Consolidated Bioprocessing: Synthetic Biology Routes to Fuels and Fine Chemicals
Abstract
1. Introduction
2. Lignocellulose as a Carbon Source
2.1. Lignocellulose: A Heterogeneous Source of Polymeric Sugars
2.2. Lignocellulose Pre-Treatments
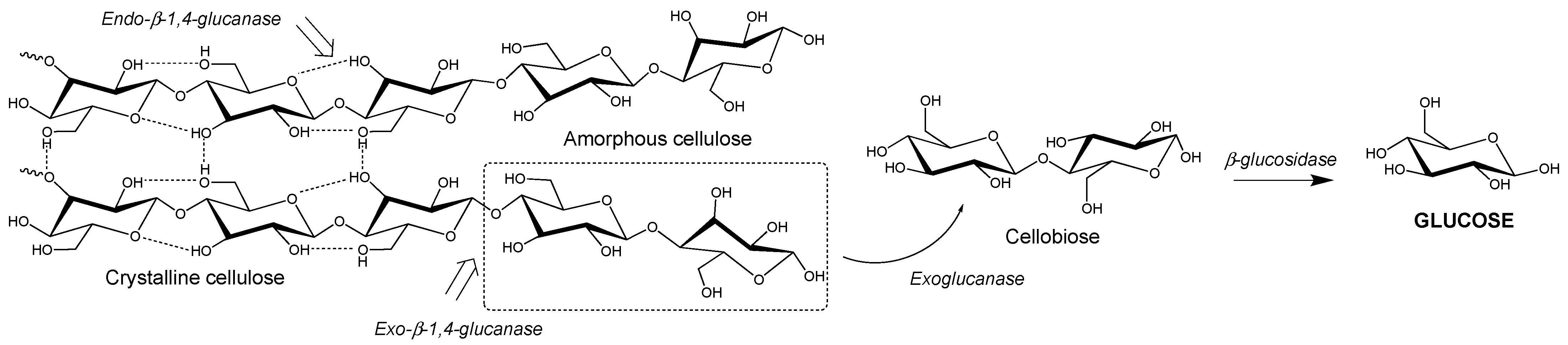
2.3. Enzymatic Lignocellulose Degradation
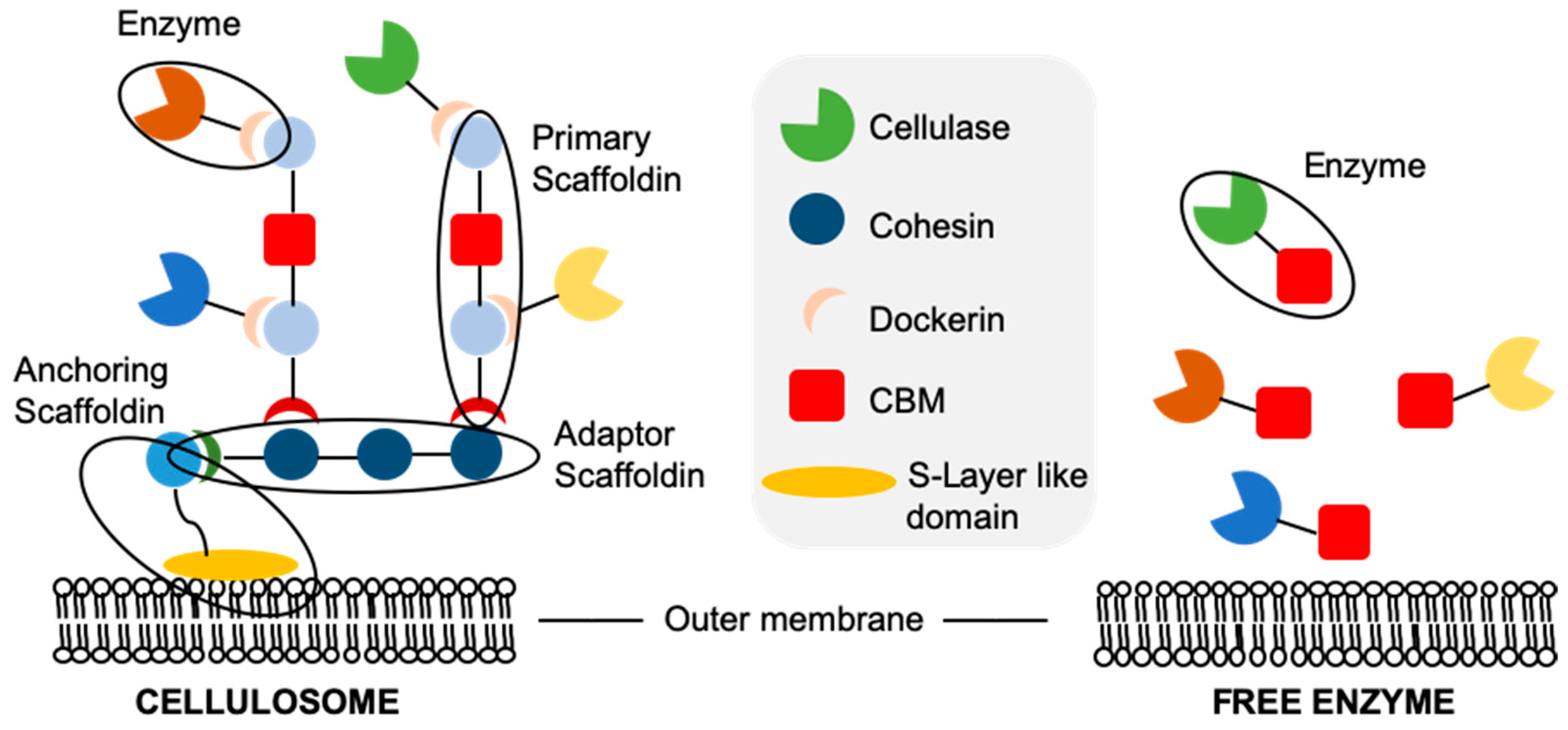
2.4. Cellulase Localisation
3. Consolidated Bioprocessing
3.1. Naturally Cellulolytic Microorganisms
3.2. Non-Cellulolytic Chemical Producers
3.3. Model Organism: E. coli
4. Looking to the Future: The Biofoundry Approach
5. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Anastas, P.; Eghbali, N. Green chemistry: Principles and practice. Chem. Soc. Rev. 2010, 39, 301–312. [Google Scholar] [CrossRef]
- Oosthoek, J.; Gills, B.K. Humanity at the crossroads: The globalization of environmental crisis. Globalizations 2005, 2, 283–291. [Google Scholar] [CrossRef]
- Baumschlager, A.; Khammash, M. Synthetic biological approaches for optogenetics and tools for transcriptional light-control in bacteria. Adv. Biol. 2021. [Google Scholar] [CrossRef]
- Carbonell, P.; Currin, A.; Dunstan, M.; Fellows, D.; Jervis, A.; Rattray, N.J.W.; Robinson, C.J.; Swainston, N.; Vinaixa, M.; Williams, A.; et al. SYNBIOCHEM-a SynBio foundry for the biosynthesis and sustainable production of fine and speciality chemicals. Biochem. Soc. Trans. 2016, 44, 675–677. [Google Scholar] [CrossRef]
- Chang, M.C.Y.; Keasling, J.D. Production of isoprenoid pharmaceuticals by engineered microbes. Nat. Chem. Biol. 2006, 2, 674–681. [Google Scholar] [CrossRef]
- Kim, I.-K.; Roldão, A.; Siewers, V.; Nielsen, J. A systems-level approach for metabolic engineering of yeast cell factories. FEMS Yeast Res. 2012, 12, 228–248. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, W. Natural products from synthetic biology. Curr. Opin. Chem. Biol. 2011, 14, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Nazhand, A.; Durazzo, A.; Lucarini, M.; Santini, A. Recent advances in metabolic engineering and synthetic biology for microbial production of isoprenoid-based biofuels: An overview. In Bioprocessing for Biofuel Production; Molina, G., Gupta, V.K., Singh, B.N., Gathergood, N., Eds.; Springer: Singapore, 2021; pp. 183–201. [Google Scholar]
- Patra, P.; Das, M.; Kundu, P.; Ghosh, A. Recent advances in systems and synthetic biology approaches for developing novel cell-factories in non-conventional yeasts. Biotechnol. Adv. 2021, 47, 107695. [Google Scholar] [CrossRef] [PubMed]
- Rollié, S.; Mangold, M.; Sundmacher, K. Designing biological systems systems engineering meets synthetic biology. Chem. Eng. Sci. 2012, 69, 1–29. [Google Scholar] [CrossRef]
- Smanski, M.J.; Zhou, H.; Claesen, J.; Ben, S.; Fischbach, M.A.; Voigt, C.A. Synthetic biology to access and expand nature’s chemical diversity. Nat. Rev. Microbiol. 2016, 14, 135–149. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, Y.; Fu, L.; Guo, E.; Wang, B.; Dai, L.; Si, T. Accelerating strain engineering in biofuel research via build and test automation of synthetic biology. Curr. Opin. Biotechnol. 2021, 67, 88–98. [Google Scholar] [CrossRef]
- Zhang, Y.H.P.; Myung, S.; You, C.; Zhu, Z.; Rollin, J.A. Toward low-cost biomanufacturing through in vitro synthetic biology: Bottom-up design. J. Mater. Chem. 2011, 21, 18877. [Google Scholar] [CrossRef]
- Liao, J.C.; Mi, L.; Pontrelli, S.; Luo, S. Fuelling the future: Microbial engineering for the production of sustainable biofuels. Nat. Rev. Microbiol. 2016, 14, 288–304. [Google Scholar] [CrossRef] [PubMed]
- Arundel, A.; Sawaya, D. The Bioeconomy to 2030: Designing a Policy Agenda 2009; OECD Publishing: Paris, France, 2009. [Google Scholar] [CrossRef]
- Ajikumar, P.K.; Xiao, W.-H.; Tyo, K.E.J.; Wang, Y.; Simeon, F.; Leonard, E.; Mucha, O.; Phon, T.H.; Pfeifer, B.; Stephanopoulos, G. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 2010, 330, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Amiri, P.; Shahpiri, A.; Asadollahi, M.A.; Momenbeik, F.; Partow, S. Metabolic engineering of Saccharomyces cerevisiae for linalool production. Biotechnol. Lett. 2016, 38, 503–508. [Google Scholar] [CrossRef]
- Hong, J.; Yang, H.; Zhang, K.; Liu, C.; Zou, S.; Zhang, M. Development of a cellulolytic Saccharomyces cerevisiae strain with enhanced cellobiohydrolase activity. World J. Microbiol. Biotechnol. 2014, 30, 2985–2993. [Google Scholar] [CrossRef]
- Vemuri, G.N.; Eiteman, M.A.; Altman, E. Succinate production in dual-phase Escherichia coli fermentations depends on the time of transition from aerobic to anaerobic conditions. J. Ind. Microbiol. Biotechnol. 2002, 28, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Yim, H.; Haselbeck, R.; Niu, W.; Pujol-Baxley, C.; Burgard, A.; Boldt, J.; Khandurina, J.; Trawick, J.D.; Osterhout, R.E.; Stephen, R.; et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 2011, 7, 445–452. [Google Scholar] [CrossRef]
- Biz, A.; Proulx, S.; Xu, Z.; Siddartha, K.; Mulet Indrayanti, A.; Mahadevan, R. Systems biology based metabolic engineering for non-natural chemicals. Biotechnol. Adv. 2019, 37, 107379. [Google Scholar] [CrossRef] [PubMed]
- Budin, I.; Keasling, J.D. Synthetic biology for fundamental biochemical discovery. Biochemistry 2019, 58, 1464–1469. [Google Scholar] [CrossRef] [PubMed]
- Lin, G.-M.; Warden-Rothman, R.; Voigt, C.A. Retrosynthetic design of metabolic pathways to chemicals not found in Nature. Curr. Opin. Syst. Biol. 2019, 14, 82–107. [Google Scholar] [CrossRef]
- Keasling, J.D. Synthetic biology and the development of tools for metabolic engineering. Metab. Eng. 2012, 14, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Siegel, J.B.; Zanghellini, A.; Lovick, H.M.; Kiss, G.; Lambert, A.R.; St Clair, J.L.; Gallaher, J.L.; Hilvert, D.; Gelb, M.H.; Stoddard, B.L.; et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels-Alder reaction. Science 2010, 329, 309–313. [Google Scholar] [CrossRef] [PubMed]
- Paddon, C.J.; Keasling, J.D. Semi-synthetic artemisinin: A model for the use of synthetic biology in pharmaceutical development. Nat. Rev. Microbiol. 2014, 12, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Meadows, A.L.; Hawkins, K.M.; Tsegaye, Y.; Antipov, E.; Kim, Y.; Raetz, L.; Dahl, R.H.; Tai, A.; Mahatdejkul-Meadows, T.; Xu, L.; et al. Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature 2016, 537, 694–697. [Google Scholar] [CrossRef]
- Karuppiah, V.; Ranaghan, K.E.; Leferink, N.G.H.; Johannissen, L.O.; Shanmugam, M.; Ní Cheallaigh, A.; Bennett, N.J.; Kearsey, L.J.; Takano, E.; Gardiner, J.M.; et al. Structural basis of catalysis in the bacterial monoterpene synthases linalool synthase and 1,8-cineole synthase. ACS Catal. 2017, 7, 6268–6282. [Google Scholar] [CrossRef] [PubMed]
- Leferink, N.G.H.; Jervis, A.J.; Zebec, Z.; Toogood, H.S.; Hay, S.; Takano, E.; Scrutton, N.S. A ‘Plug and Play’ platform for the production of diverse monoterpene hydrocarbon scaffolds in Escherichia coli. ChemistrySelect 2016, 1, 1893–1896. [Google Scholar] [CrossRef]
- Li, Y.; Li, S.; Thodey, K.; Trenchard, I.; Cravens, A.; Smolke, C.D. Complete biosynthesis of noscapine and halogenated alkaloids in yeast. Proc. Natl. Acad. Sci. USA 2018, 115, E3922–E3931. [Google Scholar] [CrossRef]
- Gaida, S.M.; Liedtke, A.; Jentges, A.H.W.; Engels, B.; Jennewein, S. Metabolic engineering of Clostridium cellulolyticum for the production of n-butanol from crystalline cellulose. Microb. Cell. Fact. 2016, 15, 6. [Google Scholar] [CrossRef]
- Turk, S.C.H.J.; Kloosterman, W.P.; Ninaber, D.K.; Kolen, K.P.A.M.; Knutova, J.; Suir, E.; Schürmann, M.; Raemakers-Franken, P.C.; Müller, M.; de Wildeman, S.M.A.; et al. Metabolic engineering toward sustainable production of Nylon-6. ACS Synth. Biol. 2016, 5, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Men, X.; Chen, H.; Li, M.; Ding, Z.; Chen, G.; Wang, F.; Liu, H.; Wang, Q.; Zhu, Y.; et al. A systematic optimization of styrene biosynthesis in Escherichia coli BL21(DE3). Biotechnol. Biofuels 2018, 11, 14. [Google Scholar] [CrossRef] [PubMed]
- De Jong, E.; Jungmeier, G. Biorefinery concepts in comparison to petrochemical refineries. In Industrial Biorefineries and White Biotechnology; Elsevier: Amsterdam, The Netherlands, 2015; pp. 3–33. [Google Scholar]
- Toogood, H.S.; Scrutton, N.S. Retooling microorganisms for the fermentative production of alcohols. Curr. Opin. Biotechnol. 2018, 50, 1–10. [Google Scholar] [CrossRef]
- Association, R.F. Annual Fuel Ethanol Production U.S. and World Ethanol Production. Available online: https://ethanolrfa.org/statistics/annual-ethanol-production/ (accessed on 20 January 2021).
- Bhatia, S.K.; Kim, S.-H.; Yoon, J.-J.; Yang, Y.-H. Current status and strategies for second generation biofuel production using microbial systems. Energy Convers. Manag. 2017, 148, 1142–1156. [Google Scholar] [CrossRef]
- Lopes, M.L.; de Lima Paulillo, S.C.; Godoy, A.; Cherubin, R.A.; Lorenzi, M.S.; Giometti, F.H.C.; Bernardino, C.D.; de Amorim Neto, H.B.; de Amorim, H.V. Ethanol production in Brazil: A bridge between science and industry. Braz. J. Microbiol. 2016, 47, 64–76. [Google Scholar] [CrossRef]
- UNCTAD. Second Generation Biofuel Markets: State of Play, Trade and Developing Country Perspectives 2016; United Nations Publication: Herndon, VA, USA, 2016; p. 69. Available online: https://unctad.org/en/PublicationsLibrary/ditcted2015d8_en.pdf (accessed on 17 May 2021).
- Alcalde, M. Engineering the ligninolytic enzyme consortium. Trends. Biotechnol. 2015, 33, 155–162. [Google Scholar] [CrossRef]
- Beig, B.; Riaz, M.; Raza Naqvi, S.; Hassan, M.; Zheng, Z.; Karimi, K.; Pugazhendhi, A.; Atabani, A.E.; Thuy Lan Chi, N. Current challenges and innovative developments in pretreatment of lignocellulosic residues for biofuel production: A review. Fuel 2021, 287, 119670. [Google Scholar] [CrossRef]
- Andlar, M.; Rezić, T.; Marđetko, N.; Kracher, D.; Ludwig, R.; Šantek, B. Lignocellulose degradation: An overview of fungi and fungal enzymes involved in lignocellulose degradation. Eng. Life Sci. 2018, 18, 768–778. [Google Scholar] [CrossRef]
- De Souza, W.R. Microbial degradation of lignocellulosic biomass. In Sustainable Degradation of Lignocellulosic Biomass—Techniques, Applications and Commercialization [Online]; Chandel, A.K., da Silva, S.S., Eds.; IntechOpen: Rijeka, Croatia, 2013; pp. 208–247. [Google Scholar]
- De Vries, R.P.; Visser, J. Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol. Rev. 2001, 65, 497–522. [Google Scholar] [CrossRef]
- Schädel, C.; Blöchl, A.; Richter, A.; Hoch, G. Quantification and monosaccharide composition of hemicelluloses from different plant functional types. Plant. Physiol. Biochem. 2010, 48, 1–8. [Google Scholar] [CrossRef]
- Gibson, L.J. The hierarchical structure and mechanics of plant materials. J. R. Soc. Interface 2012, 9, 2749–2766. [Google Scholar] [CrossRef]
- Álvarez, C.; Reyes-Sosa, F.M.; Díez, B. Enzymatic hydrolysis of biomass from wood. Microb. Biotechnol. 2016, 9, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Hahn-Hägerdal, B.; Karhumaa, K.; Fonseca, C.; Spencer-Martins, I.; Gorwa-Grauslund, M.F. Towards industrial pentose-fermenting yeast strains. Appl. Microbiol. Biotechnol. 2007, 74, 937–953. [Google Scholar] [CrossRef] [PubMed]
- Beckham, G.T.; Johnson, C.W.; Karp, E.M.; Salvachúa, D.; Vardon, D.R. Opportunities and challenges in biological lignin valorization. Curr. Opin. Biotechnol. 2016, 42, 40–53. [Google Scholar] [CrossRef] [PubMed]
- Biswas, B.; Bisht, Y.; Kumar, J.; Yenumala, S.R.; Bhaskar, T. Effects of temperature and solvent on hydrothermal liquefaction of the corncob for production of phenolic monomers. Biomass Convers. Biorefin. 2020. [Google Scholar] [CrossRef]
- Chandra, R.P.; Bura, R.; Mabee, W.E.; Berlin, A.; Pan, X.; Saddler, J.N. Substrate pretreatment: The key to effective enzymatic hydrolysis of lignocellulosics? Adv. Biochem. Eng. Biotechnol. 2007, 108, 67–93. [Google Scholar] [PubMed]
- Solarte-Toro, J.C.; Romero-García, J.M.; Martínez-Patiño, J.C.; Ruiz-Ramos, E.; Castro-Galiano, E.; Cardona-Alzate, C.A. Acid pretreatment of lignocellulosic biomass for energy vectors production: A review focused on operational conditions and techno-economic assessment for bioethanol production. Renew. Sustain. Energy Rev. 2019, 107, 587–601. [Google Scholar] [CrossRef]
- Kim, J.S.; Lee, Y.Y.; Kim, T.H. A review on alkaline pretreatment technology for bioconversion of lignocellulosic biomass. Bioresour. Technol. 2016, 199, 42–48. [Google Scholar] [CrossRef]
- Wang, F.-L.; Li, S.; Sun, Y.-X.; Han, H.-Y.; Zhang, B.-X.; Hu, B.-Z.; Gao, Y.-F.; Hu, X.-M. Ionic liquids as efficient pretreatment solvents for lignocellulosic biomass. RSC Adv. 2017, 7, 47990–47998. [Google Scholar] [CrossRef]
- Akhtar, N.; Gupta, K.; Goyal, D.; Goyal, A. Recent advances in pretreatment technologies for efficient hydrolysis of lignocellulosic biomass. Environ. Prog. Sustainable Energy 2016, 35, 489–511. [Google Scholar] [CrossRef]
- Baruah, J.; Nath, B.K.; Sharma, R.; Kumar, S.; Deka, R.C.; Baruah, D.C.; Kalita, E. Recent trends in the pretreatment of lignocellulosic biomass for value-added products. Front. Energy Res. 2018, 6, 141. [Google Scholar] [CrossRef]
- Galbe, M.; Wallberg, O. Pretreatment for biorefineries: A review of common methods for efficient utilisation of lignocellulosic materials. Biotechnol. Biofuels 2019, 12, 294. [Google Scholar] [CrossRef]
- Jönsson, L.J.; Alriksson, B.; Nilvebrant, N. Bioconversion of lignocellulose: Inhibitors and detoxification. Biotechnol. Biofuels 2013, 6, 16. [Google Scholar] [CrossRef] [PubMed]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, 490–495. [Google Scholar] [CrossRef]
- Davies, G.; Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 1995, 3, 853–859. [Google Scholar] [CrossRef]
- Wilson, D.B. Microbial diversity of cellulose hydrolysis. Curr. Opin. Microbiol. 2011, 14, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Greene, E.R.; Himmel, M.E.; Beckham, G.T.; Tan, Z. Glycosylation of cellulases: Engineering better enzymes for biofuels. In Advances in Carbohydrate Chemistry and Biochemistry; Baker, D.C., Horton, D., Eds.; Academic Press: Cambridge, MA, USA, 2015; Volume 72, pp. 63–112. [Google Scholar]
- Juturu, V.; Wu, J.C. Microbial cellulases: Engineering, production and applications. Renew. Sustain. Energy Rev. 2014, 33, 188–203. [Google Scholar] [CrossRef]
- Horn, S.J.; Vaaje-Kolstad, G.; Westereng, B.; Eijsink, V. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 2012, 5, 45. [Google Scholar] [CrossRef] [PubMed]
- Lynd, L.R.; Weimer, P.J.; van Zyl, W.H.; Pretorius, I.S. Microbial cellulose utilization: Fundamentals and biotechnology. Bioresour. Technol. 2002, 66, 506–577. [Google Scholar]
- Teeri, T.T. Crystalline cellulose degradation: New insight into the function of cellobiohydrolases. Trends Biotechnol. 1997, 15, 160–167. [Google Scholar] [CrossRef]
- Dimarogona, M.; Topakas, E.; Christakopoulos, P. Cellulose degradation by oxidative enzymes. CSBJ 2012, 2, e201209015. [Google Scholar] [CrossRef]
- Bischof, R.H.; Ramoni, J.; Seiboth, B. Cellulases and beyond: The first 70 years of the enzyme producer Trichoderma reesei. Microb. Cell. Fact. 2016, 15, 106. [Google Scholar] [CrossRef] [PubMed]
- Lopes, A.M.; Ferreira Filho, E.X.; Moreira, L.R.S. An update on enzymatic cocktails for lignocellulose breakdown. J. Appl. Microbiol. 2018, 125, 632–645. [Google Scholar] [CrossRef]
- Jabbour, D.; Borrusch, M.S.; Banerjee, G.; Walton, J.D. Enhancement of fermentable sugar yields by α-xylosidase supplementation of commercial cellulases. Biotechnol. Biofuels 2013, 6, 58. [Google Scholar] [CrossRef]
- Ye, Z.; Zheng, Y.; Li, B.; Borrusch, M.S.; Storms, R.; Walton, J.D. Enhancement of synthetic Trichoderma-based enzyme mixtures for biomass conversion with an alternative family 5 glycosyl hydrolase from Sporotrichum thermophile. PLoS ONE 2014, 9, e109885. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Zhang, J.; Bao, J. Cost evaluation of cellulase enzyme for industrial-scale cellulosic ethanol production based on rigorous Aspen Plus modeling. Bioprocess. Biosyst. Eng. 2016, 39, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Annamalai, N.; Rajeswari, M.V.; Sivakumar, N. Cellobiohydrolases: Role, mechanism, and recent developments. In Microbial Enzymes in Bioconversions of Biomass; Gupta, V.K., Ed.; Springer: Berlin/Heidelberg, Germany, 2016; pp. 29–36. [Google Scholar]
- Janusz, G.; Pawlik, A.; Sulej, J.; Świderska-Burek, U.; Jarosz-Wilkołazka, A.; Paszczyński, A. Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol. Rev. 2017, 41, 941–962. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Kubicek, C.P. Genetic engineering of Trichoderma reesei cellulases and their production. Microb. Biotechnol. 2017, 10, 1485–1499. [Google Scholar] [CrossRef] [PubMed]
- Yamada, R.; Hasunuma, T.; Kondo, A. Endowing non-cellulolytic microorganisms with cellulolytic activity aiming for consolidated bioprocessing. Biotechnol. Adv. 2013, 31, 754–763. [Google Scholar] [CrossRef]
- Shoseyov, O.; Shani, Z.; Levy, I. Carbohydrate binding modules: Biochemical properties and novel applications. Microbiol. Mol. Biol. Rev. 2006, 70, 283–295. [Google Scholar] [CrossRef]
- Bayer, E.A.; Belaich, J.-P.; Shoham, Y.; Lamed, R. The cellulosomes: Multienzyme machines for degradation of plant cell wall polysaccharides. Annu. Rev. Microbiol. 2004, 58, 521–554. [Google Scholar] [CrossRef]
- Hyeon, J.E.; Jeon, W.J.; Whang, S.Y.; Han, S.O. Production of minicellulosomes for the enhanced hydrolysis of cellulosic substrates by recombinant Corynebacterium glutamicum Enzyme. Microb. Technol. 2011, 48, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Fontes, C.M.G.A.; Gilbert, H.J. Cellulosomes: Highly efficient nanomachines designed to deconstruct plant cell wall complex carbohydrates. Annu. Rev. Biochem. 2010, 79, 655–681. [Google Scholar] [CrossRef] [PubMed]
- Costa, T.R.D.; Felisberto-Rodrigues, C.; Meir, A.; Prevost, M.S.; Redzej, A.; Trokter, M.; Waksman, G. Secretion systems in Gram-negative bacteria: Structural and mechanistic insights. Nat. Rev. Microbiol. 2015, 13, 343–359. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Lu, X.; Hu, J.; Chen, Y.; Shen, W.; Liu, L. Boosting secretion of extracellular protein by Escherichia coli via cell wall perturbation. Appl. Environ. Microbiol. 2018, 84, e01382-18. [Google Scholar] [CrossRef] [PubMed]
- Burdette, L.A.; Leach, S.A.; Wong, H.T.; Tullman-Ercek, D. Developing Gram-negative bacteria for the secretion of heterologous proteins. Microb. Cell. Fact. 2018, 17, 196. [Google Scholar] [CrossRef]
- Mergulhão, F.J.M.; Summers, D.K.; Monteiro, G.A. Recombinant protein secretion in Escherichia coli. Biotechnol. Adv. 2005, 23, 177–202. [Google Scholar] [CrossRef]
- Green, E.R.; Mecsas, J. Bacterial secretion systems—An overview. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Patel, R.; Smith, S.M.; Robinson, C. Protein transport by the bacterial Tat pathway. Biochim. Biophys. Acta Mol. Cell. Res. 2014, 1843, 1620–1628. [Google Scholar] [CrossRef] [PubMed]
- Xiao, W.; Li, H.; Xia, W.; Yang, Y.; Hu, P.; Zhou, S.; Hu, Y.; Liu, X.; Dai, Y.; Jiang, Z. Co-expression of cellulase and xylanase genes in Sacchromyces cerevisiae toward enhanced bioethanol production from corn stover. Bioengineered 2019, 10, 513–521. [Google Scholar] [CrossRef]
- Anandharaj, M.; Lin, Y.-J.; Rani, R.P.; Nadendla, E.K.; Ho, M.-C.; Huang, C.-C.; Cheng, J.-F.; Chang, J.-J.; Li, W.-H. Constructing a yeast to express the largest cellulosome complex on the cell surface. Proc. Natl. Acad. Sci. USA 2020, 117, 2385–2394. [Google Scholar] [CrossRef]
- Yan, Q.; Fong, S.S. Challenges and advances for genetic engineering of non-model bacteria and uses in consolidated bioprocessing. Front. Microbiol. 2017, 8, 2060. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Chen, D.; Wei, Y.; Wang, Q.; Li, Z.; Chen, Y.; Huang, R. Direct ethanol production from lignocellulosic sugars and sugarcane bagasse by a recombinant Trichoderma reesei strain HJ48. Sci. World J. 2014, 2014, 798683. [Google Scholar]
- Singh, N.; Mathur, A.S.; Gupta, R.P.; Barrow, C.J.; Tuli, D.; Puri, M. Enhanced cellulosic ethanol production via consolidated bioprocessing by Clostridium thermocellum ATCC 31924. Bioresour. Technol. 2018, 250, 860–867. [Google Scholar] [CrossRef] [PubMed]
- Xin, F.; Dong, W.; Zhang, W.; Ma, J.; Jiang, M. Biobutanol production from crystalline cellulose through consolidated bioprocessing. Trends Biotechnol. 2019, 37, 167–180. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.P.; Mi, L.; Morioka, A.H.; Yoshino, K.M.; Konishi, S.; Xu, S.C.; Papanek, B.A.; Riley, L.A.; Guss, A.M.; Liao, J.C. Consolidated bioprocessing of cellulose to isobutanol using Clostridium thermocellum. Metab. Eng. 2015, 31, 44–52. [Google Scholar] [CrossRef]
- Celińska, E.; Nicaud, J.-M.; Białas, W. Hydrolytic secretome engineering in Yarrowia lipolytica for consolidated bioprocessing on polysaccharide resources: Review on starch, cellulose, xylan, and inulin. Appl. Microbiol. Biotechnol. 2021, 105, 975–989. [Google Scholar] [CrossRef]
- Guo, Z.; Duquesne, S.; Bozonnet, S.; Cioci, G.; Nicaud, J.M.; Marty, A.; O’Donohue, M.J. Development of cellobiose-degrading ability in Yarrowia lipolytica strain by overexpression of endogenous genes. Biotechnol. Biofuels 2015, 8, 1–16. [Google Scholar] [CrossRef]
- Duquesne, S.; Bozonnet, S.; Bordes, F.; Dumon, C.; Nicaud, J.M.; Marty, A. Construction of a highly active xylanase displaying oleaginous yeast: Comparison of anchoring systems. PLoS ONE 2014, 9, e95128. [Google Scholar] [CrossRef]
- Inokuma, K.; Bamba, T.; Ishii, J.; Ito, Y.; Hasunuma, T.; Kondo, A. Enhanced cell-surface display and secretory production of cellulolytic enzymes with Saccharomyces cerevisiae Sed1 signal peptide. Biotechnol. Bioeng. 2016, 113, 2358–2366. [Google Scholar] [CrossRef]
- Wen, F.; Sun, J.; Zhao, H. Yeast surface display of trifunctional minicellulosomes for simultaneous saccharification and fermentation of cellulose to ethanol. Appl. Environ. Microbiol. 2010, 76, 1251–1260. [Google Scholar] [CrossRef]
- Willson, B.J.; Kovács, K.; Wilding-Steele, T.; Markus, R.; Winzer, K.; Minton, N.P. Production of a functional cell wall-anchored minicellulosome by recombinant Clostridium acetobutylicum ATCC 824. Biotechnol. Biofuels 2016, 9, 109. [Google Scholar] [CrossRef]
- Wang, C.; Pfleger, B.F.; Kim, S.W. Reassessing Escherichia coli as a cell factory for biofuel production. Curr. Opin. Biotechnol. 2017, 45, 92–103. [Google Scholar] [CrossRef] [PubMed]
- Keseler, I.M.; Mackie, A.; Santos-Zavaleta, A.; Billington, R.; Bonavides-Martínez, C.; Caspi, R.; Fulcher, C.; Gama-Castro, S.; Kothari, A.; Krummenacker, M.; et al. The EcoCyc database: Reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 2017, 45, 543–550. [Google Scholar] [CrossRef] [PubMed]
- Karp, P.D.; Ong, W.K.; Paley, S.; Billington, R.; Caspi, R.; Fulcher, C.; Kothari, A.; Krummenacker, M.; Latendresse, M.; Midford, P.E.; et al. The EcoCyc Database. EcoSal Plus 2018, 8. [Google Scholar] [CrossRef]
- Sezonov, G.; Joseleau-Petit, D.; D’Ari, R. Escherichia coli physiology in Luria-Bertani broth. J. Bacteriol. 2007, 189, 8746–8749. [Google Scholar] [CrossRef] [PubMed]
- Baeshen, M.N.; Al-Hejin, A.M.; Bora, R.S.; Ahmed, M.M.; Ramadan, H.A.; Saini, K.S.; Baeshen, N.A.; Redwan, E.M. Production of biopharmaceuticals in E. coli: Current scenario and future perspectives. J. Microbiol. Biotechnol. 2015, 25, 953–962. [Google Scholar] [CrossRef] [PubMed]
- Unden, G.; Becker, S.; Bongaerts, J.; Schirawski, J.; Six, S. Oxygen regulated gene expression in facultatively anaerobic bacteria. Antonie van Leeuwenhoek 1994, 66, 3–22. [Google Scholar] [CrossRef]
- Koppolu, V.; Vasigala, V.K. Role of Escherichia coli in biofuel production. Microbiol. Insights 2016, 9, 29–35. [Google Scholar] [CrossRef]
- Goeddel, D.V.; Kleid, D.G.; Bolivar, F.; Heyneker, H.L.; Yansura, D.G.; Crea, R.; Hirose, T.; Kraszewski, A.; Itakura, K.; Riggs, A.D. Expression in Escherichia coli of chemically synthesized genes for human insulin. Proc. Natl. Acad. Sci. USA 1979, 76, 106–110. [Google Scholar] [CrossRef]
- Nakamura, C.E.; Whited, G.M. Metabolic engineering for the microbial production of 1,3-propanediol. Curr. Opin. Biotechnol. 2003, 14, 454–459. [Google Scholar] [CrossRef]
- Ingram, L.O.; Conway, T.; Alterthum, F. Ethanol Production by Escherichia coli Strains Co-Expressing Zymomonas PDC and ADH Genes. 1988. Available online: https://patents.google.com/patent/US5000000A/en (accessed on 17 May 2021).
- Yomano, L.P.; York, S.W.; Zhou, S.; Shanmugam, K.T.; Ingram, L.O. Re-engineering Escherichia coli for ethanol production. Biotechnol. Lett. 2008, 30, 2097–2103. [Google Scholar] [CrossRef]
- Atsumi, S.; Hanai, T.; Liao, J.C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 2008, 451, 86–89. [Google Scholar] [CrossRef]
- Amer, M.; Hoeven, R.; Kelly, P.; Faulkner, M.; Smith, M.H.; Toogood, H.S.; Scrutton, N.S. Renewable and tuneable bio-LPG blends derived from amino acids. Biotechnol. Biofuels 2020, 13, 125. [Google Scholar] [CrossRef] [PubMed]
- Amer, M.; Wojcik, E.Z.; Sun, C.; Hoeven, R.; Hughes, J.M.X.; Faulkner, M.; Yunus, I.S.; Tait, S.; Johannissen, L.O.; Hardman, S.J.O.; et al. Low carbon strategies for sustainable bio-alkane gas production and renewable energy. Energy Environ. Sci. 2020, 13, 1818–1831. [Google Scholar] [CrossRef]
- Ascue Avalos, G.A.; Toogood, H.S.; Tait, S.; Messiha, H.L.; Scrutton, N.S. From bugs to bioplastics: Total (+)-dihydrocarvide biosynthesis by engineered Escherichia coli. Chem. Eur. J. Chem. Biol. 2019, 20, 785–792. [Google Scholar] [CrossRef]
- Steen, E.J.; Kang, Y.; Bokinsky, G.; Hu, Z.; Schirmer, A.; McClure, A.; del Cardayre, S.B.; Keasling, J.D. Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 2010, 463, 559–562. [Google Scholar] [CrossRef]
- Martin, V.J.J.; Pitera, D.J.; Withers, S.T.; Newman, J.D.; Keasling, J.D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 2003, 21, 796–802. [Google Scholar] [CrossRef]
- Alonso-Gutierrez, J.; Chan, R.; Batth, T.S.; Adams, P.D.; Keasling, J.D.; Petzold, C.J.; Lee, T.S. Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production. Metab. Eng. 2013, 19, 33–41. [Google Scholar] [CrossRef]
- Dunstan, M.S.; Robinson, C.J.; Jervis, A.J.; Yan, C.; Carbonell, P.; Hollywood, K.A.; Currin, A.; Swainston, N.; Le Feuvre, R.; Micklefield, J.; et al. Engineering Escherichia coli towards de novo production of gatekeeper (2S)-flavanones: Naringenin, pinocembrin, eriodictyol and homoeriodictyol. Synth. Biol. 2020, 5, ysaa012. [Google Scholar] [CrossRef]
- Yang, J.; Xian, M.; Su, S.; Zhao, G.; Nie, Q.; Jiang, X.; Zheng, Y.; Liu, W. Enhancing production of bio-isoprene using hybrid MVA pathway and isoprene synthase in E. coli. PLoS ONE 2012, 7, e33509. [Google Scholar] [CrossRef]
- Whited, G.M.; Feher, F.J.; Benko, D.A.; Cervin, M.A.; Chotani, G.K.; McAuliffe, J.C.; LaDuca, R.J.; Ben-Shoshan, E.A.; Sanford, K.J. Development of a gas-phase bioprocess for isoprene-monomer production using metabolic pathway engineering. Ind. Biotechnol. 2010, 6, 152–163. [Google Scholar] [CrossRef]
- Minami, H.; Kim, J.-S.; Ikezawa, N.; Takemura, T.; Katayama, T.; Kumagai, H.; Sato, F. Microbial production of plant benzylisoquinoline alkaloids. Proc. Natl. Acad. Sci. USA 2008, 105, 7393–7398. [Google Scholar] [CrossRef]
- Nakagawa, A.; Matsumura, E.; Koyanagi, T.; Katayama, T.; Kawano, N.; Yoshimatsu, K.; Yamamoto, K.; Kumagai, H.; Sato, F.; Minami, H. Total biosynthesis of opiates by stepwise fermentation using engineered Escherichia coli. Nat. Commun. 2016, 7, 10390. [Google Scholar] [CrossRef]
- Bokinsky, G.; Peralta-Yahya, P.P.; George, A.; Holmes, B.M.; Steen, E.J.; Dietrich, J.; Lee, T.S.; Tullman-Ercek, D.; Voigt, C.A.; Simmons, B.A.; et al. Synthesis of three advanced biofuels from ionic liquid-pretreated switchgrass using engineered Escherichia coli. Proc. Natl. Acad. Sci. USA 2011, 108, 19949–19954. [Google Scholar] [CrossRef]
- Ryu, S.; Karim, M.N. A whole cell biocatalyst for cellulosic ethanol production from dilute acid-pretreated corn stover hydrolyzates. Appl. Microbiol. Biotechnol. 2011, 91, 529–542. [Google Scholar] [CrossRef]
- Pang, J.; Liu, Z.-Y.; Hao, M.; Zhang, Y.-F.; Qi, Q.-S. An isolated cellulolytic Escherichia coli from bovine rumen produces ethanol and hydrogen from corn straw. Biotechnol. Biofuels 2017, 10, 165. [Google Scholar] [CrossRef]
- Gao, D.; Luan, Y.; Wang, Q.; Liang, Q.; Qi, Q. Construction of cellulose-utilizing Escherichia coli based on a secretable cellulase. Microb. Cell. Fact. 2015, 14, 159. [Google Scholar] [CrossRef]
- Qian, Z.-G.; Xia, X.-X.; Choi, J.H.; Lee, S.Y. Proteome-based identification of fusion partner for high-level extracellular production of recombinant proteins in Escherichia coli. Biotechnol. Bioeng. 2008, 101, 587–601. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Wang, S.; Li, H.; Yu, H.; Qi, Q. Identification of a heterologous cellulase and its N-terminus that can guide recombinant proteins out of Escherichia coli. Microb. Cell. Fact. 2015, 14, 49. [Google Scholar] [CrossRef]
- Ashiuchi, M.; Nawa, C.; Kamei, T.; Song, J.J.; Hong, S.P.; Sung, M.H.; Soda, K.; Misono, H. Physiological and biochemical characteristics of poly gamma-glutamate synthetase complex of Bacillus subtilis. Eur. J. Biochem. 2001, 268, 5321–5328. [Google Scholar] [CrossRef]
- Longwell, C.K.; Labanieh, L.; Cochran, J.R. High-throughput screening technologies for enzyme engineering. Curr. Opin. Biotechnol. 2017, 48, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Hillson, N.; Caddick, M.; Cai, Y.; Carrasco, J.A.; Chang, M.W.; Curach, N.C.; Bell, D.J.; Le Feuvre, R.; Friedman, D.C.; Fu, X.; et al. Building a global alliance of biofoundries. Nat. Commun. 2019, 10, 2040. [Google Scholar] [CrossRef]
- Young, R.; Haines, M.; Storch, M.; Freemont, P.S. Combinatorial metabolic pathway assembly approaches and toolkits for modular assembly. Metab Eng. 2021, 63, 81–101. [Google Scholar] [CrossRef]
- Carbonell, P.; Jervis, A.J.; Robinson, C.J.; Yan, C.; Dunstan, M.; Swainston, N.; Vinaixa, M.; Hollywood, K.A.; Currin, A.; Rattray, N.J.W.; et al. An automated Design-Build-Test-Learn pipeline for enhanced microbial production of fine chemicals. Commun. Biol. 2018, 1, 66. [Google Scholar] [CrossRef]
- Delépine, B.; Duigou, T.; Carbonell, P.; Faulon, J.-L. RetroPath2.0: A retrosynthesis workflow for metabolic engineers. Metab. Eng. 2018, 45, 158–170. [Google Scholar] [CrossRef]
- Carbonell, P.; Wong, J.; Swainston, N.; Takano, E.; Turner, N.J.; Scrutton, N.S.; Kell, D.B.; Breitling, R.; Faulon, J.-L. Selenzyme: Enzyme selection tool for pathway design. Bioinformatics 2018, 34, 2153–2154. [Google Scholar] [CrossRef] [PubMed]
- Moriya, Y.; Shigemizu, D.; Hattori, M.; Tokimatsu, T.; Kotera, M.; Goto, S.; Kanehisa, M. PathPred: An enzyme-catalyzed metabolic pathway prediction server. Nucleic Acids Res. 2010, 38, W138–W143. [Google Scholar] [CrossRef]
- Jervis, A.J.; Carbonell, P.; Taylor, S.; Sung, R.; Dunstan, M.S.; Robinson, C.J.; Breitling, R.; Takano, E.; Scrutton, N.S. SelProm: A queryable and predictive expression vector selection tool for Escherichia coli. ACS Synth. Biol. 2019, 8, 1478–1483. [Google Scholar] [CrossRef]
- Angermueller, C.; Pärnamaa, T.; Parts, L.; Stegle, O. Deep learning for computational biology. Mol. Syst. Biol. 2016, 12, 878. [Google Scholar] [CrossRef]
- Casini, A.; Chang, F.-Y.; Eluere, R.; King, A.M.; Young, E.M.; Dudley, Q.M.; Karim, A.; Pratt, K.; Bristol, C.; Forget, A.; et al. A pressure test to make 10 molecules in 90 days: External evaluation of methods to engineer biology. J. Am. Chem. Soc. 2018, 140, 4302–4316. [Google Scholar] [CrossRef] [PubMed]
- Robinson, C.J.; Carbonell, P.; Jervis, A.J.; Yan, C.; Hollywood, K.A.; Dunstan, M.S.; Currin, A.; Swainston, N.; Spiess, R.; Taylor, S.; et al. Rapid prototyping of microbial production strains for the biomanufacture of potential materials monomers. Metab. Eng. 2020, 60, 168–182. [Google Scholar] [CrossRef]
- Choudhari, V.G.; Odaneth, A.A.; Lali, A.M. Application of high-throughput screening for evaluating hydrolytic potential of cellulases. Biomass Convers. Biorefin. 2019, 9, 659–667. [Google Scholar] [CrossRef]
- King, B.C.; Donnelly, M.K.; Bergstrom, G.C.; Walker, L.P.; Gibson, D.M. An optimized microplate assay system for quantitative evaluation of plant cell wall-degrading enzyme activity of fungal culture extracts. Biotechnol. Bioeng. 2009, 102, 1033–1044. [Google Scholar] [CrossRef]
- Yan, S.; Wu, G. Prediction of Michaelis-Menten constant of beta-glucosidases using nitrophenyl-beta-D-glucopyranoside as substrate. Protein Pept. Lett. 2011, 18, 1053–1057. [Google Scholar] [CrossRef]
- Deshpande, M.V.; Eriksson, K.E.; Göran Pettersson, L. An assay for selective determination of exo-1,4,-β-glucanases in a mixture of cellulolytic enzymes. Anal. Biochem. 1984, 138, 481–487. [Google Scholar] [CrossRef]
- Körfer, G.; Pitzler, C.; Vojcic, L.; Martinez, R.; Schwaneberg, U. In vitro flow cytometry-based screening platform for cellulase engineering. Sci. Rep. 2016, 6, 26128. [Google Scholar] [CrossRef]
- Beneyton, T.; Wijaya, I.P.M.; Postros, P.; Najah, M.; Leblond, P.; Couvent, A.; Mayot, E.; Griffiths, A.D.; Drevelle, A. High-throughput screening of filamentous fungi using nanoliter-range droplet-based microfluidics. Sci. Rep. 2016, 6, 27223. [Google Scholar] [CrossRef]
- Ostafe, R.; Prodanovic, R.; Lloyd Ung, W.; Weitz, D.A.; Fischer, R. A high-throughput cellulase screening system based on droplet microfluidics. Biomicrofluidics 2014, 8, 041102. [Google Scholar] [CrossRef]
- Najah, M.; Calbrix, R.; Mahendra-Wijaya, I.P.; Beneyton, T.; Griffiths, A.D.; Drevelle, A. Droplet-based microfluidics platform for ultra-high-throughput bioprospecting of cellulolytic microorganisms. Chem. Biol. 2014, 21, 1722–1732. [Google Scholar] [CrossRef]

| Product | Use | Design | Yield | Ref. |
|---|---|---|---|---|
| 1,3-Propanediol | PTT production 1 | Glycerol-3-phosphate dehydrogenase (DAR1 and GPP2) from S. cerevisiae. Glycerol dehydratase (dhaB1, dhaB2 and dhaB3) from Klebsiella pneumoniae. Endogenous ene-reductase (YqhD). | 130 g/L | [108] |
| 1,4-Butanediol | Advanced biofuel Polymer | Succinate semialdehyde dehydrogenase from E. coli and Porphyromonas gingivalis. 4-hydroxybutyrate dehydrogenase and 4-hydroxybutyryl-CoA transferase from P. gingivalis. Alcohol dehydrogenase from Clostridium acetobuylicum. | 20 g/L | [20] |
| Ethanol | Biofuel | Pyruvate decarboxylase and alcohol dehydrogenase from Z. mobilis. | 46 g/L | [109,110] |
| Isobutanol | Advanced biofuel | Endogenous 2-hydroxy-3-ketol-acid reductoisomerase, dihydroxy-acid dehydratase and alcohol dehydrogenase. Acetolactate synthase from B. subtilis. Ketoisovalerate decarboxylase from L. lactis. | 22 g/L | [111] |
| Hydrocarbon gases (bio-LPG) | Advanced synthetic fuels | Multiple de novo metabolic routes based on amino acid utilisation, fatty acid biosynthesis, Clostridial butanol production and single step from butyric acid via fatty acid photodecarboxylase. | 30–180 mg/g/d 2 | [112,113] |
| (+)-Dihydrocarvide | Bioplastics | Mentha spicata route to carvone with an ene-reductase and cyclohexanone monooxygenase variant. | 6.6 mg/L | [114] |
| Linalool | Hygiene products; chemical intermediate | “Plug-and-play” monoterpenoid production platform with linalool synthase. | 363 mg/L 3 | [28,29] |
| Fatty acid esters | Biodiesel | Thioesterase (tesA) and wax-ester synthase. Pyruvate decarboxylase and alcohol dehydrogenase from Z. mobilis. | 674 mg/L | [115] |
| Limonene | Platform chemical Pharmaceutical industry | Heterologous methylerythritol 4-phosphate (MEP) pathway. Limonene synthase from Mentha spicata. | 430 mg/L | [116,117] |
| Naringenin | Pharmaceutical industry | Flavanone pathway from L-tyrosine. | 199 mg/L | [118] |
| Isopropene | Synthetic rubber | Heterologous mevalonate (MVA) pathway. Isoprene synthase from Populus alba and P. kudzu. | 60 g/L | [119,120] |
| Taxiden-5α-ol | Taxol (anti-cancer drug) | Heterologous MEP pathway. Taxidene synthase from Taxus brevifolia, taxadiene 5α-hydroxylase and cytochrome P450. | 58 mg/L | [16] |
| Succinic acid | Tetrahydrofuran | Knockdown of metabolic pyruvate drains. Pyruvate carboxylase from Rhizobium etli. | 99 g/L | [19] |
| Hydrocodone | Opiate | Thebaine 6-O-demethylase and morphinone reductase from Pseudomonas putida and (R)-reticuline biosynthesis. | 2.1 mg/L | [121,122] |
| Feedstock | Cellulases | Export Tag | Product | Yield | Ref. |
|---|---|---|---|---|---|
| Ionic liquid pre-treated switchgrass | β-Glucosidase, endoxylanase and xylobiosidase | OsmY fusion | Fatty acid ethyl esters Butane Pinene | 71 mg/L 8 mg/L 1.7 mg/L | [123] |
| Amorphous cellulose | Cel-CD and β-glucosidase | Cel-CD tag | 3-hydroxybutyrate | 0.3 g/L | [126] |
| Dilute acid pre-treated corn stover | Endoglucanase Cel5A, exoglucanase Cel9E, and β-glucosidase | PsgA | Ethanol | 0.3 g/L | [124] |
| Corn straw | Endogenous cellulase | Native | Ethanol Hydrogen | 0.36 g/L 3.3 mL/g | [125] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Banner, A.; Toogood, H.S.; Scrutton, N.S. Consolidated Bioprocessing: Synthetic Biology Routes to Fuels and Fine Chemicals. Microorganisms 2021, 9, 1079. https://doi.org/10.3390/microorganisms9051079
Banner A, Toogood HS, Scrutton NS. Consolidated Bioprocessing: Synthetic Biology Routes to Fuels and Fine Chemicals. Microorganisms. 2021; 9(5):1079. https://doi.org/10.3390/microorganisms9051079
Chicago/Turabian StyleBanner, Alec, Helen S. Toogood, and Nigel S. Scrutton. 2021. "Consolidated Bioprocessing: Synthetic Biology Routes to Fuels and Fine Chemicals" Microorganisms 9, no. 5: 1079. https://doi.org/10.3390/microorganisms9051079
APA StyleBanner, A., Toogood, H. S., & Scrutton, N. S. (2021). Consolidated Bioprocessing: Synthetic Biology Routes to Fuels and Fine Chemicals. Microorganisms, 9(5), 1079. https://doi.org/10.3390/microorganisms9051079






