Household Transmission of SARS-CoV-2: A Prospective Longitudinal Study Showing Higher Viral Load and Increased Transmissibility of the Alpha Variant Compared to Previous Strains
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Study Population
2.2. National COVID-19 Isolation and Quarantine Regulations
2.3. Sampling and Data Collection
2.4. Laboratory Testing
Detection of SARS-CoV-2 by rRT-PCR
2.5. Quantitative Analysis of SARS-CoV-2 by Droplet Digital PCR (ddPCR)
2.6. Sequencing of SARS-CoV-2
2.7. Blood Typing
2.8. Definition of Cases and Contacts
2.9. Definition of Variables
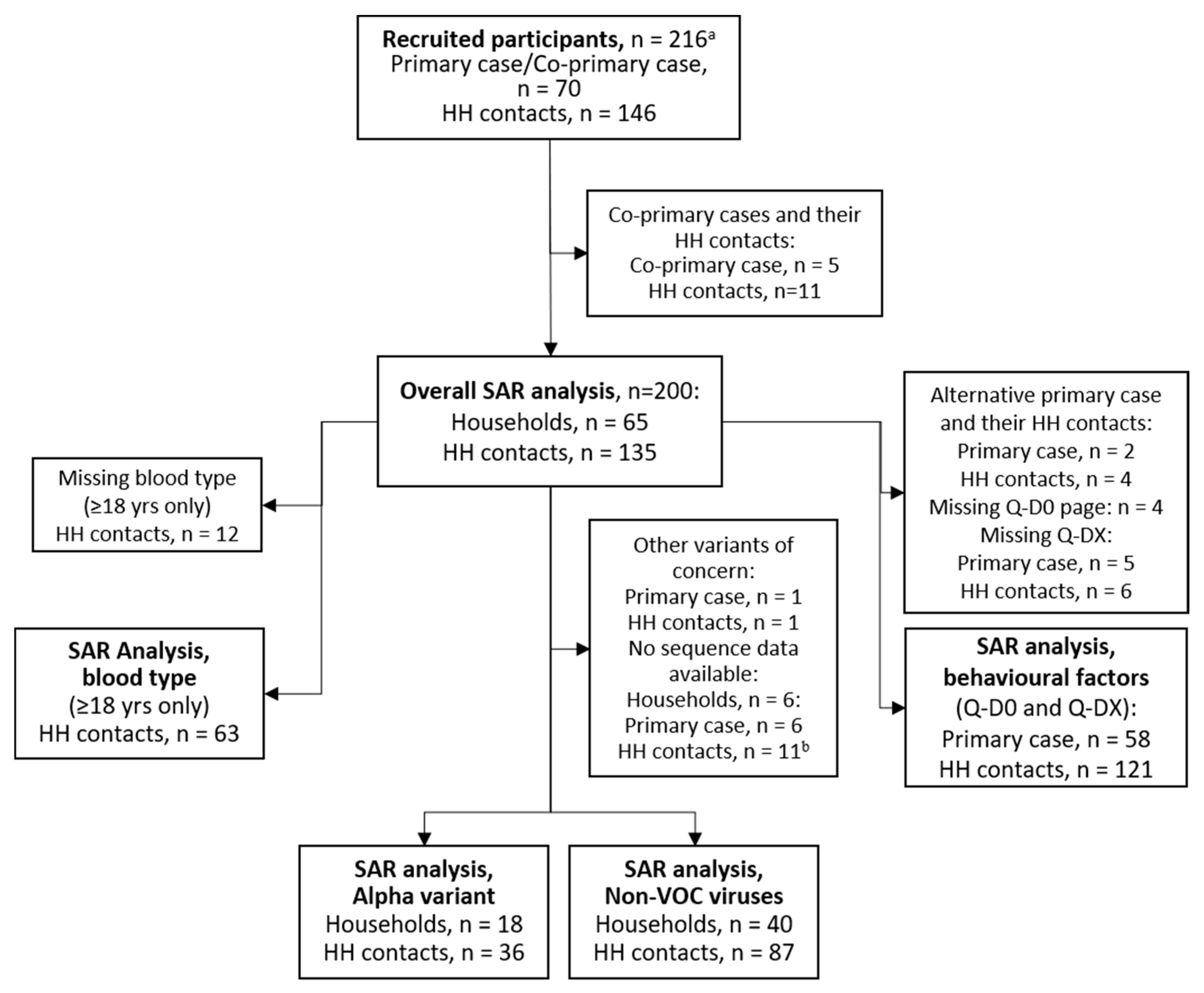
2.10. Study Samples Included in Analysis
2.11. Data Analysis and Statistical Methods
3. Results
3.1. Baseline Characteristics of Households and Participants
3.2. Secondary Transmission of COVID-19 in Households
3.3. Effect of Host and Household Characteristics on Secondary Transmission
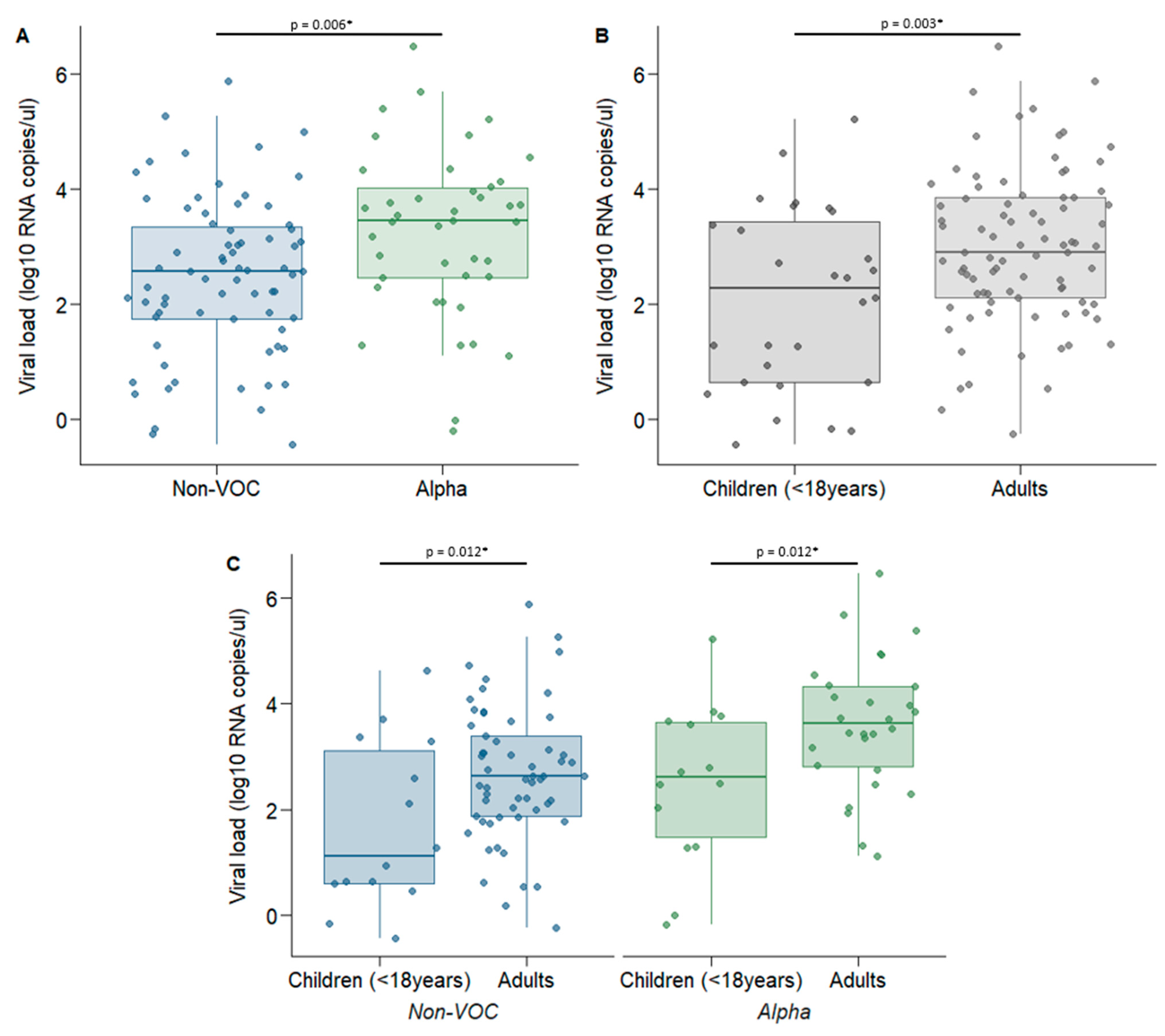
3.4. Role of Viral Load Measured by ddPCR
3.5. The Impact of Behavioral Factors and Precautionary Practices on Secondary Transmission
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HH contact | Household Contact |
| non-VOC | Non-Variant of Concern |
| VOC | Variant of Concern |
| HH | Households |
| IQR | Interquartile Range |
| SAR | Secondary Attack Rate |
| OR | Odds Ratio |
| CI | Confidence Interval |
References
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.-L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021, 19, 141–154. [Google Scholar] [CrossRef]
- Norwegian Institute of Public Health (NIPH). COVID-19 Ukesrapport-uke 34. Available online: https://www.fhi.no/contentassets/8a971e7b0a3c4a06bdbf381ab52e6157/vedlegg/2021/ukerapport-uke-34-23.08---29.08.21.pdf (accessed on 1 September 2021).
- Madewell, Z.J.; Yang, Y.; Longini, I.M., Jr.; Halloran, M.E.; Dean, N.E. Household Transmission of SARS-CoV-2: A Systematic Review and Meta-analysis. JAMA Netw. Open 2020, 3, e2031756. [Google Scholar] [CrossRef]
- Thompson, H.A.; Mousa, A.; Dighe, A.; Fu, H.; Arnedo-Pena, A.; Barrett, P.; Bellido-Blasco, J.; Bi, Q.; Caputi, A.; Chaw, L.; et al. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Setting-specific Transmission Rates: A Systematic Review and Meta-analysis. Clin. Infect. Dis. 2021, 73, e754–e764. [Google Scholar] [CrossRef]
- Oude Munnink, B.B.; Worp, N.; Nieuwenhuijse, D.F.; Sikkema, R.S.; Haagmans, B.; Fouchier, R.A.M.; Koopmans, M. The next phase of SARS-CoV-2 surveillance: Real-time molecular epidemiology. Nat. Med. 2021, 27, 1518–1524. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Loman, N.; Pybus, O.; Barclay, W.; Barrett, J.; Carabelli, A.; Connor, T.; Peacock, T.; Robertson, D.; Volz, E. Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations. Genom. Epidemiol. 2020, 1–5. [Google Scholar]
- Chand, M.; Hopkins, S.; Dabrera, G.; Achison, C.; Barclay, W.; Ferguson, N. Investigation of novel SARS-COV-2 variant Variant of Concern 202012/01. Public Health Engl. 2020, 21. [Google Scholar]
- Norwegian Institute of Public Health (NIPH). COVID-19 Ukesrapport-uke 28. Available online: https://www.fhi.no/contentassets/8a971e7b0a3c4a06bdbf381ab52e6157/vedlegg/2021/ukerapport-uke-28-12.07---18.07.21.pdf (accessed on 1 September 2021).
- Piantham, C.; Linton, N.M.; Nishiura, H.; Ito, K. Estimating the elevated transmissibility of the B.1.1.7 strain over previously circulating strains in England using GISAID sequence frequencies. medRxiv 2021. [Google Scholar] [CrossRef]
- Davies, N.G.; Abbott, S.; Barnard, R.C.; Jarvis, C.I.; Kucharski, A.J.; Munday, J.D.; Pearson, C.A.B.; Russell, T.W.; Tully, D.C.; Washburne, A.D.; et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 2021, 372, eabg3055. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.; Mishra, S.; Chand, M.; Barrett, J.C.; Johnson, R.; Geidelberg, L.; Hinsley, W.R.; Laydon, D.J.; Dabrera, G.; O’Toole, Á.; et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature 2021, 593, 266–269. [Google Scholar] [CrossRef] [PubMed]
- Cosentino, G.; Bernard, M.; Ambroise, J.; Giannoli, J.-M.; Guedj, J.; Débarre, F.; Blanquart, F. SARS-CoV-2 viral dynamics in infections with Alpha and Beta variants of concern in the French community. J. Infect. 2021. [Google Scholar] [CrossRef]
- Kissler, S.M.; Fauver, J.R.; Mack, C.; Tai, C.G.; Breban, M.I.; Watkins, A.E.; Samant, R.M.; Anderson, D.J.; Ho, D.D.; Metti, J.; et al. Densely sampled viral trajectories for SARS-CoV-2 variants alpha (B.1.1.7) and epsilon (B.1.429). medRxiv 2021. [Google Scholar] [CrossRef]
- Lyngse, F.P.; Mølbak, K.; Skov, R.L.; Christiansen, L.E.; Mortensen, L.H.; Albertsen, M.; Møller, C.H.; Krause, T.G.; Rasmussen, M.; Michaelsen, T.Y.; et al. Increased Transmissibility of SARS-CoV-2 Lineage B.1.1.7 by Age and Viral Load: Evidence from Danish Households. medRxiv 2021. [Google Scholar] [CrossRef]
- Dyavar, S.R.; Ye, Z.; Byrareddy, S.N.; Scarsi, K.K.; Winchester, L.C.; Weinhold, J.A.; Fletcher, C.V.; Podany, A.T. Normalization of cell associated antiretroviral drug concentrations with a novel RPP30 droplet digital PCR assay. Sci. Rep. 2018, 8, 3626. [Google Scholar] [CrossRef] [Green Version]
- Kim, K.B.; Choi, H.; Lee, G.D.; Lee, J.; Lee, S.; Kim, Y.; Cho, S.Y.; Lee, D.G.; Kim, M. Analytical and Clinical Performance of Droplet Digital PCR in the Detection and Quantification of SARS-CoV-2. Mol. Diagn. Ther. 2021, 25, 617–628. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Household Transmission Investigation Protocol for 2019-Novel Coronavirus (COVID-19) Infection. Available online: https://www.who.int/publications/i/item/household-transmission-investigation-protocol-for-2019-novel-coronavirus-(2019-ncov)-infection (accessed on 30 March 2020).
- World Health Organization (WHO). Protocol: Real-Time RT-PCR Assays for the Detection of SARS-CoV-2 Institut Pasteur, Paris. Geneva: WHO. Available online: https://www.who.int/docs/default-source/coronaviruse/real-time-rt-pcr-assays-for-the-detection-of-sars-cov-2-institut-pasteur-paris.pdf?sfvrsn=3662fcb6_2 (accessed on 15 June 2021).
- Basu, A.S. Digital Assays Part I: Partitioning Statistics and Digital PCR. SLAS Technol. Transl. Life Sci. Innov. 2017, 22, 369–386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quick, J. nCoV-2019 Sequencing Protocol v3 (LoCost). Protocols.io. Version Created by Josh Quick. Available online: https://protocols.io/view/ncov-2019-sequencing-protocol-v3-locost-bh42j8ye (accessed on 1 June 2021).
- Tyson, J.R.; James, P.; Stoddart, D.; Sparks, N.; Wickenhagen, A.; Hall, G.; Choi, J.H.; Lapointe, H.; Kamelian, K.; Smith, A.D.; et al. Improvements to the ARTIC multiplex PCR method for SARS-CoV-2 genome sequencing using nanopore. BioRxiv 2020. [Google Scholar] [CrossRef]
- Norwegian Institute of Public Health (NIPH). FHI Pipelines for SARS-CoV-2 Sequences Generated with the ARTIC Sequencing Protocol. Available online: https://github.com/folkehelseinstituttet/fhi-ncov-seq-pipelines (accessed on 1 June 2021).
- The Norwegian Sequencing Centre (NSC). SARS-CoV-2 Whole Genome Sequencing Based on Multiplexed Amplicon Method Using Short-Read Illumina Sequencers. Available online: https://github.com/nsc-norway/covid-seq (accessed on 1 June 2021).
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020, 5, 1403–1407. [Google Scholar] [CrossRef] [PubMed]
- Statistics Norway. Concept Variable-Crowded Dwelling. Available online: https://www.ssb.no/a/metadata/conceptvariable/vardok/3462/en (accessed on 1 June 2021).
- European Centre for Disease Prevention and Control (ECDC). SARS-CoV-2 Variants of Concern as of 24 June 2021. Available online: https://www.ecdc.europa.eu/en/covid-19/variants-concern (accessed on 1 June 2021).
- Rao, J.N.K.; Scott, A.J. On Chi-Squared Tests for Multiway Contingency Tables with Cell Proportions Estimated from Survey Data. Ann. Stat. 1984, 12, 46–60. [Google Scholar] [CrossRef]
- Fung, H.F.; Martinez, L.; Alarid-Escudero, F.; Salomon, J.A.; Studdert, D.M.; Andrews, J.R.; Goldhaber-Fiebert, J.D.; Group, S.-C.M. The household secondary attack rate of SARS-CoV-2: A rapid review. Clin. Infect. Dis 2020, 73 (Suppl. 2), S138–S145. [Google Scholar] [CrossRef] [PubMed]
- McLean, H.Q.; Grijalva, C.G.; Hanson, K.E.; Zhu, Y.; Deyoe, J.E.; Meece, J.K.; Halasa, N.B.; Chappell, J.D.; Mellis, A.; Reed, C.; et al. Household Transmission and Clinical Features of SARS-CoV-2 Infections by Age in 2 US Communities. medRxiv 2021. [Google Scholar] [CrossRef]
- Kuwelker, K.; Zhou, F.; Blomberg, B.; Lartey, S.; Brokstad, K.A.; Trieu, M.C.; Bansal, A.; Madsen, A.; Krammer, F.; Mohn, K.G.; et al. Attack rates amongst household members of outpatients with confirmed COVID-19 in Bergen, Norway: A case-ascertained study. Lancet Reg. Health Eur. 2021, 3, 100014. [Google Scholar] [CrossRef]
- Bernal, J.L.; Panagiotopoulos, N.; Byers, C.; Garcia Vilaplana, T.; Boddington, N.; Zhang, X.-S.; Charlett, A.; Elgohari, S.; Coughlan, L.; Whillock, R.; et al. Transmission dynamics of COVID-19 in household and community settings in the United Kingdom. medRxiv 2020. [Google Scholar] [CrossRef]
- Grijalva, C.G.; Rolfes, M.A.; Zhu, Y.; McLean, H.Q.; Hanson, K.E.; Belongia, E.A.; Halasa, N.B.; Kim, A.; Reed, C.; Fry, A.M.; et al. Transmission of SARS-COV-2 Infections in Households-Tennessee and Wisconsin, April-September 2020. Morb. Mortal. Wkly. Rep. 2020, 69, 1631–1634. [Google Scholar] [CrossRef] [PubMed]
- Reukers, D.F.M.; van Boven, M.; Meijer, A.; Rots, N.; Reusken, C.; Roof, I.; van Gageldonk-Lafeber, A.B.; van der Hoek, W.; van den Hof, S. High infection secondary attack rates of SARS-CoV-2 in Dutch households revealed by dense sampling. Clin. Infect. Dis. 2021, ciab237. [Google Scholar] [CrossRef] [PubMed]
- Telle, K.; Jørgensen, S.B.; Hart, R.; Greve-Isdahl, M.; Kacelnik, O. Secondary attack rates of COVID-19 in Norwegian families: A nation-wide register-based study. Eur. J. Epidemiol. 2021, 36, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Buchan, S.A.; Tibebu, S.; Daneman, N.; Whelan, M.; Vanniyasingam, T.; Murti, M.; Brown, K.A. Increased household secondary attacks rates with Variant of Concern SARS-CoV-2 index cases. Clin. Infect. Dis. 2021, ciab496. [Google Scholar] [CrossRef]
- Lindstrøm, J.C.; Engebretsen, S.; Kristoffersen, A.B.; Rø, G.Ø.I.; Palomares, A.D.-L.; Engø-Monsen, K.; Madslien, E.H.; Forland, F.; Nygård, K.M.; Hagen, F.; et al. Increased transmissibility of the alpha SARS-CoV-2 variant: Evidence from contact tracing data in Oslo, January to February 2021. Infect. Dis. 2021, 1–6. [Google Scholar] [CrossRef]
- Geismar, C.; Fragaszy, E.; Nguyen, V.; Fong, W.L.E.; Shrotri, M.; Beale, S.; Rogers, A.; Lampos, V.; Byrne, T.; Kovar, J.; et al. Serial interval of COVID-19 and the effect of Variant B.1.1.7: Analyses from a prospective community cohort study (Virus Watch). medRxiv 2021. [Google Scholar] [CrossRef]
- Miller, E.; Waight, P.A.; Andrews, N.J.; McOwat, K.; Brown, K.E.; Katja, H.; Ijaz, S.; Letley, L.; Haskins, D.; Sinnathamby, M.; et al. Transmission of SARS-CoV-2 in the household setting: A prospective cohort study in children and adults in England. J. Infect. 2021, 83, 483–489. [Google Scholar] [CrossRef]
- Spielberger, B.D.; Goerne, T.; Geweniger, A.; Henneke, P.; Elling, R. Intra-Household and Close-Contact SARS-CoV-2 Transmission among Children—A Systematic Review. Front. Pediatr. 2021, 9, 613292. [Google Scholar] [CrossRef]
- Ng, O.T.; Marimuthu, K.; Koh, V.; Pang, J.; Linn, K.Z.; Sun, J.; De Wang, L.; Chia, W.N.; Tiu, C.; Chan, M.; et al. SARS-CoV-2 seroprevalence and transmission risk factors among high-risk close contacts: A retrospective cohort study. Lancet Infect. Dis. 2021, 21, 333–343. [Google Scholar] [CrossRef]
- Goldstein, E.; Lipsitch, M.; Cevik, M. On the Effect of Age on the Transmission of SARS-CoV-2 in Households, Schools, and the Community. J. Infect. Dis. 2021, 223, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Singanayagam, A.; Hakki, S.; Dunning, J.; Madon, K.J.; Crone, M.A.; Koycheva, A.; Derqui-Fernandez, N.; Barnett, J.L.; Whitfield, M.G.; Varro, R.; et al. Community transmission and viral load kinetics of the SARS-CoV-2 delta (B.1.617.2) variant in vaccinated and unvaccinated individuals in the UK: A prospective, longitudinal, cohort study. Lancet Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Maltezou, H.C.; Vorou, R.; Papadima, K.; Kossyvakis, A.; Spanakis, N.; Gioula, G.; Exindari, M.; Metallidis, S.; Lourida, A.N.; Raftopoulos, V.; et al. Transmission dynamics of SARS-CoV-2 within families with children in Greece: A study of 23 clusters. J. Med. Virol. 2021, 93, 1414–1420. [Google Scholar] [CrossRef]
- Yan, D.; Zhang, X.; Chen, C.; Jiang, D.; Liu, X.; Zhou, Y.; Huang, C.; Zhou, Y.; Guan, Z.; Ding, C.; et al. Characteristics of Viral Shedding Time in SARS-CoV-2 Infections: A Systematic Review and Meta-Analysis. Front. Public Health 2021, 9, 652842. [Google Scholar] [CrossRef]
- Cendejas-Bueno, E.; Romero-Gómez, M.P.; Escosa-García, L.; Jiménez-Rodríguez, S.; Mingorance, J.; García-Rodríguez, J. Lower nasopharyngeal viral loads in pediatric population. The missing piece to understand SARS-CoV-2 infection in children? J. Infect. 2021, 83, e18–e19. [Google Scholar] [CrossRef]
- Madera, S.; Crawford, E.; Langelier, C.; Tran, N.K.; Thornborrow, E.; Miller, S.; DeRisi, J.L. Nasopharyngeal SARS-CoV-2 viral loads in young children do not differ significantly from those in older children and adults. Sci. Rep. 2021, 11, 3044. [Google Scholar] [CrossRef]
- Tong, J.Y.; Wong, A.; Zhu, D.; Fastenberg, J.H.; Tham, T. The Prevalence of Olfactory and Gustatory Dysfunction in COVID-19 Patients: A Systematic Review and Meta-analysis. Otolaryngol. Head Neck Surg. 2020, 163, 3–11. [Google Scholar] [CrossRef]
- Jain, A.; Pandey, A.K.; Kaur, J.; Kumar, L.; Singh, M.; Das, S.; Purohit, S. Is there a correlation between viral load and olfactory & taste dysfunction in COVID-19 patients? Am. J. Otolaryngol. 2021, 42, 102911. [Google Scholar] [CrossRef]
- Nakagawara, K.; Masaki, K.; Uwamino, Y.; Kabata, H.; Uchida, S.; Uno, S.; Asakura, T.; Funakoshi, T.; Kanzaki, S.; Ishii, M.; et al. Acute onset olfactory/taste disorders are associated with a high viral burden in mild or asymptomatic SARS-CoV-2 infections. Int. J. Infect. Dis. 2020, 99, 19–22. [Google Scholar] [CrossRef]

| Participant Characteristics a | All Participants | All Households (HH) | Alpha Variant Households | Non-VOC Virus Households | |||
|---|---|---|---|---|---|---|---|
| (N = 200) | Primary Cases (N = 65) | HH Contacts (N = 135) | Primary Cases (N = 18) | HH Contacts (N = 36) | Primary Cases (N = 40) | HH Contacts (N = 87) | |
| Age (years) | |||||||
| Median (range) | 31 (2–73) | 38 (2–71) b | 24 (2–73) | 38 (2–68) b | 22 (2–63) | 38 (15–68) | 24 (2–70) |
| (IQR) | (14.5–42) | (31–45) | (8–38) | (28–43) | (8.5–38.5) | (30.5–48) | (9–40) |
| 2–17, n (%) | 63 (31.5) | 3 (4.6) | 60 (44.4) | 2 (11.1) | 16 (44.4) | 1 (2.5) | 39 (44.8) |
| ≥18, n (%) | 137 (68.8) | 62 (95.4) | 75 (55.6) | 16 (88.9) | 20 (55.6) | 39 (97.5) | 48 (55.2) |
| Sex | |||||||
| Female, n (%) | 104 (52) | 35 (53.9) | 69 (51.1) | 10 (55.6) | 16 (44.4) | 19 (47.5) | 49 (56.3) |
| Male, n (%) | 96 (48) | 30 (46.1) | 66 (48.9) | 8 (44.4) | 20 (55.6) | 21 (52.5) | 38 (43.7) |
| Ethnicity c | |||||||
| Nordic, n (%) | 101 (50.5) | ||||||
| Part Nordic, n (%) | 24 (12) | ||||||
| Other, n (%) | 7 (3.5) | ||||||
| missing, n (%) | 68 (34) | ||||||
| Chronic illness d | |||||||
| yes, n (%) | 33 (16.5) | 13 (20.0) | 20 (14.8) | 4 (22.2) | 4 (11.1) | 7 (17.5) | 14 (16.1) |
| no, n (%) | 165 (82.5) | 52 (80.0) | 113 (83.7) | 14 (77.8) | 32 (88.9) | 33 (82.5) | 71 (81.6) |
| missing, n (%) | 2 (1.0) | 2 (1.5) | 0 (0) | 2 (2.3) | |||
| Profession (age ≥ 16 yrs) | |||||||
| Healthcare, n (%) | 28 (19.3) | 13 (20.0) | 15 (18.1) | 9 (23.1) | 10 (18.5) | 3 (18.8) | 3 (13.6) |
| Other, n (%) | 117 (80.7) | 49 (79.0) | 68 (91.9) | 30 (76.9) | 44 (81.5) | 13 (81.3) | 19 (86.4) |
| Household Characteristics | All Households | Alpha Variant Households | Non-VOC Virus Households | ||||
| Household size | |||||||
| 2 persons, n (%) | 24 (36.9) | 6 (33.3) | 15 (37.5) | ||||
| 3 persons, n (%) | 9 (13.9) | 11 (61.1) | 9 (22.5) | ||||
| 4 persons, n (%) | 22 (33.9) | 1 (5.6) | 7 (17.5) | ||||
| 5–6 persons, n (%) | 10 (15.3) | 0 (0) | 9 (22.5) | ||||
| Young children e | |||||||
| yes, n (%) | 28 (43.1) | 9 (50) | 16 (40) | ||||
| no, n (%) | 37 (56.9) | 9 (50) | 24 (60) | ||||
| Households a with Transmission | % with Transmission, 95% CI | PCR+ (n)/(N) | p-Value b | Crude OR, 95% CI | p-Value | ||
|---|---|---|---|---|---|---|---|
| All variants | 60.0 (47.4–71.4) | 39/65 | |||||
| Non-VOC viruses | 55.0 (39.8–70.1) | 22/40 | 1 (Ref) | ||||
| Alpha variant | 83.3 (55.9–95.2) | 15/18 | 0.04 | 4.24 (0–4.2 × 1032) | 0.04 | ||
| Household Contact c | SAR %, 95% CI | PCR+ (n)/(N) | p-Value b | Crude OR, 95% CI | p-Value | Adjusted OR d, 95% CI | p-Value |
| All variants | 49.6 (37.8–61.5) | 67/135 | |||||
| Non-VOC viruses | 42.5 (28.7–57.7) | 37/87 | 1 (Ref) | 1 (Ref) | |||
| Alpha variant | 77.8 (49.4–92.6) | 28/36 | 0.02 | 65.7 (1.74–2481) | 0.02 | 468 (1.8–1.2 × 105) | 0.03 |
| Characteristic | SAR %, (95% CI) | p-Value a | PCR+ (n)/Total (N) | Crude OR (95% CI) | p-Value | Adjusted OR b (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|
| PRIMARY CASE CHARACTERISTICS | |||||||
| Age (yrs) | |||||||
| 12–39 c | 47 (31–64) | 33/70 | 1 (Ref) | 1 (Ref) | |||
| ≥40 | 52 (35–69) | 0.67 | 34/65 | 2.30 (0.25–21.1) | 0.46 | 1.6 (0.15–16.9) | 0.70 |
| Sex | |||||||
| Female | 46 (30–63) | 29/63 | 1 (Ref) | 1 (Ref) | |||
| Male | 53 (35–70) | 0.58 | 38/72 | 1.51 (0.18–12.7) | 0.71 | 1.7 (0.15–18.7) | 0.68 |
| HOUSEHOLD CONTACT CHARACTERISTICS | |||||||
| Age (yrs) | |||||||
| 2–17 yr | 47 (31–63) | 28/60 | 1 (Ref) | 1 (Ref) | |||
| 18–39 | 40 (25–56) | 17/43 | 0.31 (0.05–2.04) | 0.23 | 0.18 (0.02–1.33) | 0.09 | |
| ≥40 | 69 (49–83) | 0.03 | 22/32 | 8.03 (1.15–56.2) | 0.04 | 7.53 (1.07–52.8) | 0.04 |
| Sex | |||||||
| Female | 52 (38–66) | 36/69 | 1 (Ref) | 1 (Ref) | |||
| Male | 47 (32–62) | 0.55 | 31/66 | 0.81 (0.22–3.01) | 0.76 | 0.97 (0.24–3.91) | 0.96 |
| Blood type (≥18 yrs) | |||||||
| O | 48 (27–69) | 10/21 | 1 (Ref) | 1 (Ref) | |||
| A | 56 (37–73) | 18/32 | 1.45 (0.38–5.5) | 0.59 | 1.41 (0.40–4.96) | 0.59 | |
| AB | 33 (0–100) | 1/3 | 0.49 (0.02–10.3) | 0.64 | 0.40 (0.02–6.47) | 0.52 | |
| B | 71 (22–96) | 0.61 | 5/7 | 3.03 (0.32–28.3) | 0.33 | 3.02 (0.41–22.5) | 0.28 |
| HOUSEHOLD CHARACTERISTICS | |||||||
| Household size | |||||||
| 2 pers | 54 (33–74) | 13/24 | 1 (Ref) | 1 (Ref) | |||
| 3 pers | 59 (27–84) | 10/17 | 1.72 (0.06–53.0) | 0.76 | 2.0 (0.05–88) | 0.71 | |
| 4 pers | 47 (28–68) | 28/59 | 0.48 (0.03–7.72) | 0.61 | 0.8 (0.04–17) | 0.88 | |
| 5–6 pers | 46 (20–74) | 0.87 | 16/35 | 0.48 (0.15–18.6) | 0.68 | 0.7 (0.02–29) | 0.84 |
| Overcrowding d | |||||||
| No | 52 (37–66) | 47/91 | 1 (Ref) | 1 (Ref) | |||
| Yes | 90 (24–100) | 0.01 | 9/10 | 122.7 (0.16–94,464) | 0.16 | 480.9 (0.11–2 × 106) | 0.15 |
| Number of bathrooms e | |||||||
| 1 | 58 (39–75) | 36/62 | 1 (Ref) | 1 (Ref) | |||
| ≥2 | 44 (27–61) | 0.25 | 27/62 | 0.2 (0.02–2.7) | 0.25 | 0.1 (0.01–2.2) | 0.17 |
| SAR % (95% CI) | PCR+ (n)/ Total (N) | p-Value a | Crude OR (95%CI) | p-Value | Adjusted b OR (95%CI) | p-Value | |
|---|---|---|---|---|---|---|---|
| Severity | |||||||
| Asymptomatic | 33 (3–88) | 4/12 | 1 (Ref) | 1 (Ref) | |||
| Mild | 47 (27–68) | 25/53 | 10.2 (0.17–613) | 0.27 | 8.7 (0.13–594) | 0.31 | |
| Moderate | 54 (39–69) | 38/70 | 0.54 | 12.1 (0.22–672) | 0.22 | 11.8 (0.14–974) | 0.28 |
| Loss of taste/smell | |||||||
| No | 27 (13–47) | 12/44 | 1 (Ref) | 1 (Ref) | |||
| Yes | 60 (45–74) | 55/91 | <0.01 | 29.5 (1.33–654) | 0.03 | 68.3 (1.95–2389) | 0.02 |
| Fever | |||||||
| No | 39 (24–57) | 27/69 | 1 (Ref) | 1 (Ref) | |||
| Yes | 61 (43–76) | 40/66 | 0.08 | 10.3 (0.78–136) | 0.08 | 10.4 (0.76–140) | 0.08 |
| Cough | |||||||
| No | 27 (11–53) | 7/26 | 1 (Ref) | 1 (Ref) | |||
| Yes | 55 (41–68) | 60/109 | 0.04 | 10.0 (0.54–184) | 0.12 | 10.2 (0.51–203) | 0.13 |
| Dyspnea | |||||||
| No | 46 (29–64) | 31/68 | 1 (Ref) | 1 (Ref) | |||
| Yes | 54 (38–69) | 36/67 | 0.50 | 1.40 (0.17–12) | 0.76 | 0.97 (0.1–9.51) | 0.98 |
| SAR% (95% CI) | PCR+/ HH Contacts | p-Value a | Crude OR (95% CI) | p-Value | Adjusted OR b (95% CI) | p-Value | ||
|---|---|---|---|---|---|---|---|---|
| Behavioral factors: contact with the primary case prior to confirmation of infection | ||||||||
| Cared for | No | 48 (35–61) | 51/106 | 1 (Ref) | 1 (Ref) | |||
| Yes | 53 (27–78) | 8/15 | 0.70 | 1.14 (0.31–2.74) | 0.91 | 0.54 (0.04–6.79) | 0.63 | |
| Hugged | No | 39 (21–61) | 11/28 | 1 (Ref) | 1 (Ref) | |||
| Yes | 52 (37–66) | 48/93 | 0.28 | 3.26 (0.54–9.60) | 0.20 | 3.90 (0.55–27.7) | 0.17 | |
| Kissed | No | 44 (29–60) | 26/59 | 1 (Ref) | 1 (Ref) | |||
| Yes | 53 (36–70) | 33/62 | 0.41 | 7.14 (0.86–9.10) | 0.07 | 5.27 (0.63–44.0) | 0.13 | |
| Shook/ held hands | No | 47 (30–65) | 16/34 | 1 (Ref) | 1 (Ref) | |||
| Yes | 49 (36–63) | 43/87 | 0.80 | 3.51 (0.56–22.02) | 0.18 | 4.30 (0.52–35.5) | 0.18 | |
| Ate together | No | 50 (25–75) | 6/12 | 1 (Ref) | 1 (Ref) | |||
| Yes | 49 (36–62) | 53/109 | 0.91 | 1.58 (0.20–2.78) | 0.67 | 1.72 (0.19–15.1) | 0.63 | |
| Shared a cup/glass/bottle | No | 49 (36–62) | 50/102 | 1 (Ref) | 1 (Ref) | |||
| Yes | 47 (20–77) | 9/19 | 0.92 | 0.85 (0.10–0.25) | 0.88 | 0.60 (0.06–5.64) | 0.65 | |
| Slept in the same room | No | 43 (26–61) | 25/58 | 1 (Ref) | 1 (Ref) | |||
| Yes | 54 (39–68) | 34/63 | 0.30 | 3.40 (0.79–4.66) | 0.10 | 2.58 (0.53–12.5) | 0.24 | |
| Shared a bed | No | 42 (27–60) | 25/59 | 1 (Ref) | 1 (Ref) | |||
| Yes | 55 (40–69) | 34/62 | 0.19 | 2.93 (0.68–2.54) | 0.15 | 2.18 (0.45–10.5) | 0.34 | |
| Shared a toilet | No | 20 (2–71) | 2/10 | 1 (Ref) | 1 (Ref) | |||
| Yes | 51 (38–65) | 57/111 | 0.10 | 117 (0.5–26,964) | 0.085 | 125 (0.5–29,152) | 0.08 | |
| Precautionary practices: performed by primary cases after confirmation of infection | ||||||||
| Isolated c | No | 67 (40–86) | 33/22 | 1 (Ref) | 1 (Ref) | |||
| Yes | 42 (28–57) | 37/88 | 0.10 | 0.07 (0.00–1.15) | 0.06 | 0.06 (0.00–1.19) | 0.07 | |
| Social distanced (≥2 m) | No | 59 (40–76) | 35/59 | 1 (Ref) | 1 (Ref) | |||
| Yes | 39 (23–57) | 24/62 | 0.11 | 0.11 (0.01–1.31) | 0.08 | 0.10 (0.01–1.32) | 0.08 | |
| Used face mask | No | 55 (38–70) | 46/84 | 1 (Ref) | 1 (Ref) | |||
| Yes | 35 (19–56) | 13/37 | 0.12 | 0.13 (0.01–1.62) | 0.11 | 0.12 (0.01–1.51) | 0.10 | |
| Slept in a different room | No | 67 (44–85) | 29/43 | 1 (Ref) | 1 (Ref) | |||
| Yes | 38 (25–54) | 30/78 | 0.03 | 0.08 (0.01–0.98) | 0.048 | 0.07 (0.00–0.98) | 0.048 | |
| Used separate bathroom/toilet | No | 53 (38–68) | 41/77 | 1 (Ref) | 1 (Ref) | |||
| Yes | 41 (21–64) | 18/44 | 0.35 | 0.30 (0.03–3.00) | 0.31 | 0.30 (0.02–3.63) | 0.34 | |
| Did not share a towel/items d | No | 69 (25–94) | 11/16 | NA | NA | |||
| Yes | 49 (32–67) | 30/61 | 0.34 | |||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Julin, C.H.; Robertson, A.H.; Hungnes, O.; Tunheim, G.; Bekkevold, T.; Laake, I.; Aune, I.F.; Killengreen, M.F.; Strand, T.R.; Rykkvin, R.; et al. Household Transmission of SARS-CoV-2: A Prospective Longitudinal Study Showing Higher Viral Load and Increased Transmissibility of the Alpha Variant Compared to Previous Strains. Microorganisms 2021, 9, 2371. https://doi.org/10.3390/microorganisms9112371
Julin CH, Robertson AH, Hungnes O, Tunheim G, Bekkevold T, Laake I, Aune IF, Killengreen MF, Strand TR, Rykkvin R, et al. Household Transmission of SARS-CoV-2: A Prospective Longitudinal Study Showing Higher Viral Load and Increased Transmissibility of the Alpha Variant Compared to Previous Strains. Microorganisms. 2021; 9(11):2371. https://doi.org/10.3390/microorganisms9112371
Chicago/Turabian StyleJulin, Cathinka Halle, Anna Hayman Robertson, Olav Hungnes, Gro Tunheim, Terese Bekkevold, Ida Laake, Idunn Forland Aune, Marit Fodnes Killengreen, Torunn Ramsem Strand, Rikard Rykkvin, and et al. 2021. "Household Transmission of SARS-CoV-2: A Prospective Longitudinal Study Showing Higher Viral Load and Increased Transmissibility of the Alpha Variant Compared to Previous Strains" Microorganisms 9, no. 11: 2371. https://doi.org/10.3390/microorganisms9112371
APA StyleJulin, C. H., Robertson, A. H., Hungnes, O., Tunheim, G., Bekkevold, T., Laake, I., Aune, I. F., Killengreen, M. F., Strand, T. R., Rykkvin, R., Dorenberg, D. H., Stene-Johansen, K., Berg, E. S., Bodin, J. E., Oftung, F., Steens, A., & Næss, L. M. (2021). Household Transmission of SARS-CoV-2: A Prospective Longitudinal Study Showing Higher Viral Load and Increased Transmissibility of the Alpha Variant Compared to Previous Strains. Microorganisms, 9(11), 2371. https://doi.org/10.3390/microorganisms9112371






