Cell Plasticity and Genomic Structure of a Novel Filterable Rhizobiales Bacterium that Belongs to a Widely Distributed Lineage
Abstract
1. Introduction
2. Materials and Methods
2.1. Test for Cell Morphological Change and Filterability
2.2. Genomic Sequencing, Assembly, and Annotation
2.3. Phylogenetic and Comparative Genomic Analyses
3. Results and Discussion
3.1. Temperature-Induced Cell Morphological Change
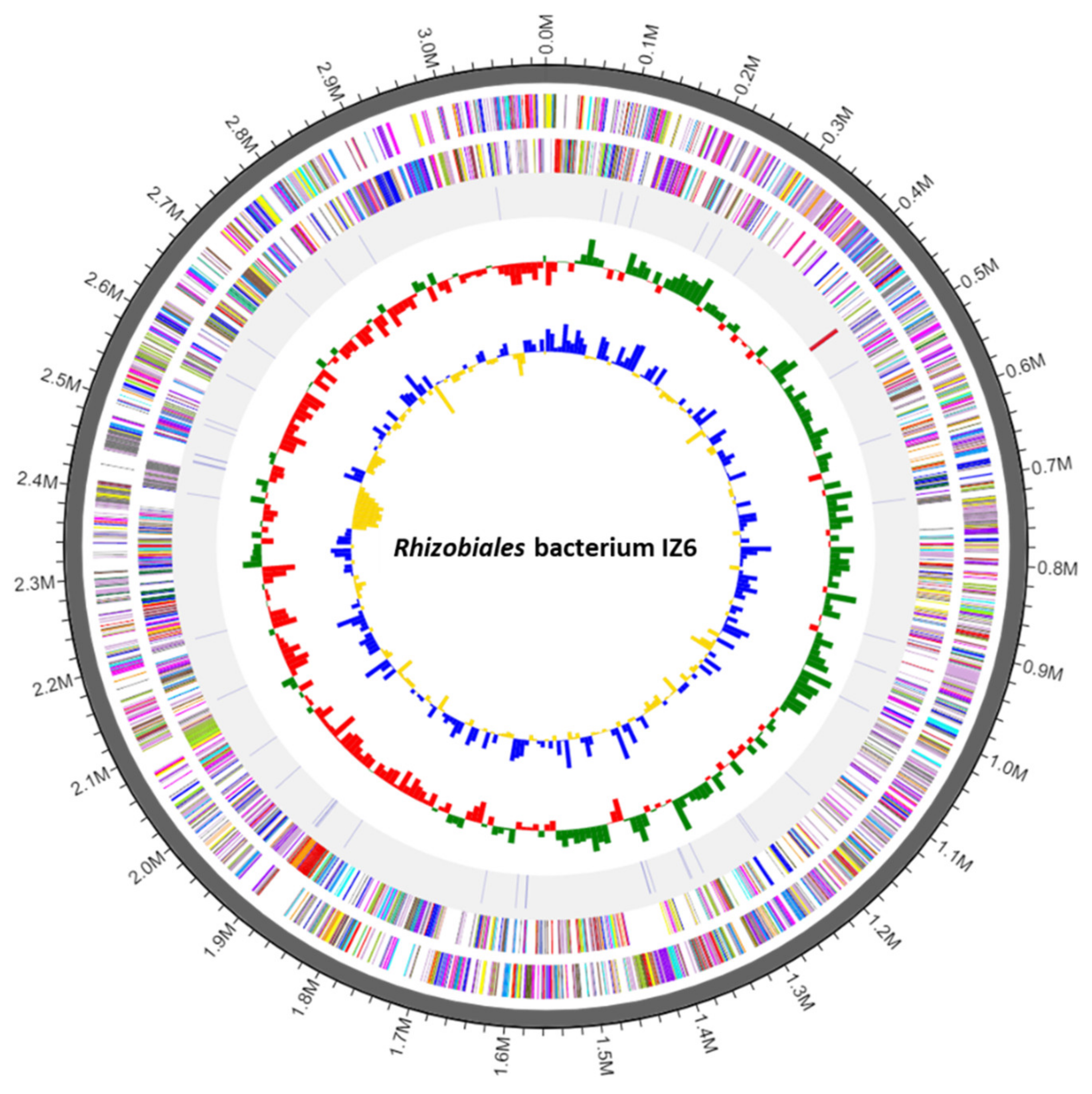
3.2. Genome Structure and Phylogenetic Relationships
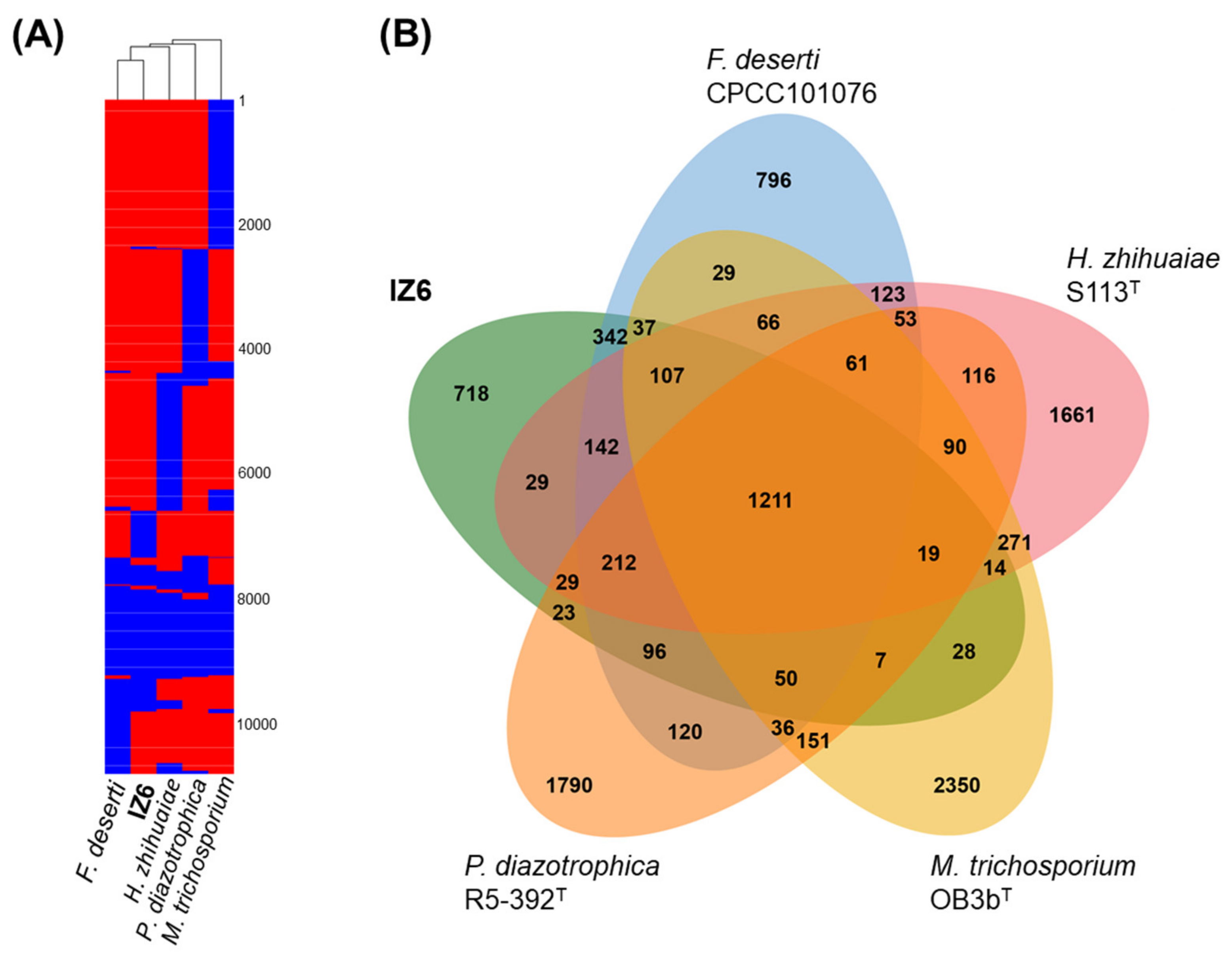
3.3. Ecophysiological Characteristics and Potential Inferred from Comparative Genomics
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Haller, C.M.; Rölleke, S.; Vybiral, D.; Witte, A.; Velimirov, B. Investigation of 0.2 μm filterable bacteria from the Western Mediterranean Sea using a molecular approach: Dominance of potential starvation forms. FEMS Microbiol. Ecol. 2000, 31, 153–161. [Google Scholar] [CrossRef]
- Lannes, R.; Olsson-Francis, K.; Lopez, P.; Bapteste, E. Carbon fixation by marine ultrasmall prokaryotes. Genome Biol. Evol. 2019, 11, 1166–1177. [Google Scholar] [CrossRef]
- Fedotova, A.V.; Belova, S.E.; Kulichevskaya, I.S.; Dedysh, S.N. Molecular identification of filterable bacteria and archaea in the water of acidic lakes of northern Russia. Microbiology 2012, 81, 281–287. [Google Scholar] [CrossRef]
- Vigneron, A.; Cruaud, P.; Langlois, V.; Lovejoy, C.; Culley, A.I.; Vincent, W.F. Ultra-small and abundant: Candidate phyla radiation bacteria are potential catalysts of carbon transformation in a thermokarst lake ecosystem. Limnol. Oceanogr. Lett. 2020, 5, 212–220. [Google Scholar] [CrossRef]
- Naganuma, T.; Miyoshi, T.; Kimura, H. Phylotype diversity of deep-sea hydrothermal vent prokaryotes trapped by 0.2- and 0.1-μm-pore-size filters. Extremophiles 2007, 11, 637–646. [Google Scholar] [CrossRef] [PubMed]
- Nakai, R.; Abe, T.; Takeyama, H.; Naganuma, T. Metagenomic analysis of 0.2-μm-passable microorganisms in deep-sea hydrothermal fluid. Mar. Biotechnol. 2011, 13, 900–908. [Google Scholar] [CrossRef]
- Miyoshi, T.; Iwatsuki, T.; Naganuma, T. Phylogenetic characterization of 16S rRNA gene clones from deep-groundwater microorganisms that pass through 0.2-micrometer-pore-size filters. Appl. Environ. Microbiol. 2005, 71, 1084–1088. [Google Scholar] [CrossRef]
- Luef, B.; Frischkorn, K.R.; Wrighton, K.C.; Holman, H.-Y.N.; Birarda, G.; Thomas, B.C.; Singh, A.; Williams, K.H.; Siegerist, C.E.; Tringe, S.G.; et al. Diverse uncultivated ultra-small bacterial cells in groundwater. Nat. Commun. 2015, 6, 6372. [Google Scholar] [CrossRef]
- Hahn, M.W.; Lünsdorf, H.; Wu, Q.; Schauer, M.; Höfle, M.G.; Boenigk, J.; Stadler, P. Isolation of novel ultramicrobacteria classified as Actinobacteria from five freshwater habitats in Europe and Asia. Appl. Environ. Microbiol. 2003, 69, 1442–1451. [Google Scholar] [CrossRef]
- Geissinger, O.; Herlemann, D.P.R.; Mörschel, E.; Maier, U.G.; Brune, A. The ultramicrobacterium “Elusimicrobium minutum” gen. nov., sp. nov., the first cultivated representative of the termite group 1 phylum. Appl. Environ. Microbiol. 2009, 75, 2831–2840. [Google Scholar] [CrossRef]
- Hahn, M.W.; Stadler, P.; Wu, Q.L.; Pöckl, M. The filtration–acclimatization method for isolation of an important fraction of the not readily cultivable bacteria. J. Microbiol. Methods 2004, 57, 379–390. [Google Scholar] [CrossRef] [PubMed]
- Duda, V.I.; Suzina, N.E.; Polivtseva, V.N.; Boronin, A.M. Ultramicrobacteria: Formation of the concept and contribution of ultramicrobacteria to biology. Microbiology 2012, 81, 379–390. [Google Scholar] [CrossRef]
- Nakai, R. Size matters: Ultra-small and filterable microorganisms in the environment. Microbes Environ. 2020, 35. [Google Scholar] [CrossRef] [PubMed]
- Nakai, R.; Shibuya, E.; Justel, A.; Rico, E.; Quesada, A.; Kobayashi, F.; Iwasaka, Y.; Shi, G.-Y.; Amano, Y.; Iwatsuki, T.; et al. Phylogeographic analysis of filterable bacteria with special reference to Rhizobiales strains that occur in cryospheric habitats. Antarct. Sci. 2013, 25, 219–228. [Google Scholar] [CrossRef]
- Andersson, A.; Larsson, U.; Hagström, Å. Size-selective grazing by a microflagellate on pelagic bacteria. Mar. Ecol. Prog. Ser. 1986, 33, 51–57. [Google Scholar] [CrossRef]
- Lee, I.; Chalita, M.; Ha, S.-M.; Na, S.-I.; Yoon, S.-H.; Chun, J. ContEst16S: An algorithm that identifies contaminated prokaryotic genomes using 16S RNA gene sequences. Int. J. Syst. Evol. Microbiol. 2017, 67, 2053–2057. [Google Scholar] [CrossRef]
- Yoon, S.-H.; Ha, S.-M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar] [CrossRef]
- Hyatt, D.; Chen, G.-L.; LoCascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 119. [Google Scholar] [CrossRef]
- Schattner, P.; Brooks, A.N.; Lowe, T.M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res. 2005, 33, W686–W689. [Google Scholar] [CrossRef]
- Nawrocki, E.P.; Burge, S.W.; Bateman, A.; Daub, J.; Eberhardt, R.Y.; Eddy, S.R.; Floden, E.W.; Gardner, P.P.; Jones, T.A.; Tate, J.; et al. Rfam 12.0: Updates to the RNA families database. Nucleic Acids Res. 2014, 43, D130–D137. [Google Scholar] [CrossRef]
- Edgar, R.C. PILER-CR: Fast and accurate identification of CRISPR repeats. BMC Bioinform. 2007, 8, 18. [Google Scholar] [CrossRef] [PubMed]
- Bland, C.; Ramsey, T.L.; Sabree, F.; Lowe, M.; Brown, K.; Kyrpides, N.C.; Hugenholtz, P. CRISPR Recognition Tool (CRT): A tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinform. 2007, 8, 209. [Google Scholar] [CrossRef] [PubMed]
- Powell, S.; Forslund, K.; Szklarczyk, D.; Trachana, K.; Roth, A.; Huerta-Cepas, J.; Gabaldón, T.; Rattei, T.; Creevey, C.; Kuhn, M.; et al. eggNOG v4.0: Nested orthology inference across 3686 organisms. Nucleic Acids Res. 2013, 42, D231–D239. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium UniProt: A hub for protein information. Nucleic Acids Res. 2014, 43, D204–D212. [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. Data, information, knowledge and principle: Back to metabolism in KEGG. Nucleic Acids Res. 2013, 42, D199–D205. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2013, 42, D206–D214. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Tanizawa, Y.; Fujisawa, T.; Nakamura, Y. DFAST: A flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 2017, 34, 1037–1039. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Ouk Kim, Y.; Park, S.-C.; Chun, J. OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 2016, 66, 1100–1103. [Google Scholar] [CrossRef] [PubMed]
- Ward, N.; Moreno-Hagelsieb, G. Quickly finding orthologs as reciprocal best hits with BLAT, LAST, and UBLAST: How much do we miss? PLoS ONE 2014, 9, e101850. [Google Scholar] [CrossRef] [PubMed]
- Chun, J.; Grim, C.J.; Hasan, N.A.; Lee, J.H.; Choi, S.Y.; Haley, B.J.; Taviani, E.; Jeon, Y.-S.; Kim, D.W.; Lee, J.-H.; et al. Comparative genomics reveals mechanism for short-term and long-term clonal transitions in pandemic Vibrio cholerae. Proc. Natl. Acad. Sci. USA 2009, 106, 15442–15447. [Google Scholar] [CrossRef] [PubMed]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. jvenn: An interactive Venn diagram viewer. BMC Bioinform. 2014, 15, 293. [Google Scholar] [CrossRef]
- Watson, S.P.; Clements, M.O.; Foster, S.J. Characterization of the starvation-survival response of Staphylococcus aureus. J. Bacteriol. 1998, 180, 1750–1758. [Google Scholar] [CrossRef]
- Monier, J.-M.; Lindow, S.E. Pseudomonas syringae responds to the environment on leaves by cell size reduction. Phytopathology 2003, 93, 1209–1216. [Google Scholar] [CrossRef]
- Schaechter, M.; MaalØe, O.; Kjeldgaard, N.O. Dependency on medium and temperature of cell size and chemical composition during balanced growth of Salmonella typhimurium. Microbiology 1958, 19, 592–606. [Google Scholar] [CrossRef]
- Velimirov, B. Nanobacteria, ultramicrobacteria and starvation forms: A search for the smallest metabolizing bacterium. Microbes Environ. 2001, 16, 67–77. [Google Scholar] [CrossRef]
- Dong, L.; Han, M.-X.; Wang, D.; Liu, F.; Asem, M.D.; Jiao, J.-Y.; Xiao, M.; Salam, N.; Li, W.-J. Flaviflagellibacter deserti gen. nov., sp. nov., a novel member of the order Rhizobiales isolated from a desert soil. Antonie Van Leeuwenhoek 2019, 112, 947–954. [Google Scholar] [CrossRef] [PubMed]
- Field, E.K.; D’Imperio, S.; Miller, A.R.; VanEngelen, M.R.; Gerlach, R.; Lee, B.D.; Apel, W.A.; Peyton, B.M. Application of molecular techniques to elucidate the influence of cellulosic waste on the bacterial community structure at a simulated low-level-radioactive-waste site. Appl. Environ. Microbiol. 2010, 76, 3106–3115. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Williamson, K.E.; Kan, J.; Polson, S.W.; Williamson, S.J. Optimizing the indirect extraction of prokaryotic DNA from soils. Soil Biol. Biochem. 2011, 43, 736–748. [Google Scholar] [CrossRef]
- Webb, H.K.; Ng, H.J.; Ivanova, E.P. The family Methylocystaceae. In The Prokaryotes: Alphaproteobacteria and Betaproteobacteria; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin, Germany, 2014; pp. 341–347. ISBN 978-3-642-30197-1. [Google Scholar]
- Kim, M.; Oh, H.-S.; Park, S.-C.; Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 346–351. [Google Scholar] [CrossRef]
- Chen, I.-M.A.; Chu, K.; Palaniappan, K.; Pillay, M.; Ratner, A.; Huang, J.; Huntemann, M.; Varghese, N.; White, J.R.; Seshadri, R.; et al. IMG/M v.5.0: An integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 2019, 47, D666–D677. [Google Scholar] [CrossRef]

| Temperature | Cell Length × Width and Volume | |
|---|---|---|
| Before Filtration | After Filtration 1 | |
| 25 °C for 2 weeks | 1.5–2.0 × 0.4–0.5 µm | no cell |
| 0.17–0.36 µm3 | ||
| 25 °C for 2 weeks followed by 4 °C for 3 weeks | 2.0–2.5 × 0.4–0.5 µm | 1.0 × 0.2–0.3 µm |
| 0.23–0.46 µm3 | 0.03–0.06 µm3 | |
| 15 °C for 3 weeks | 1.5–2.0 × 0.3–0.5 µm | 0.5–1.0 × 0.2–0.3 µm |
| 0.10–0.36 µm3 | 0.01–0.06 µm3 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakai, R.; Naganuma, T.; Tazato, N.; Morohoshi, S.; Koide, T. Cell Plasticity and Genomic Structure of a Novel Filterable Rhizobiales Bacterium that Belongs to a Widely Distributed Lineage. Microorganisms 2020, 8, 1373. https://doi.org/10.3390/microorganisms8091373
Nakai R, Naganuma T, Tazato N, Morohoshi S, Koide T. Cell Plasticity and Genomic Structure of a Novel Filterable Rhizobiales Bacterium that Belongs to a Widely Distributed Lineage. Microorganisms. 2020; 8(9):1373. https://doi.org/10.3390/microorganisms8091373
Chicago/Turabian StyleNakai, Ryosuke, Takeshi Naganuma, Nozomi Tazato, Sho Morohoshi, and Tomomi Koide. 2020. "Cell Plasticity and Genomic Structure of a Novel Filterable Rhizobiales Bacterium that Belongs to a Widely Distributed Lineage" Microorganisms 8, no. 9: 1373. https://doi.org/10.3390/microorganisms8091373
APA StyleNakai, R., Naganuma, T., Tazato, N., Morohoshi, S., & Koide, T. (2020). Cell Plasticity and Genomic Structure of a Novel Filterable Rhizobiales Bacterium that Belongs to a Widely Distributed Lineage. Microorganisms, 8(9), 1373. https://doi.org/10.3390/microorganisms8091373






