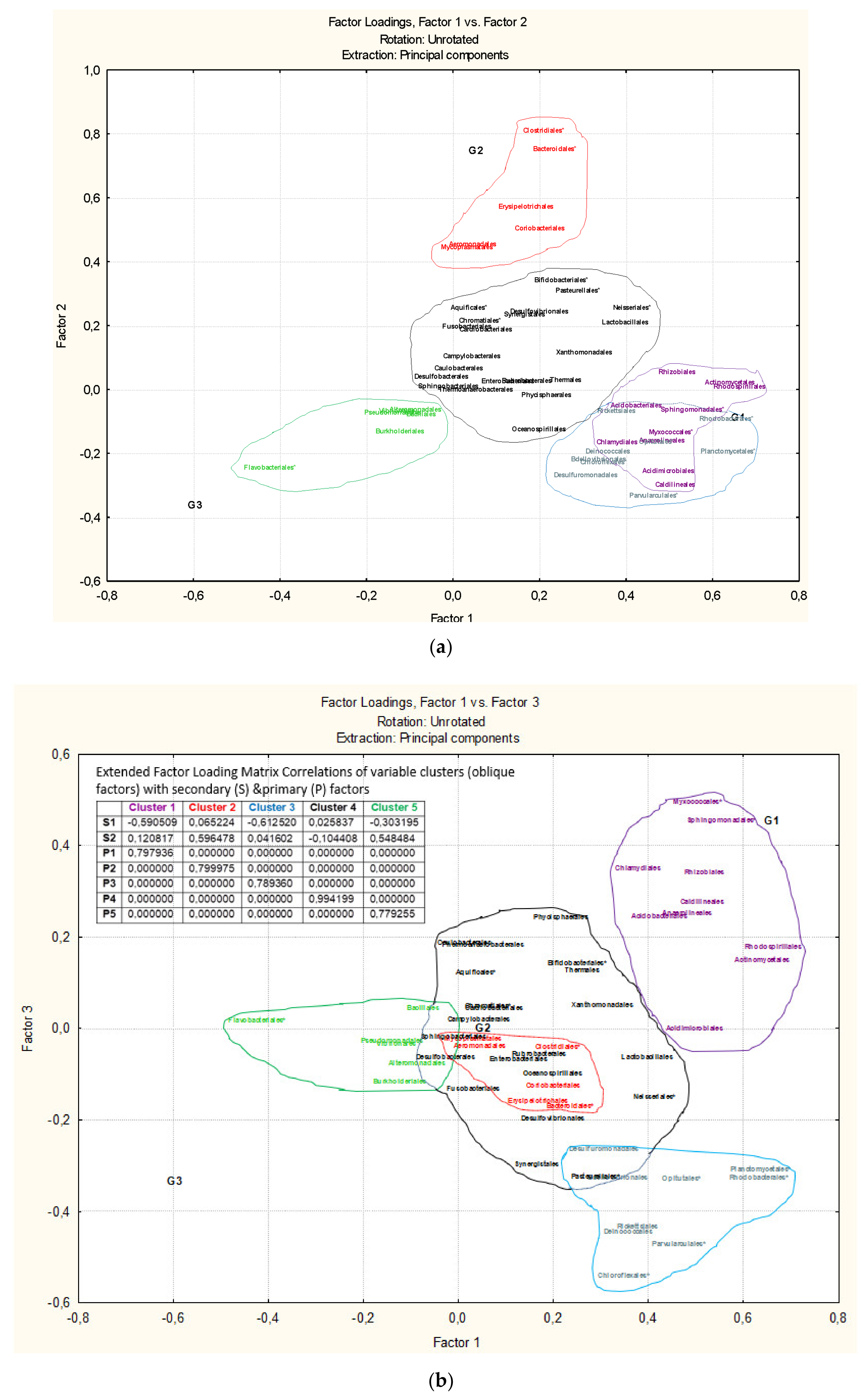
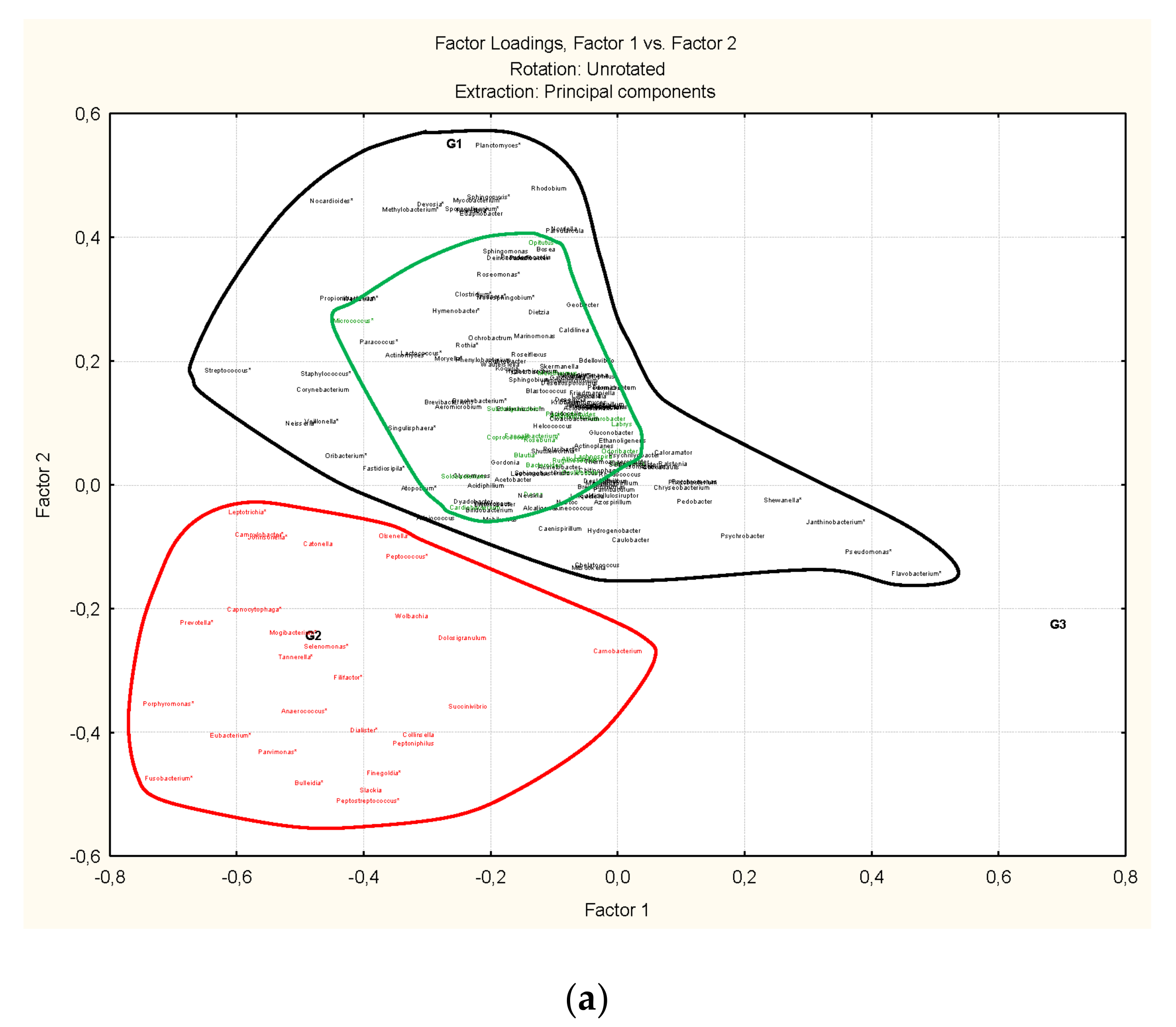
3.3. Microbiota Structure Associated to Anisakids
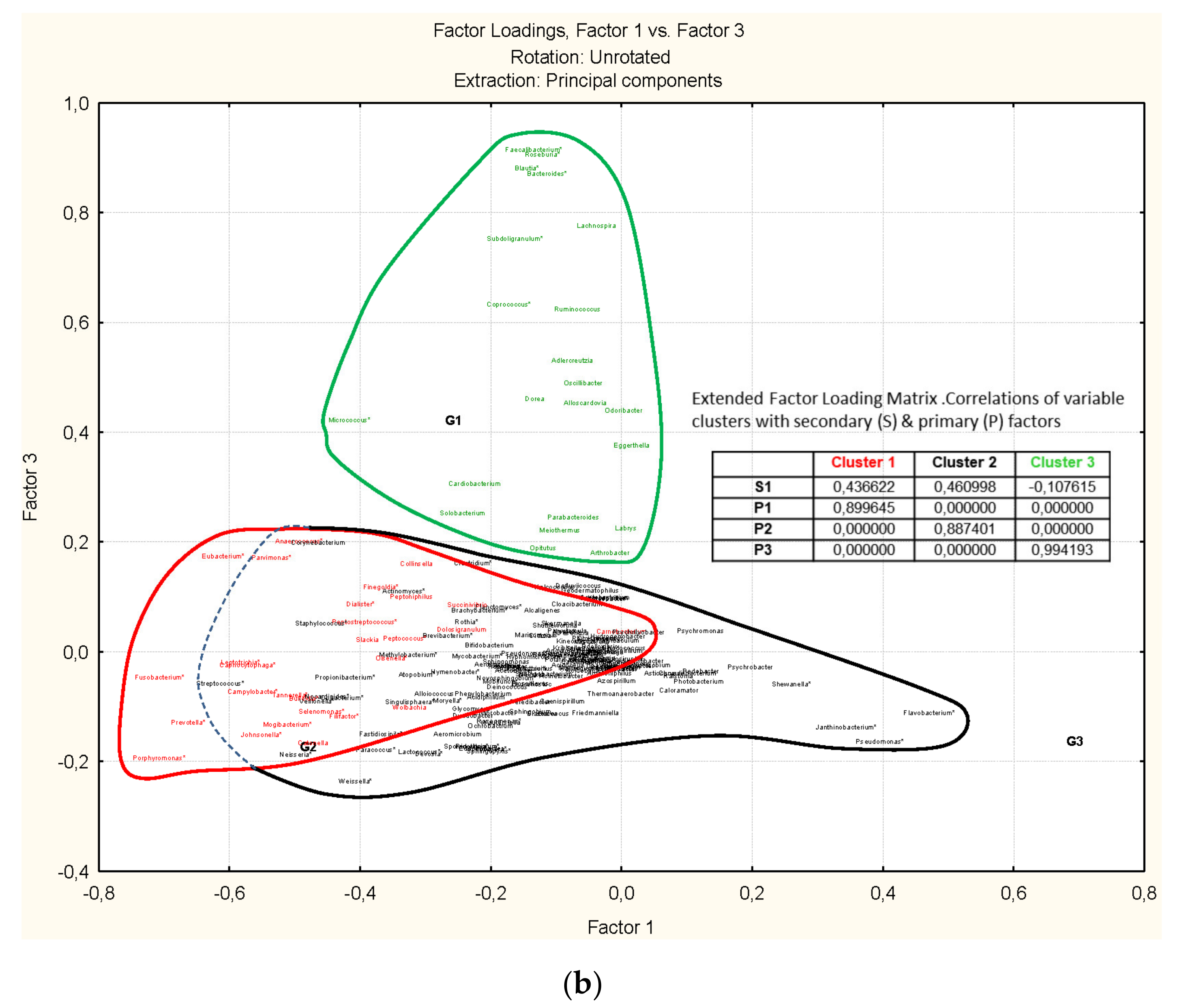
3.4. Genus-Level Associations (Clusters Description): The Ultimate Test Is the Significant Structure which Represents the Microbiota Association at Genus Level
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rohlwing, T.; Palm, H.W.; Rosenthal, H. Parasitation with Pseudoterranova decipiens (Nematoda) influences the survival rate of the European smelt Osmerus eperlanus retained by a screen wall of a nuclear power plant. Dis. Aquat. Organ. 1998, 32, 233–236. [Google Scholar] [CrossRef] [PubMed]
- Sakanari, J.A.; McKerrow, J.H. Anisakiasis. Clin. Microbiol. Rev. 1989, 2, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Llorens, C.; Arcos, S.C.; Robertson, L.; Ramos, R.; Futami, R.; Soriano, B.; Ciordia, S.; Careche, M.; González-Muñoz, M.; Jiménez-Ruiz, Y.; et al. Functional insights into the infective larval stage of Anisakis simplex s.s., Anisakis pegreffii and their hybrids based on gene expression patterns. BMC Genom. 2018, 19, 592. [Google Scholar] [CrossRef]
- Moneo, I.; Caballero, M.L.; Gómez, F.; Ortega, E.; Alonso, M.J. Isolation and characterization of a major allergen from the fish parasite Anisakis simplex. J. Allergy Clin. Immunol. 2000, 106, 177–182. [Google Scholar] [CrossRef]
- Baeza, M.L.; Matheu, V.; Rodriguez, A.; De Barrio, M.; Tornero, P.; Rubio, M.; Zubeldia, J.M. Excretory-secretory Anisakis simplex allergens: In vivo and in vitro studies. J. Allergy Clin. Immunol. 2001, 107, S184. [Google Scholar]
- Wekell, M.M.; Hungerford, J.M. Microbiological quality of seafoods: Marine toxins. In Seafood: Chemistry, Processing Technology and Quality; Shahidi, F., Botta, J.R., Eds.; Blackie: London, UK, 1994; pp. 220–232. [Google Scholar] [CrossRef]
- Van Thiel, P.H.; Kuipers, F.C.; Roskam, R.T. A nematode parasitic to herring causing acute abdominal syndromes in man. Trop. Geogr. Med. 1960, 12, 97–113. [Google Scholar]
- Køie, M.; Berland, B.; Burt, M.D.B. Development to third-stage larva occurs in eggs of Anisakis simplex and Pseudoterranova decipiens (Nematoda, Ascaridoidea, Anisakidae). Can. J. Fish Aquat. Sci. 1995, 52, 134–139. [Google Scholar] [CrossRef]
- Moneo, I.; Caballero, M.L.; Rodríguez-Pérez, R.; Rodríguez-Mahillo, A.I.; González-Muñoz, M. Sensitization to the fish parasite Anisakis simplex: Clinical and laboratory aspects. Parasitol. Res. 2007, 101, 1051–1055. [Google Scholar] [CrossRef]
- Kleter, G.A.; Prandini, A.; Filippi, L.; Marvin, H.J.P. Identification of potentially emerging food safety issues by analysis of reports published by the European Community’s Rapid Alert System for Food and Feed (RASFF) during a four-year period. Food Chem. Toxicol. 2009, 47, 932–950. [Google Scholar] [CrossRef]
- Puente, P.; Anadón, A.M.; Rodero, M.; Romaní, F.; Ubeira, F.M.; Cuellar, C. Anisakis simplex: The high prevalence in Madrid (Spain) and its relation with fish consumption. Exp. Parasitol. 2007, 118, 271–274. [Google Scholar] [CrossRef]
- Anderson, R.C.; Chabaud, A.G.; Willmott, S. Keys to the Nematodes Parasites of Vertebrates; CABI International: Cambridge, Oxfordshire, UK, 2009; 463p. [Google Scholar]
- Mattiucci, S.; Cipriani, P.; Webb, S.C.; Paoletti, M.; Marcer, F.; Bellisario, B.; Gibson, D.I.; Nascetti, G. Genetic and Morphological Approaches Distinguish the Three Sibling Species of the Anisakis simplex Species Complex, with a Species Designation as Anisakis berlandi n. sp. for A. simplex sp. C (Nematoda: Anisakidae). J. Parasitol. 2014, 100, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Audicana, M.T.; Ansotegui, I.J.; Fernández de Corres, L.; Kennedy, M.W. Anisakis simplex: Dangerous-dead and alive? Trends Parasitol. 2002, 18, 20–25. [Google Scholar] [CrossRef]
- Audicana, M.T.; Kennedy, M.W. Anisakis simplex: From Obscure Infectious Worm to Inducer of Immune Hypersensitivity. Clin. Microbiol. Rev. 2008, 21, 360–379. [Google Scholar] [CrossRef] [PubMed]
- Navas, A.; Cobas, G.; Talavera, M.; López, J.A.; Ayala, J.; Martínez, J.L. Experimental validation of Haldane’s hipótesis on the role of infection as an evolutionary force for Metazoans. Proc. Natl. Acad. Sci. USA 2007, 34, 13728–13731. [Google Scholar] [CrossRef] [PubMed]
- Cabot, G.; Zamorano, L.; Moyà, B.; Juan, C.; Navas, A.; Blázquez, J.; Oliver, A. Evolution of Pseudomonas aeruginosa antimicrobial resistance and fitness under low and high mutation supply rates. Antimicrob. Agents Chemother. 2016, 60, 1767–1778. [Google Scholar] [CrossRef]
- Ruíz Díez, B.; Sánchez, P.; Baquero, F.; Martínez, J.L.; Navas, A. Differential interaction and virulence within the Caenorhabditis elegans-Pseudomonas aeruginosa pathogenesis model. J. Theor. Biol. 2003, 225, 469–476. [Google Scholar] [CrossRef]
- Couillaut, C.; Ewbank, J.J. Diverse bacteria pathogens of Caenorhabditis elegans. Infect. Immun. 2002, 70, 4705–4707. [Google Scholar] [CrossRef] [PubMed]
- Sifri, C.D.; Mylonakis, E.; Singh, K.V.; Qin, X.; Garsin, D.A.; Murray, B.E.; Ausubel, F.M.; Calderwood, S.B. Virulence effect of Enterococcus faecalis protease genes and the quorum-sensing locus fsr in Caenorhabditis elegans and mice. Infect. Immun. 2002, 70, 56475650. [Google Scholar] [CrossRef]
- Garsin, D.A.; Villanueva, J.M.; Begun, J.; Kim, D.H.; Sifri, C.D.; Calderwood, S.B.; Ruvkun, G.; Ausubel, F.M. Long-lived C.elegans daf-2 mutants are resistant to bacterial pathogens. Science 2003, 300, 1921. [Google Scholar] [CrossRef]
- Tenor, J.L. Caenorhabdditis elegans based screen identifies Salmonella virulence factors required for conserved host-pathogens interactions. Proc. Natl. Acad. Sci. USA 2004, 96, 2408–2413. [Google Scholar]
- Mallo, G.V.; Kurz, C.L.; Couillault, C.; Pujol, N.; Granjeaud, S.; Kohara, Y.; Ewbank, J.J. Inducible antibacterial defense system in C. elegans. Curr. Biol. 2002, 12, 1209–1214. [Google Scholar] [CrossRef]
- Bae, T.; Banger, A.K.; Wallace, A.; Glass, E.M.; Aslund, F.; Schneewind, O.; Missiakas, D.M. Staphylococcus aureus virulence genes identified by bursa aurealis mutagenesis and nematode killing. Proc. Natl. Acad. Sci. USA 2004, 101, 12312–12317. [Google Scholar] [CrossRef]
- Bolm, M.; Jansen, W.T.M.; Schnabel, R.; Chhatwal, G.S. Hydrogen peroxide-mediated killing of Caenorhabditis elegans: A common feature of different streptoccal species. Infect. Immun. 2004, 72, 1192–1194. [Google Scholar] [CrossRef]
- Darby, C.; Hsu, J.W.; Ghori, N.; Falkow, S. Caenorhabditis elegans: Plague bacteria biofilm blocks food intake. Nature 2002, 417, 243–244. [Google Scholar] [CrossRef] [PubMed]
- González Quijada, S.; González Escudero, R.; Arias García, L.; Gil Martín, A.R.; Vicente Serrano, J.; Corral Fernández, E. Anisakiasis gastrointestinal manifestations: Description of 42 cases. Rev. Clin. Esp. 2005, 205, 311–315. [Google Scholar] [CrossRef] [PubMed]
- Valero, A.; Lopez-Cuello, M.; Benítez, R.; Adroher, F.J. Anisakis spp. in European hake, Merluccius merluccius (L.) from the Atlantic off north-west Africa and the Mediterranean off southern Spain. Acta Parasitol. 2006, 51, 209–212. [Google Scholar] [CrossRef]
- Iglesias, L.; Valero, A.; Adroher, F.J. Some factors which influence the in vitro maintenance of Anisakis simplex (Nematoda). Folia Parasitol. 1997, 44, 297–301. [Google Scholar]
- Zhu, X.Q.; Podolska, M.; Liu, J.S.; Yu, H.Q.; Chen, H.H.; Lin, Z.X.; Luo, C.B.; Song, H.Q.; Lin, R.Q. Identification of anisakid nematodes with zoonotic potential from Europe and China by single-strand conformation polymorphism analysis of nuclear ribosomal DNA. Parasitol. Res. 2007, 101, 1703–1707. [Google Scholar] [CrossRef] [PubMed]
- Abollo, E.; Paggi, L.; Pascual, S.; D’Amelio, S. Occurrence of recombinant genotypes of Anisakis simplex s. s. and Anisakis pegreffii (Nematoda, Anisakidae) in an area of sympatry. Infect. Genet. Evol. 2003, 3, 175–181. [Google Scholar] [CrossRef]
- D’Amelio, S.; Mathiopoulos, K.D.; Santos, C.P.; Pugachev, O.N.; Webb, S.C.; Picanco, M.; Paggi, L. Genetic markers in ribosomal DNA for the identification of members of the genus Anisakis (Nematoda: Ascaridoidea) defined by polymerase-chain-reaction-based restriction fragment length polymorphism. Int. J. Parasitol. 2000, 30, 223–226. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing Mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. App. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Andrews, S. FastQC a Quality Control Tool for High Throughput Sequence Data [Online]. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 8 January 2017).
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schlossa, P.D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. App. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Narciso-da-Rocha, C.; Rocha, J.; Vaz-Moreira, I.; Lira, F.; Tamames, J.; Henriques, I.; Martinez, J.L.; Manaia, C.M. Bacterial lineages putatively associated with the dissemination of antibiotic resistance genes in a full-scale urban wastewater treatment plant. Environ. Int. 2018, 118, 179–188. [Google Scholar] [CrossRef]
- Schloss, P.D. Secondary structure improves OTU assignments of 16S rRNA rRNA gene sequences. ISME J. 2013, 7, 457–460. [Google Scholar] [CrossRef]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.G.; Peplies, J.; Glöckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucl. Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef]
- Pruesse, E.; Joerg, P.; Gloeckner, F.O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 2012, 28, 1823–1829. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Henderson, P.A. Practical Methods in Ecology; Blackwell Publishing Company: Oxford, UK, 2006. [Google Scholar]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAlaeontological STatistics, ver.1.9 Users Manual. 2009. Available online: http://folk.uio.no/ohammer/past/past.pdf (accessed on 4 April 2020).
- StatSoft, Inc. STATISTICA (Data Analysis Software System), Version 6. 2001. Available online: https://www.statsoft.com (accessed on 19 July 2019).
- Werren, J.H.; Baldo, L.; Clark, M.E. Wolbachia: Master manipulators of invertebrate biology. Nat. Rev. Microbiol. 2008, 6, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Picolos, D.K.; Lerche-Sehm, J.; Abron, A.; Fine James, B.; Papapanou, P.N. Infection patterns in chronic and aggressive periodontitis. J. Clin. Periodontol. 2005, 32, 1055–1061. [Google Scholar] [CrossRef]
- Goodson, J.M.; Groppo, D.; Halem, S.; Carpino, E. Is Obesity an Oral Bacterial Disease? J. Dent. Res. 2009, 88, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.; Pikis, A. Metabolism of sugars by genetically diverse species of oral Leptotrichia. Mol. Oral Microbiol. 2012, 27, 34–44. [Google Scholar] [CrossRef]
- Downes, J.; Olsvik, B.; Hiom, S.J.; Spratt, D.A.; Cheeseman, S.L.; Olsen, I.; Weightman, A.J.; Wade, W.G. Bulleidia extructa gen. nov., sp. nov., isolated from the oral cavity. Int. J. Syst. Evol. Microbiol. 2000, 50, 979–983. [Google Scholar] [CrossRef][Green Version]
- Nakazawa, F.; Sato, M.; Poco, S.E.; Hashimura, T.; Ikeda, T.; Kalfas, S.; Sundqvist, G.; Hoshino, E. Description of Mogibacterium pumilum gen. nov., sp. nov. and Mogibacterium vescum gen. nov., sp. nov., and reclassification of Eubacterium timidum (Holdeman et al., 1980) as Mogibacterium timidum gen. nov., comb. nov. Int. J. Syst. Evol. Microbiol. 2000, 50, 679–688. [Google Scholar] [CrossRef]
- Moore, L.V.; Moore, W.E.C. Oribaculum catoniae gen. nov., sp. nov.; Catonella morbi gen. nov., sp. nov.; Hallella seregens gen. nov., sp. nov.; Johnsonella ignava gen. nov., sp. nov.; and Dialister pneumosintes gen. nov., comb. nov., nom. rev., Anaerobic Gram-Negative Bacilli from the Human Gingival Crevice. Int. J. Syst. Bacteriol. 1994, 44, 187–192. [Google Scholar] [CrossRef]
- Collins, M.D.; Lawson, P.A.; Willems, A.; Cordoba, J. J.; Fernandez-Garayzabal, J.; Garcia, P.; Cai, J.; Hippe, H.; Farrow, J.A.E. The Phylogeny of the Genus Clostridium: Proposal of Five New Genera and Eleven New Species Combinations. Int. J. Syst. Bacteriol. 1994, 44, 812–826. [Google Scholar] [CrossRef]
- Tindall, B.J.; Euzéby, J.P. Proposal of Parvimonas gen. nov. and Quatrionicoccus gen. nov. as replacements for the illegitimate, prokaryotic, generic names Micromonas Murdoch and Shah 2000 and Quadricoccus Maszenan et al., 2002, respectively. Int. J. Syst. Evol. Microbiol. 2006, 56, 2711–2713. [Google Scholar] [CrossRef]
- Kumar, P.S. Smoking and the subgingival ecosystem: A pathogen-enriched community. Future Microbiol. 2012, 7, 917–919. [Google Scholar] [CrossRef]
- Leisner, J.J.; Laursen, B.G.; Prévost, H.; Drider, D.; Dalgaard, P. Carnobacterium: Positive and negative effects in the environment and in foods. FEMS Microbiol. Rev. 2007, 31, 592–613. [Google Scholar] [CrossRef] [PubMed]
- Hespell, R.B. The Genera Succinivibrio and Succinimonas. In The Prokaryotes; Balows, A., Trüper, H.G., Dworkin, M., Harder, W., Schleifer, K.H., Eds.; Springer: New York, NY, USA, 1992. [Google Scholar]
- Ricaboni, D.; Mailhe, M.; Vitton, V.; Bittar, F.; Raoult, D.; Fournier, P.E. Olsenella provencensis sp. nov., Olsenella phocaeensis sp. nov., and Olsenella mediterranea sp. nov. isolated from the human colon. Hum. Microbiome J. 2017, 4, 22–23. [Google Scholar] [CrossRef]
- Wade, W.G.; Downes, J.; Dymock, D.; Hiom, S.J.; Weightman, A.J.; Dewhirst, F.E.; Paster, B.J.; Tzellas, N.; Coleman, B. The family Coriobacteriaceae: Reclassification of Eubacterium exiguum (Poco et al., 1996) and Peptostreptococcus heliotrinreducens (Lanigan 1976) as Slackia exigua gen. nov., comb. nov. and Slackia heliotrinireducens gen. nov., comb. nov., and Eubacterium lentum (Prevot 1938) as Eggerthella lenta gen. nov., comb. nov. Int. J. Syst. Bacteriol. 1999, 49, 595–600. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, A.; Benno, Y.; Nakase, T. Phylogenetic and phenotypic evidence for the transfer of Eubacterium aerofaciens to the genus Collinsella as Collinsella aerofaciens gen. nov., comb. nov. Int. J. Syst. Bacteriol. 1999, 49, 557–565. [Google Scholar] [CrossRef] [PubMed]
- Shkoporov, A.N.; Efimov, B.A.; Kondova, I.; Ouwerling, B.; Chaplin, A.V.; Shcherbakova, V.A.; Langermans, J.A.M. Peptococcus simiae sp. nov., isolated from Rhesus macaque faeces and emended description of the genus Peptococcus. Int. J. Syst. Evol. Microbiol. 2016, 66, 5187–5191. [Google Scholar] [CrossRef] [PubMed]
- Ezaki, T.; Kawamura, Y.; Li, N.; Li, Z.Y.; Zhao, L.; Shu, S. Proposal of the genera Anaerococcus gen. nov., Peptoniphilus gen. nov. and Gallicola gen. nov. for members of the genus Peptostreptococcus. Int. J. Syst. Evol. Microbiol. 2001, 51, 1521–1528. [Google Scholar] [CrossRef]
- De Moreuil, C.; Héry-Arnaud, G.; David, C.H.; Provost, B.; Mondine, P.; Alavi, Z.; de Saint Martin, L.; Bezon, E.; Berre, R.L. Finegoldia magna, not a well-known infectious agent of bacteriemic post-sternotomy mediastinitis. Anaerobe 2015, 32, 32–33. [Google Scholar] [CrossRef]
- Brown, K.; Church, D.; Lynch, T.; Gregson, D. Bloodstream infections due to Peptoniphilus spp.: Report of 15 cases. Clin. Microbiol. Infec. 2014, 20, O857–O860. [Google Scholar] [CrossRef]
- Lecuyer, H.; Audibert, J.; Bobigny, A.; Eckert, C.; Janniere-Nartey, C.; Buu-Hoi, A.; Mainardi, J.L.; Podglajen, I. Dolosigranulum pigrum Causing Nosocomial Pneumonia and Septicemia. J. Clin. Microbiol. 2007, 45, 3474–3475. [Google Scholar] [CrossRef]
- Neuvonen, M.M.; Tamarit, D.; Näslund, K.; Liebig, J.; Feldhaar, H.; Moran, N.A.; Guy, L.; Andersson, S.G.E. The genome of Rhizobiales bacteria in predatory ants reveals urease gene functions but no genes for nitrogen fixation. Sci. Rep. 2016, 6, 39197. [Google Scholar] [CrossRef]
- Vannini, C.; Rosati, G.; Verni, F.; Petroni, G. Identification of the bacterial endosymbiontsof the marine ciliate Euplotes magnicirratus (Ciliophora, Hypotrichia) and proposal of Candidatus Devosia euplotis. Int. J. Syst. Evol. Microbiol. 2004, 54, 1151–1156. [Google Scholar] [CrossRef][Green Version]
- Erlacher, A.; Cernava, T.; Cardinale, M.; Soh, J.; Sensen, C.W.; Grube, M.; Berg, G. Rhizobiales as functional and endosymbiontic members in the lichen symbiosis of Lobaria pulmonaria L. Front. Microbiol. 2015, 6, 53. [Google Scholar] [CrossRef]
- Rainey, F.A.; Ward-Rainey, N.; Gliesche, C.G.; Stackebrandtl, E. Phylogenetic analysis and intrageneric structure of the genus Hyphornicrobium and the related genus Filornicrobium. Int. J. Syst. Bacteriol. 1998, 48, 635–639. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gallego, V.; Garcia, M.T.; Ventosa, A. Methylobacterium adhaesivum sp. nov., a methylotrophic bacterium isolated from drinking water. Int. J. Syst. Evol. Microbiol. 2006, 56, 339–342. [Google Scholar] [CrossRef] [PubMed]
- Garrity, G.M.; Winters, M.; Searles, D.B. Taxonomic Outline of the Procaryotic Genera Bergey’s Manual of Systematic Bacteriology, 2nd ed.; Release 1.0; Springer: New York, NY, USA, 2001; pp. 1–39. [Google Scholar]
- Hiraishi, A.; Urata, K.; Satoh, T. A new genus of marine budding phototrophic bacteria, Rhodobium gen. nov., which includes Rhodobium orientis sp. nov. and Rhodobium marinum comb. nov. Int. J. Syst. Evol. Microbiol. 1995, 45, 226–234. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yoon, J.H.; Kang, S.J.; Im, W.T.; Lee, S.T.; Oh, T.K. Chelatococcus daeguensis sp. nov., isolated from wastewater of a textile dye works, and emended description of the genus Chelatococcus. Int. J. Syst. Evol. Microbiol. 2008, 58, 2224–2228. [Google Scholar] [CrossRef]
- Schleheck, D.; Tindall, B.J.; Rosselló-Mora, R.; Cook, A.M. Parvibaculum lavamentivorans gen. nov., sp. nov., a novel heterotroph that initiates catabolism of linear alkylbenzenesulfonate. Int. J. Syst. Evol. Microbiol. 2004, 54, 1489–1497. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B.; Barrassi, L.; Raoult, D. A novel alpha-Proteobacterium, Nordella oligomobilis gen. nov., sp. nov., isolated by using amoebal co-cultures. Res. Microbiol. 2004, 155, 47–51. [Google Scholar] [CrossRef]
- Holmes, B.; Popoff, M.; Kiredjian, M.; Kersters, K. Ochrobactrum anthropi gen. nov., sp. nov. from human clinical specimens and previously known as Group Vd. Int. J. Syst. Bacteriol. 1988, 38, 408. [Google Scholar] [CrossRef]
- Kern, W.V.; Oethinger, M.; Kaufhold, A.; Rozdizinski, E.; Marre, R. Ochobactrum anthropi bacteremia: Report of four cases and short review. Infection 1993, 21, 306–310. [Google Scholar] [CrossRef]
- Abraham, W.R.; Macedo, A.J.; Lünsdorf, H.; Fischer, R.; Pawelczyk, S.; Smith, J.; Vancanneyt, M. Phylogeny by a polyphasic approach of the order Caulobacterales, proposal of Caulobacter mirabilis sp. nov., Phenylobacterium haematophilum sp. nov. and Phenylobacterium conjunctum sp. nov., and emendation of the genus Phenylobacterium. Int. J. Syst. Evol. Microbiol. 2008, 58, 1939–1949. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.C.; Giovannoni, S.J. Parvularcula bermudensis gen. nov., sp. nov., a marine bacterium that forms a deep branch in the α-Proteobacteria. Int. J. Syst. Evol. Bacteriol. 2003, 53, 1031–1036. [Google Scholar] [CrossRef] [PubMed]
- Buchan, A.; González, J.M.; Moran, M.A. Overview of the marine roseobacter lineage. Appl. Environ. Microb. 2005, 71, 5665–5677. [Google Scholar] [CrossRef]
- Bryant, D.A.; Frigaard, N.U. Prokaryotic photosynthesis and phototrophy illuminated. Trends Microbiol. 2006, 14, 488–496. [Google Scholar] [CrossRef]
- Dé, I.; Rolston, K.V.; Han, X.Y. Clinical significance of Roseomonas species isolated from catheter and blood samples: Analysis of 36 cases in patients with cancer. Clin. Infect. Dis. 2004, 38, 1579–1584. [Google Scholar] [CrossRef] [PubMed]
- Walker, D.H. Rickettsiae. In Barron’s Medical Microbiology, 4th ed.; Barron, S., Ed.; University of Texas Medical Branch: Galveston, TX, USA, 1996. [Google Scholar]
- Kertesz, M.A.; Kawasaki, A.; Stolz, A. Aerobic Hydrocarbon-Degrading Alphaproteobacteria: Sphingomonadales. In Taxonomy, Genomics and Ecophysiology of Hydrocarbon-Degrading Microbes. Handbook of Hydrocarbon and Lipid Microbiology; McGenity, T., Ed.; Springer: Cham, Switzerland, 2019. [Google Scholar] [CrossRef]
- Knapp, J.S. Historical perspectives and identification of Neisseria and related species. Clin. Microbiol. Rev. 1988, 1, 415–431. [Google Scholar] [CrossRef]
- Kavuncuoglu, F.; Unal, A.; Oguzhan, N.; Tokgoz, B.; Oymak, O.; Utas, C. First Reported Case of Alcaligenes faecalis Peritonitis. J. Int. Soc. Periton. Dial. 2010, 30, 118–119. [Google Scholar] [CrossRef]
- Saddler, G.S. Management of bacterial wilt disease. In Bacterial wilt Disease and the Ralstonia Solanacearum Species Complex; Allen, C., Prior, P., Hayward, A.C., Paul, M.N., Eds.; APS Press: St Paul, MN, USA, 2005; pp. 121–132. [Google Scholar]
- Brucker, R.M.; Harris, R.N.; Schwantes, C.R.; Gallaher, T.N.; Flaherty, D.C.; Lam, B.A.; Minbiole, K.P.C. Amphibian chemical defense: Antifungal metabolites of the microsymbiont Janthinobacterium lividum on the salamander Plethodon cinereus. J. Chem. Ecol. 2008, 34, 1422–1429. [Google Scholar] [CrossRef]
- Fera, M.T.; Maugeri, T.L.; Gugliandolo, C.; Beninati, C.; Giannone, M.; La Camera, E.; Carbone, M. Detection of Arcobacter spp. in the Coastal Environment of the Mediterranean Sea. App. Environ. Microbiol. 2004, 70, 1271–1276. [Google Scholar] [CrossRef]
- Kruse, S.; Goris, T.; Westermann, M.; Adrian, L.; Dieker, G. Hydrogen production by Sulfurospirillum species enables syntrophic interactions of Epsilonproteobacteria. Nat. Commun. 2018, 9, 4872. [Google Scholar] [CrossRef] [PubMed]
- Dworkin, M. (Editor-in-Chief); The Prokaryotes. A Handbook on the Biology of Bacteria Third EditionVolume 7: Proteobacteria: Delta and Epsilon Subclasses. Deeply Rooting Bacteria; Falkow, S., Rosenberg, E., Schleifer, K.H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; pp. 5–191. [Google Scholar]
- Tena, D.; González-Praetorius, A.; Gimeno, C.; Pérez-Pomata, M.T.; Bisquer, J. Infección extraintestinal por Aeromonas spp.: Revisión de 38 casos. Enferm. Infecc. Microbiol. Clin. 2007, 25, 235–241. [Google Scholar] [CrossRef]
- Ivanova, E.P.; Flavier, S.; Christen, R. Phylogenetic relationships among marine Alteromonas-like proteobacteria: Emended description of the family Alteromonadaceae and proposal of Pseudoalteromonadaceae fam. nov., Colwelliaceae fam. nov., Shewanellaceae fam. nov., Moritellaceae fam. nov., Ferrimonadaceae fam. nov., Idiomarinaceae fam. nov. and Psychromonadaceae fam. nov. Int. J. Syst. Evol. Microbiol. 2004, 54, 1773–1788. [Google Scholar] [PubMed]
- Mountfort, D.O.; Rainey, F.A.; Burghardt, J.; Kaspar, H.F.; Stackebrandt, E. Psychromonas antarctica gen. nov., sp. nov., a new aerotolerant anaerobic, halophilic psychrophile isolated from pond sediment of the McMurdo Ice Shelf, Antarctica. Arch. Microbiol. 1998, 169, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Wertz, J.E.; Goldstone, C.; Gordon, D.M.; Riley, M.A. A molecular phylogeny of enteric bacteria and implications for a bacterial species concept. J. Evol. Biol. 2003, 16, 1236–1248. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, E.; Marco-Noales, E.; Gómez, D.; Lucas-Elío, P.; Ordax, M.; Garcias-Bonet, N.; Duarte, C.M.; Sanchez-Amat, A. Taxonomic study of Marinomonas strains isolated from the seagrass Posidonia oceanica, with descriptions of Marinomonas balearica sp. nov. and Marinomonas pollencensis sp. nov. Int. J. Syst. Evol. Microbiol. 2010, 60, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Romanenko, L.A. Psychrobacter submarinus sp. nov. and Psychrobacter marincola sp. nov., psychrophilic halophiles from marine environments. Int. J. Syst. Evol. Microbiol. 2002, 52, 1291–1297. [Google Scholar] [PubMed]
- Páèová, Z.; Urbanová, E.; Durnová, E. Psychrobacter immobilis isolated from foods: Characteristics and identification. Vet. Med. 2001, 46, 95–100. [Google Scholar] [CrossRef]
- Dijkshoorn, L. The Diversity of the Genus Acinetobacter. In Acinetobacter Molecular Biology; Gerischer, U., Ed.; Caister Academic Press: Poole, UK, 2008. [Google Scholar]
- Anzai, Y.; Kim, H.; Park, J.Y.; Wakabayashi, H. Phylogenetic affiliation of the pseudomonads based on 16S rRNA rRNA sequence. Int. J. Syst. Evol. Microbiol. 2000, 50, 1563–1589. [Google Scholar] [CrossRef]
- Rahal, J. Novel antibiotic combinations against infections with almost completely resistant Pseudomonas aeruginosa and Acinetobacter species. Clin. Infect. Dis. 2006, 43 (Suppl. 2), S95–S99. [Google Scholar] [CrossRef]
- Madigan, M.; Martinko, J.; Brock, T.; Dunlap, P.C. Brock Biology of Microorganisms, 12nd ed.; Benjamin Cummings: San Francisco, CA, USA, 2009. [Google Scholar]
- Austin, B.; Austin, D. A. Bacterial Fish Pathogens: Disease of Farmed and Wild Fish, 6th ed.; eBook; Springer: Berlin/Heidelberg, Germany, 2016; 732p. [Google Scholar]
- Stürmeyer, H.; Overmann, J.; Babenzien, H.D.; Cypionka, H. Ecophysiological and Phylogenetic Studies of Nevskia ramosa in Pure Culture. Appl. Environ. Microbiol. 1998, 64, 1890–1894. [Google Scholar] [CrossRef]
- Naushad, S.; Adeolu, M.; Wong, S.; Sohail, M.; Schellhorn, H.E.; Gupta, R.S. A phylogenomic and molecular marker based taxonomic framework for the order Xanthomonadales: Proposal to transfer the families Algiphilaceae and Solimonadaceae to the order Nevskiales ord. nov. and to create a new family within the order Xanthomonadales, the family Rhodanobacteraceae fam. nov., containing the genus Rhodanobacter and its closest relatives. Antonie Leeuwenhoek 2015, 107, 467–485. [Google Scholar] [PubMed]
- Suzuki, K.; Collins, M.D.; Iijima, E.; Komagata, K. Chemotaxonomic characterization of a radiotolerant bacterium Arthrobacter radiotolerans: Description of Rubrobacter radiotolerans gen. nov., comb. nov. FEMS Microbiol. Lett. 1988, 52, 33–40. [Google Scholar] [CrossRef]
- Collins, M.D.; Wallbanks, S. Comparative sequence analyses of the 16S rRNA rRNA genes of Lactobacillus minutus, Lactobacillus rimae and Streptococcus parvulus: Proposal for the creation of a new genus Atopobium. FEMS Microbiol. Lett. 1992, 95, 235–240. [Google Scholar] [CrossRef]
- Rodriguez Jovita, M.; Collins, M.D.; Sjödén, B.; Falsen, E. Characterization of a novel Atopobium isolate from the human vagina: Description of Atopobium vaginae sp. nov. Int. J. Syst. Bacteriol. 1999, 49, 1573–1576. [Google Scholar] [CrossRef]
- Cools, P.; Oyaert, M.; Vaneechoutte, M.; De Laere, E.; Vervaeke, S. Atopobium deltae sp. nov., isolated from the blood of a patient with Fournier’s gangrene. Int. J. Syst. Evol. Microbiol. 2014, 64, 3140–3145. [Google Scholar] [CrossRef]
- Waksman, S.A.; Schatz, A.; Reynolds, D.M. Production of antibiotic substances by Actinomycetes. Ann. N. Y. Acad. Sci. 2010, 1213, 112–124. [Google Scholar] [CrossRef]
- Kaur, G.; Kumar, N.; Mual, P.; Kumar, A.; Kumar, R.M.; Mayilraj, S. Brachybacterium aquaticum sp. nov., a novel actinobacterium isolated from seawater. Int. J. Syst. Evol. Microbiol. 2016, 66, 4705–4710. [Google Scholar] [CrossRef]
- Koerner, R.J.; Goodfellow, M.; Jones, A.L. The genus Dietzia: A new home for some known and emerging opportunist pathogens. FEMS Immunol. Med. Microbiol. 2009, 55, 296–305. [Google Scholar] [CrossRef]
- Ramanan, P.; Barreto, J.N.; Osmon, D.R.; Tosh, P.K. Rothia Bacteremia: A 10-Year Experience at Mayo Clinic, Rochester, Minnesota. J. Clin. Microbiol. 2014, 52, 3184–3189. [Google Scholar] [CrossRef]
- Murray, P.R.; Rosenthal, K.S.; Pfaller, M.A. Capítulo 28: Mycobacterium. In Microbiología Médica, 6th ed.; Murray, P.R., Ed.; Elsevier-Mosby: Madrid, España, 2009; pp. 277–290. [Google Scholar]
- Ryan, K.J.; Ray, C.G. (Eds.) Sherris Medical Microbiology, 4th ed.; McGraw Hill: Madrid, Spain, 2004; Volume xiii, 979p. [Google Scholar]
- Yang, J.A.; Kwon, K.K.; Oh, H.M. Complete Genome Sequence of Flavobacteriales Bacterium Strain UJ101 Isolated from a Xanthid Crab. Genome Announc. 2017, 5, e01551-16. [Google Scholar] [CrossRef]
- Noga, E.J. Fish Disease: Diagnosis and Treatment, 2nd ed.; Wiley-Blackwell: Ames, IA, USA, 2010; pp. 13–48. [Google Scholar]
- Krieg, N.R.; Staley, J.T.; Brown, D.R.; Hedlund, B.P.; Paster, B.J.; Ward, N.L.; Brown, D.; Parte, A. (Eds.) Bergey’s Manual of Systematic Bacteriology, 2nd ed.; The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes; Springer: New York, NY, USA, 2010. [Google Scholar]
- Grégoire, P.; Bohli, M.; Cayol, J.L.; Joseph, M.; Guasco, S.; Dubourg, K.; Cambar, J.; Michotey, V.; Bonin, P.; Fardeau, M.L.; et al. Caldilinea tarbellica sp. nov., a filamentous, thermophilic, anaerobic bacterium isolated from a deep hot aquifer in the Aquitaine Basin. Int. J. Syst. Evol. Microbiol. 2011, 61, 1436–1441. [Google Scholar] [CrossRef]
- Hanada, S.; Takaichi, S.; Matsuura, K.; Nakamura, K. Roseiflexus castenholzii gen. nov., sp. nov., a thermophilic, filamentous, photosynthetic bacterium that lacks chlorosomes. Int. J. Syst. Evol. Microbiol. 2002, 52, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Cavalier-Smith, T. Rooting the tree of life by transition analyses. Biol. Direct. 2006, 1, 19. [Google Scholar] [CrossRef][Green Version]
- Edelson, B.T.; Unanue, E.R. Immunity to Listeria infection. Curr. Opin. Imunol. 2000, 12, 425–431. [Google Scholar] [CrossRef]
- Zaitsev, G.M.; Tsitko, I.V.; Rainey, F.A.; Trotsenko, Y.A.; Uotila, J.S.; Stackebrandt, E.; Salkinoja-Salonen, M.S. New aerobic ammonium-dependent obligately oxalotrophic bacteria: Description of Ammoniphilus oxalaticus gen. nov., sp. nov. and Ammoniphilus oxalivorans gen. nov., sp. nov. Int. J. Syst. Bacteriol. 1998, 48, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Mei, Y. Screening and distributing features of bacteria with hydantoinase and carbamoylase. Microbiol. Res. 2009, 3, 322–329. [Google Scholar] [CrossRef]
- Foster, T. Staphylococcus. In Medical Microbiology, 4th ed.; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996; Chapter 12. Available online: https://www.ncbi.nlm.nih.gov/books/NBK8448 (accessed on 24 September 2019).
- Yoon, J.H.; Kim, I.G.; Shin, Y.K.; Park, Y.H. Proposal of the genus Thermoactinomyces sensu stricto and three new genera, Laceyella, Thermoflavimicrobium and Seinonella, on the basis of phenotypic, phylogenetic and chemotaxonomic analyses. Int. J. Syst. Evol. Microbiol. 2005, 55, 395–400. [Google Scholar] [CrossRef] [PubMed]
- Gatesoupe, F.J. Updating the importance of lactic acid bacteria in fish farming: Natural occurrence and probiotic treatments. J. Mol. Microb. Biotech. 2008, 14, 107–114. [Google Scholar] [CrossRef]
- Harimaya, A.; Takada, R.; Hendolin, P.H.; Fujii, N.; Ylikoski, J.; Himi, T. High Incidence of Alloiococcus otitidis in Children with Otitis Media, Despite Treatment with Antibiotics. J. Clin. Microbiol. 2006, 44, 946–949. [Google Scholar] [CrossRef]
- Lorenzo, J.M.; Munekata, P.E.; Dominguez, R.; Pateiro, M.; Saraiva, J.A.; Franco, D. Main Groups of Microorganisms of Relevance for Food Safety and Stability: General Aspects and Overall Description. In Innovative Technologies for Food Preservation. Inactivation of Spoilage and Pathogenic Microorganisms; Barba, F.J., Sant’Ana, A.S., Orlien, V., Koubaa, M., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 53–107. [Google Scholar]
- Hogan, C.M. Extremophile. In Encyclopedia of Earth; Monosson, E., Cleveland, C., Eds.; National Council for Science and the Environment: Washington, DC, USA, 2010. [Google Scholar]
- Boutard, M.; Cerisy, T.; Nogue, P.Y. Functional diversity of carbohydrate-active enzymes enabling a bacterium to ferment plant biomass. PLoS Genet. 2014, 10, e1004773. [Google Scholar] [CrossRef] [PubMed]
- Labus, J.S.; Hsiao, E.; Tap, J.; Derrien, M.; Gupta, A.; Le Nevé, B.; Brazeilles, R.; Grinsvall, C.; Ohman, L.; Törnblom, H.; et al. Clostridia from the Gut Microbiome are Associated with Brain Functional Connectivity and Evoked Symptoms in IBS. Gastroenterology 2017, 152, S40. [Google Scholar] [CrossRef]
- Kuske, C.R.; Barns, S.M.; Busch, J.D. Diverse uncultivated bacterial groups from soils of the arid southwestern United States that are present in many geographic regions. Appl. Environ. Microbiol. 1997, 63, 3614–3621. [Google Scholar] [CrossRef]
- Quaiser, A.; Ochsenreiter, T.; Lanz, C.; Schuster, S.C.; Treusch, A.H.; Eck, J.; Schleper, C. Acidobacteria form a coherent but highly diverse group within the bacterial domain: Evidence from environmental genomics». Mol. Microbiol. 2003, 50, 563–575. [Google Scholar] [CrossRef]
- Kawasumi, T.; Igarashi, Y.; Kodama, T.; Minoda, Y. Hydrogenobacter thermophilus gen. nov., sp. nov., an extremely thermophilic, aerobic, hydrogen-oxidizing bacterium. Int. J. Syst. Bacteriol. 1984, 34, 5–10. [Google Scholar] [CrossRef]
- Huber, R.; Wilharm, T.; Huber, D.; Trincone, A.; Burggaf, S.; Konig, H.; Reinhard, R.; Rockinger, I.; Fricke, H.; Stetter, K.O. Aquifex pyrophilus gen. nov. sp. nov., represents a novel group of marine hyperthermophilic hydrogen-oxidizing bacteria. Syst. Appl. Microbiol. 1992, 15, 340–351. [Google Scholar] [CrossRef]
- Griffiths, E.; Gupta, R.S. Identification of signature proteins that are distinctive of the Deinococcus–Thermus phylum. Int. Microbiol. 2007, 10, 201–208. [Google Scholar] [PubMed]
- Albuquerque, L.; Simões, C.; Nobre, M.F.; Pino, N.M.; Battista, J.; Silva, M.T.; Rainey, F.A.; da Costa, M.S. Truepera radiovictrix gen. nov., sp. nov., a new radiation resistant species and the proposal of Trueperaceae fam. nov. FEMS Microbiol. Lett. 2005, 247, 161–169. [Google Scholar] [CrossRef]
- Yang, Y.; Itoh, T.; Yokobori, S.; Itahashi, S.; Shimada, H.; Satoh, K.; Ohba, H.; Narumi, I.; Yamagishi, A. Deinococcus aerius sp. nov., isolated from the high atmosphere. Int. J. Syst. Evol. Microbiol. 2009, 59, 1862–1866. [Google Scholar] [CrossRef]
- Zhao, J.S.; Manno, D.; Hawari, J. Psychrilyobacter atlanticus gen. nov., sp. nov., a marine member of the phylum Fusobacteriathatproduces H2and degrades nitramine explosivesunder low temperature condition. Int. J. Syst. Evol. Microbiol. 2009, 59, 491–497. [Google Scholar] [CrossRef]
- Harwich, M.D.; Serrano, M.G.; Fettweis, J.M.; Alves, J.M.P.; Reimers, M.A.; Vaginal Microbiome Consortium (additional members); Buck, G.A.; Jefferson, K.K. Genomic sequence analysis and characterization of Sneathia amnii sp. nov. BMC Genom. 2012, 13, S4. [Google Scholar] [CrossRef] [PubMed]
- Glöckner, F.O.; Kube, M.; Bauer, M.; Teeling, H.; Lombardot, T.; Ludwig, W.; Gade, D.; Beck, A.; Borzym, K.; Heitman, K.; et al. Complete genome sequence of the marine planctomycete Pirellula sp. strain 1. Proc. Natl. Acad. Sci. USA 2003, 100, 8298–8303. [Google Scholar] [CrossRef] [PubMed]
- Kulichevskaya, I.S.; Ivanova, A.O.; Baulina, O.I.; Bodelier, P.L.; Damsté, J.S.; Dedysh, S.N. Singulisphaera acidiphila gen. nov., sp. nov., a non-filamentous, Isosphaera-like planctomycete from acidic northern wetlands. Int. J. Syst. Evol. Microbiol. 2008, 58, 1186–1193. [Google Scholar] [CrossRef] [PubMed]
- Fuerst, J.; Sagulenko, E. Beyond the bacterium: Planctomycetes challenge our concepts of microbial structure and function. Nat. Rev. Microbiol. 2011, 9, 403–413. [Google Scholar] [CrossRef] [PubMed]
- Jumas-Bilak, E.; Carlier, J.P.; Jean-Pierre, H.; Citron, D.; Bernard, K.; Damay, A.; Gay, B.; Teyssier, C.; Campos, J.; Marchandin, H. Jonquetella anthropi gen. nov., sp. nov., the first member of the candidate phylum ‘Synergistetes’ isolated from man. Int. J. Syst. Bacteriol. 2007, 57, 2743–2748. [Google Scholar] [CrossRef]
- Albert, R.A.; Waas, N.E.; Langer, S.; Pavlons, S.C.; Feldner, J.L.; Rosselló-Mora, R.; Busse, H.J. Labrys wisconsinensis sp. nov., a budding bacterium isolated from Lake Michigan water, and emended description of the genus Labrys. Int. J. Syst. Evol. Microbiol. 2010, 60, 1570–1576. [Google Scholar] [CrossRef]
- Han, X.Y.; Falsen, E. Characterization of oral strains of Cardiobacterium valvarum and emended description of the organism. J. Clin. Microbiol. 2005, 43, 2370–2374. [Google Scholar] [CrossRef]
- Albuquerque, L.; Ferreira, C.; Tomaz, D.; Tiago, T.; Veríssimo, A.; da Costa, M.S.; Nobre, M.F. Meiothermus rufus sp. nov., a new slightly thermophilic red-pigmentedspecies and emended description of the genus Meiothermus. Syst. App. Microbiol. 2009, 32, 306–313. [Google Scholar] [CrossRef] [PubMed]
- Chin, K.J.; Liesack, W.; Janssen, P.H. Opitutus terrae gen. nov., sp. nov., to accommodate novel strains of the division Verrucomicrobia isolated from rice paddy soil. Int. J. Syst. Evol. Microbiol. 2001, 51, 1965–1968. [Google Scholar] [CrossRef]
- Kocur, M.; Kloos, W.E.; Schleifer, K.H. The Genus Micrococcus. In The Prokaryotes; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; pp. 961–971. [Google Scholar] [CrossRef]
- Mattarelli, P.; Brandi, G.; Calabrese, C.; Fornari, F.; Prati, G.M.; Biavati, B.; Sgorbati, B. Occurrence of Bifidobacteriaceae in human hypochlorhydria stomach. Microb. Ecol. Health Dis. 2014, 25. [Google Scholar] [CrossRef]
- Maruo, T.; Sakamoto, M.; Ito, C.; Toda, T.; Benno, Y. Adlercreutzia equolifaciens gen. Nov., sp. Nov., an equol-producing bacterium isolated from human faeces, and emended description of the genus Eggerthella. Int. J. Syst. Evol. Microbiol. 2008, 58, 1221–1227. [Google Scholar] [CrossRef]
- Ormerod, K.L.; Wood, D.L.A.; Lachner, N.; Gellatly, S.L.; Daly, J.N.; Parsons, J.D.; Dal’Molin, C.G.O.; Palfreyman, R.W.; Nielsen, L.K.; Cooper, M.A.; et al. Genomic characterization of the uncultured Bacteroidales family S24-7 inhabiting the guts of homeothermic animals. Microbiome 2016, 4, 36. [Google Scholar] [CrossRef]
- Wang, K.; Lu, W.; Tu, Q.; Ge, Y.; He, J.; Zhou, Y.; Gou, Y.; Van Nostrand, J.D.; Qin, Y.; Li, J.; et al. Preliminary analysis of salivary microbiome and their potential roles in oral lichen planus. Sci. Rep. 2016, 6, 22943. [Google Scholar] [CrossRef] [PubMed]
- Seshadri, R.; Leahy, S.C.; Attwood, G.T.; Teh, K.H.; Lambie, S.C.; Cookson, A.L.; Eloe-Fadrosh, E.A.; Pavlopoulos, G.A.; Hadjithomas, M.; Varghese, N.J.; et al. Cultivation and sequencing of rumen microbiome members from the Hungate1000 Collection. Nat. Biotechnol. 2018, 36, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Kanki, P.; Grimes, D.J. (Eds.) Infectious Diseases Selected Entries from the Encyclopedia of Sustainability Science and Technology; Springer: New York, NY, USA, 2013. [Google Scholar]
- Stanton, T.B.; Savage, D.C. Roseburia cecicola gen. nov., sp. nov., a motile, obligately anaerobic bacterium from a mouse cecum. Int. J. Syst. Bacteriol. 1983, 33, 618–627. [Google Scholar] [CrossRef]
- Duncan, S.H.; Aminov, R.I.; Scott, K.P.; Louis, P.; Stanton, T.B.; Flint, H.J. Proposal of Roseburia faecis sp. nov., Roseburia hominis sp. nov. and Roseburia inulinivorans sp. nov., based on isolates from human faeces. Int. J. Syst. Evol. Microbiol. 2006, 56, 2437–2441. [Google Scholar] [CrossRef] [PubMed]
- Holdeman, L.V.; Moore, W.E.C. New genus, Coprococcus, twelve new species, and emended descriptions of four previously described species of bacteria from human feces. Int. J. Syst. Bacteriol. 1974, 24, 260–277. [Google Scholar] [CrossRef]
- McLellan, S.L.; Newton, R.J.; Vandewalle, J.L.; Shanks, O.C.; Huse, S.M.; Eren, A.M.; Sogin, M.L. Sewage reflects the distribution of human faecal Lachnospiraceae. Environ. Microbiol. 2013, 15, 2213–2222. [Google Scholar] [CrossRef] [PubMed]
- Taras, D.; Simmering, R.; Collins, M.D.; Lawson, P.A.; Blaut, M. Reclassification of Eubacterium formicigenerans Holdeman and Moore 1974 as Dorea formicigenerans gen. nov., comb. nov., and description of Dorea longicatena sp. nov., isolated from human faeces. Int. J. Syst. Evol. Microbiol. 2002, 52, 423–428. [Google Scholar] [CrossRef]
- Bryant, M.P.; Small, N. Characteristics of two new genera of anaerobic curved rods isolated from the rumen of cattle. J. Bacteriol. 1956, 72, 22–26. [Google Scholar] [CrossRef]
- Cornick, N.A.; Jensen, N.S.; Stahl, D.A.; Hartman, P.A.; Allison, M.J. Lachnospira pectinoschiza sp. nov., an anaerobic pectinophile from the pig intestine. Int. J. Syst. Bacteriol. 1994, 44, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Rainey, F.A. Ruminococcaceae fam. nov. In Bergey’s Manual of Systematics of Archaea and Bacteria; Whitman, W.B., Ed.; in association with Bergey’s Manual Trust; Wiley and Sons, Inc.: Hoboken, NJ, USA, 2015; Volume 2. [Google Scholar] [CrossRef]
- Khan, M.T.; Duncan, S.H.; Stams, A.J.; van Dijl, J.M.; Flint, H.J.; Harmsen, H.J. The gut anaerobe Faecalibacterium prausnitzii uses an extracellular electron shuttle to grow at oxic-anoxic interphases. ISME J. 2012, 6, 1578–1585. [Google Scholar] [CrossRef]
- Rajilić-Stojanović, M.; de Vos, W.M. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol. Rev. 2014, 38, 996–1047. [Google Scholar] [CrossRef] [PubMed]
- Holmstrøm, K.; Collins, M.D.; Møller, T.; Falsen, E.; Lawson, P.A. Subdoligranulum variabile gen. nov., sp. nov. from human feces. Anaerobe 2004, 10, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Iino, T.; Mori, K.; Tanaka, K.; Suzuki, K.I.; Harayama, S. Oscillibacter valericigenes gen. nov., sp. nov., a valerate-producing anaerobic bacterium isolated from the alimentary canal of a Japanese corbicula clam. Int. J. Syst. Evol. Microbiol. 2007, 57, 1840–1845. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.H.; Rhee, M.S.; Chang, D.H.; Lee, J.; Kim, S.; Yoon, M.H.; Kim, B.C. Oscillibacter ruminantium sp. nov., isolated from the rumen of Korean native cattle. Int. J. Syst. Evol. Microbiol. 2013, 63, 1942–1946. [Google Scholar] [CrossRef] [PubMed]
- Bosch, T.C.G.; McFall-Ngai, M.J. Metaorganisms as the new frontier. Zoology 2011, 114, 185–190. [Google Scholar] [CrossRef]
- Bordenstein, S.R.; Theis, K.R. Host biology in light of the microbiome: Ten principles of holobionts and hologenomes. PLoS Biol. 2015, 13, e1002226. [Google Scholar] [CrossRef]
- Zilber-Rosenberg, I.; Rosenberg, E. Role of microorganisms in the evolution of animals and plants: The hologenome theory of evolution. FEMS Microbiol. Rev. 2008, 32, 723–735. [Google Scholar] [CrossRef]
- Dirksen, P.; Arnaud Marsh, S.; Braker, I.; Heitland, N.; Wagner, S.; Nakad, R.; Mader, S.; Petersen, C.; Kowallik, V.; Rosenstiel, P.; et al. The native microbiome of the nematode Caenorhabditis elegans: Gateway to a new host-microbiome model. BMC Biol. 2016, 14, 38. [Google Scholar] [CrossRef]
- Palomares-Rius, J.E.; Archidona-Yuste, A.; Cantalapiedra-Navarrete, C.; Prieto, P.; Castillo, P. Molecular diversity of bacterial endosymbionts associated with dagger nematodes of the genus Xiphinema (Nematoda: Longidoridae) reveals a high degree of phylogenetic congruence with their host. Mol. Ecol. 2016, 25, 6225–6247. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Mylonakis, E. Caenorhabditis elegans immune conditioning with the probiotic bacterium Lactobacillus acidophilus strain NCFM enhances grampositive immune responses. Infect. Immun. 2012, 80, 2500–2508. [Google Scholar] [CrossRef] [PubMed]
- Danna, P.L.; Urban, C.; Rahal, J.J.; Bellin, E. Role of Candida in pathogenesis of antibiotic-associated diarrhoea in elderly inpatients. Lancet 1991, 337, 511–514. [Google Scholar] [CrossRef]
- Morran, L.T.; Penley, M.J.; Byrd, V.S.; Meyer, A.J.; O’Sullivan, T.S.; Bashey, F.; Goodrich-Blair, H.; Lively, C.M. Nematode-Bacteria Mutualism: Selection Within the Mutualism Supersedes Selection Outside of the Mutualism. Evolution 2016, 70, 687–695. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zeppilli, D.; Bellec, L.; Cambon-Bonavita, M.A.; Decraemer, W.; Fontaneto, D.; Fuchs, S.; Gayet, N.; Mandon, P.; Michel, L.N.; Portail, M.; et al. Ecology and trophic role of Oncholaimus dyvae sp. nov. (Nematoda: Oncholaimidae) from the lucky strike hydrothermal vent field (Mid-Atlantic Ridge). BMC Zool. 2019, 4, 6. [Google Scholar] [CrossRef]
- Bellec, L.; Cambon-Bonavita, M.A.; Cueff-Gauchard, V.; Durand, L.; Gayet, N.; Zeppilli, D.A. Nematode of the Mid-Atlantic Ridge Hydrothermal Vents Harbors a Possible Symbiotic Relationship. Front. Microbiol. 2018, 9, 2246. [Google Scholar] [CrossRef]
- Sanders, H.L. Benthic marine diversity and the stability-time hypothesis. Brookhaven Symp. Biol. 1969, 22, 71–81. [Google Scholar]
- Hughes, J.B.; Bohannan, B.J.M. Application of ecological diversity statistics in microbial ecology. Mol. Microb. Ecol. Manual 2004, 1321–1344. [Google Scholar] [CrossRef]
- Kim, B.R.; Shin, J.; Guevarra, R.B.; Lee, J.H.; Wan Kim, D.; Seol, K.H.; Lee, J.H.; Kim, H.B.; Isaacson, R. Deciphering Diversity Indices for a Better Understanding of MicrobialCommunitie. J. Microbiol. Biotechnol. 2017, 27, 2089–2093. [Google Scholar] [CrossRef]
- Rosenfeld, J.S. Functional redundancy in Ecology and Conservation. OIKOS 2002, 98, 156–162. [Google Scholar] [CrossRef]
- Xue, Z.; Zhang, W.; Wang, L.; Hou, R.; Zhang, M.; Fei, L.; Zhang, X.; Huang, H.; Bridgewater, L.C.; Jiang, Y.; et al. The Bamboo-Eating Giant Panda Harbors a Carnivore-Like Gut Microbiota, with Excessive Seasonal Variations. mBio 2015, 6, e00022-15. [Google Scholar] [CrossRef] [PubMed]
- Ettema, C.H. Soil nematode diversity: Species coexistence and ecosystem function. J. Nematol. 1998, 30, 159–169. [Google Scholar]
- De Ley, P. A quick tour of nematode diversity and the backbone of nematode phylogeny. WormBook 2006, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Borgonie, G.; Garcia-Moyano, A.; Litthauer, D.; Bert, W.; Bester, A.; van Heerden, E.; Moller, C.; Erasmus, M.; Onstott, T.C. Nematoda from the terrestrial deep subsurface of South Africa. Nature 2011, 474, 79–82. [Google Scholar] [CrossRef]
- Poinar, G.O., Jr.; Hansen, E.L. Associations between nematodes and bacteria. Helminthol. Abst. B 1986, 55, 61–79. [Google Scholar]
- Goodrich-Blair, H. They’ve got a ticket to ride: Xenorhabdus nematophila-Steinernema carpocapsae symbiosis. Curr. Opin. Microbiol. 2007, 10, 225–230. [Google Scholar] [CrossRef]
- Gram, L.; Huss, H.H. Microbiological spoilage of fish and fish products. In. J. Food Microbiol. 1996, 33, 121–137. [Google Scholar] [CrossRef]
- Svanevik, C.S.; Lunestad, B.T. Characterisation of the microbiota of Atlantic mackerel (Scomber scombrus). Int. J. Food Microbiol. 2011, 151, 164–170. [Google Scholar] [CrossRef]
- Huss, H. Quality and quality changes in fresh fish. In FAO Fisheries; Technical Paper No. 348; FAO: Rome, Italy, 1995. [Google Scholar]
- Svanevik, C.S.; Levsen, A.; Lunestad, B.T. The role of muscle-invading anisakid larvae on bacterial contamination of the flesh of post-harvest blue whiting (Micromesistius poutassou). Food Control 2013, 30, 526–530. [Google Scholar] [CrossRef]
- Svanevik, C.S.; Lunestad, B.T.; Levsen, A. Effect of Anisakis simplex (sl) larvae on the spoilage rate and shelf-life of fish mince prod ucts under laboratory conditions. Food Control 2014, 46, 121–126. [Google Scholar] [CrossRef]
- Novotny, L.; Dvorska, L.; Lorencova, A.; Beran, V.; Pavlik, I. Fish: A potential source of bacterial pathogens for human beings. Vet. Med. 2004, 49, 343–358. [Google Scholar] [CrossRef]
- Svanevik, C.S.; Roiha, I.S.; Levsen, A.; Lunestad, B.T. Microbiological assessment along the fish production chain of the Norwegian pelagic fisheries sector results from a spot sampling programme. Food Microbiol. 2015, 51, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Butt, A.A.; Aldridge, K.E.; Sanders, C.V. Infections related to the ingestion of seafood Part I: Viral and bacterial infections. Lancet Infec. Dis. 2004, 4, 201–212. [Google Scholar] [CrossRef]
- Chaillou, S.; Chaulot-Talmon, A.; Caekebeke, H.; Cardinal, M.; Christieans, S.; Denis, C.; Desmonts, M.H.; Dousset, X.; Feurer, C.; Hamon, E.; et al. Origin and ecological selection of core and food-specific bacterial communities associated with meat and seafood spoilage. ISME J. 2015, 9, 1105–1118. [Google Scholar] [CrossRef] [PubMed]
| Acidobacteria | Actinobacteria* | Aquificae* | Bacteroidetes* | Chlamydiae | Chloroflexi* | Deinococcus-Thermus* | Firmicutes* | Fusobacteria* | Planctomycetes* | Proteobacteria* | Synergistetes* | Verrucomicrobia | Cyanobacteria* | Unclassified (105 seqs) | ||
| Acidobacteria | Acidobacteriales | (75.2%), 59.80 | ||||||||||||||
| Rubrobacteria | Rubrobacterales | (13.3%), 8.93 | ||||||||||||||
| Actinobacteria | Acidimicrobiales Actinomycetales Bifidobacteriales * | (54%), 75.79 (100%), 2625.95 (57.52%), 79.78 | ||||||||||||||
| Coriobacteriia | Coriobacteriales | (65.48%), 76.01 | ||||||||||||||
| Aquificae | Aquificales* | (2.65%), 76.00 | ||||||||||||||
| Bacteroidia | Bacteroidales* | (97.34%), 211.62 | ||||||||||||||
| Flavobacteria | Flavobacteriales* | (99.11%), 254.03 | ||||||||||||||
| Sphingobacteria | Sphingobacteriales | (91.15%), 129.55 | ||||||||||||||
| Chlamydiae | Chlamydiales | (15.93%), 9.55 | ||||||||||||||
| Anaerolineae | Anaerolineales | (13.27%), 21.13 | ||||||||||||||
| Caldilineae | Caldilineales | (7.08%), 23.87 | ||||||||||||||
| Chloroflexia | Chloroflexales* | (10.62%), 5.41 | ||||||||||||||
| Deinococci | Thermales Deinoccocales | (6.19%), 30.14 (59.30%), 21.50 | ||||||||||||||
| Bacilli | Bacillales Lactobacillales | (99.11%), 112,41 (99.11%), 323.06 | ||||||||||||||
| Mollicutes | Mycoplasmatales | (8.10%), 16.78 | ||||||||||||||
| Erysipelotrichia | Erysipelothrichales | (52.21%), 22.76 | ||||||||||||||
| Clostridia | Clostridiales* Thermoanaerobacteriales | (100%), 4450.98 (8.85%), 21.90 | ||||||||||||||
| Fusobacteria | Fusobacteriales | (100%), 755.18 | ||||||||||||||
| Planctomycetya | Planctomycetales* | (45.13%), 16.43 | ||||||||||||||
| Phycisphaerae | Phycisphaerales | (5.31%), 32.00 | ||||||||||||||
| Alphaproteobacteria | Caulobacterales Parvularculales* Rhizobiales Rhodobacterales* Rhodospirillales Rickettsiales Sphingomonadales* | (94.70%), 667.46 (0.89%), 31.00 (100%), 2472.23 (91,15%), 192.87 (96,46%), 197.50 (7,68%), 15.11 (100%), 2692.78 | ||||||||||||||
| Betaproteobacteria | Burkholderiales Neisseriales* | (100%), 189.26(83,18%), 28.25 | ||||||||||||||
| Epsiloproteobacteria | Campylobacteriales | (95.60%), 540.42 | ||||||||||||||
| Deltaproteobacteria | Bdellovibrionales Desulfobacterales Desulfovibrionales Desulfuromonadales Myxococcales | (14.16%), 25.31 (1.76%), 15,50 (6.2%), 6.28 (7.96%), 7.22 (25.66%), 10.93 | ||||||||||||||
| Gammaproteobacteria | Aeromonadales Alteromonadales Cardiobacteriales Chromatiales* Enterobacteriales Oceanospirillales Pasteurellales* Pseudomonadales Vibrionales Xanthomonadales | (44.24%), 77.56 (53,09%), 22.10 (19,46%), 11.77 (8.85%), 7.30 (87.61%), 35,56 (30.97%), 7.65(80.53%), 37,61 (100%), 301.72 (82,30%), 199.86 (57,52%), 10.41 | ||||||||||||||
| Synergistia | Synergistales | |||||||||||||||
| Opitutae | Opitutales | |||||||||||||||
| unclassified | unclassified | |||||||||||||||
| MLE1-12 | unclassified | |||||||||||||||
| Subsection IV | unclassified | |||||||||||||||
| Subsection III | unclassified | |||||||||||||||
| (Phyla % in total samples) Mean of sequences | (76%) 59.60 | (100%) 2838.28 | (2.65%) 76.00 | (100%) 705.41 | (15.93%) 9.55 | (74.33%) 66.13 | (61.94%) 23.87 | (100%) 5659.03 | (100%) 755.18 | (63.72%) 35.56 | (100%) 9291.94 | (10.62%) 73.75 | (23.9%) 17.59 | (100%) 297.74 |
| Alphaproteobacteria | Betaproteobacteria | Epsiloproteobacteria | Deltaproteobacteria | Gammaproteobacteria | |||||||||||||||||||||
| Caulobacterales | Parvularculales | Rhizobiales | Rhodobacterales | Rhodospirillales | Rickettsiales | Sphingomonadales | Burkholderiales | Neisseriales | Campylobacterales | Bdellovibrionales | Desulfobacterales | Desulfovibrionales | Desulfuromonadales | Myxococcales | Aeromonadales | Alteromonadales | Cardiobacteriales | Chromatiales | Enterobacteriales | Oceanospirillales | Pasteurellales | Pseudomonadales | Vibrionales | Xanthomonadales | |
| Caulobacteraceae | Caulobacter Brevundimonas Phenylobacterium Asticcacaulis | ||||||||||||||||||||||||
| Parvularculaceae | Parvularcula | ||||||||||||||||||||||||
| Bradyrhizobiaceae | Bradyrhizobium unclassified Bosea | ||||||||||||||||||||||||
| Methylobacteriaceae | Methylobacterium* | ||||||||||||||||||||||||
| unclassified | unclassified | ||||||||||||||||||||||||
| Hyphomicrobiaceae | Devosia* unclassified Hyphomicrobium Angulomicrobium Pedomicrobium | ||||||||||||||||||||||||
| Rhodobiaceae | RhodobiuM* | ||||||||||||||||||||||||
| Phyllobacteriaceae | Mesorhizobium unclassified | ||||||||||||||||||||||||
| Xanthobacteraceae | Labrys | ||||||||||||||||||||||||
| Beijerinckiaceae | Chelatococcus | ||||||||||||||||||||||||
| Methylocystaceae | unclassified | ||||||||||||||||||||||||
| Rhodobiaceae | Parvibaculum | ||||||||||||||||||||||||
| Brucellaceae | Ochrobactrum | ||||||||||||||||||||||||
| Nordella cluster | Nordella | ||||||||||||||||||||||||
| Rhizobiaceae | unclassified | ||||||||||||||||||||||||
Rhodobacteraceae | Paracoccus* unclassified Roseobacter_clade | ||||||||||||||||||||||||
| Acetobacteraceae | Gluconobacter unclassified Roseomonas* Acetobacter Acidocella Acidiphilium | ||||||||||||||||||||||||
| Rhodospirillaceae | Skermanella Azospirillum Caenispirillum Magnetospirillum unclassified Defluviicoccus | ||||||||||||||||||||||||
| unclassified | unclassified | ||||||||||||||||||||||||
| wr0007 | unclassified | ||||||||||||||||||||||||
| Candidatus_Alysiosphaera cluster | unclassified | ||||||||||||||||||||||||
| Anaplasmataceae | Wolbachia | ||||||||||||||||||||||||
| Rickettsiaceae | Rickettsia* | ||||||||||||||||||||||||
| Candidatus_Odyssella cluster | unclassified | ||||||||||||||||||||||||
| SM2D12 | unclassified | ||||||||||||||||||||||||
| Sphingomonadaceae | Sphingomonas* Novosphingobium* unclassified Sphingobium Sphingopyxis* | ||||||||||||||||||||||||
| unclassified | unclassified | ||||||||||||||||||||||||
| Erythrobacteraceae | unclassified Erythrobacter | ||||||||||||||||||||||||
| GOBB3-C201 | unclassified | ||||||||||||||||||||||||
| Oxalobacteraceae | Janthinobacterium* | ||||||||||||||||||||||||
| Alcaligenaceae | Alcaligenes | ||||||||||||||||||||||||
| Comamonadaceae | unclassified | ||||||||||||||||||||||||
| Burkholderiaceae | Ralstonia | ||||||||||||||||||||||||
| Neisseriaceae | Neisseria* unclassified | ||||||||||||||||||||||||
Campylobacteraceae | Arcobacter Campylobacter* Sulfurospirillum | ||||||||||||||||||||||||
| Bacteriovoraceae | Peredibacter | ||||||||||||||||||||||||
| Bdellovibrionacea | Bdellovibrio | ||||||||||||||||||||||||
| Desulfobulbaceae | Desulfobulbus | ||||||||||||||||||||||||
| Desulfovibrionaceae | unclassified | ||||||||||||||||||||||||
| Geobacteraceae | Geobacter | ||||||||||||||||||||||||
| Polyangiaceae | unclassified | ||||||||||||||||||||||||
| Haliangiaceae | Haliangium | ||||||||||||||||||||||||
| Unclassified | unclassified | ||||||||||||||||||||||||
| Aeromonadaceae | Aeromonas | ||||||||||||||||||||||||
| Succinivibrionaceae | Succinivibrio | ||||||||||||||||||||||||
| Shewanellaceae | Shewanella* | ||||||||||||||||||||||||
| Psychromonadaceae | Psychromonas | ||||||||||||||||||||||||
| Cardiobacteriaceae | unclassified Cardiobacterium | ||||||||||||||||||||||||
| Chromatiaceae | Unclassified | ||||||||||||||||||||||||
| Enterobacteriaceae | unclassified Enteric_Bacteria_cluster | ||||||||||||||||||||||||
| Oceanospirillaceae | Marinomonas | ||||||||||||||||||||||||
| Pasteurellaceae | unclassified | ||||||||||||||||||||||||
| Pseudomonadaceae | Pseudomonas* unclassified | ||||||||||||||||||||||||
Moraxellaceae | Psychrobacter Acinetobacter unclassified | ||||||||||||||||||||||||
| Vibrionaceae | unclassified Photobacterium | ||||||||||||||||||||||||
| Sinobacteraceae | Nevskia | ||||||||||||||||||||||||
| Xanthomonadaceae | unclassified | ||||||||||||||||||||||||
| Actinobacteria | Bacteroidetes | Chloroflexi | Firmicutes | ||||||||||||||
| Rubrobacteria | Actinobacteria | Coriobacteria | Bacteroidia | Flavobacteria | Sphingobacteria | Anaerolineae | Caldilineae | Chloroflexia | Bacilli | Mollicutes | Erysipelotrichia | Clostridia | |||||
| Rubrobacterales | Acidimicrobiales | Actinomycetales | Bifidobacteriales | Coriobacteriales | Bacteroidales | Flavobacteriales | Sphingobacteriales | Anaerolineales | Caldilineales | Chloroflexales | Bacillales | Lactobacillales | Mycoplasmatales | Erysipelotrichales | Clostridiales | Thermoanaerobacterales | |
| Rubrobacteriaceae | Rubrobacter | ||||||||||||||||
| Acidimicrobiaceae (5) | unclassified | ||||||||||||||||
| Actinomycetaceae (2) | Actinomyces* Mobiluncus | ||||||||||||||||
| Brevibacteriaceae (2) | Brevibacterium* | ||||||||||||||||
| Cellulomonadaceae | unclassified | ||||||||||||||||
| Corynebacteriaceae | Corynebacterium* | ||||||||||||||||
| Dermabacteraceae | Brachybacterium* | ||||||||||||||||
| Dermatophilaceae | unclassified | ||||||||||||||||
| Dietziaceae | Dietzia | ||||||||||||||||
| Geodermatophilaceae | Blastococcus Geodermatophilus | ||||||||||||||||
| Glycomycetaceae | Glycomyces | ||||||||||||||||
| Intrasporangiaceae | Ornithinicoccus | ||||||||||||||||
| Kineosporiaceae | Kineococcus | ||||||||||||||||
| Microbacteriaceae | unclassified | ||||||||||||||||
| Micrococcaceae | Micrococcus* Rothia * Kocuria Arthrobacter | ||||||||||||||||
| Micromonosporaceae | Actinoplanes | ||||||||||||||||
| Mycobacteriaceae | Mycobacterium* | ||||||||||||||||
Nocardiaceae | Rhodococcus Gordonia Nocardia | ||||||||||||||||
Nocardioidaceae | Nocardioides* Aeromicrobium Kribbella | ||||||||||||||||
| Propionibacteriaceae | Propionibacterium* Friedmanniella | ||||||||||||||||
Pseudonocardiaceae | Pseudonocardia Amycolatopsis | ||||||||||||||||
| Sporichthyaceae | hgcI_clade | ||||||||||||||||
| Streptomycetaceae | Streptomyces | ||||||||||||||||
| Streptosporangiaceae | Nonomuraea | ||||||||||||||||
| Thermomonosporaceae | Actinomadura | ||||||||||||||||
| Bifidobacteriaceae | Alloscardovia Bifidobacterium Gardnerella | ||||||||||||||||
| Coriobacteriaceae | Atopobium* Olsenella* Slackia Unclassified Collinsella Adlercreutzia Eggerthella | ||||||||||||||||
| Prevotellaceae | Prevotella* | ||||||||||||||||
| Bacteroidaceae | Bacteroides* | ||||||||||||||||
Porphyromonadaceae | Porphyromonas* Parabacteroides Tannerella* Odoribacter | ||||||||||||||||
| Flavobacteriaceae | Flavobacterium* unclassified Cloacibacterium Capnocytophaga* Chryseobacterium Polaribacter Wautersiella Amoebinatus | ||||||||||||||||
Sphingobacteriaceae | Pedobacter unclassified Sphingobacterium | ||||||||||||||||
Cytophagaceae | Hymenobacter* Dyadobacter | ||||||||||||||||
| unclassified | unclassified | ||||||||||||||||
Chitinophagaceae | unclassified Chitinophaga | ||||||||||||||||
| Anaerolineaceae | Unclassified | ||||||||||||||||
| Caldilineaceae | Caldilinea | ||||||||||||||||
| Chloroflexaceae | Roseiflexus | ||||||||||||||||
| Listeriaceae | Listeria | ||||||||||||||||
| Paenibacillaceae | Ammoniphilus | ||||||||||||||||
| Planococcaceae | Kurthia | ||||||||||||||||
| Staphylococcaceae | Staphylococcus* | ||||||||||||||||
| Thermoactinomycetaceae | Laceyella | ||||||||||||||||
| unclassified | unclassified | ||||||||||||||||
| Carnobacteriaceae | unclassified Desemzia Dolosigranulum Alloiococcus Carnobacterium | ||||||||||||||||
Streptococcaceae | Streptococcus* Lactococcus* | ||||||||||||||||
Leuconostocaceae | Leuconostoc Weissella* | ||||||||||||||||
Lactobacillaceae | unclassified Lactobacillus | ||||||||||||||||
Aerococcaceae | Flacklamia Aerococcus | ||||||||||||||||
| Mycoplasmataceae | unclassified | ||||||||||||||||
Erysipelotrichaceae | Bulleidia* Solobacterium unclassified | ||||||||||||||||
| Family_XI_Incertae_Sedis | Anaerococcus* Finegoldia * Peptoniphilus Parvimonas* Mogibacterium* Helcococcus unclassified Eubacterium Sedimentibacter | ||||||||||||||||
| Lachnospiraceae | Roseburia* Johnsonella* Catonella* unclassified Incertae_Sedis Coprococcus* Shuttleworthia Blautia* Oribacterium* Moryella* Butyrivibrio-Pseudobutyrivibrio Dorea Lachnospira | ||||||||||||||||
| Ruminococcaceae | Faecalibacterium* Subdoligranulum unclassified Fastidiosipila* Incertae_Sedis Ruminococcus Ethanoligenens Acetivibrio Oscillibacter | ||||||||||||||||
| Peptostreptococc aceae | Incertae_Sedis Peptostreptococcus* Filifactor* Peptococcus* Sporacetigenium* Desulfosporosinus | ||||||||||||||||
| Veillonellaceae | Veillonella* Selenomonas* unclassified Mitsuokella Megamonas Dialister* | ||||||||||||||||
Clostridiaceae | unclassified Clostridium* Caloramator | ||||||||||||||||
| unclassified | unclassified | ||||||||||||||||
| Eubacteriaceae | Eubacterium* Acetobacterium | ||||||||||||||||
Family_III_Incertae_Sedis | Caldicellulosiruptor Thermoanaerobacter | ||||||||||||||||
| Acidobacteria | Aquificae | Chamydiae | Deinococcus-Thermus | Fusobacteria | Planctomycetes | Synergistetes | Verrucomicrobia | |||
| Acidobacteria | Aquificae | Chlamydiae | Deinococci | Fusobacteria | Planctomyceta | Phycisphaerae | Synergistia | Opitutae | ||
| Acidobacteriales | Aquificales | Chlamydiales | Thermales | Deinococcales | Fusobacteriales | Planctomycetales | Phycisphaerales | Synergistales | Opitutales | |
| Acidobacteriaceae | Chloroacidobacterium Acidobacterium Edaphobacter* Koribacter Solibacter | |||||||||
| Aquificaceae | Hydrogenobacter | |||||||||
| Unclassified | unclassified | |||||||||
| Thermaceae | Meiothermus | |||||||||
| Trueperaceae | Truepera* | |||||||||
| Deinococcaceae | Deinococcus* | |||||||||
| Fusobacteriaceae | Fusobacterium* unclassified Psychrilyobacter | |||||||||
| Leptotrichiaceae | Leptotrichia* Sneathia unclassified | |||||||||
| Planctomycetaceae | Singulisphaera* Planctomyces* Unclassified | |||||||||
| Phycisphaeraceae | CL500-3 Phycisphaera | |||||||||
| Synergistaceae | unclassified Jonquetella | |||||||||
| Opitutaceae | Opitutus* | |||||||||
| F (28.194) = 20.998 p < 0.0000 | F-Remove (2.97) | p-Level | Sample Group 1 | Sample Group 2 | Sample Group 3 | All Groups |
| Actinobacteria | 6.0925 ** | 0.003218 | 5324.208 | 2505.167 | 2115.364 | 2838.283 |
| Aquificae | 4.4817 ** | 0.013752 | 0.000 | 18.000 | 0.1558442 | 2.017699 |
| Firmicutes | 126.8766 *** | 0.000000 | 5412.583 | 24247.08 | 2839.013 | 5659.035 |
| Proteobacteria | 76.6841 *** | 0.000000 | 23561.5 | 5097.167 | 5498.013 | 9291.938 |
| Percent correct classification | 87.50 | 91.66 | 100.00 | 96.46 | ||
| (a) | ||||||
| F (104.118) = 5.7435 p < 0.0000 | F-Remove (2.59) | p-Level | Sample Group 1 | Sample Group 2 | Sample Group 3 | All Groups |
| Actinomycetales | 6.736752 *** | 0.002467 | 5007.208 | 2131 | 1960.87 | 2625.947 |
| Anaerolineales | 3.30950 * | 0.043427 | 8.833333 | 0.00000 | 1.363636 | 2.80531 |
| Bacillales | 3.98242 * | 0.023859 | 91.08334 | 158.8333 | 110.3636 | 111.4159 |
| Bacteroidales | 6.13735 ** | 0.003789 | 177.4167 | 858.75 | 113.1948 | 206.0089 |
| Caldilineales | 3.22566 * | 0.046832 | 7.75 | 0.08333334 | 0.05194805 | 1.690266 |
| Campylobacterales | 9.84095 *** | 0.000205 | 1903.167 | 95.58334 | 149.9091 | 516.5132 |
| Caulobacterales | 8.75310 *** | 0.000469 | 1726.917 | 109.6667 | 372.1688 | 632.0266 |
| Clostridiales | 6.00333 * | 0.004235 | 4161.167 | 20972.33 | 1966.558 | 4450.982 |
| Sphingomonadales | 13.72837 *** | 0.000013 | 60.54167 | 184.8333 | 125.6234 | 118.0885 |
| Percent correct classification | 91.66 | 91.66 | 100 | 97.34 | ||
| (b) | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arcos, S.C.; Lira, F.; Robertson, L.; González, M.R.; Carballeda-Sangiao, N.; Sánchez-Alonso, I.; Zamorano, L.; Careche, M.; Jiménez-Ruíz, Y.; Ramos, R.; et al. Metagenomics Analysis Reveals an Extraordinary Inner Bacterial Diversity in Anisakids (Nematoda: Anisakidae) L3 Larvae. Microorganisms 2021, 9, 1088. https://doi.org/10.3390/microorganisms9051088
Arcos SC, Lira F, Robertson L, González MR, Carballeda-Sangiao N, Sánchez-Alonso I, Zamorano L, Careche M, Jiménez-Ruíz Y, Ramos R, et al. Metagenomics Analysis Reveals an Extraordinary Inner Bacterial Diversity in Anisakids (Nematoda: Anisakidae) L3 Larvae. Microorganisms. 2021; 9(5):1088. https://doi.org/10.3390/microorganisms9051088
Chicago/Turabian StyleArcos, Susana C., Felipe Lira, Lee Robertson, María Rosa González, Noelia Carballeda-Sangiao, Isabel Sánchez-Alonso, Laura Zamorano, Mercedes Careche, Yolanda Jiménez-Ruíz, Ricardo Ramos, and et al. 2021. "Metagenomics Analysis Reveals an Extraordinary Inner Bacterial Diversity in Anisakids (Nematoda: Anisakidae) L3 Larvae" Microorganisms 9, no. 5: 1088. https://doi.org/10.3390/microorganisms9051088
APA StyleArcos, S. C., Lira, F., Robertson, L., González, M. R., Carballeda-Sangiao, N., Sánchez-Alonso, I., Zamorano, L., Careche, M., Jiménez-Ruíz, Y., Ramos, R., Llorens, C., González-Muñoz, M., Oliver, A., Martínez, J. L., & Navas, A. (2021). Metagenomics Analysis Reveals an Extraordinary Inner Bacterial Diversity in Anisakids (Nematoda: Anisakidae) L3 Larvae. Microorganisms, 9(5), 1088. https://doi.org/10.3390/microorganisms9051088








