Extrinsic Factors Shaping the Skin Microbiome
Abstract
1. Introduction
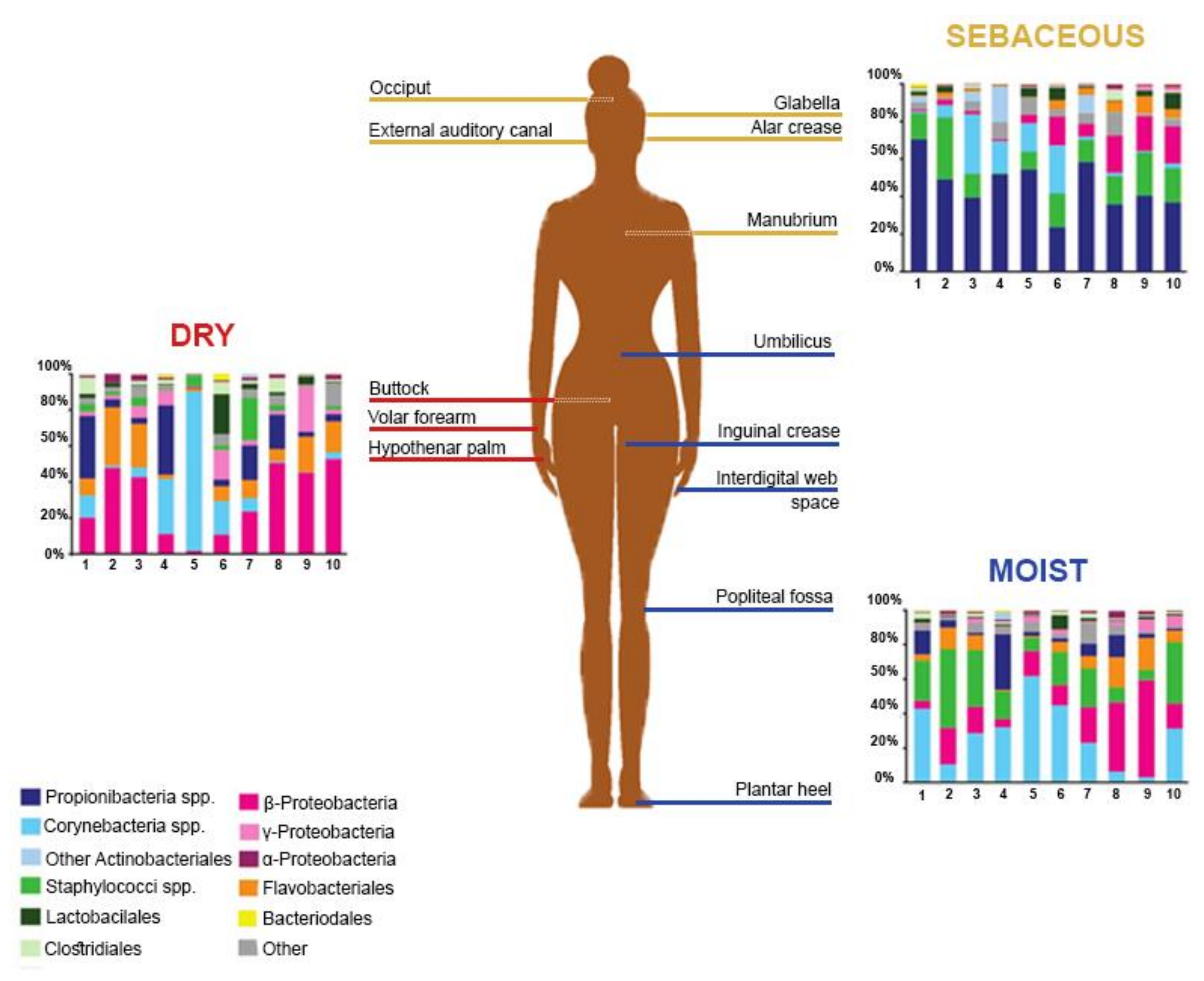
2. The Healthy Skin Microbiome
3. Microbiome Dysbiosis
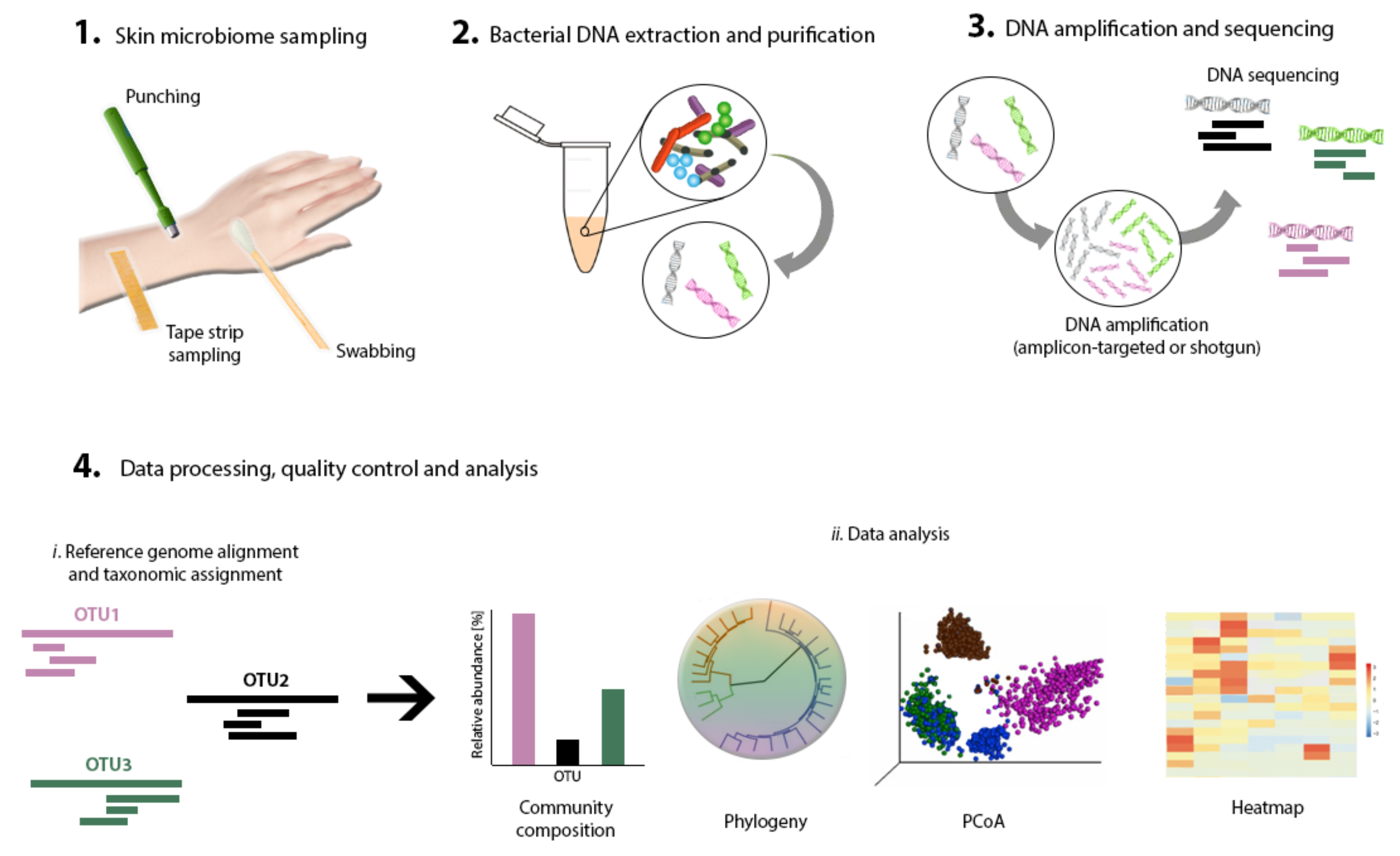
4. Strategies for Skin Microbiome Research
5. Extrinsic Influences Shaping the Skin Microbiome
5.1. Early Life Exposures
5.2. Cosmetics
5.3. Environment and Nature
5.4. Antimicrobial Agents
5.5. Bacterium-Based Agents
6. Future Perspectives
Funding
Acknowledgments
Conflicts of Interest
References
- Ursell, L.K.; Metcalf, J.L.; Parfrey, L.W.; Knight, R. Defining the human microbiome. Nutr. Rev. 2012, 70, S38–S44. [Google Scholar] [CrossRef] [PubMed]
- Marchesi, J.R.; Ravel, J. The Vocabulary of Microbiome Research: A Proposal; BioMed Central: London, UK, 2015. [Google Scholar]
- National Academies of Sciences, Engineering and Medicine. Environmental Chemicals, the Human Microbiome, and Health Risk: A Research Strategy; National Academies Press: Washington, DC, USA, 2018. [Google Scholar]
- Cho, I.; Blaser, M.J. The human microbiome: At the interface of health and disease. Nat. Rev. Genet. 2012, 13, 260. [Google Scholar] [CrossRef] [PubMed]
- Huttenhower, C.; Gevers, D.; Knight, R.; Abubucker, S.; Badger, J.H.; Chinwalla, A.T.; Creasy, H.H.; Earl, A.M.; FitzGerald, M.G.; Fulton, R.S. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207. [Google Scholar]
- Meisel, J.S.; Sfyroera, G.; Bartow-McKenney, C.; Gimblet, C.; Bugayev, J.; Horwinski, J.; Kim, B.; Brestoff, J.R.; Tyldsley, A.S.; Zheng, Q. Commensal microbiota modulate gene expression in the skin. Microbiome 2018, 6, 20. [Google Scholar] [CrossRef]
- Hooper, L.V.; Gordon, J.I. Commensal Host-Bacterial Relationships in the Gut. Science 2001, 292, 1115–1118. [Google Scholar] [CrossRef]
- Levy, M.; Kolodziejczyk, A.A.; Thaiss, C.A.; Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 2017, 17, 219. [Google Scholar] [CrossRef]
- Uhr, G.T.; Dohnalová, L.; Thaiss, C.A. The dimension of time in host-microbiome interactions. MSystems 2019, 4, e00216–e00218. [Google Scholar] [CrossRef]
- Capone, K.A.; Dowd, S.E.; Stamatas, G.N.; Nikolovski, J. Diversity of the human skin microbiome early in life. J. Investig. Dermatol. 2011, 131, 2026–2032. [Google Scholar] [CrossRef]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244. [Google Scholar] [CrossRef]
- Byrd, A.L.; Belkaid, Y.; Segre, J.A. The human skin microbiome. Nat. Rev. Microbiol. 2018, 16, 143. [Google Scholar] [CrossRef]
- Belkaid, Y.; Segre, J.A. Dialogue between skin microbiota and immunity. Science 2014, 346, 954–959. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.; Byrd, A.L.; Deming, C.; Conlan, S.; Barnabas, B.; Blakesley, R.; Bouffard, G.; Brooks, S.; Coleman, H.; Dekhtyar, M. Biogeography and individuality shape function in the human skin metagenome. Nature 2014, 514, 59. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; Green, E.D. Topographical and temporal diversity of the human skin microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef] [PubMed]
- Balato, A.; Cacciapuoti, S.; Di Caprio, R.; Marasca, C.; Masarà, A.; Raimondo, A.; Fabbrocini, G. Human microbiome: Composition and role in inflammatory skin diseases. Arch. Immunol. Et Ther. Exp. 2019, 67, 1–18. [Google Scholar] [CrossRef]
- Grice, E.A.; Kong, H.H.; Renaud, G.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Wolfsberg, T.G.; Turner, M.L.; Segre, J.A.; NISC Comparative Sequencing Program. A diversity profile of the human skin microbiota. Genome Res. 2008. [Google Scholar] [CrossRef]
- Perez, G.I.P.; Gao, Z.; Jourdain, R.; Ramirez, J.; Gany, F.; Clavaud, C.; Demaude, J.; Breton, L.; Blaser, M.J. Body site is a more determinant factor than human population diversity in the healthy skin microbiome. PLoS ONE 2016, 11, e0151990. [Google Scholar] [CrossRef]
- Gao, Z.; Tseng, C.-H.; Pei, Z.; Blaser, M.J. Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 2007, 104, 2927–2932. [Google Scholar] [CrossRef]
- Oh, J.; Conlan, S.; Polley, E.C.; Segre, J.A.; Kong, H.H. Shifts in human skin and nares microbiota of healthy children and adults. Genome Med. 2012, 4, 77. [Google Scholar] [CrossRef]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 2012, 22, 850–859. [Google Scholar] [CrossRef]
- Iwase, T.; Uehara, Y.; Shinji, H.; Tajima, A.; Seo, H.; Takada, K.; Agata, T.; Mizunoe, Y. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 2010, 465, 346. [Google Scholar] [CrossRef]
- Kaiser, C.; Franklin, O.; Dieckmann, U.; Richter, A. Microbial community dynamics alleviate stoichiometric constraints during litter decay. Ecol. Lett. 2014, 17, 680–690. [Google Scholar] [CrossRef]
- Foster, J.A.; Krone, S.M.; Forney, L.J. Application of ecological network theory to the human microbiome. Interdiscip. Perspect. Infect. Dis. 2008, 2008. [Google Scholar] [CrossRef] [PubMed]
- Prosser, J.I.; Bohannan, B.J.; Curtis, T.P.; Ellis, R.J.; Firestone, M.K.; Freckleton, R.P.; Green, J.L.; Green, L.E.; Killham, K.; Lennon, J.J. The role of ecological theory in microbial ecology. Nat. Rev. Microbiol. 2007, 5, 384–392. [Google Scholar] [CrossRef] [PubMed]
- Prescott, S.L.; Larcombe, D.-L.; Logan, A.C.; West, C.; Burks, W.; Caraballo, L.; Levin, M.; Van Etten, E.; Horwitz, P.; Kozyrskyj, A. The skin microbiome: Impact of modern environments on skin ecology, barrier integrity, and systemic immune programming. World Allergy Organ. J. 2017, 10, 29. [Google Scholar] [CrossRef] [PubMed]
- Ursell, L.K.; Clemente, J.C.; Rideout, J.R.; Gevers, D.; Caporaso, J.G.; Knight, R. The interpersonal and intrapersonal diversity of human-associated microbiota in key body sites. J. Allergy Clin. Immunol. 2012, 129, 1204–1208. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, C. Out of your skin. Nat. Biotechnol. 2020, 38, 392–397. [Google Scholar] [CrossRef]
- Niehues, H.; Bouwstra, J.A.; El Ghalbzouri, A.; Brandner, J.M.; Zeeuwen, P.L.; van den Bogaard, E.H. 3D skin models for 3R research: The potential of 3D reconstructed skin models to study skin barrier function. Exp. Dermatol. 2018, 27, 501–511. [Google Scholar] [CrossRef]
- Kim, J.; Hegde, M.; Jayaraman, A. Co-culture of epithelial cells and bacteria for investigating host–pathogen interactions. Lab A Chip 2010, 10, 43–50. [Google Scholar] [CrossRef]
- Fritz, J.V.; Desai, M.S.; Shah, P.; Schneider, J.G.; Wilmes, P. From meta-omics to causality: Experimental models for human microbiome research. Microbiome 2013, 1, 14. [Google Scholar] [CrossRef]
- Kong, H.H.; Andersson, B.; Clavel, T.; Common, J.E.; Jackson, S.A.; Olson, N.D.; Segre, J.A.; Traidl-Hoffmann, C. Performing skin microbiome research: A method to the madness. J. Investig. Dermatol. 2017, 137, 561–568. [Google Scholar] [CrossRef]
- Bjerre, R.D.; Hugerth, L.W.; Boulund, F.; Seifert, M.; Johansen, J.D.; Engstrand, L. Effects of sampling strategy and DNA extraction on human skin microbiome investigations. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chng, K.R.; Tay, A.S.L.; Li, C.; Ng, A.H.Q.; Wang, J.; Suri, B.K.; Matta, S.A.; McGovern, N.; Janela, B.; Wong, X.F.C.C. Whole metagenome profiling reveals skin microbiome-dependent susceptibility to atopic dermatitis flare. Nat. Microbiol. 2016, 1, 16106. [Google Scholar] [CrossRef] [PubMed]
- Grogan, M.D.; Bartow-McKenney, C.; Flowers, L.; Knight, S.A.; Uberoi, A.; Grice, E.A. Research Techniques Made Simple: Profiling the Skin Microbiota. J. Investig. Dermatol. 2019, 139, 747–752.e741. [Google Scholar] [CrossRef] [PubMed]
- Kuczynski, J.; Lauber, C.L.; Walters, W.A.; Parfrey, L.W.; Clemente, J.C.; Gevers, D.; Knight, R. Experimental and analytical tools for studying the human microbiome. Nat. Rev. Genet. 2012, 13, 47. [Google Scholar] [CrossRef] [PubMed]
- Videnska, P.; Smerkova, K.; Zwinsova, B.; Popovici, V.; Micenkova, L.; Sedlar, K.; Budinska, E. Stool sampling and DNA isolation kits affect DNA quality and bacterial composition following 16S rRNA gene sequencing using MiSeq Illumina platform. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Knudsen, B.E.; Bergmark, L.; Munk, P.; Lukjancenko, O.; Priemé, A.; Aarestrup, F.M.; Pamp, S.J. Impact of sample type and DNA isolation procedure on genomic inference of microbiome composition. MSystems 2016, 1, e00095-00016. [Google Scholar] [CrossRef]
- Wang, W.-L.; Xu, S.-Y.; Ren, Z.-G.; Tao, L.; Jiang, J.-W.; Zheng, S.-S. Application of metagenomics in the human gut microbiome. World J. Gastroenterol. Wjg 2015, 21, 803. [Google Scholar] [CrossRef]
- Rosenthal, M.; Goldberg, D.; Aiello, A.; Larson, E.; Foxman, B. Skin microbiota: Microbial community structure and its potential association with health and disease. Infect. Genet. Evol. 2011, 11, 839–848. [Google Scholar] [CrossRef]
- Malla, M.A.; Dubey, A.; Kumar, A.; Yadav, S.; Hashem, A.; Abd_Allah, E.F. Exploring the human microbiome: The potential future role of next-generation sequencing in disease diagnosis and treatment. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef]
- Sergaki, C.; Lagunas, B.; Lidbury, I.; Gifford, M.L.; Schäfer, P. Challenges and approaches in microbiome research: From fundamental to applied. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Ranjan, R.; Rani, A.; Metwally, A.; McGee, H.S.; Perkins, D.L. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 2016, 469, 967–977. [Google Scholar] [CrossRef]
- Chakravorty, S.; Helb, D.; Burday, M.; Connell, N.; Alland, D. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods 2007, 69, 330–339. [Google Scholar] [CrossRef] [PubMed]
- Castelino, M.; Eyre, S.; Moat, J.; Fox, G.; Martin, P.; Ho, P.; Upton, M.; Barton, A. Optimisation of methods for bacterial skin microbiome investigation: Primer selection and comparison of the 454 versus MiSeq platform. Bmc Microbiol. 2017, 17, 23. [Google Scholar] [CrossRef] [PubMed]
- Allaband, C.; McDonald, D.; Vázquez-Baeza, Y.; Minich, J.J.; Tripathi, A.; Brenner, D.A.; Loomba, R.; Smarr, L.; Sandborn, W.J.; Schnabl, B. Microbiome 101: Studying, analyzing, and interpreting gut microbiome data for clinicians. Clin. Gastroenterol. Hepatol. 2019, 17, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.K.; Di Rienzi, S.C.; Poole, A.C.; Koren, O.; Walters, W.A.; Caporaso, J.G.; Knight, R.; Ley, R.E. Conducting a microbiome study. Cell 2014, 158, 250–262. [Google Scholar] [CrossRef]
- Mardis, E.R. Next-Generation Sequencing Platforms. Annu. Rev. Anal. Chem. 2013, 6, 287–303. [Google Scholar] [CrossRef]
- Lu, H.; Giordano, F.; Ning, Z. Oxford Nanopore MinION sequencing and genome assembly. Genom. Proteom. Bioinform. 2016, 14, 265–279. [Google Scholar] [CrossRef]
- Dimitriu, P.A.; Iker, B.; Malik, K.; Leung, H.; Mohn, W.; Hillebrand, G.G. New Insights into the Intrinsic and Extrinsic Factors That Shape the Human Skin Microbiome. mBio 2019, 10, e00839-00819. [Google Scholar] [CrossRef]
- Krieger, Y.; Horev, A.; Wainstock, T.; Sheiner, E.; Walfisch, A. Meconium stained amniotic fluid as a protective factor against childhood dermatitis and skin rash related hospitalization in the offspring–a population based cohort analysis. J. Eur. Acad. Dermatol. Venereol. 2019. [Google Scholar] [CrossRef]
- Zheng, Y.; Wang, Q.; Ma, L.; Chen, Y.; Gao, Y.; Zhang, G.; Cui, S.; Liang, H.; Song, L.; He, C. Shifts in the skin microbiome associated with diaper dermatitis and emollient treatment amongst infants and toddlers in China. Exp. Dermatol. 2019. [Google Scholar] [CrossRef]
- Lee, H.J.; Jeong, S.E.; Lee, S.; Kim, S.; Han, H.; Jeon, C.O. Effects of cosmetics on the skin microbiome of facial cheeks with different hydration levels. MicrobiologyOpen 2018, 7, e00557. [Google Scholar] [CrossRef] [PubMed]
- Bojar, R.A.; Holland, K.T. Acne and Propionibacterium acnes. Clin. Dermatol. 2004, 22, 375–379. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Liu, X.; Niu, Y.; Shen, C.; Heminger, K.; Moulton, L.; Yu, A.; Allen, T.; Zhang, L.; Yue, F. Skin benefits of moisturising body wash formulas for children with atopic dermatitis: A randomised controlled clinical study in China. Australas. J. Dermatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- van Mierlo, M.M.; Totté, J.E.; Fieten, K.B.; van den Broek, T.J.; Schuren, F.H.; Pardo, L.M.; Pasmans, S.G. The influence of treatment in alpine and moderate maritime climate on the composition of the skin microbiome in patients with difficult to treat atopic dermatitis. Clin. Exp. Allergy 2019. [Google Scholar] [CrossRef]
- Patra, V.; Wagner, K.; Arulampalam, V.; Wolf, P. Skin Microbiome Modulates the Effect of Ultraviolet Radiation on Cellular Response and Immune Function. iScience 2019, 15, 211–222. [Google Scholar] [CrossRef]
- Vandegrift, R.; Fahimipour, A.K.; Muscarella, M.; Bateman, A.C.; Van Den Wymelenberg, K.; Bohannan, B.J. Moving microbes: The dynamics of transient microbial residence on human skin. bioRxiv 2019, 586008. Available online: https://www.biorxiv.org/content/10.1101/586008v1 (accessed on 18 February 2020).
- Chien, A.L.; Tsai, J.; Leung, S.; Mongodin, E.F.; Nelson, A.M.; Kang, S.; Garza, L.A. Association of systemic antibiotic treatment of acne with skin microbiota characteristics. Jama Dermatol. 2019, 155, 425–434. [Google Scholar] [CrossRef]
- Zhang, M.; Jiang, Z.; Li, D.; Jiang, D.; Wu, Y.; Ren, H.; Peng, H.; Lai, Y. Oral antibiotic treatment induces skin microbiota dysbiosis and influences wound healing. Microb. Ecol. 2015, 69, 415–421. [Google Scholar] [CrossRef]
- Dellacecca, E.R.; Cosgrove, C.; Mukhatayev, Z.; Akhtar, S.; Engelhard, V.H.; Rademaker, A.W.; Knight, K.; Le Poole, I.C. Antibiotics drive microbial imbalance and vitiligo development in mice. J. Investig. Dermatol. 2019. [Google Scholar] [CrossRef]
- Zeng, J.; Dou, J.; Gao, L.; Xiang, Y.; Huang, J.; Ding, S.; Chen, J.; Zeng, Q.; Luo, Z.; Tan, W. Topical ozone therapy restores microbiome diversity in atopic dermatitis. Int. Immunopharmacol. 2020, 80, 106191. [Google Scholar] [CrossRef]
- Kalliomäki, M.; Salminen, S.; Poussa, T.; Isolauri, E. Probiotics during the first 7 years of life: A cumulative risk reduction of eczema in a randomized, placebo-controlled trial. J. Allergy Clin. Immunol. 2007, 119, 1019. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeldt, V.; Benfeldt, E.; Valerius, N.H.; Pærregaard, A.; Michaelsen, K.F. Effect of probiotics on gastrointestinal symptoms and small intestinal permeability in children with atopic dermatitis. J. Pediatrics 2004, 145, 612–616. [Google Scholar] [CrossRef] [PubMed]
- Jung, G.W.; Tse, J.E.; Guiha, I.; Rao, J. Prospective, randomized, open-label trial comparing the safety, efficacy, and tolerability of an acne treatment regimen with and without a probiotic supplement and minocycline in subjects with mild to moderate acne. J. Cutan. Med. Surg. 2013, 17, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Karska-Wysocki, B.; Bazo, M.; Smoragiewicz, W. Antibacterial activity of Lactobacillus acidophilus and Lactobacillus casei against methicillin-resistant Staphylococcus aureus (MRSA). Microbiol. Res. 2010, 165, 674–686. [Google Scholar] [CrossRef]
- Venosi, S.; Ceccarelli, G.; de Angelis, M.; Laghi, L.; Bianchi, L.; Martinelli, O.; Maruca, D.; Cavallari, E.N.; Toscanella, F.; Vassalini, P. Infected chronic ischemic wound topically treated with a multi-strain probiotic formulation: A novel tailored treatment strategy. J. Transl. Med. 2019, 17, 364. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Chen, T.H.; Narala, S.; Chun, K.A.; Two, A.M.; Yun, T.; Shafiq, F.; Kotol, P.F.; Bouslimani, A.; Melnik, A.V. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. 2017, 9, eaah4680. [Google Scholar] [CrossRef]
- Myles, I.A.; Earland, N.J.; Anderson, E.D.; Moore, I.N.; Kieh, M.D.; Williams, K.W.; Saleem, A.; Fontecilla, N.M.; Welch, P.A.; Darnell, D.A. First-in-human topical microbiome transplantation with Roseomonas mucosa for atopic dermatitis. Jci Insight 2018, 3. [Google Scholar] [CrossRef]
- Schoch, J.J.; Monir, R.L.; Satcher, K.G.; Harris, J.; Triplett, E.; Neu, J. The infantile cutaneous microbiome: A review. Pediatric Dermatol. 2019, 36, 574–580. [Google Scholar] [CrossRef]
- Milani, C.; Duranti, S.; Bottacini, F.; Casey, E.; Turroni, F.; Mahony, J.; Belzer, C.; Palacio, S.D.; Montes, S.A.; Mancabelli, L. The first microbial colonizers of the human gut: Composition, activities, and health implications of the infant gut microbiota. Microbiol. Mol. Biol. Rev. 2017, 81, e00036-00017. [Google Scholar] [CrossRef]
- Chu, D.M.; Ma, J.; Prince, A.L.; Antony, K.M.; Seferovic, M.D.; Aagaard, K.M. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 2017, 23, 314–326. [Google Scholar] [CrossRef]
- Ardissone, A.N.; Diomel, M.; Davis-Richardson, A.G.; Rechcigl, K.T.; Li, N.; Drew, J.C.; Murgas-Torrazza, R.; Sharma, R.; Hudak, M.L.; Triplett, E.W. Meconium microbiome analysis identifies bacteria correlated with premature birth. PLoS ONE 2014, 9, e90784. [Google Scholar] [CrossRef]
- Salem, I.; Ramser, A.; Isham, N.; Ghannoum, M.A. The Gut Microbiome as a Major Regulator of the Gut-Skin Axis. Front Microbiol. 2018, 9, 1459. [Google Scholar] [CrossRef] [PubMed]
- Schommer, N.N.; Gallo, R.L. Structure and function of the human skin microbiome. Trends Microbiol. 2013, 21, 660–668. [Google Scholar] [CrossRef] [PubMed]
- Ferrazzini, G.; Kaiser, R.; Cheng, S.-K.H.; Wehrli, M.; Della Casa, V.; Pohlig, G.; Gonser, S.; Graf, F.; Jörg, W. Microbiological aspects of diaper dermatitis. Dermatology 2003, 206, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Verdier-Sévrain, S.; Bonté, F. Skin hydration: A review on its molecular mechanisms. J. Cosmet. Dermatol. 2007, 6, 75–82. [Google Scholar]
- Moissl-Eichinger, C.; Probst, A.J.; Birarda, G.; Auerbach, A.; Koskinen, K.; Wolf, P.; Holman, H.-Y.N. Human age and skin physiology shape diversity and abundance of Archaea on skin. Sci. Rep. 2017, 7, 4039. [Google Scholar] [CrossRef]
- Nutten, S. Atopic dermatitis: Global epidemiology and risk factors. Ann. Nutr. Metab. 2015, 66, 8–16. [Google Scholar] [CrossRef]
- Woo, T.E.; Sibley, C.D. The emerging utility of the cutaneous microbiome in the treatment of acne and atopic dermatitis. J. Am. Acad. Dermatol. 2019. [Google Scholar] [CrossRef]
- Vargason, A.M.; Anselmo, A.C. Clinical translation of microbe-based therapies: Current clinical landscape and preclinical outlook. Bioeng. Transl. Med. 2018, 3, 124–137. [Google Scholar] [CrossRef]
- Prescott, S.L.; Logan, A.C. Transforming Life: A Broad View of the Developmental Origins of Health and Disease Concept from an Ecological Justice Perspective. Int. J. Environ. Res. Public Health 2016, 13, 1075. [Google Scholar] [CrossRef]
- Clausen, M.-L.; Agner, T.; Lilje, B.; Edslev, S.M.; Johannesen, T.B.; Andersen, P.S. Association of disease severity with skin microbiome and filaggrin gene mutations in adult atopic dermatitis. Jama Dermatol. 2018, 154, 293–300. [Google Scholar] [CrossRef]
- Patra, V.; Byrne, S.N.; Wolf, P. The skin microbiome: Is it affected by UV-induced immune suppression? Front. Microbiol. 2016, 7, 1235. [Google Scholar] [CrossRef] [PubMed]
- Nishijima, S.; Kurokawa, I.; Katoh, N.; Watanabe, K. The bacteriology of acne vulgaris and antimicrobial susceptibility of Propionibacterium acnes and Staphylococcus epidermidis isolated from acne lesions. J. Dermatol. 2000, 27, 318–323. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, A.M.; Gallo, R.L. Host-microbiome interactions and recent progress into understanding the biology of acne vulgaris. Microbiome 2018, 6, 177. [Google Scholar] [CrossRef] [PubMed]
- Prince, T.; McBain, A.J.; O’Neill, C.A. Lactobacillus reuteri protects epidermal keratinocytes from Staphylococcus aureus-induced cell death by competitive exclusion. Appl. Environ. Microbiol. 2012, 78, 5119–5126. [Google Scholar] [CrossRef] [PubMed]
- Mohammedsaeed, W.; McBain, A.J.; Cruickshank, S.M.; O’Neill, C.A. Lactobacillus rhamnosus GG inhibits the toxic effects of Staphylococcus aureus on epidermal keratinocytes. Appl. Environ. Microbiol. 2014, 80, 5773–5781. [Google Scholar] [CrossRef]
- Panduru, M.; Panduru, N.; Sălăvăstru, C.; Tiplica, G.S. Probiotics and primary prevention of atopic dermatitis: A meta-analysis of randomized controlled studies. J. Eur. Acad. Dermatol. Venereol. 2015, 29, 232–242. [Google Scholar] [CrossRef]
- Nole, K.L.B.; Yim, E.; Keri, J.E. Probiotics and prebiotics in dermatology. J. Am. Acad. Dermatol. 2014, 71, 814–821. [Google Scholar] [CrossRef]
- Imanishi, I.; Uchiyama, J.; Tsukui, T.; Hisatsune, J.; Ide, K.; Matsuzaki, S.; Sugai, M.; Nishifuji, K. Therapeutic potential of an endolysin derived from kayvirus S25-3 for staphylococcal impetigo. Viruses 2019, 11, 769. [Google Scholar] [CrossRef]
- Smits, J.P.; Ederveen, T.H.; Rikken, G.; van den Brink, N.J.; van Vlijmen-Willems, I.M.; Boekhorst, J.; Kamsteeg, M.; Schalkwijk, J.; van Hijum, S.A.; Zeeuwen, P.L. TARgeting the cutaneous microbiota in atopic dermatitis by coal tar via AHR-dependent induction of antimicrobial peptides. J. Investig. Dermatol. 2019. [Google Scholar] [CrossRef]
- Lai, Y.; Li, D.; Li, C.; Muehleisen, B.; Radek, K.A.; Park, H.J.; Jiang, Z.; Li, Z.; Lei, H.; Quan, Y. The antimicrobial protein REG3A regulates keratinocyte proliferation and differentiation after skin injury. Immunity 2012, 37, 74–84. [Google Scholar] [CrossRef] [PubMed]
- Elvis, A.M.; Ekta, J.S. Ozone therapy: A clinical review. J. Nat. Sci. Biol. Med. 2011, 2, 66–70. [Google Scholar] [CrossRef] [PubMed]
- Gasbarrini, G.; Bonvicini, F.; Gramenzi, A. Probiotics history. J. Clin. Gastroenterol. 2016, 50, S116–S119. [Google Scholar] [CrossRef] [PubMed]
- Joint FAO/WHO Working Group. Guidelines for the Evaluation of Probiotics in Food. Fao/Wholondonon; Food and Agriculture Organization of the United Nations: Rome, Italy; World Health Organization: Geneva, Switzerland, 2002. [Google Scholar]
- Lolou, V.; Panayiotidis, M.I. Functional role of probiotics and prebiotics on skin health and disease. Fermentation 2019, 5, 41. [Google Scholar] [CrossRef]
- Knackstedt, R.; Knackstedt, T.; Gatherwright, J. The role of topical probiotics on skin conditions: A systematic review of animal and human studies and implications for future therapies. Exp. Dermatol. 2019. [Google Scholar] [CrossRef]
- Dimarzio, L.; Cinque, B.; Cupelli, F.; De Simone, C.; Cifone, M.G.; Giuliani, M. Increase of skin-ceramide levels in aged subjects following a short-term topical application of bacterial sphingomyelinase from Streptococcus thermophilus. Int. J. Immunopathol. Pharmacol. 2008, 21, 137–143. [Google Scholar] [CrossRef]
- Kimoto-Nira, H.; Aoki, R.; Sasaki, K.; Suzuki, C.; Mizumachi, K. Oral intake of heat-killed cells of Lactococcus lactis strain H61 promotes skin health in women. J. Nutr. Sci. 2012, 1. [Google Scholar] [CrossRef]
- Krutmann, J. Pre-and probiotics for human skin. J. Dermatol. Sci. 2009, 54, 1–5. [Google Scholar] [CrossRef]
- Hendricks, A.J.; Mills, B.W.; Shi, V.Y. Skin bacterial transplant in atopic dermatitis: Knowns, unknowns and emerging trends. J. Dermatol. Sci. 2019. [Google Scholar] [CrossRef]
- ClinicalTrials.gov. Targeted Microbiome Transplantation in Atopic Dermatitis (NCT03151148). Available online: https://www.clinicaltrials.gov/ (accessed on 23 March 2020).
- Integrative HMP (iHMP) Research Network Consortium. The integrative human microbiome project. Nature 2019, 569, 641–648. [Google Scholar]
| Classification | Factor | Skin Condition | Effect on Microbiome | Clinical Outcome | Reference |
|---|---|---|---|---|---|
| Early life exposures | Meconium-stained amniotic fluid | Healthy | ↑ gut microbiome diversity | Prevention of skin inflammation-related diseases | [51] |
| Diapers | Diaper dermatitis | ↑ Enterococcus and S. aureus abundances | Not described | [52] | |
| ↓ Lactobacillus and Bifidobacterium abundances | |||||
| Cosmetics | Skin hydration set | Healthy | ↑ bacterial diversity | Not described | [53] |
| ↓ Propionibacterium abundance | |||||
| Madecassoside | Healthy | ↓ P. acnes | Enhanced skin hydration | [54] | |
| Suppressed inflammation | |||||
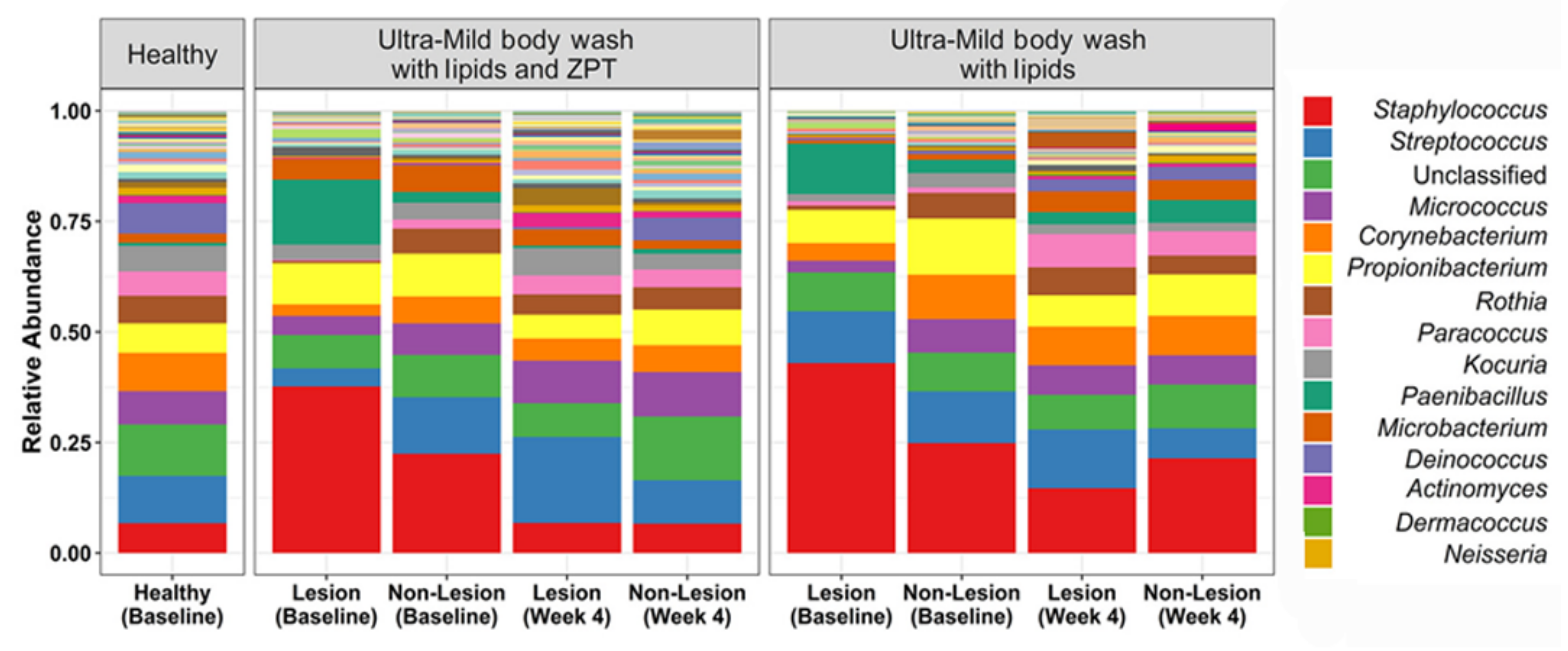
| Lipidic body wash with ZPT | Atopic dermatitis | Bacterial shift to healthy controls | No effect on AD severity | [55] | |
| ↓ S. aureus abundance | |||||
| Environment and nature | Alpine climate | Atopic dermatitis | ↓ S. aureus abundance | Decrease in disease severity | [56] |
| UV-B radiation | Healthy | Immune response intermediate | Protection against immuno-suppression | [57] | |
| Soil and plant leaves | Healthy | Altered microbiome diversity | Transient bacterial shift dependent on donor bacterial biomass | [58] | |
| Resemblance of human bacterial composition to that of the donor (soil/leaf) | |||||
| Antimicrobial agents | Minocycline | Acne | ↓ Cutibacterium, Corynebacterium, Prevotella, Lactobacillus, and Porphyromonas | Reduced abundance of skin-protective bacteria | [59] |
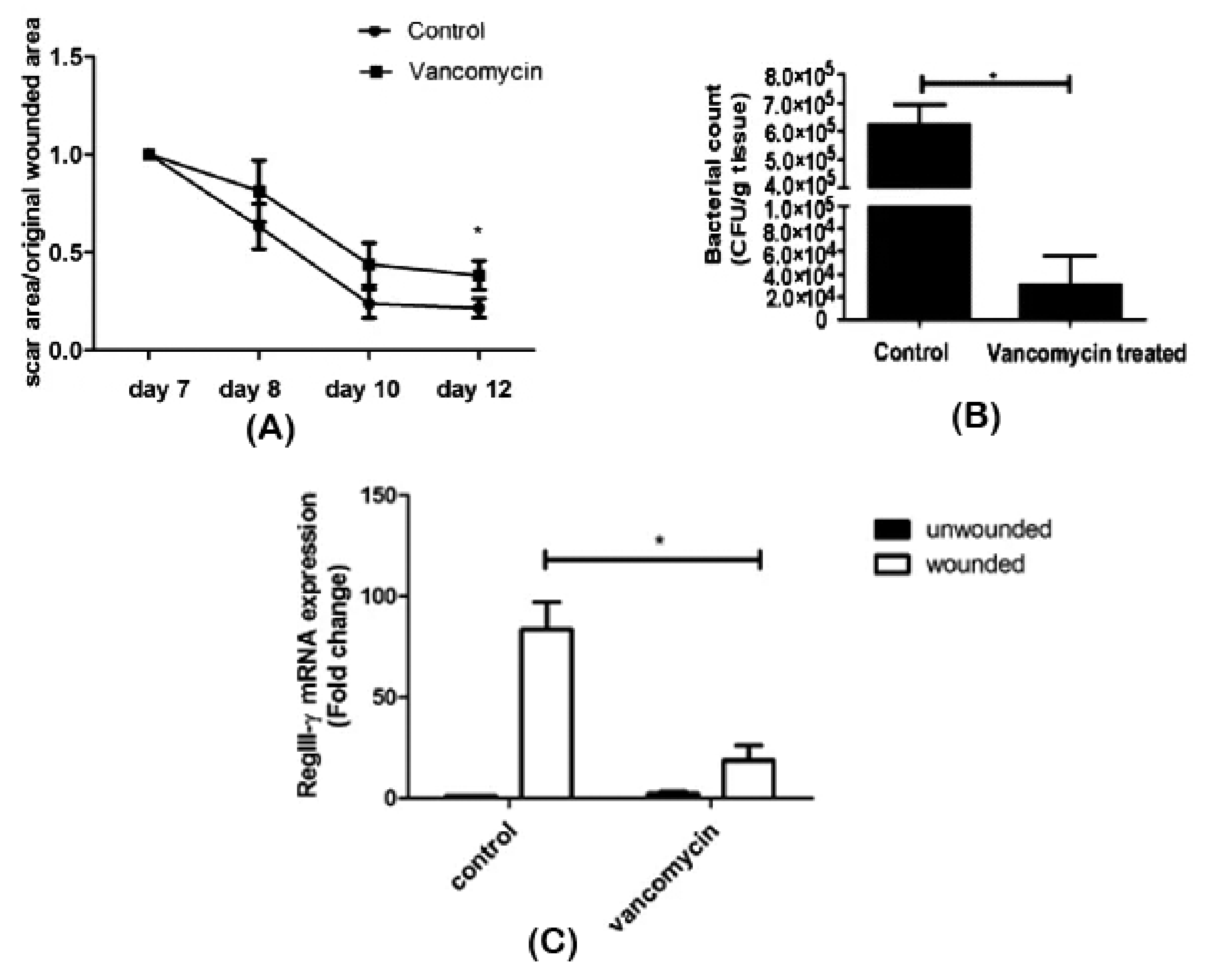
| Vancomycin | Wounded skin | ↓ Staphylococcaceae | Delayed wound repair | [60] | |
| ↑ Lactobacillaceae | |||||
| ↓ RegIII-γ | |||||
| Ampicillin | Vitiligo | ↓ gut bacterial abundance | Accelerated depigmentation | [61] | |
| Ozone | AD | ↑ skin microbiome diversity | Mitigation of AD lesions | [62] | |
| ↓ Staphylococcus, Acinetobacter, Lactobacillus, Streptococcus, and Propionibacterium abundances | |||||
| Bacterium-based agents | L. rhamnosus | AD | No mechanism described | Reduced risk of disease development in children | [63] |
| L. rhamnosus and L. reuteri | AD | No mechanism described | Reduced disease severity in children | [64] | |
| L. acidophilus, L. delbrueckii, and B. bifidum | Acne | No mechanism described | Reduced lesion number | [65] | |
| L. acidophilus and L casei | Wounded skin | ↓MRSA | Reduced risk of infection | [66] | |
| K. pneumoniae, E. faecalis, and P. mirabilis | Chronic ischemic wounds | ↓Pathogen growth | Complete wound healing | [67] | |
| Immune response modulation | |||||
| S. hominis | AD | ↓S. aureus abundance | Improved disease symptoms | [68] | |
| R. mucosa | AD | ↓S. aureus abundance | Reduced disease severity | [69] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moskovicz, V.; Gross, A.; Mizrahi, B. Extrinsic Factors Shaping the Skin Microbiome. Microorganisms 2020, 8, 1023. https://doi.org/10.3390/microorganisms8071023
Moskovicz V, Gross A, Mizrahi B. Extrinsic Factors Shaping the Skin Microbiome. Microorganisms. 2020; 8(7):1023. https://doi.org/10.3390/microorganisms8071023
Chicago/Turabian StyleMoskovicz, Veronica, Adi Gross, and Boaz Mizrahi. 2020. "Extrinsic Factors Shaping the Skin Microbiome" Microorganisms 8, no. 7: 1023. https://doi.org/10.3390/microorganisms8071023
APA StyleMoskovicz, V., Gross, A., & Mizrahi, B. (2020). Extrinsic Factors Shaping the Skin Microbiome. Microorganisms, 8(7), 1023. https://doi.org/10.3390/microorganisms8071023





