The Composition and Diversity of the Gut Microbiota in Children Is Modifiable by the Household Dogs: Impact of a Canine-Specific Probiotic
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Fecal Samples and Microbiota Analyses
2.3. Data Analysis
3. Results
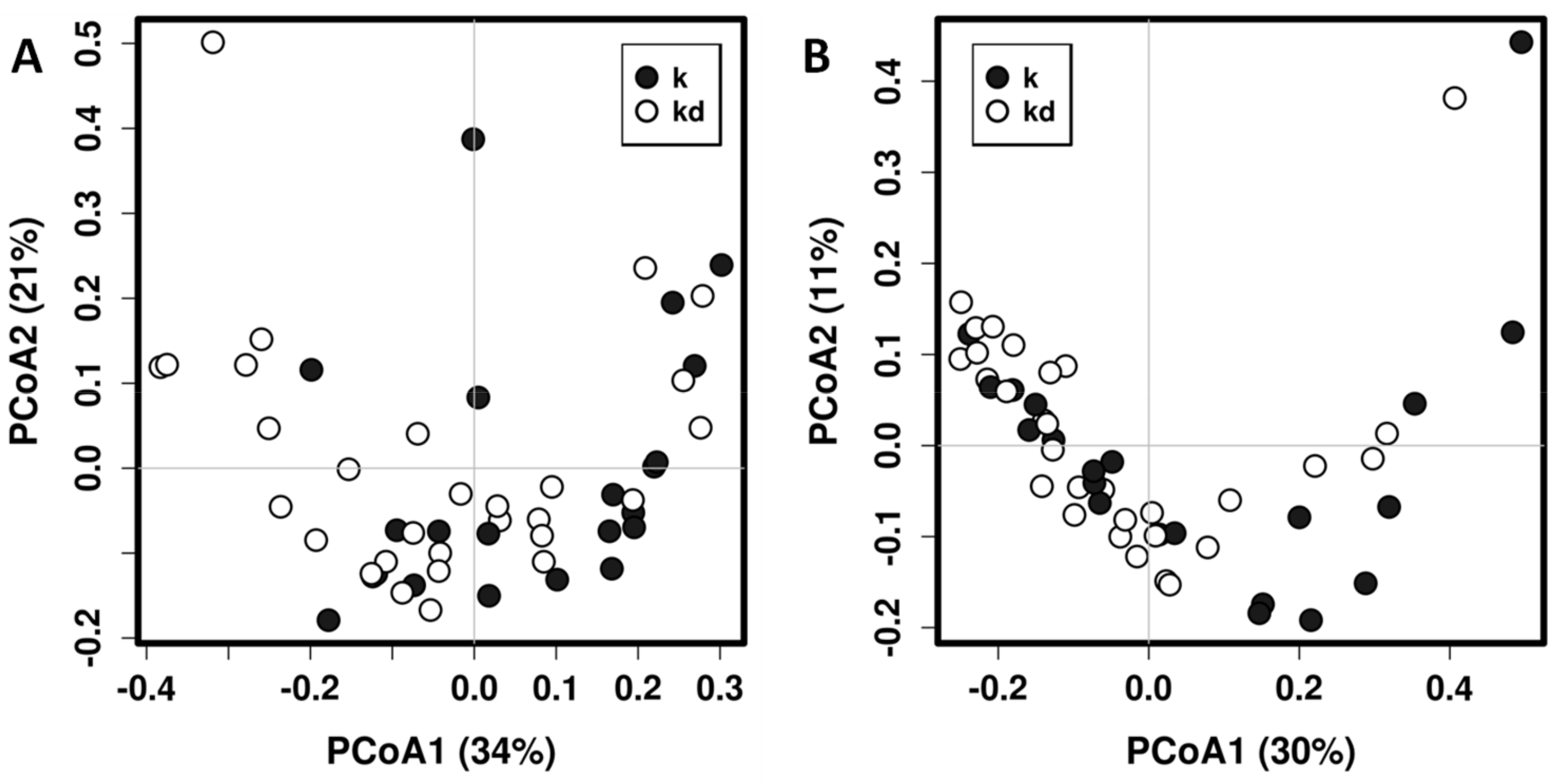
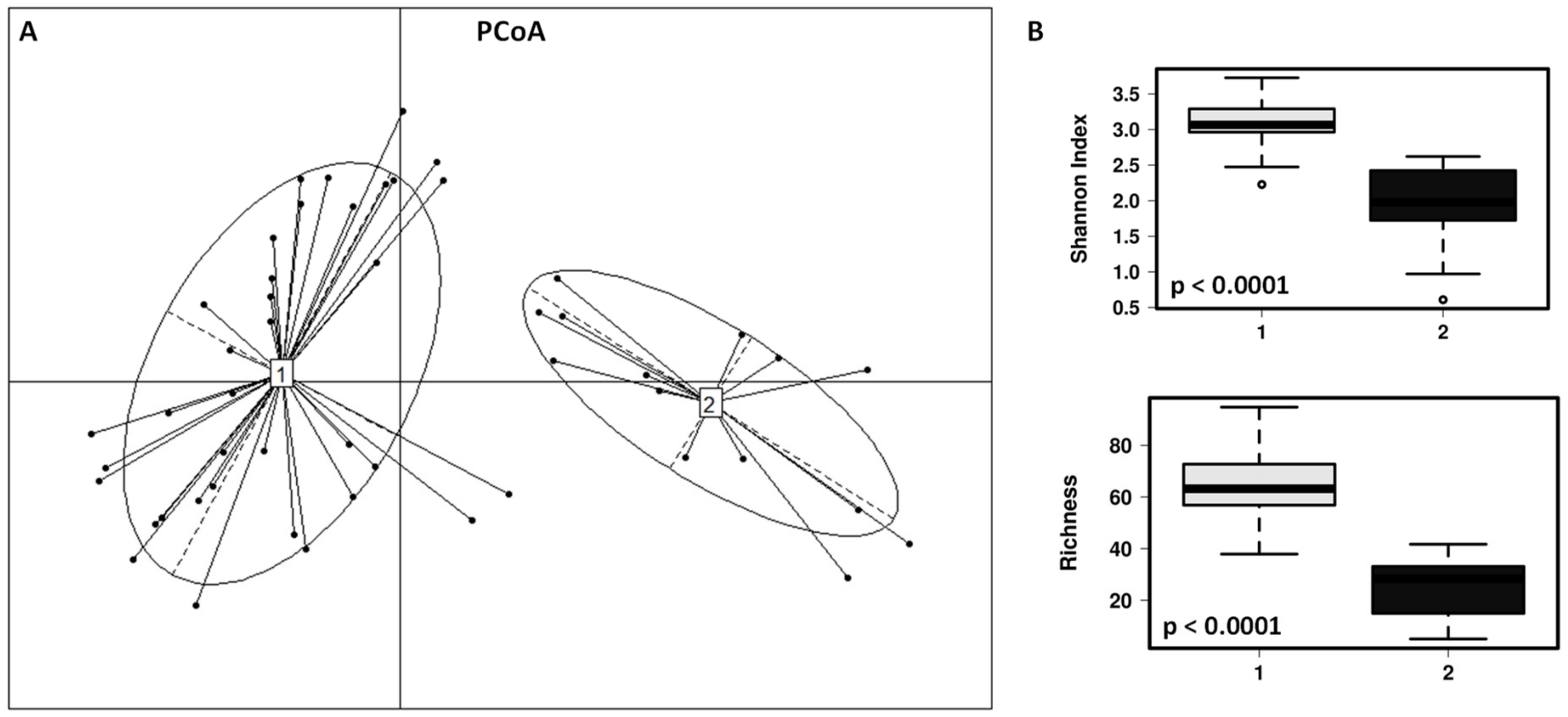
3.1. Impact of the Dog on the Gut Microbiota of Children
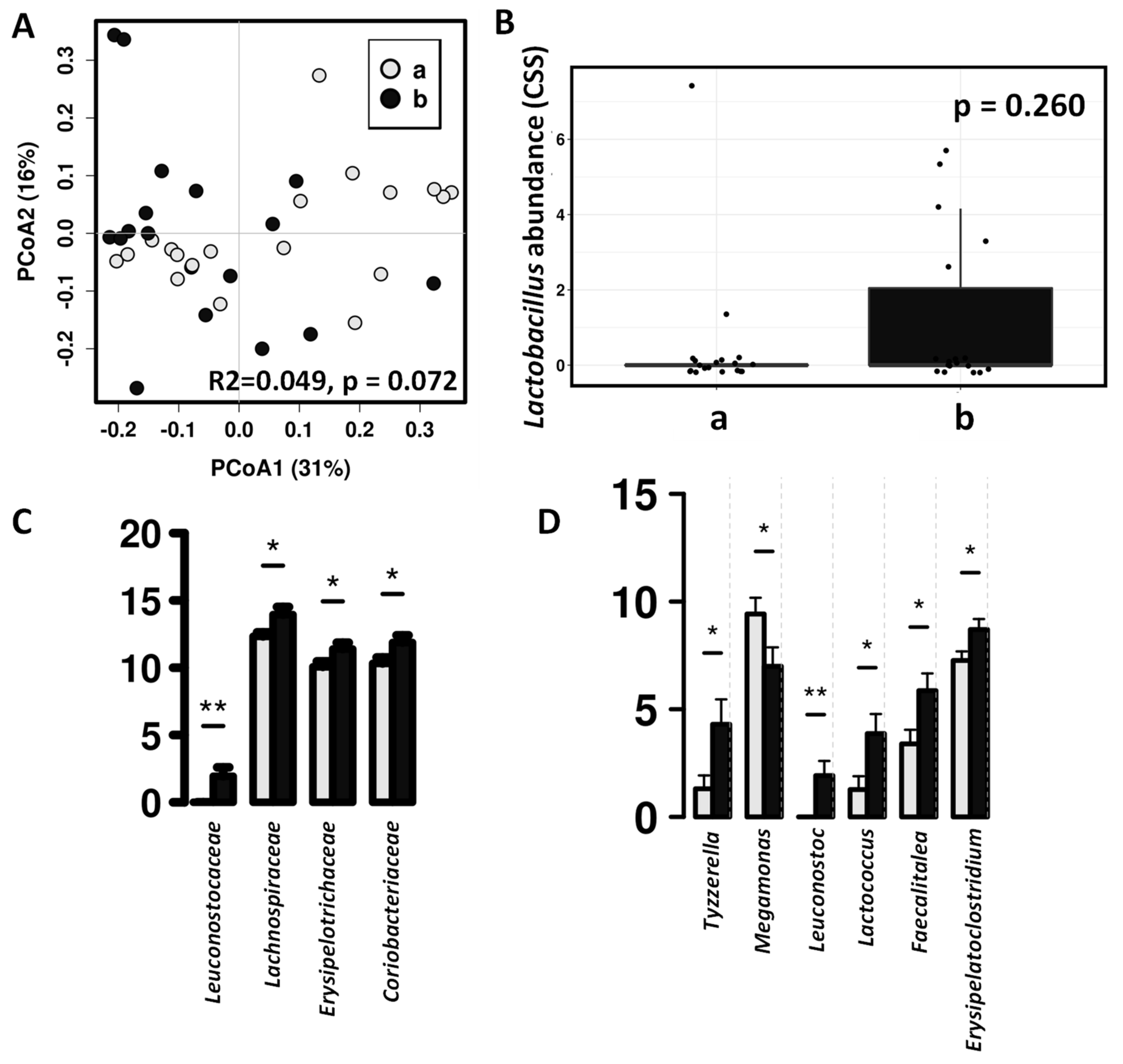
3.2. Impact of the Canine-Specific Lactic Acid Bacteria Product on Dogs’ Fecal Microbiota
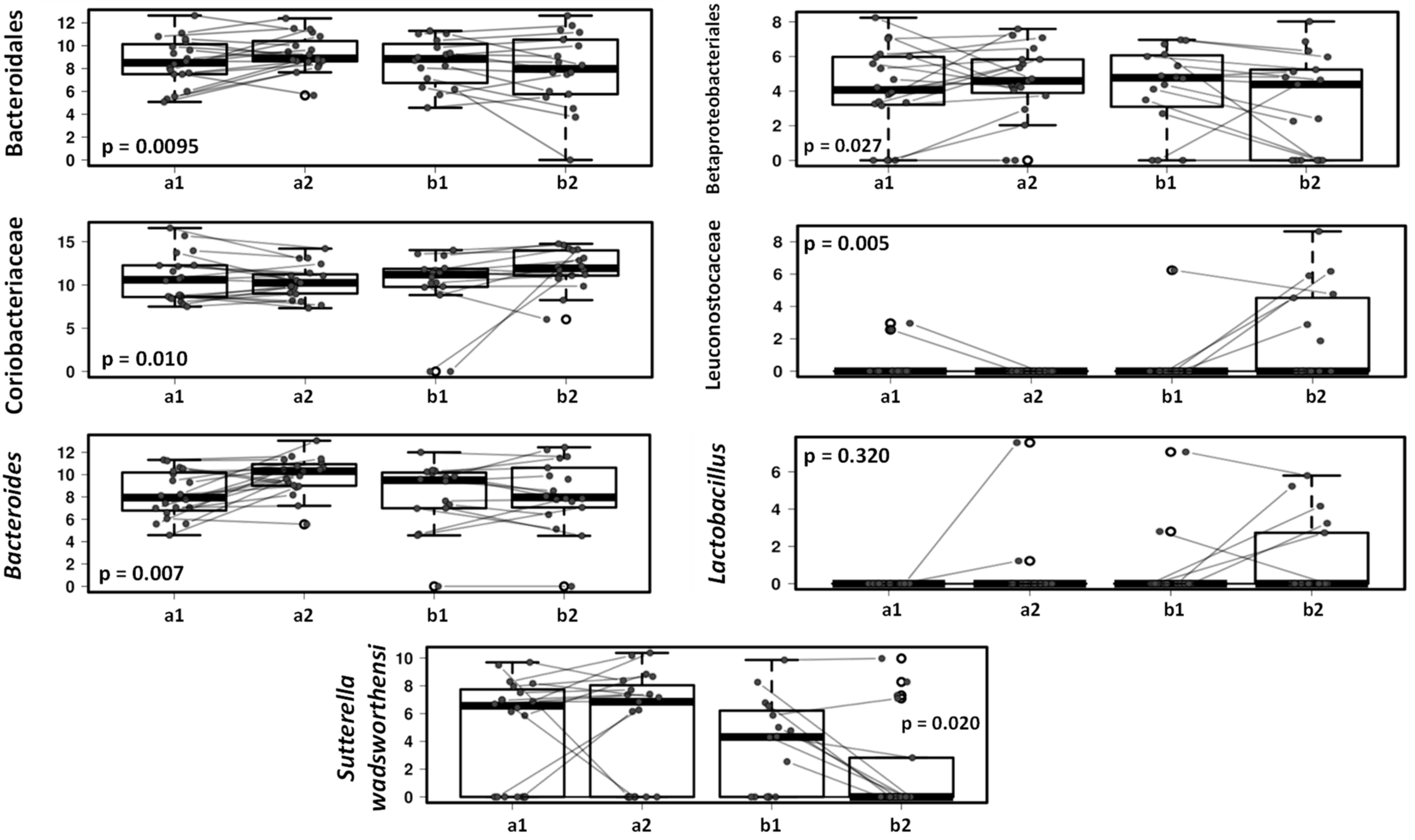
3.3. Effect of the Canine-Specific Product on the Gut Microbiota of Children with Dog
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Moloney, R.D.; Desbonnet, L.; Clarke, G.; Dinan, T.G.; Cryan, J.F. The microbiome: Stress, health and disease. Mamm. Genome 2014, 25, 49–74. [Google Scholar] [CrossRef]
- Milani, C.; Duranti, S.; Bottacini, F.; Casey, E.; Turroni, F.; Mahony, J.; Belzer, C.; Delgado Palacio, S.; Arboleya Montes, S.; Mancabelli, L.; et al. The First Microbial Colonizers of the Human Gut: Composition, Activities, and Health Implications of the Infant Gut Microbiota. Microbiol. Mol. Biol. Rev. 2017, 81. [Google Scholar] [CrossRef]
- Lehtimäki, J.; Karkman, A.; Laatikainen, T.; Paalanen, L.; Von Hertzen, L.; Haahtela, T.; Hanski, I.; Ruokolainen, L. Patterns in the skin microbiota differ in children and teenagers between rural and urban environments. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef]
- De Filippo, C.; Di Paola, M.; Ramazzotti, M.; Albanese, D.; Pieraccini, G.; Banci, E.; Miglietta, F.; Cavalieri, D.; Lionetti, P. Diet, environments, and gut microbiota. A preliminary investigation in children living in rural and Urban Burkina Faso and Italy. Front. Microbiol. 2017, 8, 1979. [Google Scholar] [CrossRef]
- Schröder, P.C.; Li, J.; Wong, G.W.K.; Schaub, B. The rural-urban enigma of allergy: What can we learn from studies around the world? Pediatr. Allergy Immunol. 2015, 26, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Zhao, J.; Han, Z.R.; Chen, Y.; Leung, T.F.; Wong, G.W.K. Very low prevalence of asthma and allergies in schoolchildren from rural Beijing, China. Pediatr. Pulmonol. 2009, 44, 793–799. [Google Scholar] [CrossRef]
- Nermes, M.; Endo, A.; Aarnio, J.; Salminen, S.; Isolauri, E. Furry pets modulate gut microbiota composition in infants at risk for allergic disease. J. Allergy Clin. Immunol. 2015, 136, 1688–1690.e1. [Google Scholar] [CrossRef]
- Oh, C.; Lee, K.; Cheong, Y.; Lee, S.W.; Park, S.Y.; Song, C.S.; Choi, I.S.; Lee, J.B. Comparison of the oral microbiomes of canines and their owners using next- generation sequencing. PLoS ONE 2015, 10, e0131468. [Google Scholar] [CrossRef]
- Manninen, T.J.K.; Rinkinen, M.L.; Beasley, S.S.; Saris, P.E.J. Alteration of the canine small-intestinal lactic acid bacterium microbiota by feeding of potential probiotics. Appl. Environ. Microbiol. 2006, 72, 6539–6543. [Google Scholar] [CrossRef] [PubMed]
- Grześkowiak, L.; Collado, M.C.; Beasley, S.; Salminen, S. Pathogen exclusion properties of canine probiotics are influenced by the growth media and physical treatments simulating industrial processes. J. Appl. Microbiol. 2014, 116, 1308–1314. [Google Scholar] [CrossRef]
- Gómez-Gallego, C.; Junnila, J.; Männikkö, S.; Hämeenoja, P.; Valtonen, E.; Salminen, S.; Beasley, S. A canine-specific probiotic product in treating acute or intermittent diarrhea in dogs: A double-blind placebo-controlled efficacy study. Vet. Microbiol. 2016, 197, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Nylund, L.; Heilig, H.G.H.J.; Salminen, S.; de Vos, W.M.; Satokari, R. Semi-automated extraction of microbial DNA from feces for qPCR and phylogenetic microarray analysis. J. Microbiol. Methods 2010, 83, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Díaz, J.; García-Mantrana, I.; Vila-Vicent, S.; Gozalbo-Rovira, R.; Buesa, J.; Monedero, V.; Collado, M.C. Relevance of secretor status genotype and microbiota composition in susceptibility to rotavirus and norovirus infections in humans. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- RStudio Team. RStudio: Integrated Development for R. Available online: http://www.rstudio.com/ (accessed on 2 October 2020).
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef]
- Zakrzewski, M.; Proietti, C.; Ellis, J.J.; Hasan, S.; Brion, M.J.; Berger, B.; Krause, L. Calypso: A user-friendly web-server for mining and visualizing microbiome-environment interactions—PubMed. Bioinforma. 2017, 33, 782–783. [Google Scholar] [CrossRef]
- Paulson, J.N.; Colin Stine, O.; Bravo, H.C.; Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 2013, 10, 1200–1202. [Google Scholar] [CrossRef]
- Selma-Royo, M.; García-Mantrana, I.; Calatayud, M.; Parra-Llorca, A.; Martínez-Costa, C.; Collado, M.C. Maternal Microbiota, Cortisol Concentration, and Post-Partum Weight Recovery are Dependent on Mode of Delivery. Nutrients 2020, 12, 1779. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Maechler, M.; Rousseeuw, P.; Struyf, A.; Hubert, M.; Hornik, K. Cluster: Cluster Analysis Basics and Extensions. R Package Version 2.1.0. Available online: https://cran.r-project.org/web/packages/cluster/cluster.pdf (accessed on 14 July 2020).
- Venables, W.N.; Ripley, B.D. Modern Applied Statistics with S.; Springer: New York, NY, USA, 2002. [Google Scholar]
- Walesiak Marek; Dudek Andrzej clusterSim: Searching for Optimal Clustering Procedure for a Data Set Version 0.48-3 from CRAN. Available online: https://rdrr.io/cran/clusterSim/ (accessed on 14 July 2020).
- Dray, S.; Dufour, A.B. The ade4 package: Implementing the duality diagram for ecologists. J. Stat. Softw. 2007, 22, 1–20. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Sitarik, A.R.; Woodcroft, K.; Johnson, C.C.; Zoratti, E. Birth Mode, Breastfeeding, Pet Exposure, and Antibiotic Use: Associations with the Gut Microbiome and Sensitization in Children. Curr. Allergy Asthma Rep. 2019, 19, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tun, H.M.; Konya, T.; Takaro, T.K.; Brook, J.R.; Chari, R.; Field, C.J.; Guttman, D.S.; Becker, A.B.; Mandhane, P.J.; Turvey, S.E.; et al. Exposure to household furry pets influences the gut microbiota of infants at 3–4 months following various birth scenarios. Microbiome 2017, 5, 1–14. [Google Scholar] [CrossRef]
- Azad, M.B.; Konya, T.; Maughan, H.; Guttman, D.S.; Field, C.J.; Sears, M.R.; Becker, A.B.; Scott, J.A.; Kozyrskyj, A.L. Infant gut microbiota and the hygiene hypothesis of allergic disease: Impact of household pets and siblings on microbiota composition and diversity. Allergy Asthma Clin. Immunol. 2013, 9, 15. [Google Scholar] [CrossRef]
- Song, S.J.; Lauber, C.; Costello, E.K.; Lozupone, C.A.; Humphrey, G.; Berg-Lyons, D.; Gregory Caporaso, J.; Knights, D.; Clemente, J.C.; Nakielny, S.; et al. Cohabiting family members share microbiota with one another and with their dogs. Elife 2013, 2013, e00458. [Google Scholar] [CrossRef]
- Gergen, P.J.; Mitchell, H.E.; Calatroni, A.; Sever, M.L.; Cohn, R.D.; Salo, P.M.; Thorne, P.S.; Zeldin, D.C. Sensitization and Exposure to Pets: The Effect on Asthma Morbidity in the US Population. J. Allergy Clin. Immunol. Pract. 2018, 6, 101–107.e2. [Google Scholar] [CrossRef] [PubMed]
- Perzanowski, M.S.; Ronmark, E.; James, H.R.; Hedman, L.; Schuyler, A.J.; Bjerg, A.; Lundback, B.; Platts-Mills, T.A.E. Relevance of specific IgE antibody titer to the prevalence, severity, and persistence of asthma among 19-year-olds in northern Sweden. J. Allergy Clin. Immunol. 2016, 138, 1582–1590. [Google Scholar] [CrossRef]
- Fall, T.; Lundholm, C.; Örtqvist, A.K.; Fall, K.; Fang, F.; Hedhammar, Å.; Kämpe, O.; Ingelsson, E.; Almqvist, C. Early exposure to dogs and farm animals and the risk of childhood asthma. JAMA Pediatr. 2015, 169, e153219. [Google Scholar] [CrossRef]
- Hesselmar, B.; Åberg, N.; Åberg, B.; Eriksson, B.; Björkstén, B. Does early exposure to cat or dog protect against later allergy development? Clin. Exp. Allergy 1999, 29, 611–617. [Google Scholar] [CrossRef]
- Krzych-Fałta, E.; Furmańczyk, K.; Piekarska, B.; Raciborski, F.; Tomaszewska, A.; Walkiewicz, A.; Samel-Kowalik, P.; Borowicz, J.; Namysłowski, A.; Samoliński, B. Extent of protective or allergy-inducing effects in cats and dogs. Ann. Agric. Environ. Med. 2018, 25, 268–273. [Google Scholar] [CrossRef]
- Fall, T.; Ekberg, S.; Lundholm, C.; Fang, F.; Almqvist, C. Dog characteristics and future risk of asthma in children growing up with dogs. Sci. Rep. 2018, 8, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hesselmar, B.; Hicke-Roberts, A.; Lundell, A.C.; Adlerberth, I.; Rudin, A.; Saalman, R.; Wennergren, G.; Wold, A.E. Pet-keeping in early life reduces the risk of allergy in a dose-dependent fashion. PLoS ONE 2018, 13, e0208472. [Google Scholar] [CrossRef]
- Gschwendtner, S.; Kang, H.; Thiering, E.; Kublik, S.; Fösel, B.; Schulz, H.; Krauss-Etschmann, S.; Heinrich, J.; Schöler, A.; Schloter, M.; et al. Early life determinants induce sustainable changes in the gut microbiome of six-year-old children. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How colonization by microbiota in early life shapes the immune system. Science 2016, 352, 539–544. [Google Scholar] [CrossRef]
- Bergström, A.; Skov, T.H.; Bahl, M.I.; Roager, H.M.; Christensen, L.B.; Ejlerskov, K.T.; Mølgaard, C.; Michaelsen, K.F.; Licht, T.R. Establishment of intestinal microbiota during early life: A longitudinal, explorative study of a large cohort of Danish infants. Appl. Environ. Microbiol. 2014, 80, 2889–2900. [Google Scholar] [CrossRef]
- Kuang, Y.S.; Li, S.H.; Guo, Y.; Lu, J.H.; He, J.R.; Luo, B.J.; Jiang, F.J.; Shen, H.; Papasian, C.J.; Pang, H.; et al. Composition of gut microbiota in infants in China and global comparison. Sci. Rep. 2016, 6, 1–10. [Google Scholar] [CrossRef]
- Zhong, H.; Penders, J.; Shi, Z.; Ren, H.; Cai, K.; Fang, C.; Ding, Q.; Thijs, C.; Blaak, E.E.; Stehouwer, C.D.A.; et al. Impact of early events and lifestyle on the gut microbiota and metabolic phenotypes in young school-age children. Microbiome 2019, 7, 2. [Google Scholar] [CrossRef]
- Crost, E.H.; Le Gall, G.; Laverde-Gomez, J.A.; Mukhopadhya, I.; Flint, H.J.; Juge, N. Mechanistic insights into the cross-feeding of Ruminococcus gnavus and Ruminococcus bromii on host and dietary carbohydrates. Front. Microbiol. 2018, 9, 2558. [Google Scholar] [CrossRef]
- Dabard, J.; Bridonneau, C.; Phillipe, C.; Anglade, P.; Molle, D.; Nardi, M.; Ladiré, M.; Girardin, H.; Marcille, F.; Gomez, A.; et al. Ruminococcin A, a New Lantibiotic Produced by a Ruminococcus gnavus Strain Isolated from Human Feces. Appl. Environ. Microbiol. 2001, 67, 4111–4118. [Google Scholar] [CrossRef] [PubMed]
- Lynch, S.V. Gut microbiota and allergic disease: New insights. Ann. Am. Thorac. Soc. 2016, 13, S51–S54. [Google Scholar]
- Könönen, E.; Wade, W.G. Actinomyces and related organisms in human infections. Clin. Microbiol. Rev. 2015, 28, 419–442. [Google Scholar] [CrossRef]
- Truax, A.D.; Chen, L.; Tam, J.W.; Cheng, N.; Guo, H.; Koblansky, A.A.; Chou, W.C.; Wilson, J.E.; Brickey, W.J.; Petrucelli, A.; et al. The Inhibitory Innate Immune Sensor NLRP12 Maintains a Threshold against Obesity by Regulating Gut Microbiota Homeostasis. Cell Host Microbe 2018, 24, 364–378.e6. [Google Scholar] [CrossRef] [PubMed]
- Forbes, J.D.; Van Domselaar, G.; Bernstein, C.N. The gut microbiota in immune-mediated inflammatory diseases. Front. Microbiol. 2016, 7, 1081. [Google Scholar] [CrossRef] [PubMed]
- Clavel, T.; Lepage, P.; Charrier, C. The family Coriobacteriaceae. In The Prokaryotes: Actinobacteria; Springer: Berlin/Heidelberg, Germany, 2014; pp. 201–238. ISBN 9783642301384. [Google Scholar]
- Molitoris, E.; Wexler, H.M.; Finegold, S.M. Sources and Antimicrobial Susceptibilities of Campylobacter gracilis and Sutterella wadsworthensis. Clin. Infect. Dis. 1997, 25, S264–S265. [Google Scholar] [CrossRef]
- Wang, L.; Christophersen, C.T.; Sorich, M.J.; Gerber, J.P.; Angley, M.T.; Conlon, M.A. Increased abundance of Sutterella spp. and Ruminococcus torques in feces of children with autism spectrum disorder. Mol. Autism 2013, 4, 42. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gómez-Gallego, C.; Forsgren, M.; Selma-Royo, M.; Nermes, M.; Collado, M.C.; Salminen, S.; Beasley, S.; Isolauri, E. The Composition and Diversity of the Gut Microbiota in Children Is Modifiable by the Household Dogs: Impact of a Canine-Specific Probiotic. Microorganisms 2021, 9, 557. https://doi.org/10.3390/microorganisms9030557
Gómez-Gallego C, Forsgren M, Selma-Royo M, Nermes M, Collado MC, Salminen S, Beasley S, Isolauri E. The Composition and Diversity of the Gut Microbiota in Children Is Modifiable by the Household Dogs: Impact of a Canine-Specific Probiotic. Microorganisms. 2021; 9(3):557. https://doi.org/10.3390/microorganisms9030557
Chicago/Turabian StyleGómez-Gallego, Carlos, Mira Forsgren, Marta Selma-Royo, Merja Nermes, Maria Carmen Collado, Seppo Salminen, Shea Beasley, and Erika Isolauri. 2021. "The Composition and Diversity of the Gut Microbiota in Children Is Modifiable by the Household Dogs: Impact of a Canine-Specific Probiotic" Microorganisms 9, no. 3: 557. https://doi.org/10.3390/microorganisms9030557
APA StyleGómez-Gallego, C., Forsgren, M., Selma-Royo, M., Nermes, M., Collado, M. C., Salminen, S., Beasley, S., & Isolauri, E. (2021). The Composition and Diversity of the Gut Microbiota in Children Is Modifiable by the Household Dogs: Impact of a Canine-Specific Probiotic. Microorganisms, 9(3), 557. https://doi.org/10.3390/microorganisms9030557





