Outer Membrane Vesicles Derived from Klebsiella pneumoniae Influence the miRNA Expression Profile in Human Bronchial Epithelial BEAS-2B Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Cell Culture
2.2. OMV Purification
2.3. OMV Characterization
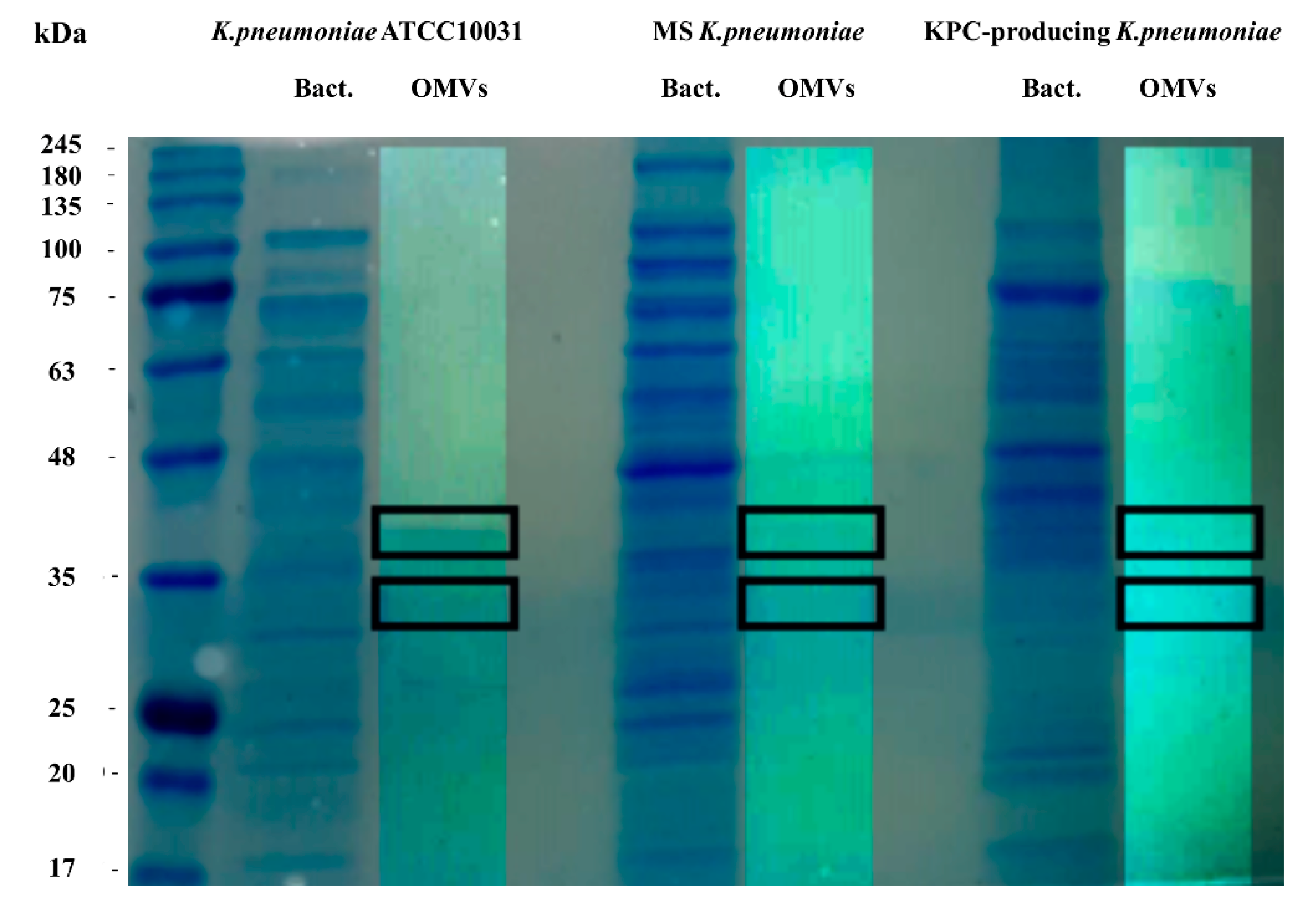
2.4. Sodium Dodecyl Sulphate Poly-Acrylamide Gel Electrophoresis (SDS-PAGE)
2.5. Proteomic Analysis of OMVs and Protein Extraction
2.6. Cellular Exposure to OMVs
2.7. RNA Extraction and miRNome Profiling
2.8. Statistical Analyses
2.9. Prediction and Function of miRNA Target Genes
2.10. miRNA Expression Analysis
3. Results
3.1. Characterization of K. pneumoniae-Derived OMVs
3.2. SDS-PAGE and LC-MS/MS Analysis of OMVs
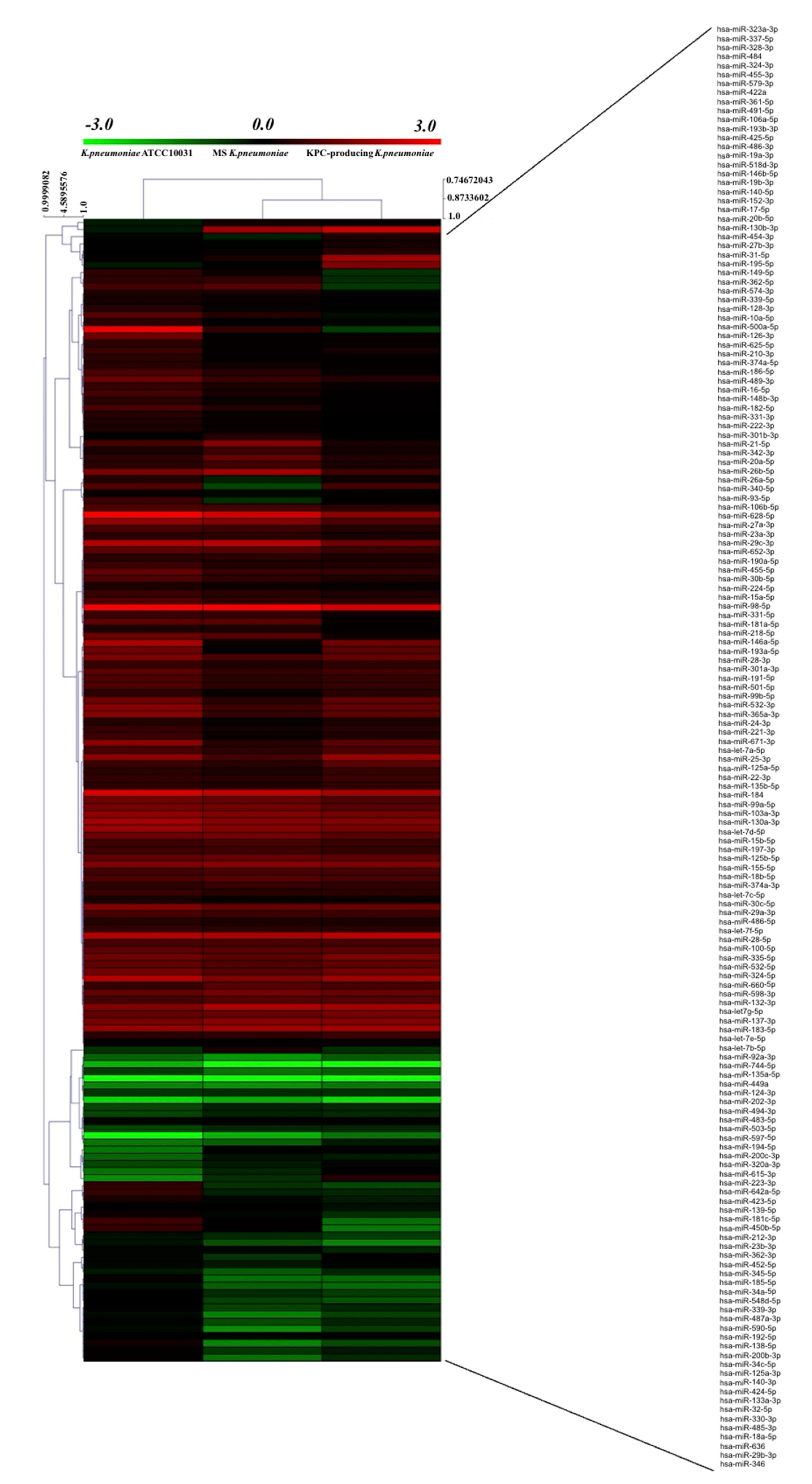
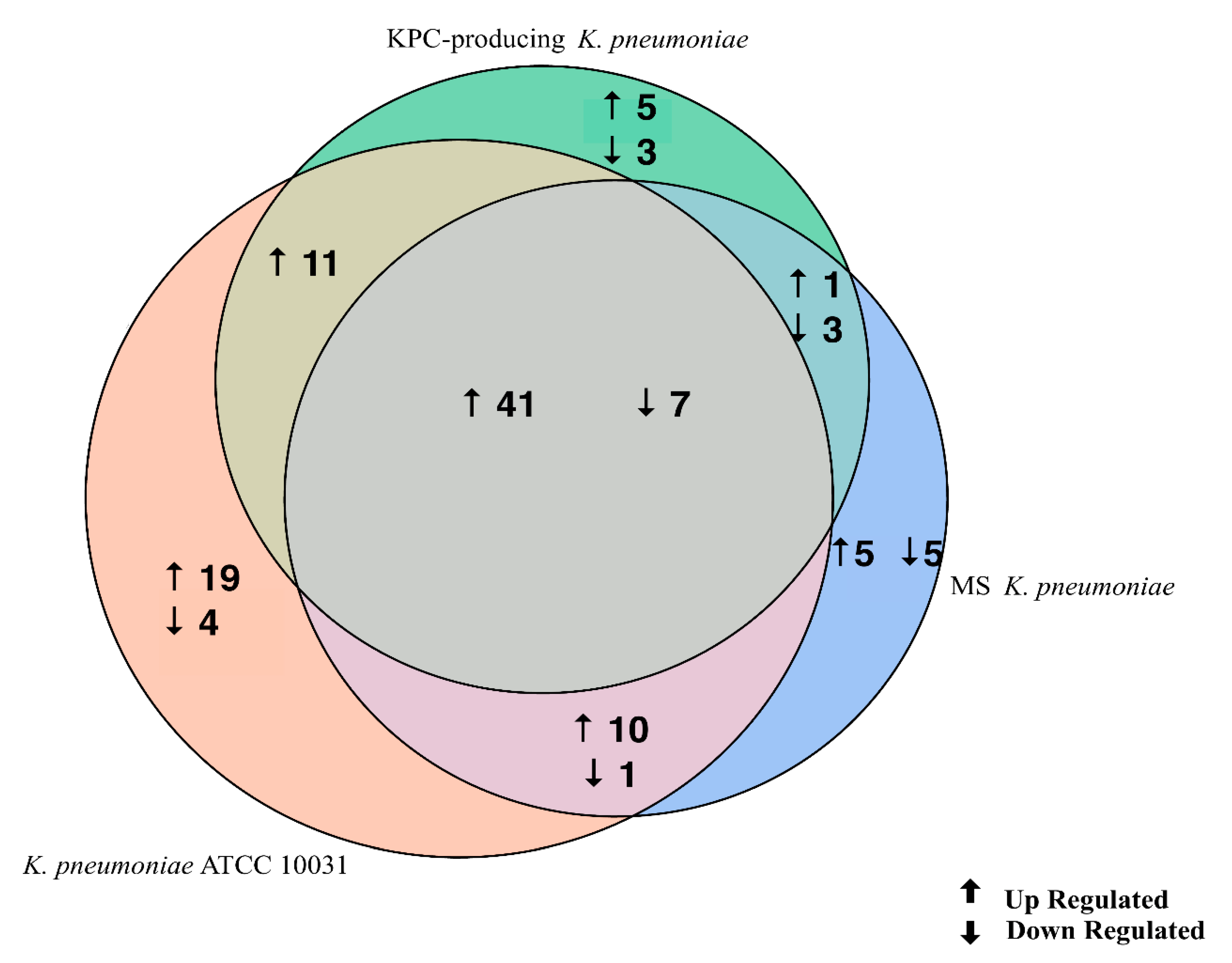
3.3. K. pneumoniae-Derived OMVs Affect miRNA Expression Profile in BEAS-2B Cells
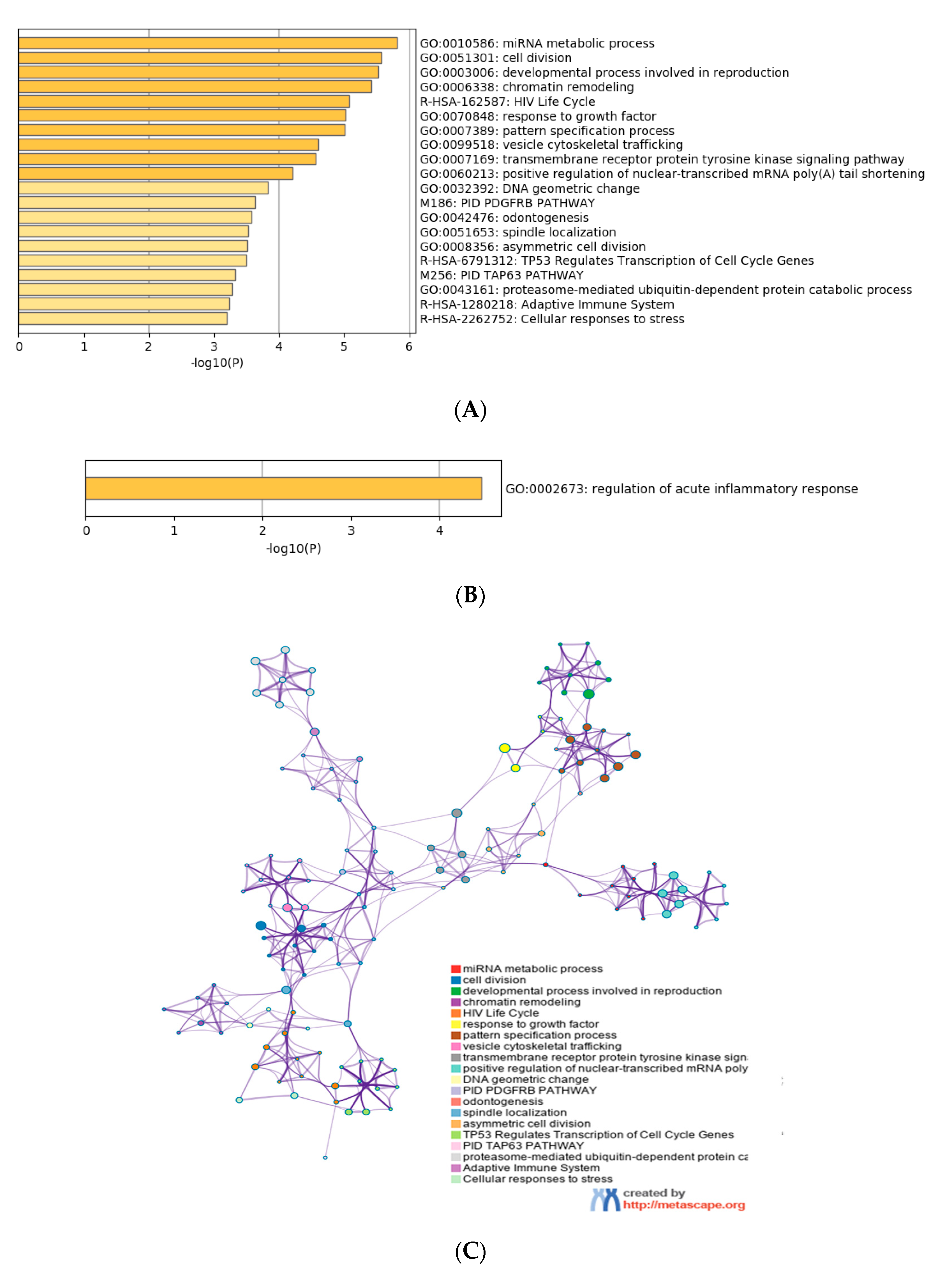
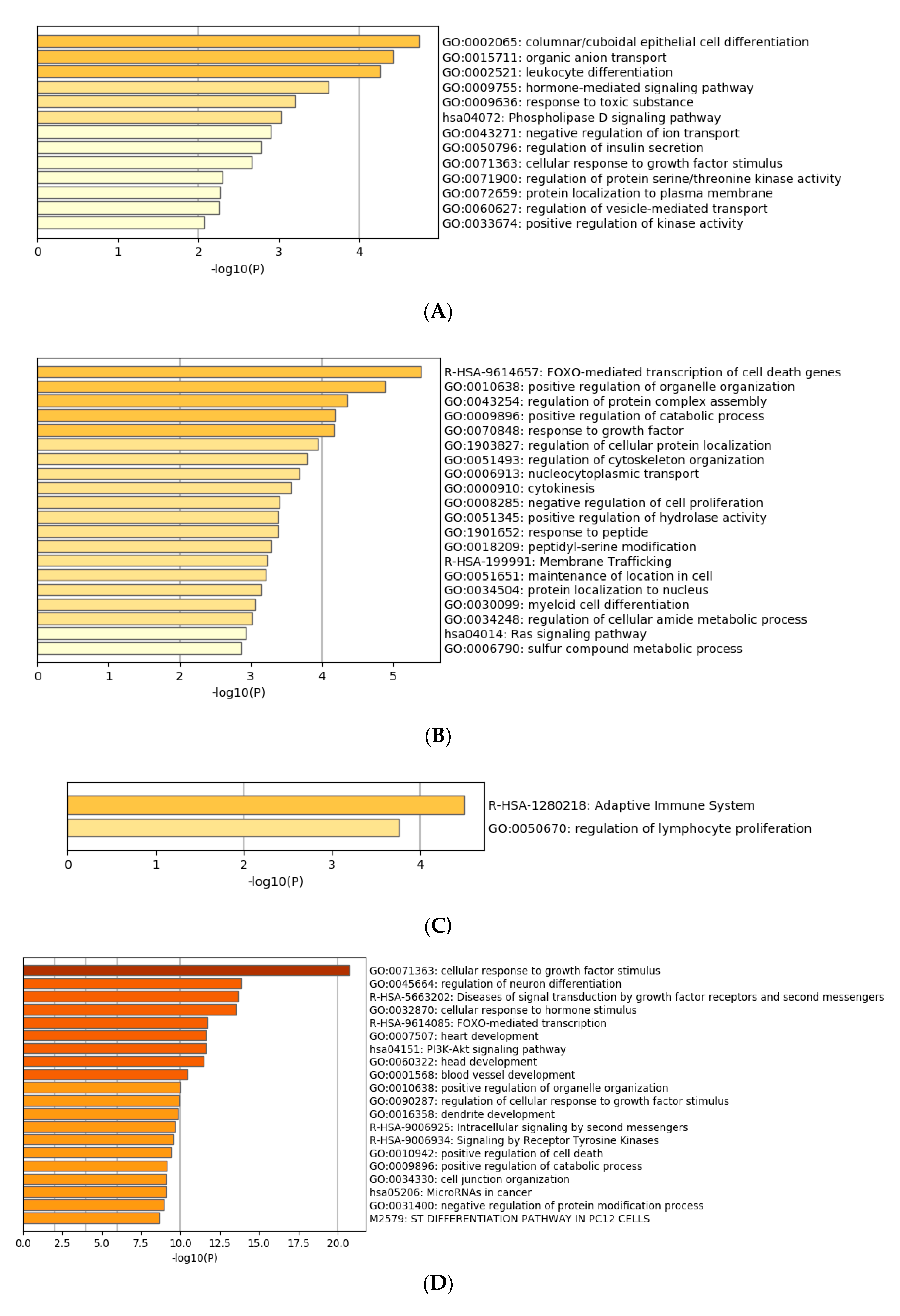
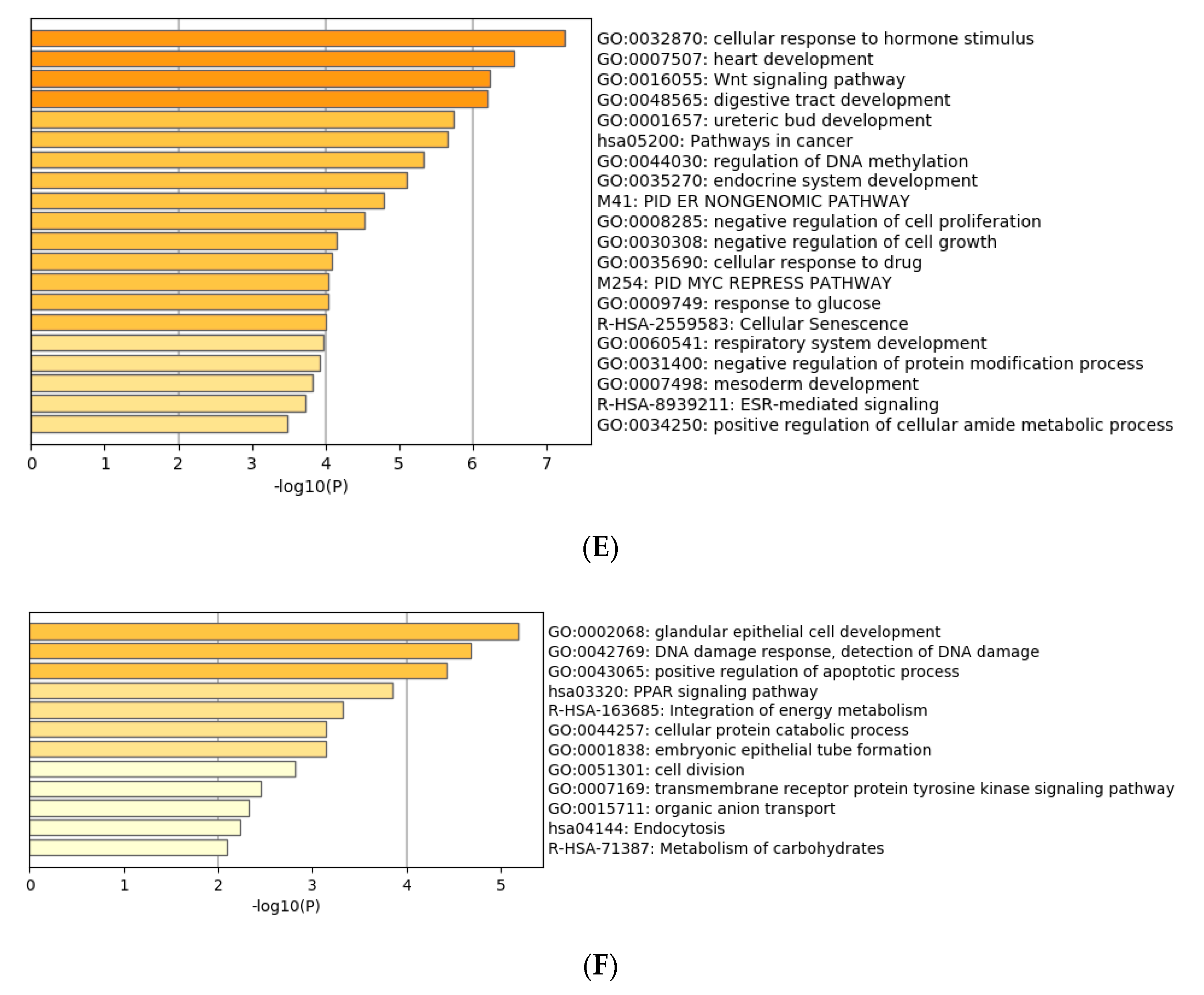
3.4. Functional Characterization of Target Genes
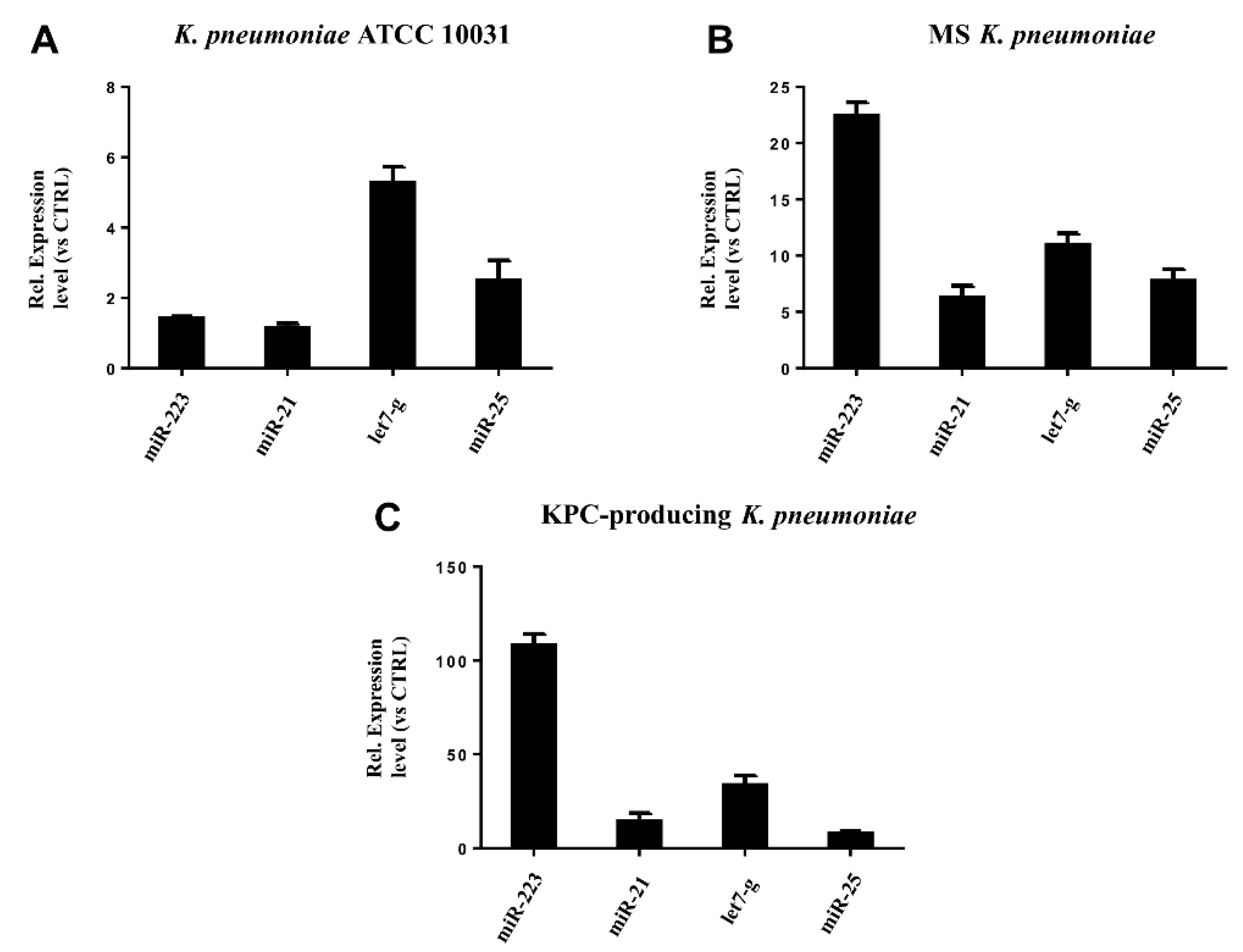
3.5. miRNAs Validation
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Caneiras, C.; Lito, L.; Melo-Cristino, J.; Duarte, A. Community- and Hospital-Acquired Klebsiella pneumoniae Urinary Tract Infections in Portugal: Virulence and Antibiotic Resistance. Microorganisms 2019, 7, 138. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.R.; Lee, J.H.; Park, K.S.; Jeon, J.H.; Kim, Y.B.; Cha, C.J.; Jeong, B.C.; Lee, S.H. Antimicrobial Resistance of Hypervirulent Klebsiella pneumoniae: Epidemiology, Hypervirulence-Associated Determinants, and Resistance Mechanisms. Front. Cell. Infect. Microbiol. 2017, 7, 483. [Google Scholar] [CrossRef] [PubMed]
- Podschun, R.; Ullmann, U. Klebsiella spp. as nosocomial pathogens: Epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 1998, 11, 589–603. [Google Scholar] [CrossRef] [PubMed]
- Bengoechea, J.A.; Sa Pessoa, J. Klebsiella pneumoniae infection biology: Living to counteract host defences. FEMS Microbiol. Rev. 2019, 43, 123–144. [Google Scholar] [CrossRef] [PubMed]
- Paczosa, M.K.; Mecsas, J. Klebsiella pneumoniae: Going on the Offense with a Strong Defense. Microbiol. Mol. Biol. Rev. 2016, 80, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.E.; Valentino, M.; Marszalowicz, G.; Magee, M.S.; Li, P.; Snook, A.E.; Stoecker, B.A.; Chang, C.; Waldman, S.A. Bacterial heat-stable enterotoxins: Translation of pathogenic peptides into novel targeted diagnostics and therapeutics. Toxins 2010, 2, 2028–2054. [Google Scholar] [CrossRef]
- Wang, H.; Zhong, Z.; Luo, Y.; Cox, E.; Devriendt, B. Heat-Stable Enterotoxins of Enterotoxigenic Escherichia coli and Their Impact on Host Immunity. Toxins 2019, 11, 24. [Google Scholar] [CrossRef]
- Buonocore, C.; Tedesco, P.; Vitale, G.A.; Esposito, F.P.; Giugliano, R.; Monti, M.C.; D’Auria, M.V.; de Pascale, D. Characterization of a New Mixture of Mono-Rhamnolipids Produced by Pseudomonas gessardii Isolated from Edmonson Point (Antarctica). Mar. Drugs 2020, 18, 269. [Google Scholar] [CrossRef]
- Morrill, H.J.; Pogue, J.M.; Kaye, K.S.; LaPlante, K.L. Treatment Options for Carbapenem-Resistant Enterobacteriaceae Infections. Open Forum Infect. Dis. 2015, 2, ofv050. [Google Scholar] [CrossRef]
- Nordmann, P.; Poirel, L. Epidemiology and Diagnostics of Carbapenem Resistance in Gram-negative Bacteria. Clin. Infect. Dis. 2019, 69, S521–S528. [Google Scholar] [CrossRef]
- Qureshi, Z.A.; Syed, A.; Clarke, L.G.; Doi, Y.; Shields, R.K. Epidemiology and clinical outcomes of patients with carbapenem-resistant Klebsiella pneumoniae bacteriuria. Antimicrob. Agents Chemother. 2014, 58, 3100–3104. [Google Scholar] [CrossRef] [PubMed]
- Geng, T.T.; Xu, X.; Huang, M. High-dose tigecycline for the treatment of nosocomial carbapenem-resistant Klebsiella pneumoniae bloodstream infections: A retrospective cohort study. Medicine 2018, 97, e9961. [Google Scholar] [CrossRef] [PubMed]
- Martora, F.; Pinto, F.; Folliero, V.; Cammarota, M.; Dell’Annunziata, F.; Squillaci, G.; Galdiero, M.; Morana, A.; Schiraldi, C.; Giovane, A.; et al. Isolation, characterization and analysis of pro-inflammatory potential of Klebsiella pneumoniae outer membrane vesicles. Microb. Pathog. 2019, 136, 103719. [Google Scholar] [CrossRef] [PubMed]
- Toyofuku, M.; Nomura, N.; Eberl, L. Types and origins of bacterial membrane vesicles. Nat. Rev. Microbiol. 2019, 17, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Gao, J.; Wang, Z. Outer membrane vesicles for vaccination and targeted drug delivery. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2019, 11, e1523. [Google Scholar] [CrossRef]
- Kohl, P.; Zingl, F.G.; Eichmann, T.O.; Schild, S. Isolation of Outer Membrane Vesicles Including Their Quantitative and Qualitative Analyses. Methods Mol. Biol. 2018, 1839, 117–134. [Google Scholar] [CrossRef] [PubMed]
- Schwechheimer, C.; Kuehn, M.J. Outer-membrane vesicles from Gram-negative bacteria: Biogenesis and functions. Nat. Rev. Microbiol. 2015, 13, 605–619. [Google Scholar] [CrossRef]
- Zavan, L.; Bitto, N.J.; Johnston, E.L.; Greening, D.W.; Kaparakis-Liaskos, M. Helicobacter pylori Growth Stage Determines the Size, Protein Composition, and Preferential Cargo Packaging of Outer Membrane Vesicles. Proteomics 2019, 19, e1800209. [Google Scholar] [CrossRef]
- Gerritzen, M.J.H.; Martens, D.E.; Wijffels, R.H.; van der Pol, L.; Stork, M. Bioengineering bacterial outer membrane vesicles as vaccine platform. Biotechnol. Adv. 2017, 35, 565–574. [Google Scholar] [CrossRef]
- Lynch, J.B.; Schwartzman, J.A.; Bennett, B.D.; McAnulty, S.J.; Knop, M.; Nyholm, S.V.; Ruby, E.G. Ambient pH Alters the Protein Content of Outer Membrane Vesicles, Driving Host Development in a Beneficial Symbiosis. J. Bacteriol. 2019, 201, e00319-19. [Google Scholar] [CrossRef]
- Ellis, T.N.; Kuehn, M.J. Virulence and immunomodulatory roles of bacterial outer membrane vesicles. Microbiol. Mol. Biol. Rev. 2010, 74, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Mandujano, A.; Hernandez-Cortez, C.; Ibarra, J.A.; Castro-Escarpulli, G. The outer membrane vesicles: Secretion system type zero. Traffic 2017, 18, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Langlete, P.; Krabberod, A.K.; Winther-Larsen, H.C. Vesicles From Vibrio cholerae Contain AT-Rich DNA and Shorter mRNAs That Do Not Correlate With Their Protein Products. Front. Microbiol. 2019, 10, 2708. [Google Scholar] [CrossRef] [PubMed]
- Bauwens, A.; Kunsmann, L.; Marejkova, M.; Zhang, W.; Karch, H.; Bielaszewska, M.; Mellmann, A. Intrahost milieu modulates production of outer membrane vesicles, vesicle-associated Shiga toxin 2a and cytotoxicity in Escherichia coli O157:H7 and O104:H4. Environ. Microbiol. Rep. 2017, 9, 626–634. [Google Scholar] [CrossRef]
- Jan, A.T. Outer Membrane Vesicles (OMVs) of Gram-negative Bacteria: A Perspective Update. Front. Microbiol. 2017, 8, 1053. [Google Scholar] [CrossRef]
- Li, M.; Zhou, H.; Yang, C.; Wu, Y.; Zhou, X.; Liu, H.; Wang, Y. Bacterial outer membrane vesicles as a platform for biomedical applications: An update. J. Control. Release 2020, 323, 253–268. [Google Scholar] [CrossRef]
- Bomberger, J.M.; Maceachran, D.P.; Coutermarsh, B.A.; Ye, S.; O’Toole, G.A.; Stanton, B.A. Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog. 2009, 5, e1000382. [Google Scholar] [CrossRef]
- Cooke, A.C.; Nello, A.V.; Ernst, R.K.; Schertzer, J.W. Analysis of Pseudomonas aeruginosa biofilm membrane vesicles supports multiple mechanisms of biogenesis. PLoS ONE 2019, 14, e0212275. [Google Scholar] [CrossRef]
- Koeppen, K.; Barnaby, R.; Jackson, A.A.; Gerber, S.A.; Hogan, D.A.; Stanton, B.A. Tobramycin reduces key virulence determinants in the proteome of Pseudomonas aeruginosa outer membrane vesicles. PLoS ONE 2019, 14, e0211290. [Google Scholar] [CrossRef]
- Fiocca, R.; Necchi, V.; Sommi, P.; Ricci, V.; Telford, J.; Cover, T.L.; Solcia, E. Release of Helicobacter pylori vacuolating cytotoxin by both a specific secretion pathway and budding of outer membrane vesicles. Uptake of released toxin and vesicles by gastric epithelium. J. Pathol. 1999, 188, 220–226. [Google Scholar] [CrossRef]
- Ronci, M.; Del Prete, S.; Puca, V.; Carradori, S.; Carginale, V.; Muraro, R.; Mincione, G.; Aceto, A.; Sisto, F.; Supuran, C.T.; et al. Identification and characterization of the alpha-CA in the outer membrane vesicles produced by Helicobacter pylori. J. Enzym. Inhib. Med. Chem. 2019, 34, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Cecil, J.D.; Sirisaengtaksin, N.; O’Brien-Simpson, N.M.; Krachler, A.M. Outer Membrane Vesicle-Host Cell Interactions. Microbiol. Spectr. 2019, 7, 201–214. [Google Scholar] [CrossRef]
- Atkins, D.; Furuta, G.T. Mucosal immunology, eosinophilic esophagitis, and other intestinal inflammatory diseases. J. Allergy Clin. Immunol. 2010, 125, S255–S261. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.K.; Guo, M.K.; Bai, X.H.; Chen, L.Q.; Su, S.M.; Li, L.; Li, J.Q. Astragaloside IV ameliorates intermittent hypoxia-induced inflammatory dysfunction by suppressing MAPK/NF-kappaB signalling pathways in Beas-2B cells. Sleep Breath. 2020, 24, 1237–1245. [Google Scholar] [CrossRef] [PubMed]
- Clark, H.R.; Powell, A.B.; Simmons, K.A.; Ayubi, T.; Kale, S.D. Endocytic Markers Associated with the Internalization and Processing of Aspergillus fumigatus Conidia by BEAS-2B Cells. mSphere 2019, 4. [Google Scholar] [CrossRef] [PubMed]
- Giudice, A.; D’Arena, G.; Crispo, A.; Tecce, M.F.; Nocerino, F.; Grimaldi, M.; Rotondo, E.; D’Ursi, A.M.; Scrima, M.; Galdiero, M.; et al. Role of Viral miRNAs and Epigenetic Modifications in Epstein-Barr Virus-Associated Gastric Carcinogenesis. Oxid Med. Cell. Longev. 2016, 2016, 6021934. [Google Scholar] [CrossRef]
- Powell, C.D.; Quain, D.E.; Smart, K.A. The Impact of Ageing on CHITIN Scars in Saccharomyces cerevisiae. Sci. World J. 2001, 1, 145. [Google Scholar] [CrossRef]
- Mens, M.M.J.; Ghanbari, M. Cell Cycle Regulation of Stem Cells by MicroRNAs. Stem. Cell. Rev. Rep. 2018, 14, 309–322. [Google Scholar] [CrossRef]
- Jung, N.; Schenten, V.; Bueb, J.L.; Tolle, F.; Brechard, S. miRNAs Regulate Cytokine Secretion Induced by Phosphorylated S100A8/A9 in Neutrophils. Int. J. Mol. Sci. 2019, 20, 5699. [Google Scholar] [CrossRef]
- McCoy, C.E. The role of miRNAs in cytokine signaling. Front. Biosci. 2011, 16, 2161–2171. [Google Scholar] [CrossRef]
- Nejad, C.; Stunden, H.J.; Gantier, M.P. A guide to miRNAs in inflammation and innate immune responses. FEBS J. 2018, 285, 3695–3716. [Google Scholar] [CrossRef] [PubMed]
- Salvi, V.; Gianello, V.; Tiberio, L.; Sozzani, S.; Bosisio, D. Cytokine Targeting by miRNAs in Autoimmune Diseases. Front. Immunol. 2019, 10, 15. [Google Scholar] [CrossRef] [PubMed]
- Folliero, V.; Caputo, P.; Della Rocca, M.T.; Chianese, A.; Galdiero, M.; Iovene, M.R.; Hay, C.; Franci, G.; Galdiero, M. Prevalence and Antimicrobial Susceptibility Patterns of Bacterial Pathogens in Urinary Tract Infections in University Hospital of Campania "Luigi Vanvitelli" between 2017 and 2018. Antibiotics 2020, 9, 215. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.C.; Yu, H.R.; Hsu, T.Y.; Chen, I.L.; Huang, H.C.; Chang, J.C.; Yang, K.D. MicroRNA-142-3p and let-7g Negatively Regulates Augmented IL-6 Production in Neonatal Polymorphonuclear Leukocytes. Int. J. Biol. Sci. 2017, 13, 690–700. [Google Scholar] [CrossRef]
- Ilisso, C.P.; Delle Cave, D.; Mosca, L.; Pagano, M.; Coppola, A.; Mele, L.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. S-Adenosylmethionine regulates apoptosis and autophagy in MCF-7 breast cancer cells through the modulation of specific microRNAs. Cancer Cell Int. 2018, 18, 197. [Google Scholar] [CrossRef]
- Lee, J.C.; Lee, E.J.; Lee, J.H.; Jun, S.H.; Choi, C.W.; Kim, S.I.; Kang, S.S.; Hyun, S. Klebsiella pneumoniae secretes outer membrane vesicles that induce the innate immune response. FEMS Microbiol. Lett. 2012, 331, 17–24. [Google Scholar] [CrossRef]
- Cahill, B.K.; Seeley, K.W.; Gutel, D.; Ellis, T.N. Klebsiella pneumoniae O antigen loss alters the outer membrane protein composition and the selective packaging of proteins into secreted outer membrane vesicles. Microbiol. Res. 2015, 180, 1–10. [Google Scholar] [CrossRef]
- Turner, K.L.; Cahill, B.K.; Dilello, S.K.; Gutel, D.; Brunson, D.N.; Alberti, S.; Ellis, T.N. Porin Loss Impacts the Host Inflammatory Response to Outer Membrane Vesicles of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2015, 60, 1360–1369. [Google Scholar] [CrossRef]
- Yuan, X.; Berg, N.; Lee, J.W.; Le, T.T.; Neudecker, V.; Jing, N.; Eltzschig, H. MicroRNA miR-223 as regulator of innate immunity. J. Leukoc. Biol. 2018, 104, 515–524. [Google Scholar] [CrossRef]
- Wu, J.; Niu, P.; Zhao, Y.; Cheng, Y.; Chen, W.; Lin, L.; Lu, J.; Cheng, X.; Xu, Z. Impact of miR-223-3p and miR-2909 on inflammatory factors IL-6, IL-1ss, and TNF-alpha, and the TLR4/TLR2/NF-kappaB/STAT3 signaling pathway induced by lipopolysaccharide in human adipose stem cells. PLoS ONE 2019, 14, e0212063. [Google Scholar] [CrossRef]
- Hsieh, C.H.; Rau, C.S.; Jeng, J.C.; Chen, Y.C.; Lu, T.H.; Wu, C.J.; Wu, Y.C.; Tzeng, S.L.; Yang, J.C.S. Whole blood-derived microRNA signatures in mice exposed to lipopolysaccharides. J. Biomed. Sci. 2012, 19, 69. [Google Scholar] [CrossRef] [PubMed]
- Dang, C.P.; Leelahavanichkul, A. Over-expression of miR-223 induces M2 macrophage through glycolysis alteration and attenuates LPS-induced sepsis mouse model, the cell-based therapy in sepsis. PLoS ONE 2020, 15, e0236038. [Google Scholar] [CrossRef] [PubMed]
- Ajuyah, P.; Hill, M.; Ahadi, A.; Lu, J.; Hutvagner, G.; Tran, N. MicroRNA (miRNA)-to-miRNA Regulation of Programmed Cell Death 4 (PDCD4). Mol. Cell. Biol. 2019, 39, e00086-19. [Google Scholar] [CrossRef]
- Zhou, W.; Su, L.; Duan, X.; Chen, X.; Hays, A.; Upadhyayula, S.; Shivde, J.; Wang, H.; Li, Y.; Huang, D.; et al. MicroRNA-21 down-regulates inflammation and inhibits periodontitis. Mol. Immunol. 2018, 101, 608–614. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Li, Z.; Gao, C.; Chen, P.; Chen, J.; Liu, W.; Xiao, S.; Lu, H. miR-21 plays a pivotal role in gastric cancer pathogenesis and progression. Lab. Investig. 2008, 88, 1358–1366. [Google Scholar] [CrossRef] [PubMed]
- Mei, Z.; Chen, S.; Chen, C.; Xiao, B.; Li, F.; Wang, Y.; Tao, Z. Interleukin-23 Facilitates Thyroid Cancer Cell Migration and Invasion by Inhibiting SOCS4 Expression via MicroRNA-25. PLoS ONE 2015, 10, e0139456. [Google Scholar] [CrossRef]
- Schulte, L.N.; Eulalio, A.; Mollenkopf, A.J.; Reinhardt, R.; Vogel, J. Analysis of the host microRNA response to Salmonella uncovers the control of major cytokines by the let-7 family. EMBO J. 2011, 30, 1977–1989. [Google Scholar] [CrossRef]
| Target Name | GeneGlobe ID |
|---|---|
| RNU6-6P | MS00033740 |
| hsa-miR-223 | MS00009184 |
| hsa-miR-21 | MS00009079 |
| hsa-miR-25 | MS00003227 |
| hsa-let-7g | MS00008337 |
| Bacterial Strain | Z-Ave (d.nm) | PDI |
|---|---|---|
| K. pneumoniae ATCC 10031 | 273.3 ± 1.3 | 0.329 ± 0.021 |
| MS K. pneumoniae | 427.1 ± 0.9 | 0.417 ± 0.017 |
| KPC-producing K. pneumoniae | 483.3 ± 1.7 | 0.333 ± 0.132 |
| Bacterial Strain | Protein Concentration [mg/mL] |
|---|---|
| K. pneumoniae ATCC 10031 | 0.08 ± 0.06 |
| MS K. pneumoniae | 0.14 ± 0.03 |
| KPC-producing K. pneumoniae | 0.21 ± 0.01 |
| Protein | Function | Theoretical MW (Da) | Score |
|---|---|---|---|
| Outer membrane protein A | Action of colicins K and L | 37,152 | 1501 |
| Outer membrane porin C | Passive diffusion across the outer membrane | 39,639 | 1298 |
| Glyceraldehyde-3-phosphate dehydrogenase (Fragment) | Enzyme in glycolysis | 32,457 | 680 |
| Nucleoside-specific channel-forming protein | Receptor for colicin K | 33,486 | 348 |
| Malate dehydrogenase | Enzyme in Krebs cycle | 32,549 | 180 |
| Glucokinase | Enzyme in glycolysis | 34,756 | 106 |
| 2-dehydro-3-deoxyphosphooctonate aldolase | Enzyme in aminoacids synthesis | 31,033 | 97 |
| Aminomethyltransferase | enzyme in the glycine cleavage complex | 39,904 | 88 |
| L-threonine 3-dehydrogenase | Enzyme in L-threonine catabolism | 37,559 | 33 |
| Elongation factor Ts | Protein biosynthesis | 30,545 | 18 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dell’Annunziata, F.; Ilisso, C.P.; Dell’Aversana, C.; Greco, G.; Coppola, A.; Martora, F.; Dal Piaz, F.; Donadio, G.; Falanga, A.; Galdiero, M.; et al. Outer Membrane Vesicles Derived from Klebsiella pneumoniae Influence the miRNA Expression Profile in Human Bronchial Epithelial BEAS-2B Cells. Microorganisms 2020, 8, 1985. https://doi.org/10.3390/microorganisms8121985
Dell’Annunziata F, Ilisso CP, Dell’Aversana C, Greco G, Coppola A, Martora F, Dal Piaz F, Donadio G, Falanga A, Galdiero M, et al. Outer Membrane Vesicles Derived from Klebsiella pneumoniae Influence the miRNA Expression Profile in Human Bronchial Epithelial BEAS-2B Cells. Microorganisms. 2020; 8(12):1985. https://doi.org/10.3390/microorganisms8121985
Chicago/Turabian StyleDell’Annunziata, Federica, Concetta Paola Ilisso, Carmela Dell’Aversana, Giuseppe Greco, Alessandra Coppola, Francesca Martora, Fabrizio Dal Piaz, Giuliana Donadio, Annarita Falanga, Marilena Galdiero, and et al. 2020. "Outer Membrane Vesicles Derived from Klebsiella pneumoniae Influence the miRNA Expression Profile in Human Bronchial Epithelial BEAS-2B Cells" Microorganisms 8, no. 12: 1985. https://doi.org/10.3390/microorganisms8121985
APA StyleDell’Annunziata, F., Ilisso, C. P., Dell’Aversana, C., Greco, G., Coppola, A., Martora, F., Dal Piaz, F., Donadio, G., Falanga, A., Galdiero, M., Altucci, L., Galdiero, M., Porcelli, M., Folliero, V., & Franci, G. (2020). Outer Membrane Vesicles Derived from Klebsiella pneumoniae Influence the miRNA Expression Profile in Human Bronchial Epithelial BEAS-2B Cells. Microorganisms, 8(12), 1985. https://doi.org/10.3390/microorganisms8121985











