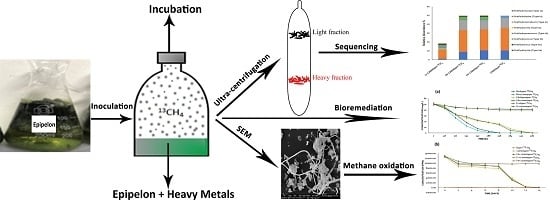
Enhanced Adsorptive Bioremediation of Heavy Metals (Cd2+, Cr6+, Pb2+) by Methane-Oxidizing Epipelon
Abstract
1. Introduction
2. Materials and Methods
2.1. Epipelon Isolation and Growth conditions
2.2. DNA-SIP and Heavy Metals Bioremediation Analysis
2.3. Morphology and Activity of Microbes in Epipelon Biofilms
2.4. DNA Extraction and SIP Gradient Fractionation
2.5. Real-Time Quantitative PCR of pmoA Genes
2.6. MiSeq Sequencing of 16S rRNA Genes
2.7. MiSeq Sequencing of pmoA Genes
2.8. Statistical and Network Analysis
3. Results and Discussion
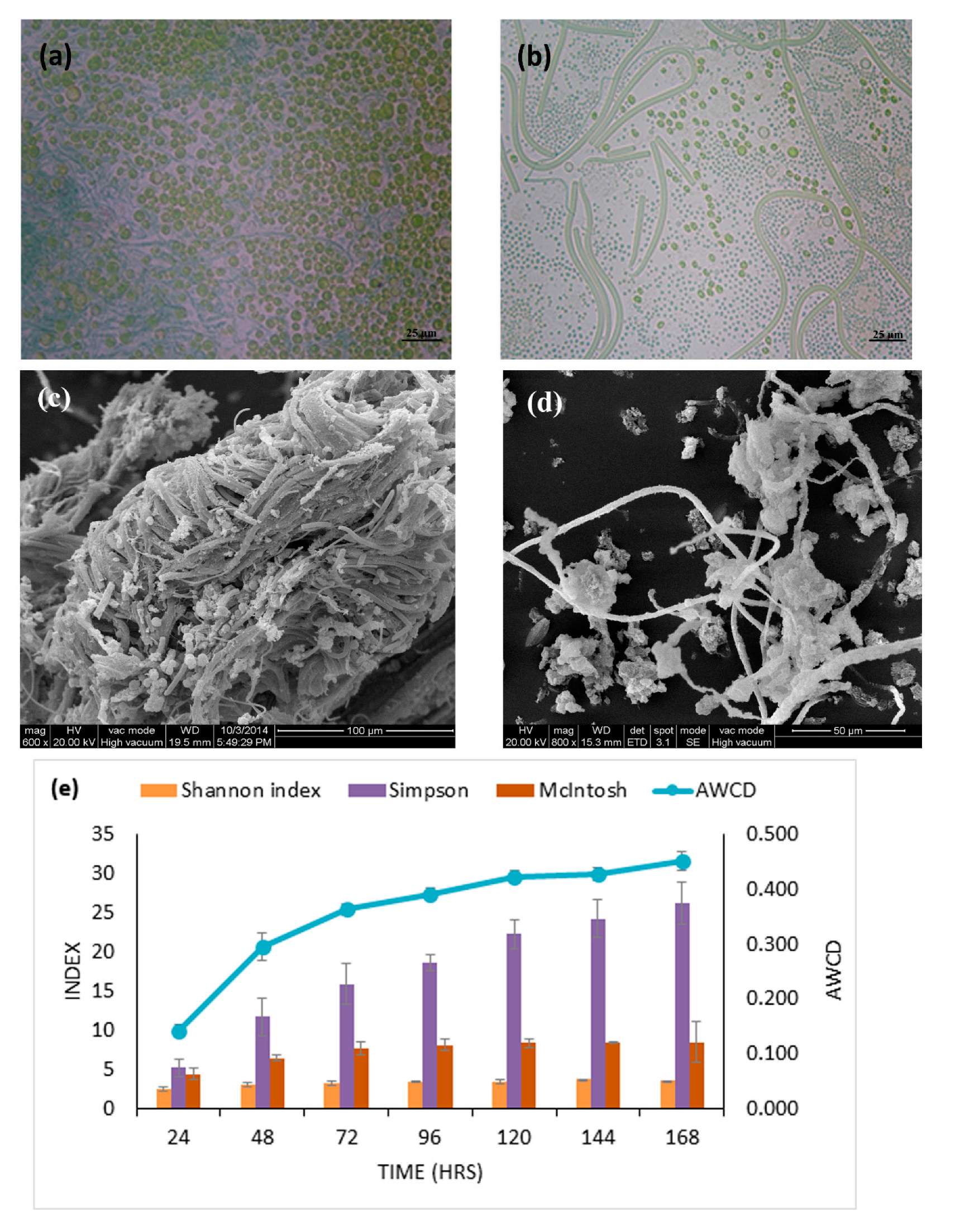
3.1. Physiochemistry, Cell Morphology and Activity of Epipelon Biofilm
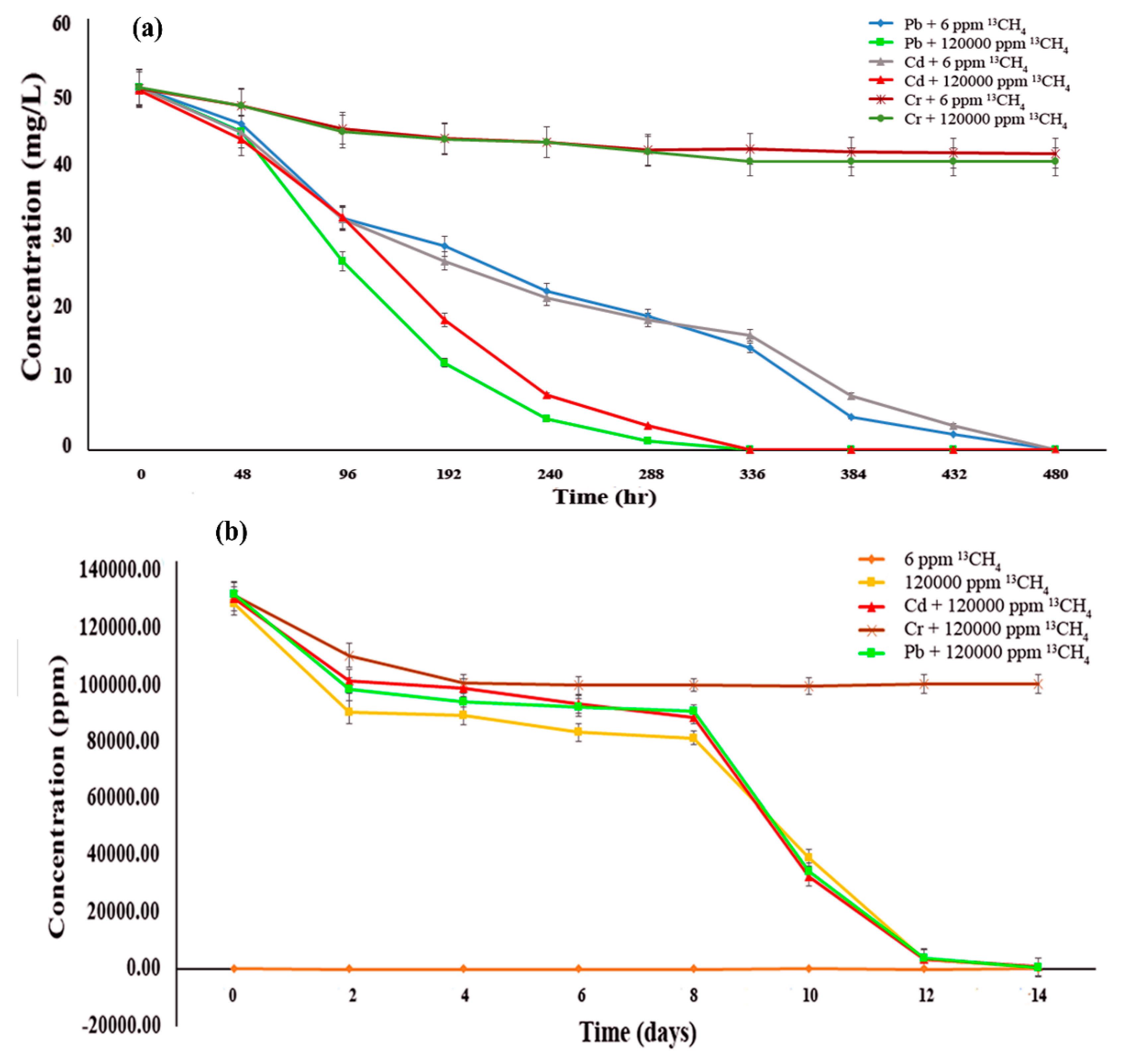
3.2. Effect of Methane Oxidation on Heavy Metal Bioremediation
3.3. Potential of Epipelon for Methane Oxidation and 13C Assimilation
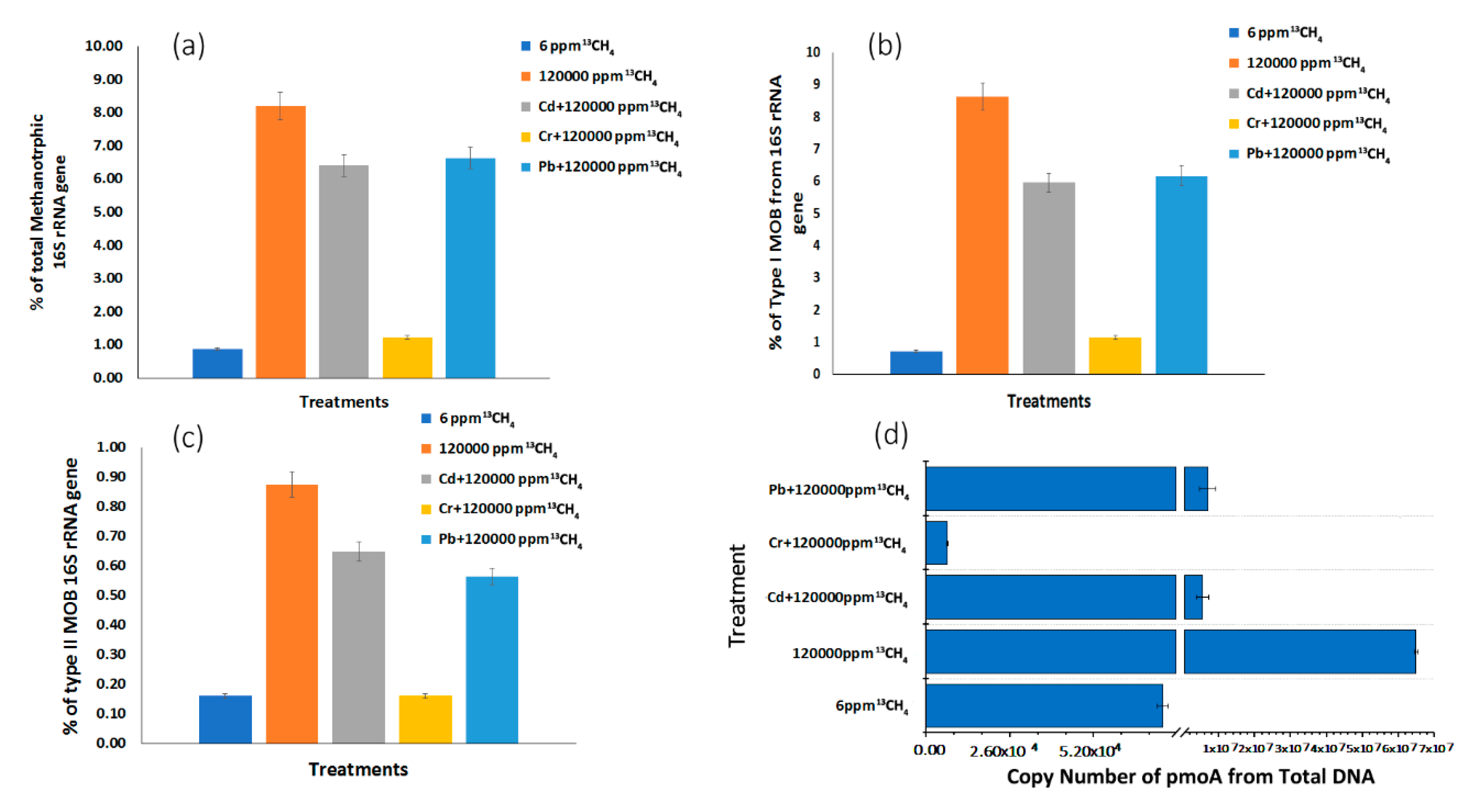
3.4. Methanotrophic Community Abundance in Response to Heavy Metals
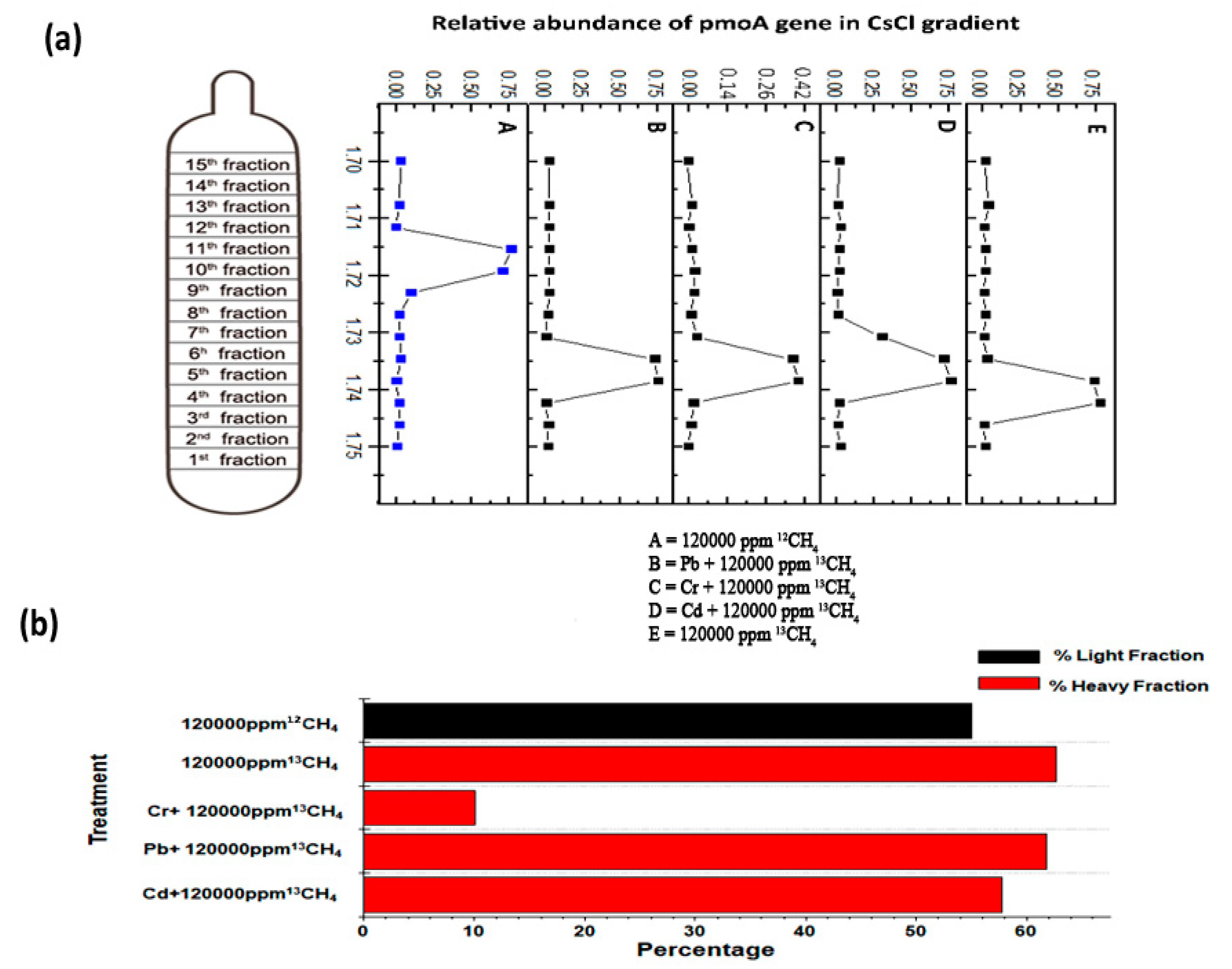
3.5. SIP for Identification of Active Methanotrophs
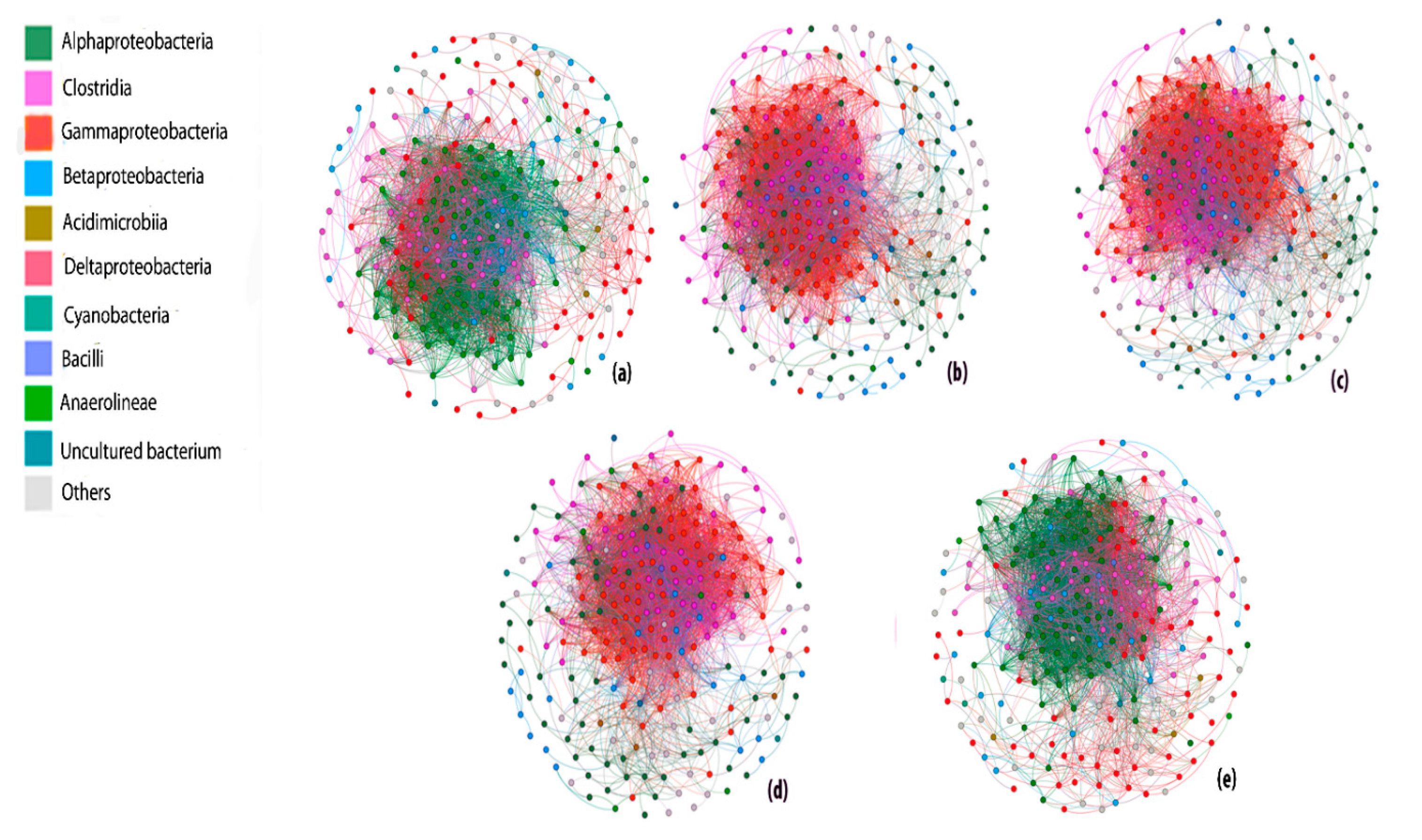
3.6. Microbial Community Response to Heavy Metals Bioremediation and Methane Oxidation
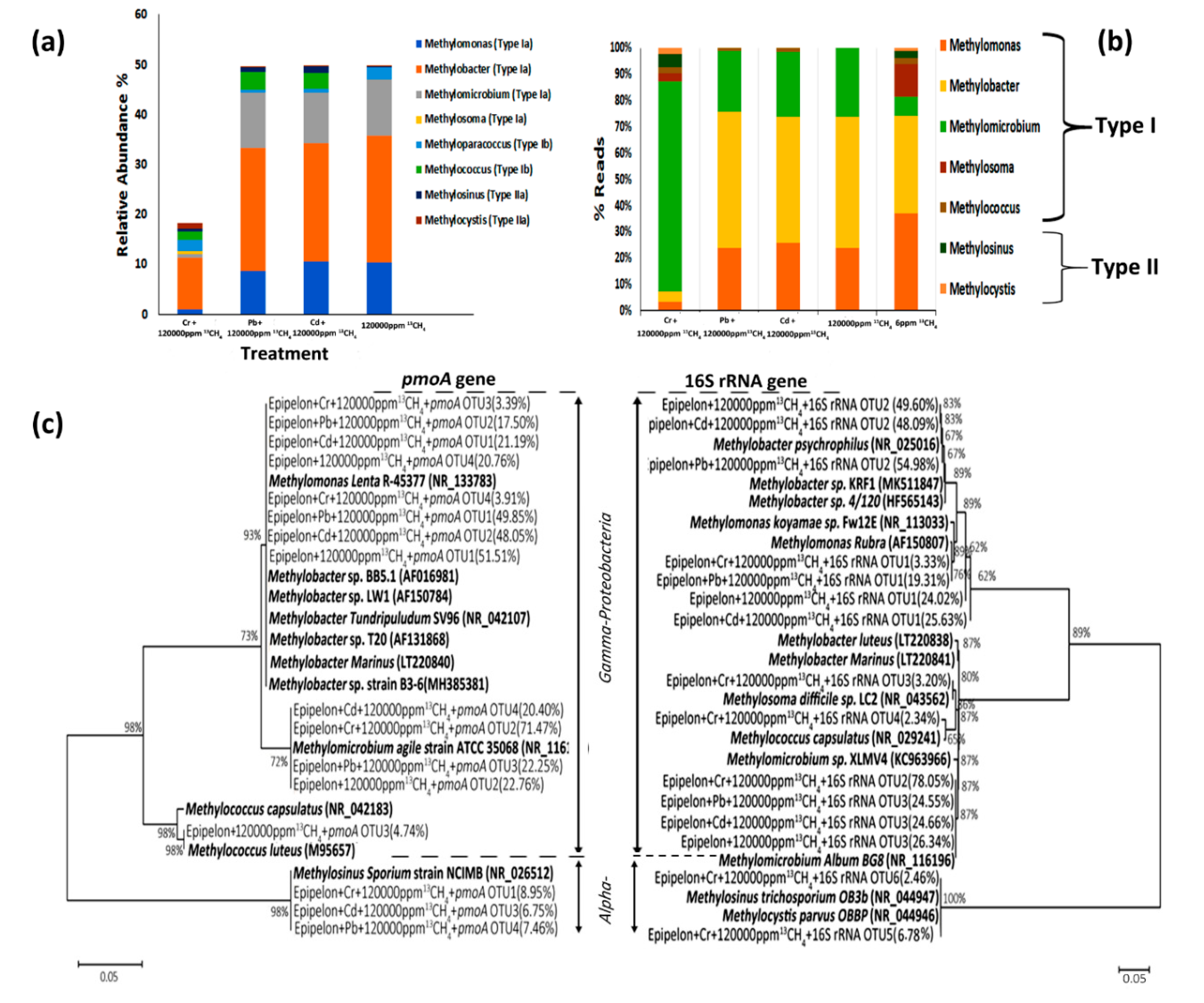
3.7. Methanotrophic Community Composition
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Duren, R.M.; Thorpe, A.K.; Foster, K.T.; Rafiq, T.; Hopkins, F.M.; Yadav, V.; Bue, B.D.; Thompson, D.R.; Conley, S.; Colombi, N.K.; et al. California’s methane super-emitters. Nature 2019, 575, 180–184. [Google Scholar] [CrossRef] [PubMed]
- Rai, P.K.; Lee, S.S.; Zhang, M.; Tsang, Y.F.; Kim, K.-H. Heavy metals in food crops: Health risks, fate, mechanisms, and management. Environ. Int. 2019, 125, 365–385. [Google Scholar] [CrossRef] [PubMed]
- Iguchi, H.; Umeda, R.; Taga, H.; Oyama, T.; Yurimoto, H.; Sakai, Y. Community composition and methane oxidation activity of methanotrophs associated with duckweeds in a fresh water lake. J. Biosci. Bioeng. 2019, 128, 450–455. [Google Scholar] [CrossRef] [PubMed]
- Landrigan, P.J.; Fuller, R.; Acosta, N.J.R.; Adeyi, O.; Arnold, R.; Basu, N.; Baldé, A.B.; Bertollini, R.; Bose-O’Reilly, S.; Boufford, J.I.; et al. The Lancet Commission on pollution and health. Lancet 2018, 391, 462–512. [Google Scholar] [CrossRef]
- Laxmi, V.; Kaushik, G. Toxicity of Hexavalent Chromium in Environment, Health Threats, and Its Bioremediation and Detoxification from Tannery Wastewater for Environmental Safety. In Bioremediation of Industrial Waste for Environmental Safety: Volume I: Industrial Waste and Its Management; Saxena, G., Bharagava, R.N., Eds.; Springer: Singapore, 2020; pp. 223–243. [Google Scholar] [CrossRef]
- Fu, Z.; Xi, S. The effects of heavy metals on human metabolism. Toxicol. Mech. Methods 2020, 30, 167–176. [Google Scholar] [CrossRef]
- Bhakta, J.; Munekage, Y.; Ohnishi, K.; Jana, B. Isolation and identification of cadmium-and lead-resistant lactic acid bacteria for application as metal removing probiotic. Int. J. Environ. Sci.Technol. 2012, 9, 433–440. [Google Scholar] [CrossRef]
- Doggaz, A.; Attoura, A.; Mostefa, M.L.P.; Côme, K.; Tlili, M.; Lapicque, F. Removal of heavy metals by electrocoagulation from hydrogenocarbonate-containing waters: Compared cases of divalent iron and zinc cations. J. Water Process Eng. 2019, 29, 100796. [Google Scholar] [CrossRef]
- Abdullah, N.; Yusof, N.; Lau, W.; Jaafar, J.; Ismail, A. Recent trends of heavy metal removal from water/wastewater by membrane technologies. J. Ind. Eng. Chem. 2019, 76, 17–38. [Google Scholar] [CrossRef]
- Soto, J.; Ortiz, J.; Herrera, H.; Fuentes, A.; Almonacid, L.; Charles, T.C.; Arriagada, C. Enhanced Arsenic Tolerance in Triticum aestivum Inoculated with Arsenic-Resistant and Plant Growth Promoter Microorganisms from a Heavy Metal-Polluted Soil. Microorganisms 2019, 7, 348. [Google Scholar] [CrossRef]
- Wang, Y.; Zhong, B.; Shafi, M.; Ma, J.; Guo, J.; Wu, J.; Ye, Z.; Liu, D.; Jin, H. Effects of biochar on growth, and heavy metals accumulation of moso bamboo (Phyllostachy pubescens), soil physical properties, and heavy metals solubility in soil. Chemosphere 2019, 219, 510–516. [Google Scholar] [CrossRef]
- Khatiwada, B.; Hasan, M.T.; Sun, A.; Kamath, K.S.; Mirzaei, M.; Sunna, A.; Nevalainen, H. Probing the Role of the Chloroplasts in Heavy Metal Tolerance and Accumulation in Euglena gracilis. Microorganisms 2020, 8, 115. [Google Scholar] [CrossRef] [PubMed]
- Hazen, T. Consequences of Microbial Interactions with Hydrocarbons, Oils, and Lipids: Biodegradation and Bioremediation; Cometabolic Bioremediation; Springer-Verlag: Berlin/Heidelberg, Germany, 2018; pp. 1–15. [Google Scholar]
- Chan, S.I.; Lee, S.J. The Biochemistry of Methane Monooxygenases. In Methanotrophs: Microbiology Fundamentals and Biotechnological Applications. Lee, E.Y., Ed.; Springer International Publishing: Cham, Germany, 2019; pp. 71–120. [Google Scholar] [CrossRef]
- Lai, C.-Y.; Zhong, L.; Zhang, Y.; Chen, J.-X.; Wen, L.-L.; Shi, L.-D.; Sun, Y.-P.; Ma, F.; Rittmann, B.E.; Zhou, C.; et al. Bioreduction of Chromate in a Methane-Based Membrane Biofilm Reactor. Environ. Sci. Technol. 2016, 50, 5832–5839. [Google Scholar] [CrossRef]
- Shi, L.-D.; Chen, Y.-S.; Du, J.-J.; Hu, Y.-Q.; Shapleigh, J.P.; Zhao, H.-P. Metagenomic Evidence for a Methylocystis Species Capable of Bioremediation of Diverse Heavy Metals. Front. Microbiol. 2019, 9, 3297. [Google Scholar] [CrossRef] [PubMed]
- Al Hasin, A.; Gurman, S.J.; Murphy, L.M.; Perry, A.; Smith, T.J.; Gardiner, P.H.E. Remediation of Chromium(VI) by a Methane-Oxidizing Bacterium. Environ. Sci. Technol. 2010, 44, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Tavares, D.A.; Lambrecht, R.W.; de Almeida Castilho, M.C.; Henry, R.; Ferragut, C. Epipelon responses to N and P enrichment and the relationships with phytoplankton and zooplankton in a mesotrophic reservoir. Aquat. Ecol. 2019, 53, 303–314. [Google Scholar] [CrossRef]
- Shabbir, S.; Faheem, M.; Ali, N.; Kerr, P.G.; Wu, Y. Evaluating role of immobilized periphyton in bioremediation of azo dye amaranth. Bioresour. Technol. 2017, 225, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, J.; Wu, C.; Kerr, P.G.; Wong, P.-K.; Wu, Y. Bioremediation of agricultural solid waste leachates with diverse species of Cu (II) and Cd (II) by periphyton. Bioresour. Technol. 2016, 221, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wang, F.; Wu, W.; Wan, J.; Yang, J.; Xiang, S.; Wu, Y. Biosorption of high-concentration Cu (II) by periphytic biofilms and the development of a fiber periphyton bioreactor (FPBR). Bioresour. Technol. 2018, 248, 127–134. [Google Scholar] [CrossRef]
- Tang, C.; Sun, P.; Yang, J.; Huang, Y.; Wu, Y. Kinetics simulation of Cu and Cd removal and the microbial community adaptation in a periphytic biofilm reactor. Bioresour. Technol. 2019, 276, 199–203. [Google Scholar] [CrossRef]
- Lu, H.; Dong, Y.; Feng, Y.; Bai, Y.; Tang, X.; Li, Y.; Yang, L.; Liu, J. Paddy periphyton reduced cadmium accumulation in rice (Oryza sativa) by removing and immobilizing cadmium from the water–soil interface. Environ. Pollut. 2020, 114103. [Google Scholar] [CrossRef]
- Li, Y.; Song, S.; Xia, L.; Yin, H.; García Meza, J.V.; Ju, W. Enhanced Pb(II) removal by algal-based biosorbent cultivated in high-phosphorus cultures. Chem. Eng. J. 2019, 361, 167–179. [Google Scholar] [CrossRef]
- Jasrotia, P.; Ogram, A. Diversity of nifH Genotypes in Floating Periphyton Mats Along a Nutrient Gradient in the Florida Everglades. Curr. Microbiol. 2008, 56, 563–568. [Google Scholar] [CrossRef][Green Version]
- Cano, M.G.; Casco, M.A.; Solari, L.C.; Mac Donagh, M.E.; Gabellone, N.A.; Claps, M.C. Implications of rapid changes in chlorophyll-a of plankton, epipelon, and epiphyton in a Pampean shallow lake: An interpretation in terms of a conceptual model. Hydrobiologia 2008, 614, 33–45. [Google Scholar] [CrossRef]
- Shabbir, S.; Faheem, M.; Wu, Y. Decolorization of high concentration crystal violet by periphyton bioreactors and potential of effluent reuse for agricultural purposes. J. Clean. Prod. 2018, 170, 425–436. [Google Scholar] [CrossRef]
- Pniewski, F.F.; Richard, P.; Latała, A.; Blanchard, G. Long- and short-term photoacclimation in epipsammon from non-tidal coastal shallows compared to epipelon from intertidal mudflat. J. Sea Res. 2018, 136, 1–9. [Google Scholar] [CrossRef]
- Sultana, N.; Zhao, J.; Zheng, Y.; Cai, Y.; Faheem, M.; Peng, X.; Wang, W.; Jia, Z. Stable isotope probing of active methane oxidizers in rice field soils from cold regions. Biol. Fertil. Soils 2019, 55, 243–250. [Google Scholar] [CrossRef]
- Ziadat, A.H.; Jiries, A.; Alojail, I. Accumulation of Heavy Metals on Soil Irrigated with Treated Wastewater at Al al-Bayt University-Jordan. In Proceedings of the 2019 Advances in Science and Engineering Technology International Conferences (ASET), Dubai, UAE, 26 March–10 April 2019; pp. 1–6. [Google Scholar]
- Shabbir, S.; Faheem, M.; Ali, N.; Kerr, P.G.; Wang, L.-F.; Kuppusamy, S.; Li, Y. Periphytic biofilm: An innovative approach for biodegradation of microplastics. Sci. Total Environ. 2020, 717, 137064. [Google Scholar] [CrossRef]
- Jia, Z.; Conrad, R. Bacteria rather than Archaea dominate microbial ammonia oxidation in an agricultural soil. Environ. Microbiol. 2009, 11, 1658–1671. [Google Scholar] [CrossRef]
- Cai, Y.; Zheng, Y.; Bodelier, P.L.E.; Conrad, R.; Jia, Z. Conventional methanotrophs are responsible for atmospheric methane oxidation in paddy soils. Nat. Commun. 2016, 7, 11728. [Google Scholar] [CrossRef]
- Vestergaard, G.; Schulz, S.; Schöler, A.; Schloter, M. Making big data smart—how to use metagenomics to understand soil quality. Biol. Fertil. Soils 2017, 53, 479–484. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Lan, Y.; Wang, Q.; Cole, J.R.; Rosen, G.L. Using the RDP Classifier to Predict Taxonomic Novelty and Reduce the Search Space for Finding Novel Organisms. PLoS ONE 2012, 7, e32491. [Google Scholar] [CrossRef] [PubMed]
- Barberán, A.; Bates, S.T.; Casamayor, E.O.; Fierer, N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 2012, 6, 343–351. [Google Scholar] [CrossRef]
- Hall, B.G. Building Phylogenetic Trees from Molecular Data with MEGA. Mol. Biol. Evol. 2013, 30, 1229–1235. [Google Scholar] [CrossRef]
- Schöler, A.; Jacquiod, S.; Vestergaard, G.; Schulz, S.; Schloter, M. Analysis of soil microbial communities based on amplicon sequencing of marker genes. Biol. Fertil. Soils 2017, 53, 485–489. [Google Scholar] [CrossRef]
- Douglas, G.M.; Beiko, R.G.; Langille, M.G.I. Predicting the Functional Potential of the Microbiome from Marker Genes Using PICRUSt. In Microbiome Analysis: Methods and Protocols; Beiko, R.G., Hsiao, W., Parkinson, J., Eds.; Springer: New York, NY, USA, 2018; pp. 169–177. [Google Scholar] [CrossRef]
- Shabbir, S.; Faheem, M.; Ali, N.; Kerr, P.G.; Wu, Y. Periphyton biofilms: A novel and natural biological system for the effective removal of sulphonated azo dye methyl orange by synergistic mechanism. Chemosphere 2017, 167, 236–246. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xie, Q.; Chen, S.; Xing, M.; Guan, T.; Wu, D. Inactivation of phosphorus in the sediment of the Lake Taihu by lanthanum modified zeolite using laboratory studies. Environ. Pollut. 2019, 247, 9–17. [Google Scholar] [CrossRef]
- Bere, T.; Chia, M.A.; Tundisi, J.G. Effects of Cr III and Pb on the bioaccumulation and toxicity of Cd in tropical periphyton communities: Implications of pulsed metal exposures. Environ. Pollut. 2012, 163, 184–191. [Google Scholar] [CrossRef]
- Sultana, M.-Y.; Akratos, C.S.; Pavlou, S.; Vayenas, D.V. Chromium removal in constructed wetlands: A review. Int. Biodeterior. Biodegrad. 2014, 96, 181–190. [Google Scholar] [CrossRef]
- Tiwari, S.; Patel, A.; Prasad, S.M. Kinetin alleviates chromium toxicity on growth and PS II photochemistry in Nostoc muscorum by regulating antioxidant system. Ecotoxicol. Environ. Saf. 2018, 161, 296–304. [Google Scholar] [CrossRef]
- Espinoza-Quiñones, F.; Silva, E.; Almeida Rizzutto, M.; Palácio, S.; Módenes, A.; Szymanski, N.; Martin, N.; Kroumov, A. Chromium ions phytoaccumulation by three floating aquatic macrophytes from a nutrient medium. World J. Microbiol. Biotechnol. 2008, 24, 3063–3070. [Google Scholar]
- Maine, M.A.; Suñe, N.; Hadad, H.; Sánchez, G.; Bonetto, C. Influence of vegetation on the removal of heavy metals and nutrients in a constructed wetland. J. Environ. Manag. 2009, 90, 355–363. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.-Z.; Chen, G.-J.; Bai, Y.-N.; Fu, L.; Qin, L.-P.; Zeng, R.J. Chromium isotope fractionation during Cr(VI) reduction in a methane-based hollow-fiber membrane biofilm reactor. Water Res. 2018, 130, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Bere, T.; Tundisi, J.G. Toxicity and sorption kinetics of dissolved cadmium and chromium III on tropical freshwater phytoperiphyton in laboratory mesocosm experiments. Sci. Total Environ. 2011, 409, 4772–4780. [Google Scholar] [CrossRef]
- Wnuk, E.; Walkiewicz, A.; Bieganowski, A. Methane oxidation in lead-contaminated mineral soils under different moisture levels. Environ. Sci. Pollut. Res. 2017, 24, 25346–25354. [Google Scholar] [CrossRef]
- Bowman, J.P.; Sly, L.I.; Hayward, A.C. Patterns of tolerance to heavy metals among methane-utilizing bacteria. Lett. Appl. Microbiol. 1990, 10, 85–87. [Google Scholar] [CrossRef]
- De Marco, P.; Pacheco, C.C.; Figueiredo, A.R.; Moradas-Ferreira, P. Novel pollutant-resistant methylotrophic bacteria for use in bioremediation. FEMS Microbiol. Lett. 2004, 234, 75–80. [Google Scholar] [CrossRef]
- Alam, M.A.; Wan, C.; Zhao, X.-Q.; Chen, L.-J.; Chang, J.-S.; Bai, F.-W. Enhanced removal of Zn2+ or Cd2+ by the flocculating Chlorella vulgaris JSC-7. J. Hazard. Mater. 2015, 289, 38–45. [Google Scholar] [CrossRef]
- Pandey, V.C.; Singh, J.S.; Singh, D.P.; Singh, R.P. Methanotrophs: Promising bacteria for environmental remediation. Int. J. Environ. Sci. Technol. 2014, 11, 241–250. [Google Scholar] [CrossRef]
- Choi, D.W.; Do, Y.S.; Zea, C.J.; McEllistrem, M.T.; Lee, S.-W.; Semrau, J.D.; Pohl, N.L.; Kisting, C.J.; Scardino, L.L.; Hartsel, S.C. Spectral and thermodynamic properties of Ag (I), Au (III), Cd (II), Co (II), Fe (III), Hg (II), Mn (II), Ni (II), Pb (II), U (IV), and Zn (II) binding by methanobactin from Methylosinus trichosporium OB3b. J. Inorg. Biochem. 2006, 100, 2150–2161. [Google Scholar] [CrossRef]
- Tsutsumi, M.; Kojima, H.; Fukui, M. Vertical profiles of abundance and potential activity of methane-oxidizing bacteria in sediment of Lake Biwa, Japan. Microbes Environ. 2012, 27, 67–71. [Google Scholar] [CrossRef]
- Mayr, M.J.; Zimmermann, M.; Guggenheim, C.; Brand, A.; Bürgmann, H. Niche partitioning of methane-oxidizing bacteria along the oxygen–methane counter gradient of stratified lakes. ISME J. 2020, 14, 274–287. [Google Scholar] [CrossRef] [PubMed]
- Van Grinsven, S.; Sinninghe Damsté, J.S.; Abdala Asbun, A.; Engelmann, J.C.; Harrison, J.; Villanueva, L. Methane oxidation in anoxic lake water stimulated by nitrate and sulfate addition. Environ. Microbiol. 2019, 22, 766–782. [Google Scholar] [CrossRef]
- Xavier, J.C.; Costa, P.E.S.; Hissa, D.C.; Melo, V.M.M.; Falcão, R.M.; Balbino, V.Q.; Mendonça, L.A.R.; Lima, M.G.S.; Coutinho, H.D.M.; Verde, L.C.L. Evaluation of the microbial diversity and heavy metal resistance genes of a microbial community on contaminated environment. Appl. Geochem. 2019, 105, 1–6. [Google Scholar] [CrossRef]
- Su, J.F.; Zhang, H.; Huang, T.L.; Wei, L.; Li, M.; Wang, Z. A new process for simultaneous nitrogen and cadmium(Cd(II)) removal using iron-reducing bacterial immobilization system. Chem. Eng. Process.—Process Intensif. 2019, 144, 107623. [Google Scholar] [CrossRef]
- Zhang, B.; Zhang, J.; Liu, Y.; Shi, P.; Wei, G. Co-occurrence patterns of soybean rhizosphere microbiome at a continental scale. Soil Biol. Biochem. 2018, 118, 178–186. [Google Scholar] [CrossRef]
- Wu, H.; Li, Y.; Zhang, W.; Wang, C.; Wang, P.; Niu, L.; Du, J.; Gao, Y. Bacterial community composition and function shift with the aggravation of water quality in a heavily polluted river. J. Environ. Manag. 2019, 237, 433–441. [Google Scholar] [CrossRef]
- Radková, A.B. The role of secondary oxides in potentially toxic elements migration. Int. Multidiscip. Sci. GeoConf. SGEM 2019, 19, 751–758. [Google Scholar]
- Zolgharnein, H.; Karami, K.; Assadi, M.M.; Sohrab, A.D. Molecular characterization and phylogenetic analyses of heavy metal removal bacteria from the Persian Gulf. Biotechnology 2010, 9, 1–8. [Google Scholar]
- Chen, L.-X.; Li, J.-T.; Chen, Y.-T.; Huang, L.-N.; Hua, Z.-S.; Hu, M.; Shu, W.-S. Shifts in microbial community composition and function in the acidification of a lead/zinc mine tailings. Environ. Microbiol. 2013, 15, 2431–2444. [Google Scholar] [CrossRef]
- Vorobev, A.; Jagadevan, S.; Baral, B.S.; DiSpirito, A.A.; Freemeier, B.C.; Bergman, B.H.; Bandow, N.L.; Semrau, J.D. Detoxification of mercury by methanobactin from Methylosinus trichosporium OB3b. Appl. Environ. Microbiol. 2013, 79, 5918–5926. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Faheem, M.; Shabbir, S.; Zhao, J.; Kerr, P.G.; Sultana, N.; Jia, Z. Enhanced Adsorptive Bioremediation of Heavy Metals (Cd2+, Cr6+, Pb2+) by Methane-Oxidizing Epipelon. Microorganisms 2020, 8, 505. https://doi.org/10.3390/microorganisms8040505
Faheem M, Shabbir S, Zhao J, Kerr PG, Sultana N, Jia Z. Enhanced Adsorptive Bioremediation of Heavy Metals (Cd2+, Cr6+, Pb2+) by Methane-Oxidizing Epipelon. Microorganisms. 2020; 8(4):505. https://doi.org/10.3390/microorganisms8040505
Chicago/Turabian StyleFaheem, Muhammad, Sadaf Shabbir, Jun Zhao, Philip G. Kerr, Nasrin Sultana, and Zhongjun Jia. 2020. "Enhanced Adsorptive Bioremediation of Heavy Metals (Cd2+, Cr6+, Pb2+) by Methane-Oxidizing Epipelon" Microorganisms 8, no. 4: 505. https://doi.org/10.3390/microorganisms8040505
APA StyleFaheem, M., Shabbir, S., Zhao, J., Kerr, P. G., Sultana, N., & Jia, Z. (2020). Enhanced Adsorptive Bioremediation of Heavy Metals (Cd2+, Cr6+, Pb2+) by Methane-Oxidizing Epipelon. Microorganisms, 8(4), 505. https://doi.org/10.3390/microorganisms8040505