Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments
Abstract
1. Introduction
Environmental Factors Driving Microbial Life in Cold Marine Environments
2. Microalgae
2.1. Salinity
2.2. Light
2.3. Nutrient
2.4. pH
2.5. Temperature
2.6. -Omics Studies on Microalgae from Cold Environments
3. Bacteria
3.1. Physiological Responses to Environmental Stressors
3.2. -Omics Studies on Bacteria from Cold Environments
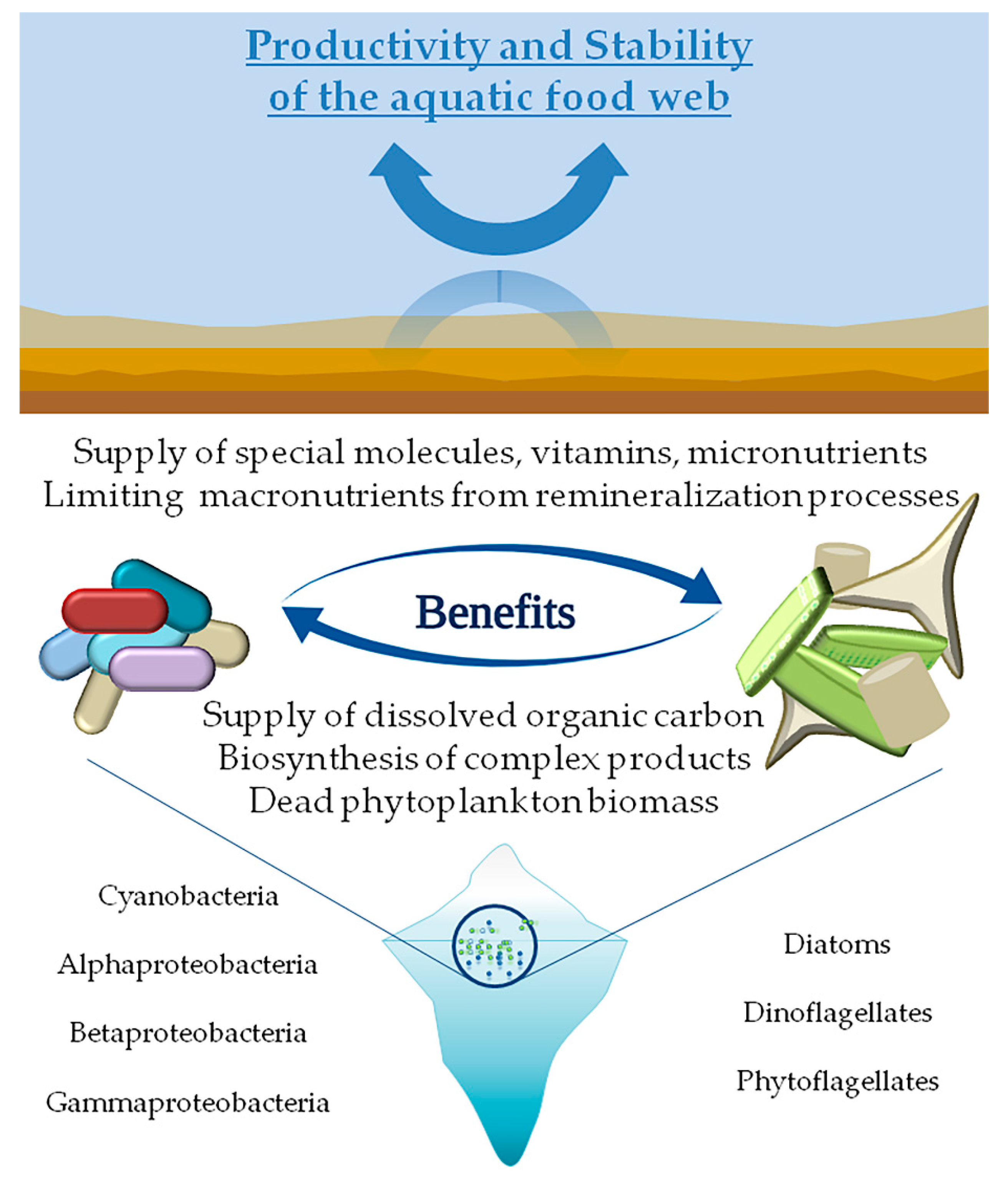
4. Phyto-Bacterioplankton Interactions in Cold Environments
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lauritano, C.; Ferrante, M.I.; Rogato, A. Marine Natural Products from Microalgae: An -Omics Overview. Mar. Drugs 2019, 17, 269. [Google Scholar] [CrossRef]
- Mocali, S.; Chiellini, C.; Fabiani, A.; Decuzzi, S.; de Pascale, D.; Parrilli, E.; Tutino, M.L.; Perrin, E.; Bosi, E.; Fondi, M.; et al. Ecology of cold environments: New insights of bacterial metabolic adaptation through an integrated genomic-phenomic approach. Sci. Rep. 2017, 7, 839. [Google Scholar] [CrossRef]
- Vitale, G.A.; Sciarretta, M.; Palma Esposito, F.; January, G.G.; Giaccio, M.; Bunk, B.; Spröer, C.; Bajerski, F.; Power, D.; Festa, C.; et al. Genomics–Metabolomics Profiling Disclosed Marine Vibrio spartinae 3.6 as a Producer of a New Branched Side Chain Prodigiosin. J. Nat. Prod. 2020, 83, 1495–1504. [Google Scholar] [CrossRef]
- Elagoz, A.M.; Ambrosino, L.; Lauritano, C. De novo transcriptome of the diatom Cylindrotheca closterium identifies genes involved in the metabolism of anti-inflammatory compounds. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef]
- Lauritano, C.; Ianora, A. Grand Challenges in Marine Biotechnology: Overview of Recent EU-Funded Projects. In Grand Challenges in Marine Biotechnology; Rampelotto, P.H., Trincone, A., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 425–449. ISBN 978-3-319-69074-2. [Google Scholar]
- Lauritano, C.; De Luca, D.; Ferrarini, A.; Avanzato, C.; Minio, A.; Esposito, F.; Ianora, A. De novo transcriptome of the cosmopolitan dinoflagellate Amphidinium carterae to identify enzymes with biotechnological potential. Sci. Rep. 2017, 7, 11701. [Google Scholar] [CrossRef]
- Lauritano, C.; De Luca, D.; Amoroso, M.; Benfatto, S.; Maestri, S.; Racioppi, C.; Esposito, F.; Ianora, A. New molecular insights on the response of the green alga Tetraselmis suecica to nitrogen starvation. Sci. Rep. 2019, 9, 3336. [Google Scholar] [CrossRef]
- Vingiani, G.M.; Štālberga, D.; De Luca, P.; Ianora, A.; De Luca, D.; Lauritano, C. De novo Transcriptome of the Non-saxitoxin Producing Alexandrium tamutum Reveals New Insights on Harmful Dinoflagellates. Mar. Drugs 2020, 18, 386. [Google Scholar] [CrossRef]
- Vingiani, G.M.; De Luca, P.; Ianora, A.; Dobson, A.D.W.; Lauritano, C. Microalgal Enzymes with Biotechnological Applications. Mar. Drugs 2019, 17, 459. [Google Scholar] [CrossRef]
- De Luca, D.; Lauritano, C. In Silico Identification of Type III PKS Chalcone and Stilbene Synthase Homologs in Marine Photosynthetic Organisms. Biology 2020, 9, 110. [Google Scholar] [CrossRef]
- Riccio, G.; De Luca, D.; Lauritano, C. Monogalactosyldiacylglycerol and Sulfolipid Synthesis in Microalgae. Mar. Drugs 2020, 18, 237. [Google Scholar] [CrossRef]
- Field, C.B.; Behrenfeld, M.J.; Randerson, J.T.; Falkowski, P. Primary Production of the Biosphere: Integrating Terrestrial and Oceanic Components. Science 1998, 281, 237–240. [Google Scholar] [CrossRef]
- Pomeroy, L.; leB Williams, P.; Azam, F.; Hobbie, J. The Microbial Loop. Oceanography 2007, 20, 28–33. [Google Scholar] [CrossRef]
- Hopes, A.; Thomas, D.N.; Mock, T. Polar Microalgae: Functional Genomics, Physiology, and the Environment. In Psychrophiles: From Biodiversity to Biotechnology; Margesin, R., Ed.; Springer International Publishing: Cham, Switzerland, 2017; pp. 305–344. ISBN 978-3-319-57057-0. [Google Scholar]
- Sarmento, H.; Gasol, J.M. Use of phytoplankton-derived dissolved organic carbon by different types of bacterioplankton. Environ. Microbiol. 2012, 14, 2348–2360. [Google Scholar] [CrossRef]
- Landa, M.; Blain, S.; Christaki, U.; Monchy, S.; Obernosterer, I. Shifts in bacterial community composition associated with increased carbon cycling in a mosaic of phytoplankton blooms. ISME J. 2016, 10, 39–50. [Google Scholar] [CrossRef]
- Seymour, J.R.; Amin, S.A.; Raina, J.-B.; Stocker, R. Zooming in on the phycosphere: The ecological interface for phytoplankton–bacteria relationships. Nat. Microbiol. 2017, 2, 17065. [Google Scholar] [CrossRef]
- Durham, B.P.; Sharma, S.; Luo, H.; Smith, C.B.; Amin, S.A.; Bender, S.J.; Dearth, S.P.; Van Mooy, B.A.S.; Campagna, S.R.; Kujawinski, E.B.; et al. Cryptic carbon and sulfur cycling between surface ocean plankton. Proc. Natl. Acad. Sci. USA 2015, 112, 453–457. [Google Scholar] [CrossRef]
- Amin, S.A.; Green, D.H.; Hart, M.C.; Küpper, F.C.; Sunda, W.G.; Carrano, C.J. Photolysis of iron–siderophore chelates promotes bacterial–algal mutualism. Proc. Natl. Acad. Sci. USA 2009, 106, 17071–17076. [Google Scholar] [CrossRef]
- Pearce, D.A. Extremophiles in Antarctica: Life at low temperatures. In Adaption of Microbial Life to Environmental Extremes: Novel Research Results and Application; Stan-Lotter, H., Fendrihan, S., Eds.; Springer: Vienna, Austria, 2012; pp. 87–118. ISBN 978-3-211-99691-1. [Google Scholar]
- Dinniman, M.S.; Klinck, J.M. A model study of circulation and cross-shelf exchange on the west Antarctic Peninsula continental shelf. Deep Sea Res. Part II Top. Stud. Oceanogr. 2004, 51, 2003–2022. [Google Scholar] [CrossRef]
- Vaughan, D.G.; Marshall, G.J.; Connolley, W.M.; Parkinson, C.; Mulvaney, R.; Hodgson, D.A.; King, J.C.; Pudsey, C.J.; Turner, J. Recent Rapid Regional Climate Warming on the Antarctic Peninsula. Clim. Chang. 2003, 60, 243–274. [Google Scholar] [CrossRef]
- Moline, M.A.; Claustre, H.; Frazer, T.K.; Schofield, O.; Vernet, M. Alteration of the food web along the Antarctic Peninsula in response to a regional warming trend. Glob. Chang. Biol. 2004, 10, 1973–1980. [Google Scholar] [CrossRef]
- Venables, H.J.; Clarke, A.; Meredith, M.P. Wintertime controls on summer stratification and productivity at the western Antarctic Peninsula. Limnol. Oceanogr. 2013, 58, 1035–1047. [Google Scholar] [CrossRef]
- Ducklow, H.; Fraser, W.; Meredith, M.; Stammerjohn, S.; Doney, S.; Martinson, D.; Sailley, S.; Schofield, O.; Steinberg, D.; Venables, H.; et al. West Antarctic Peninsula: An Ice-Dependent Coastal Marine Ecosystem in Transition. Oceanography 2013, 26, 190–203. [Google Scholar] [CrossRef]
- Hardge, K.; Peeken, I.; Neuhaus, S.; Lange, B.A.; Stock, A.; Stoeck, T.; Weinisch, L.; Metfies, K. The importance of sea ice for exchange of habitat-specific protist communities in the Central Arctic Ocean. J. Mar. Syst. 2017, 165, 124–138. [Google Scholar] [CrossRef]
- Torstensson, A.; Jiménez, C.; Nilsson, A.K.; Wulff, A. Elevated temperature and decreased salinity both affect the biochemical composition of the Antarctic sea-ice diatom Nitzschia lecointei, but not increased pCO2. Polar Biol. 2019, 42, 2149–2164. [Google Scholar] [CrossRef]
- Wiencke, C.; Clayton, M.N.; Gómez, I.; Iken, K.; Lüder, U.H.; Amsler, C.D.; Karsten, U.; Hanelt, D.; Bischof, K.; Dunton, K. Life strategy, ecophysiology and ecology of seaweeds in polar waters. Rev. Environ. Sci. Biotechnol. 2007, 6, 95–126. [Google Scholar] [CrossRef]
- Wulff, A.; Iken, K.; Quartino, M.L.; Al-Handal, A.; Wiencke, C.; Clayton, M.N. Biodiversity, biogeography and zonation of marine benthic micro- and macroalgae in the Arctic and Antarctic. Bot. Mar. 2009, 52, 491–507. [Google Scholar] [CrossRef]
- Glud, R.N.; Kühl, M.; Wenzhöfer, F.; Rysgaard, S. Benthic diatoms of a high Arctic fjord (Young Sound, NE Greenland): Importance for ecosystem primary production. Mar. Ecol. Prog. Ser. 2002, 238, 15–29. [Google Scholar] [CrossRef]
- Arrigo, K.R. Sea ice as a habitat for primary producers. In Sea Ice; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2016; pp. 352–369. ISBN 978-1-118-77837-1. [Google Scholar]
- Cota, G.F.; Legendre, L.; Gosselin, M.; Ingram, R.G. Ecology of bottom ice algae: I. Environmental controls and variability. J. Mar. Syst. 1991, 2, 257–277. [Google Scholar] [CrossRef]
- Mock, T.; Thomas, D.N. Microalgae in Polar Regions: Linking Functional Genomics and Physiology with Environmental Conditions. In Psychrophiles: From Biodiversity to Biotechnology; Margesin, R., Schinner, F., Marx, J.-C., Gerday, C., Eds.; Springer: Berlin/Heidelberg, Germany, 2008; pp. 285–312. ISBN 978-3-540-74335-4. [Google Scholar]
- Leventer, A. Sediment trap diatom assemblages from the northern Antarctic Peninsula region. Deep Sea Res. Part I Oceanogr. Res. Pap. 1991, 38, 1127–1143. [Google Scholar] [CrossRef]
- DiTullio, G.R.; Smith, W.O., Jr. Spatial patterns in phytoplankton biomass and pigment distributions in the Ross Sea. J. Geophys. Res. Ocean. 1996, 101, 18467–18477. [Google Scholar] [CrossRef]
- Peloquin, J.A.; Smith, W.O., Jr. Phytoplankton blooms in the Ross Sea, Antarctica: Interannual variability in magnitude, temporal patterns, and composition. J. Geophys. Res. Ocean. 2007, 112. [Google Scholar] [CrossRef]
- Saggiomo, M.; Poulin, M.; Mangoni, O.; Lazzara, L.; De Stefano, M.; Sarno, D.; Zingone, A. Spring-time dynamics of diatom communities in landfast and underlying platelet ice in Terra Nova Bay, Ross Sea, Antarctica. J. Mar. Syst. 2017, 166, 26–36. [Google Scholar] [CrossRef]
- Peterson, B.J.; Holmes, R.M.; McClelland, J.W.; Vörösmarty, C.J.; Lammers, R.B.; Shiklomanov, A.I.; Shiklomanov, I.A.; Rahmstorf, S. Increasing River Discharge to the Arctic Ocean. Science 2002, 298, 2171–2173. [Google Scholar] [CrossRef]
- Bluhm, B.A.; Gradinger, R. Regional variability in food availability for arctic marine mammals. Ecol. Appl. 2008, 18, S77–S96. [Google Scholar] [CrossRef]
- Pivovarov, S.; Schlitzer, R.; Novikhin, A. River run-off influence on the water mass formation in the Kara Sea. In Proceedings in Marine Science: Siberian River Run-Off in the Kara SEA: Characterisation, Quantification, Variability and Environmantal Significance; Stein, R., Fahl, K., Fütterer, D.K., Galimov, E., Eds.; Elsevier: Amsterdam, The Netherlands, 2003; pp. 9–26. [Google Scholar]
- Turner, J.; Overland, J.E.; Walsh, J.E. An Arctic and antarctic perspective on recent climate change. Int. J. Climatol. 2007, 27, 277–293. [Google Scholar] [CrossRef]
- Weeks, W.F.; Ackley, S.F. The Growth, Structure, and Properties of Sea Ice. In The Geophysics of Sea Ice; NATO ASI Series; Untersteiner, N., Ed.; Springer: Boston, MA, USA, 1986; pp. 9–164. ISBN 978-1-4899-5352-0. [Google Scholar]
- Ryan, K.G.; Ralph, P.; McMinn, A. Acclimation of Antarctic bottom-ice algal communities to lowered salinities during melting. Polar Biol. 2004, 27, 679–686. [Google Scholar] [CrossRef]
- Torstensson, A.; Fransson, A.; Currie, K.; Wulff, A.; Chierici, M. Microalgal photophysiology and macronutrient distribution in summer sea ice in the Amundsen and Ross Seas, Antarctica. PLoS ONE 2018, 13, e0195587. [Google Scholar] [CrossRef]
- Ducklow, H.; Clarke, A.; Dickhut, R.; Doney, S.; Geisz, H.; Huang, K.; Martinson, D.; Meredith, M.; Moeller, H.; Montes, M.; et al. The Marine Ecosystem of the West Antarctic Peninsula. In Antarctic Ecosystems: An Extreme Environment in a Changing World, 1st ed.; Rogers, A.D., Johnston, N.M., Murphy, E.J., Clarke, A., Eds.; Blackwell Publishing Ltd.: Hoboken, NJ, USA, 2012; pp. 121–159. [Google Scholar]
- Durham, B.P.; Boysen, A.K.; Carlson, L.T.; Groussman, R.D.; Heal, K.R.; Cain, K.R.; Morales, R.L.; Coesel, S.N.; Morris, R.M.; Ingalls, A.E.; et al. Sulfonate-based networks between eukaryotic phytoplankton and heterotrophic bacteria in the surface ocean. Nat. Microbiol. 2019, 4, 1706–1715. [Google Scholar] [CrossRef]
- Dawson, H.M.; Heal, K.R.; Boysen, A.K.; Carlson, L.T.; Ingalls, A.E.; Young, J.N. Potential of temperature-and salinity-driven shifts in diatom compatible solute concentrations to impact biogeochemical cycling within sea ice. Elem. Sci. Anthr. 2020, 8. [Google Scholar] [CrossRef]
- Shetty, P.; Gitau, M.M.; Maróti, G. Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae. Cells 2019, 8, 1657. [Google Scholar] [CrossRef]
- Alderkamp, A.-C.; Kulk, G.; Buma, A.G.J.; Visser, R.J.W.; Van Dijken, G.L.; Mills, M.M.; Arrigo, K.R. The effect of iron limitation on the photophysiology of Phaeocystis antarctica (Prymnesiophyceae) and Fragilariopsis cylindrus (Bacillariophyceae) under dynamic irradiance. J. Phycol. 2012, 48, 45–59. [Google Scholar] [CrossRef]
- Cassar, N.; DiFiore, P.; Barnett, B.; Bender, M.; Bowie, A.; Tilbrook, B.; Petrou, K.; Westwood, K.; Wright, S.; Lefevre, D. The influence of iron and light on net community production in the Subantarctic and Polar Frontal Zones. Biogeosciences 2011, 8. [Google Scholar] [CrossRef]
- Villafañe, V.E.; Sundback, K.; Lopez Figueroa, F.; Helbling, E.W. Photosynthesis in the aquatic environment as affected by UVR. In UV Effects in Aquatic Organisms and Ecosystems; Helbling, E.W., Zagarese, H., Eds.; Royal Society of Chemistry: London, UK, 2003; pp. 357–397. [Google Scholar]
- Goes, J.; Handa, N.; Suzuki, K. Ultraviolet radiation induced changes in the production of organic compounds in antarctic marine phytoplankton. Proc. NIPR Symp. Polar Biol. 1997, 10, 25–38. [Google Scholar]
- Russo, A.D.P.G.; de Souza, M.S.; Mendes, C.R.B.; Jesus, B.; Tavano, V.M.; Garcia, C.A.E. Photophysiological effects of Fe concentration gradients on diatom-dominated phytoplankton assemblages in the Antarctic Peninsula region. J. Exp. Mar. Biol. Ecol. 2015, 466, 49–58. [Google Scholar] [CrossRef]
- Duprat, L.; Corkill, M.; Genovese, C.; Townsend, A.T.; Moreau, S.; Meiners, K.M.; Lannuzel, D. Nutrient distribution in East Antarctic summer sea ice: A potential iron contribution from glacial basal melt. J. Geophys. Res. Ocean. 2020, e2020JC016130. [Google Scholar] [CrossRef]
- Lauritano, C.; Orefice, I.; Procaccini, G.; Romano, G.; Ianora, A. Key genes as stress indicators in the ubiquitous diatom Skeletonema marinoi. BMC Genom. 2015, 16, 411. [Google Scholar] [CrossRef]
- Suffrian, K.; Schulz, K.G.; Gutowska, M.A.; Riebesell, U.; Bleich, M. Cellular pH measurements in Emiliania huxleyi reveal pronounced membrane proton permeability. New Phytol. 2011, 190, 595–608. [Google Scholar] [CrossRef]
- Morgan-Kiss, R.M.; Priscu, J.C.; Pocock, T.; Gudynaite-Savitch, L.; Huner, N.P.A. Adaptation and acclimation of photosynthetic microorganisms to permanently cold environments. Microbiol. Mol. Biol. Rev. 2006, 70, 222–252. [Google Scholar] [CrossRef]
- Mock, T.; Otillar, R.P.; Strauss, J.; McMullan, M.; Paajanen, P.; Schmutz, J.; Salamov, A.; Sanges, R.; Toseland, A.; Ward, B.J.; et al. Evolutionary genomics of the cold-adapted diatom Fragilariopsis cylindrus. Nature 2017, 541, 536–540. [Google Scholar] [CrossRef]
- Mock, T.; Valentin, K. Photosynthesis and cold acclimation: Molecular evidence from a polar diatom. J. Phycol. 2004, 40, 732–741. [Google Scholar] [CrossRef]
- Krell, A.; Beszteri, B.; Dieckmann, G.; Glöckner, G.; Valentin, K.; Mock, T. A new class of ice-binding proteins discovered in a salt-stress-induced cDNA library of the psychrophilic diatom Fragilariopsis cylindrus (Bacillariophyceae). Eur. J. Phycol. 2008, 43, 423–433. [Google Scholar] [CrossRef]
- Liu, C.; Huang, X. Transcriptome-wide analysis of DEAD-box RNA helicase gene family in an Antarctic psychrophilic alga Chlamydomonas sp. ICE-L. Extremophiles 2015, 19, 921–931. [Google Scholar] [CrossRef]
- Liu, C.; Wang, X.; Wang, X.; Sun, C. Acclimation of Antarctic Chlamydomonas to the sea-ice environment: A transcriptomic analysis. Extremophiles 2016, 20, 437–450. [Google Scholar] [CrossRef]
- Hwang, Y.; Jung, G.; Jin, E. Transcriptome analysis of acclimatory responses to thermal stress in Antarctic algae. Biochem. Biophys. Res. Commun. 2008, 367, 635–641. [Google Scholar] [CrossRef]
- Park, S.; Jung, G.; Hwang, Y.; Jin, E. Dynamic response of the transcriptome of a psychrophilic diatom, Chaetoceros neogracile, to high irradiance. Planta 2009, 231, 349. [Google Scholar] [CrossRef]
- Gwak, I.G.; sic Jung, W.; Kim, H.J.; Kang, S.-H.; Jin, E. Antifreeze Protein in Antarctic Marine Diatom, Chaetoceros neogracile. Mar. Biotechnol. 2010, 12, 630–639. [Google Scholar] [CrossRef]
- Wang, N.; Qian, Z.; Luo, M.; Fan, S.; Zhang, X.; Zhang, L. Identification of salt stress responding genes using transcriptome analysis in green alga Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2018, 19, 3359. [Google Scholar] [CrossRef]
- Palmisano, A.C.; Beeler SooHoo, J.; Sullivan, C.W. Effects of four environmental variables on photosynthesis-irradiance relationships in Antarctic sea-ice microalgae. Mar. Biol. 1987, 94, 299–306. [Google Scholar] [CrossRef]
- Kirst, G.O.; Wiencke, C. Ecophysiology of polar algae. J. Phycol. 1995, 31, 181–199. [Google Scholar] [CrossRef]
- Torstensson, A.; Young, J.N.; Carlson, L.T.; Ingalls, A.E.; Deming, J.W. Use of exogenous glycine betaine and its precursor choline as osmoprotectants in Antarctic sea-ice diatoms1. J. Phycol. 2019, 55, 663–675. [Google Scholar] [CrossRef]
- Läuchli, A.; Lüttge, U. (Eds.) Salinity: Environment—Plants—Molecules; Springer: Dordrecht, The Netherlands, 2002; ISBN 978-1-4020-0492-6. [Google Scholar]
- Siegenthaler, U.; Sarmiento, J.L. Atmospheric carbon dioxide and the ocean. Nature 1993, 365, 119–125. [Google Scholar] [CrossRef]
- Häder, D.-P.; Sinha, R.P. Solar ultraviolet radiation-induced DNA damage in aquatic organisms: Potential environmental impact. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2005, 571, 221–233. [Google Scholar] [CrossRef]
- Wong, C.-Y.; Teoh, M.-L.; Phang, S.-M.; Lim, P.-E.; Beardall, J. Interactive Effects of Temperature and UV Radiation on photosynthesis of Chlorella strains from polar, temperate and tropical environments: Differential impacts on damage and repair. PLoS ONE 2015, 10, e0139469. [Google Scholar] [CrossRef]
- Van Leeuwe, M.; Tedesco, L.; Arrigo, K.R.; Assmy, P.; Campbell, K.; Meiners, K.M.; Rintala, J.-M.; Selz, V.; Thomas, D.N.; Stefels, J. Microalgal community structure and primary production in Arctic and Antarctic sea ice: A synthesis. Elem. Sci. Anthr. 2018, 6, 4. [Google Scholar] [CrossRef]
- Van Oijen, T.; van Leeuwe, M.A.; Granum, E.; Weissing, F.J.; Bellerby, R.G.J.; Gieskes, W.W.C.; de Baar, H.J.W. Light rather than iron controls photosynthate production and allocation in Southern Ocean phytoplankton populations during austral autumn. J. Plankton Res. 2004, 26, 885–900. [Google Scholar] [CrossRef]
- Falkowski, P.G. The role of phytoplankton photosynthesis in global biogeochemical cycles. Photosynth. Res. 1994, 39, 235–258. [Google Scholar] [CrossRef]
- McMinn, A.; Müller, M.N.; Martin, A.; Ryan, K.G. The response of antarctic sea ice algae to changes in pH and CO2. PLoS ONE 2014, 9, e86984. [Google Scholar] [CrossRef]
- Shadwick, E.H.; Trull, T.W.; Thomas, H.; Gibson, J.A.E. Vulnerability of polar oceans to anthropogenic acidification: Comparison of arctic and antarctic seasonal cycles. Sci. Rep. 2013, 3, 2339. [Google Scholar] [CrossRef]
- Rivaro, P.; Ianni, C.; Langone, L.; Ori, C.; Aulicino, G.; Cotroneo, Y.; Saggiomo, M.; Mangoni, O. Physical and biological forcing of mesoscale variability in the carbonate system of the Ross Sea (Antarctica) during summer 2014. J. Mar. Syst. 2017, 166, 144–158. [Google Scholar] [CrossRef]
- Reinfelder, J.R. Carbon concentrating mechanisms in eukaryotic marine phytoplankton. Ann. Rev. Mar. Sci. 2011, 3, 291–315. [Google Scholar] [CrossRef]
- Teoh, M.-L.; Phang, S.-M.; Chu, W.-L. Response of Antarctic, temperate, and tropical microalgae to temperature stress. J. Appl. Phycol. 2013, 25, 285–297. [Google Scholar] [CrossRef]
- Montes, M.; Doney, S.; Ducklow, H.; Fraser, W.; Martinson, D.; Stammerjohn, S.; Schofield, O. Recent changes in phytoplankton communities associated with rapid regional climate change along the Western Antarctic Peninsula. Science 2009, 323, 1470–1473. [Google Scholar] [CrossRef]
- Converti, A.; Casazza, A.; Ortiz, E.; Perego, P.; Del Borghi, A. Effect of temperature and nitrogen concentration on the growth and lipid content of Nannochloropsis oculata and Chlorella vulgaris for biodiesel production. Chem. Eng. Process 2009, 1146–1151. [Google Scholar] [CrossRef]
- Yun, M.; Lee, S.; Chung, I.K. Photosynthetic activity of benthic diatoms in response to different temperatures. J. Appl. Phycol. 2010, 22, 559–562. [Google Scholar] [CrossRef]
- Chong, G.-L.; Chu, W.-L.; Othman, R.Y.; Phang, S.-M. Differential gene expression of an Antarctic Chlorella in response to temperature stress. Polar Biol. 2011, 34, 637–645. [Google Scholar] [CrossRef]
- Hu, H.; Gao, K. Response of growth and fatty acid compositions of Nannochloropsis sp. to environmental factors under elevated CO2 concentration. Biotechnol. Lett. 2006, 28, 987–992. [Google Scholar] [CrossRef]
- Blanc, G.; Agarkova, I.; Grimwood, J.; Kuo, A.; Brueggeman, A.; Dunigan, D.D.; Gurnon, J.; Ladunga, I.; Lindquist, E.; Lucas, S.; et al. The genome of the polar eukaryotic microalga Coccomyxa subellipsoidea reveals traits of cold adaptation. Genome Biol. 2012, 13, R39. [Google Scholar] [CrossRef]
- Mock, T.; Krell, A.; Glöckner, G.; Kolukisaoglu, Ü.; Valentin, K. Analysis of expressed sequence tags (ESTs) from the polar diatom Fragilariopsis cylindrus. J. Phycol. 2006, 42, 78–85. [Google Scholar] [CrossRef]
- Cho, S.M.; Kim, S.; Cho, H.; Lee, H.; Lee, J.H.; Lee, H.; Park, H.; Kang, S.; Choi, H.-G.; Lee, J. Type II Ice-binding proteins isolated from an arctic microalga are similar to adhesin-like proteins and increase freezing tolerance in transgenic plants. Plant Cell Physiol. 2019, 60, 2744–2757. [Google Scholar] [CrossRef]
- Morita, R.Y. Psychrophilic bacteria. Bacteriol. Rev. 1975, 39, 144–167. [Google Scholar]
- Feller, G.; Gerday, C. Psychrophilic enzymes: Hot topics in cold adaptation. Nat. Rev. Microbiol. 2003, 1, 200–208. [Google Scholar] [CrossRef]
- Cavicchioli, R. On the concept of a psychrophile. ISME J. 2016, 10, 793–795. [Google Scholar] [CrossRef]
- Sul, W.J.; Oliver, T.A.; Ducklow, H.W.; Amaral-Zettler, L.A.; Sogin, M.L. Marine bacteria exhibit a bipolar distribution. Proc. Natl. Acad. Sci. USA 2013, 110, 2342–2347. [Google Scholar] [CrossRef]
- Sunagawa, S.; Coelho, L.P.; Chaffron, S.; Kultima, J.R.; Labadie, K.; Salazar, G.; Djahanschiri, B.; Zeller, G.; Mende, D.R.; Alberti, A.; et al. Structure and function of the global ocean microbiome. Science 2015, 348. [Google Scholar] [CrossRef]
- Verde, C.; Giordano, D.; Bellas, C.; di Prisco, G.; Anesio, A. Polar marine microorganisms and climate change. Adv. Microb. Physiol. 2016, 69, 187–215. [Google Scholar]
- Ghiglione, J.-F.; Murray, A. Pronounced summer to winter differences and higher wintertime richness in coastal Antarctic marine bacterioplankton. Environ. Microbiol. 2012, 14, 617–629. [Google Scholar] [CrossRef]
- Grzymski, J.J.; Riesenfeld, C.S.; Williams, T.J.; Dussaq, A.M.; Ducklow, H.; Erickson, M.; Cavicchioli, R.; Murray, A.E. A metagenomic assessment of winter and summer bacterioplankton from Antarctica Peninsula coastal surface waters. ISME J. 2012, 6, 1901–1915. [Google Scholar] [CrossRef]
- Williams, T.J.; Wilkins, D.; Long, E.; Evans, F.; DeMaere, M.Z.; Raftery, M.J.; Cavicchioli, R. The role of planktonic Flavobacteria in processing algal organic matter in coastal East Antarctica revealed using metagenomics and metaproteomics. Environ. Microbiol. 2013, 15, 1302–1317. [Google Scholar] [CrossRef]
- Kirchman, D.L.; Cottrell, M.T.; Lovejoy, C. The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ. Microbiol. 2010, 12, 1132–1143. [Google Scholar] [CrossRef]
- Kellogg, C.; Deming, J. Comparison of free-living, suspended particle, and aggregate-associated Bacterial and Archaeal communities in the Laptev Sea. Aquat. Microb. Ecol. 2009, 57, 1–18. [Google Scholar] [CrossRef]
- Malmstrom, R.; Straza, T.; Cottrell, M.T.; Kirchman, D. Diversity, abundance, and biomass production of bacterial groups in the western Arctic Ocean. Aquat. Microb. Ecol. 2007, 47, 45–55. [Google Scholar] [CrossRef]
- Collins, R.E.; Rocap, G.; Deming, J.W. Persistence of bacterial and archaeal communities in sea ice through an Arctic winter. Environ. Microbiol. 2010, 12, 1828–1841. [Google Scholar] [CrossRef]
- Galand, P.; Lovejoy, C.; Pouliot, J.; Garneau, M.-È.; Vincent, W. Microbial community diversity and heterotrophic production in a coastal Arctic ecosystem: A stamukhi lake and its source waters. Limnol. Oceanogr. 2008, 53, 813–823. [Google Scholar] [CrossRef]
- Han, D.; Kang, I.; Ha, H.K.; Kim, H.C.; Kim, O.-S.; Lee, B.Y.; Cho, J.-C.; Hur, H.-G.; Lee, Y.K. Bacterial communities of surface mixed layer in the Pacific sector of the Western Arctic Ocean during sea-ice melting. PLoS ONE 2014, 9, e86887. [Google Scholar] [CrossRef]
- Bowman, J.S.; Rasmussen, S.; Blom, N.; Deming, J.W.; Rysgaard, S.; Sicheritz-Ponten, T. Microbial community structure of Arctic multiyear sea ice and surface seawater by 454 sequencing of the 16S RNA gene. ISME J. 2012, 6, 11–20. [Google Scholar] [CrossRef]
- Brinkmeyer, R.; Knittel, K.; Jürgens, J.; Weyland, H.; Amann, R.; Helmke, E. Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl. Environ. Microbiol. 2003, 69, 6610–6619. [Google Scholar] [CrossRef]
- Brown, M.V.; Bowman, J.P. A molecular phylogenetic survey of sea-ice microbial communities (SIMCO). FEMS Microbiol. Ecol. 2001, 35, 267–275. [Google Scholar] [CrossRef]
- Grossmann, S. Bacterial activity in sea ice and open water of the Weddell Sea, Antarctica: A microautoradiographic study. Microb. Ecol. 1994, 28, 1–18. [Google Scholar] [CrossRef]
- Bowman, J.P.; McCuaig, R.D. Biodiversity, community structural shifts, and biogeography of prokaryotes within Antarctic continental shelf sediment. Appl. Environ. Microbiol. 2003, 69, 2463. [Google Scholar] [CrossRef]
- Li, Y.; Liu, Q.; Li, C.; Dong, Y.; Zhang, W.; Zhang, W.; Xiao, T. Bacterial and archaeal community structures in the Arctic deep-sea sediment. Acta Oceanol. Sin. 2015, 34, 93–113. [Google Scholar] [CrossRef]
- Tribelli, P.M.; López, N.I. Reporting key features in cold-adapted bacteria. Life 2018, 8, 8. [Google Scholar] [CrossRef]
- Sajjad, W.; Din, G.; Rafiq, M.; Iqbal, A.; Khan, S.; Zada, S.; Ali, B.; Kang, S. Pigment production by cold-adapted bacteria and fungi: Colorful tale of cryosphere with wide range applications. Extremophiles 2020, 24, 447–473. [Google Scholar] [CrossRef]
- Chintalapati, S.; Kiran, M.D.; Shivaji, S. Role of membrane lipid fatty acids in cold adaptation. Cell Mol. Biol. 2004, 50, 631–642. [Google Scholar]
- De Maayer, P.; Anderson, D.; Cary, C.; Cowan, D.A. Some like it cold: Understanding the survival strategies of psychrophiles. EMBO Rep. 2014, 15, 508–517. [Google Scholar] [CrossRef]
- Mykytczuk, N.C.S.; Foote, S.J.; Omelon, C.R.; Southam, G.; Greer, C.W.; Whyte, L.G. Bacterial growth at −15 °C; molecular insights from the permafrost bacterium Planococcus halocryophilus Or1. ISME J. 2013, 7, 1211–1226. [Google Scholar] [CrossRef]
- Collins, T.; Margesin, R. Psychrophilic lifestyles: Mechanisms of adaptation and biotechnological tools. Appl. Microbiol. Biotechnol. 2019, 103, 2857–2871. [Google Scholar] [CrossRef]
- Corsaro, M.M.; Casillo, A.; Parrilli, E.; Tutino, M.L. Molecular structure of lipopolysaccharides of cold-adapted bacteria. In Psychrophiles: From Biodiversity to Biotechnology; Margesin, R., Ed.; Springer International Publishing: Cham, Switzerland, 2017; pp. 285–303. ISBN 978-3-319-57057-0. [Google Scholar]
- Casillo, A.; Ziaco, M.; Lindner, B.; Parrilli, E.; Schwudke, D.; Holgado, A.; Verstrepen, L.; Sannino, F.; Beyaert, R.; Lanzetta, R.; et al. Unusual Lipid A from a Cold-Adapted Bacterium: Detailed Structural Characterization. ChemBioChem 2017, 18, 1845–1854. [Google Scholar] [CrossRef]
- Santiago, M.; Ramírez-Sarmiento, C.A.; Zamora, R.A.; Parra, L.P. Discovery, molecular mechanisms, and industrial applications of cold-active enzymes. Front. Microbiol. 2016, 7, 1408. [Google Scholar] [CrossRef]
- Celik, Y.; Drori, R.; Pertaya-Braun, N.; Altan, A.; Barton, T.; Bar-Dolev, M.; Groisman, A.; Davies, P.L.; Braslavsky, I. Microfluidic experiments reveal that antifreeze proteins bound to ice crystals suffice to prevent their growth. Proc. Natl. Acad. Sci. USA 2013, 110, 1309–1314. [Google Scholar] [CrossRef]
- Bar Dolev, M.; Braslavsky, I.; Davies, P.L. Ice-binding proteins and their function. Annu. Rev. Biochem. 2016, 85, 515–542. [Google Scholar] [CrossRef]
- Feng, S.; Powell, S.M.; Wilson, R.; Bowman, J.P. Extensive gene acquisition in the extremely psychrophilic bacterial species Psychroflexus torquis and the link to sea-ice ecosystem specialism. Genome Biol. Evol. 2014, 6, 133–148. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, S.; Li, J.; Gu, X.; Lin, X. Complete genome sequences of a H2O2-resistant psychrophilic bacterium Colwellia sp. Arc7-D isolated from Arctic Ocean sediment. Mar. Genom. 2019, 43, 65–67. [Google Scholar] [CrossRef]
- Goh, Y.J.; Klaenhammer, T.R. Insights into glycogen metabolism in Lactobacillus acidophilus: Impact on carbohydrate metabolism, stress tolerance and gut retention. Microb. Cell Factories 2014, 13, 94. [Google Scholar] [CrossRef]
- Goordial, J.; Raymond-Bouchard, I.; Zolotarov, Y.; de Bethencourt, L.; Ronholm, J.; Shapiro, N.; Woyke, T.; Stromvik, M.; Greer, C.W.; Bakermans, C.; et al. Cold adaptive traits revealed by comparative genomic analysis of the eurypsychrophile Rhodococcus sp. JG3 isolated from high elevation McMurdo Dry Valley permafrost, Antarctica. FEMS Microbiol. Ecol. 2016, 92. [Google Scholar] [CrossRef]
- Zhao, J.-S.; Deng, Y.; Manno, D.; Hawari, J. Shewanella spp. genomic evolution for a cold marine lifestyle and in-situ explosive biodegradation. PLoS ONE 2010, 5, e9109. [Google Scholar] [CrossRef]
- Raymond-Bouchard, I.; Whyte, L. From transcriptomes to metatranscriptomes: Cold adaptation and active metabolisms of psychrophiles from cold environments. In Psychrophiles: From Biodiversity to Biotechnology, 2nd ed.; Springer: Cham, Switzerland, 2017; pp. 437–457. ISBN 978-3-319-57056-3. [Google Scholar]
- Tribelli, P.M.; Venero, E.C.S.; Ricardi, M.M.; Gómez-Lozano, M.; Iustman, L.J.R.; Molin, S.; López, N.I. Novel essential role of ethanol oxidation genes at low temperature revealed by transcriptome analysis in the Antarctic bacterium Pseudomonas extremaustralis. PLoS ONE 2015, 10, e0145353. [Google Scholar] [CrossRef]
- Aliyu, H.; De Maayer, P.; Cowan, D. The genome of the Antarctic polyextremophile Nesterenkonia sp. AN1 reveals adaptive strategies for survival under multiple stress conditions. FEMS Microbiol. Ecol. 2016, 92, fiw032. [Google Scholar] [CrossRef]
- Gao, H.; Yang, Z.K.; Wu, L.; Thompson, D.K.; Zhou, J. Global transcriptome analysis of the cold shock response of Shewanella oneidensis MR-1 and mutational analysis of its classical cold shock proteins. J. Bacteriol. 2006, 188, 4560. [Google Scholar] [CrossRef]
- Bergholz, P.W.; Bakermans, C.; Tiedje, J.M. Psychrobacter arcticus 273-4 uses resource efficiency and molecular motion adaptations for subzero temperature Growth. J. Bacteriol. 2009, 191, 2340–2352. [Google Scholar] [CrossRef]
- Koh, H.Y.; Park, H.; Lee, J.H.; Han, S.J.; Sohn, Y.C.; Lee, S.G. Proteomic and transcriptomic investigations on cold-responsive properties of the psychrophilic Antarctic bacterium Psychrobacter sp. PAMC 21119 at subzero temperatures. Environ. Microbiol. 2017, 19, 628–644. [Google Scholar] [CrossRef]
- Parrilli, E.; Tedesco, P.; Fondi, M.; Tutino, M.L.; Lo Giudice, A.; de Pascale, D.; Fani, R. The art of adapting to extreme environments: The model system Pseudoalteromonas. Phys. Life Rev. 2019. [Google Scholar] [CrossRef]
- Gentile, G.; Bonasera, V.; Amico, C.; Giuliano, L.; Yakimov, M.M. Shewanella sp. GA-22, a psychrophilic hydrocarbonoclastic antarctic bacterium producing polyunsaturated fatty acids. J. Appl. Microbiol. 2003, 95, 1124–1133. [Google Scholar] [CrossRef]
- Jadhav, V.V.; Jamle, M.M.; Pawar, P.D.; Devare, M.N.; Bhadekar, R.K. Fatty acid profiles of PUFA producing Antarctic bacteria: Correlation with RAPD analysis. Ann. Microbiol. 2010, 60, 693–699. [Google Scholar] [CrossRef]
- Sato, S.; Kurihara, T.; Kawamoto, J.; Hosokawa, M.; Sato, S.B.; Esaki, N. Cold adaptation of eicosapentaenoic acid-less mutant of Shewanella livingstonensis Ac10 involving uptake and remodeling of synthetic phospholipids containing various polyunsaturated fatty acids. Extremophiles 2008, 12, 753–761. [Google Scholar] [CrossRef]
- Jøstensen, J.-P.; Landfald, B. High prevalence of polyunsaturated-fatty-acid producing bacteria in arctic invertebrates. FEMS Microbiol. Lett. 1997, 151, 95–101. [Google Scholar] [CrossRef][Green Version]
- Yoshida, K.; Hashimoto, M.; Hori, R.; Adachi, T.; Okuyama, H.; Orikasa, Y.; Nagamine, T.; Shimizu, S.; Ueno, A.; Morita, N. Bacterial long-chain polyunsaturated fatty acids: Their biosynthetic genes, functions, and practical use. Mar. Drugs 2016, 14, 94. [Google Scholar] [CrossRef]
- Chattopadhyay, M.K.; Jagannadham, M.V.; Vairamani, M.; Shivaji, S. Carotenoid pigments of an Antarctic psychrotrophic bacterium Micrococcus roseus: Temperature dependent biosynthesis, structure, and interaction with synthetic membranes. Biochem. Biophys. Res. Commun. 1997, 239, 85–90. [Google Scholar] [CrossRef]
- Jagannadham, M.V.; Chattopadhyay, M.K.; Subbalakshmi, C.; Vairamani, M.; Narayanan, K.; Mohan Rao, C.; Shivaji, S. Carotenoids of an Antarctic psychrotolerant bacterium, Sphingobacterium antarcticus, and a mesophilic bacterium, Sphingobacterium multivorum. Arch. Microbiol. 2000, 173, 418–424. [Google Scholar] [CrossRef]
- Dieser, M.; Greenwood, M.; Foreman, C.M. Carotenoid pigmentation in antarctic heterotrophic bacteria as a strategy to withstand environmental stresses. Arct. Antarct. Alp. Res. 2010, 42, 396–405. [Google Scholar] [CrossRef]
- Singh, A.; Krishnan, K.P.; Prabaharan, D.; Sinha, R.K. Lipid membrane modulation and pigmentation: A cryoprotection mechanism in Arctic pigmented bacteria. J. Basic Microbiol. 2017, 57, 770–780. [Google Scholar] [CrossRef]
- Collins, T.; Gerday, C. Enzyme Catalysis in Psychrophiles. In Psychrophiles: From Biodiversity to Biotechnology, 2nd ed.; Springer: Cham, Switzerland, 2017; pp. 209–235. ISBN 978-3-319-57057-0. [Google Scholar]
- Raymond, J.A.; Christner, B.C.; Schuster, S.C. A bacterial ice-binding protein from the Vostok ice core. Extremophiles 2008, 12, 713–717. [Google Scholar] [CrossRef]
- Lorv, J.S.H.; Rose, D.R.; Glick, B.R. Bacterial ice crystal controlling proteins. Scientifica 2014, 2014, 976895. [Google Scholar] [CrossRef]
- Poli, A.; Anzelmo, G.; Nicolaus, B. Bacterial exopolysaccharides from extreme marine habitats: Production, characterization and biological activities. Mar. Drugs 2010, 8, 1779–1802. [Google Scholar] [CrossRef]
- Deming, J.W.; Young, J.N. The Role of exopolysaccharides in microbial adaptation to cold habitats. In Psychrophiles: From Biodiversity to Biotechnology; Margesin, R., Ed.; Springer International Publishing: Cham, Switzerland, 2017; pp. 259–284. ISBN 978-3-319-57057-0. [Google Scholar]
- Caruso, C.; Rizzo, C.; Mangano, S.; Poli, A.; Donato, P.D.; Nicolaus, B.; Finore, I.; Marco, G.D.; Michaud, L.; Giudice, A.L. Isolation, characterization and optimization of EPSs produced by a cold-adapted Marinobacter isolate from Antarctic seawater. Antarct. Sci. 2019, 31, 69–79. [Google Scholar] [CrossRef]
- Caruso, C.; Rizzo, C.; Mangano, S.; Poli, A.; Donato, P.D.; Finore, I.; Nicolaus, B.; Marco, G.D.; Michaud, L.; Giudice, A.L. Production and biotechnological potential of extracellular polymeric substances from sponge-associated antarctic bacteria. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Lo Giudice, A.; Poli, A.; Finore, I.; Rizzo, C. Peculiarities of extracellular polymeric substances produced by Antarctic bacteria and their possible applications. Appl. Microbiol. Biotechnol. 2020, 104, 2923–2934. [Google Scholar] [CrossRef]
- Bowman, J. Genomics of Psychrophilic Bacteria and Archaea. In Psychrophiles: From Biodiversity to Biotechnology, 2nd ed.; Springer: Cham, Switzerland, 2017; pp. 345–387. ISBN 978-3-319-57056-3. [Google Scholar]
- Vollmers, J.; Voget, S.; Dietrich, S.; Gollnow, K.; Smits, M.; Meyer, K.; Brinkhoff, T.; Simon, M.; Daniel, R. Poles apart: Arctic and Antarctic Octadecabacter strains share high genome plasticity and a new type of xanthorhodopsin. PLoS ONE 2013, 8, e63422. [Google Scholar] [CrossRef]
- Qin, Q.-L.; Xie, B.-B.; Yu, Y.; Shu, Y.-L.; Rong, J.-C.; Zhang, Y.-J.; Zhao, D.-L.; Chen, X.-L.; Zhang, X.-Y.; Chen, B.; et al. Comparative genomics of the marine bacterial genus Glaciecola reveals the high degree of genomic diversity and genomic characteristic for cold adaptation. Environ. Microbiol. 2014, 16, 1642–1653. [Google Scholar] [CrossRef]
- Methé, B.A.; Nelson, K.E.; Deming, J.W.; Momen, B.; Melamud, E.; Zhang, X.; Moult, J.; Madupu, R.; Nelson, W.C.; Dodson, R.J.; et al. The psychrophilic lifestyle as revealed by the genome sequence of Colwellia psychrerythraea 34H through genomic and proteomic analyses. Proc. Natl. Acad. Sci. USA 2005, 102, 10913–10918. [Google Scholar] [CrossRef]
- Riley, M.; Staley, J.T.; Danchin, A.; Wang, T.Z.; Brettin, T.S.; Hauser, L.J.; Land, M.L.; Thompson, L.S. Genomics of an extreme psychrophile, Psychromonas ingrahamii. BMC Genom. 2008, 9, 210. [Google Scholar] [CrossRef]
- Chaudhary, D.K.; Khulan, A.; Kim, D.-U.; Kim, J. Flavobacterium cellulosilyticum sp. nov., a novel psychrophilic bacterium isolated from Arctic soil. Int. J. Syst. Evol. Microbiol. 2020, 70, 44–50. [Google Scholar] [CrossRef]
- Dall’Agnol, H.P.; Baraúna, R.A.; de Sá, P.H.; Ramos, R.T.; Nóbrega, F.; Nunes, C.I.; das Graças, D.A.; Carneiro, A.R.; Santos, D.M.; Pimenta, A.M.; et al. Omics profiles used to evaluate the gene expression of Exiguobacterium antarcticum B7 during cold adaptation. BMC Genom. 2014, 15, 986. [Google Scholar] [CrossRef]
- Rodrigues, D.F.; Ivanova, N.; He, Z.; Huebner, M.; Zhou, J.; Tiedje, J.M. Architecture of thermal adaptation in an Exiguobacterium sibiricum strain isolated from 3 million year old permafrost: A genome and transcriptome approach. BMC Genom. 2008, 9, 547. [Google Scholar] [CrossRef]
- Singh, A.K.; Pindi, P.K.; Dube, S.; Sundareswaran, V.R.; Shivaji, S. Importance of trmE for growth of the psychrophile Pseudomonas syringae at low temperatures. Appl. Environ. Microbiol. 2009, 75, 4419. [Google Scholar] [CrossRef]
- Sundareswaran, V.R.; Singh, A.K.; Dube, S.; Shivaji, S. Aspartate aminotransferase is involved in cold adaptation in psychrophilic Pseudomonas syringae. Arch. Microbiol. 2010, 192, 663–672. [Google Scholar] [CrossRef]
- Sakamoto, T.; Higashi, S.; Wada, H.; Murata, N.; Bryant, D.A. Low-temperature-induced desaturation of fatty acids and expression of desaturase genes in the cyanobacterium Synechococcus sp. PCC 7002. FEMS Microbiol. Lett. 1997, 152, 313–320. [Google Scholar] [CrossRef]
- Sakamoto, T.; Shen, G.; Higashi, S.; Murata, N.; Bryant, D. Alteration of low-temperature susceptibility of the cyanobacterium Synechococcus sp. PCC 7002 by genetic manipulation of membrane lipid unsaturation. Arch. Microbiol. 1997, 169, 20–28. [Google Scholar] [CrossRef]
- Thomas, D.N.; Dieckmann, G.S. Antarctic sea ice--a habitat for extremophiles. Science 2002, 295, 641–644. [Google Scholar] [CrossRef]
- Bowman, J.P.; McCammon, S.A.; Brown, M.V.; Nichols, D.S.; McMeekin, T.A. Diversity and association of psychrophilic bacteria in Antarctic sea ice. Appl. Environ. Microbiol. 1997, 63, 3068–3078. [Google Scholar] [CrossRef]
- Bowman, J.P.; McCammon, S.A.; Lewis, T.; Skerratt, J.H.; Brown, J.L.; Nichols, D.S.; McMeekin, T.A. Psychroflexus torquis gen. nov., sp. nov., a psychrophilic species from Antarctic sea ice, and reclassification of Flavobacterium gondwanense (Dobson et al. 1993) as Psychroflexus gondwanense gen. nov., comb. nov. Microbiology 1998, 144, 1601–1609. [Google Scholar] [CrossRef]
- Rapp, J.Z.; Fernández-Méndez, M.; Bienhold, C.; Boetius, A. Effects of ice-algal aggregate export on the connectivity of bacterial communities in the central Arctic Ocean. Front. Microbiol. 2018, 9, 1035. [Google Scholar] [CrossRef]
- Teeling, H.; Fuchs, B.M.; Becher, D.; Klockow, C.; Gardebrecht, A.; Bennke, C.M.; Kassabgy, M.; Huang, S.; Mann, A.J.; Waldmann, J.; et al. Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 2012, 336, 608–611. [Google Scholar] [CrossRef]
- Ramanan, R.; Kim, B.-H.; Cho, D.-H.; Oh, H.-M.; Kim, H.-S. Algae-bacteria interactions: Evolution, ecology and emerging applications. Biotechnol. Adv. 2016, 34, 14–29. [Google Scholar] [CrossRef]
- Amin, S.A.; Hmelo, L.R.; van Tol, H.M.; Durham, B.P.; Carlson, L.T.; Heal, K.R.; Morales, R.L.; Berthiaume, C.T.; Parker, M.S.; Djunaedi, B.; et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 2015, 522, 98–101. [Google Scholar] [CrossRef]
- Kim, B.-H.; Ramanan, R.; Cho, D.-H.; Oh, H.-M.; Kim, H.-S. Role of Rhizobium, a plant growth promoting bacterium, in enhancing algal biomass through mutualistic interaction. Biomass Bioenergy 2014, 69, 95–105. [Google Scholar] [CrossRef]
- Van Tol, H.M.; Amin, S.A.; Armbrust, E.V. Ubiquitous marine bacterium inhibits diatom cell division. ISME J. 2017, 11, 31–42. [Google Scholar] [CrossRef]
- Buchan, A.; LeCleir, G.R.; Gulvik, C.A.; González, J.M. Master recyclers: Features and functions of bacteria associated with phytoplankton blooms. Nat. Rev. Microbiol. 2014, 12, 686–698. [Google Scholar] [CrossRef]
- Bell, W.; Mitchell, R. Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol. Bull. 1972, 143, 265–277. [Google Scholar] [CrossRef]
- Zhen, M.; Zongjun, D.; Huirong, L.; Yanying, L.; Wei, L. Analysis of bacterial diversity in the phycosphere of five Arctic microalgae. Acta Ecol. Sin. 2015, 35. [Google Scholar] [CrossRef]
- Sapp, M.; Schwaderer, A.S.; Wiltshire, K.H.; Hoppe, H.-G.; Gerdts, G.; Wichels, A. Species-specific bacterial communities in the phycosphere of microalgae? Microb. Ecol. 2007, 53, 683–699. [Google Scholar] [CrossRef]
- Grossart, H.-P.; Levold, F.; Allgaier, M.; Simon, M.; Brinkhoff, T. Marine diatom species harbour distinct bacterial communities. Environ. Microbiol. 2005, 7, 860–873. [Google Scholar] [CrossRef]
- Schäfer, H.; Abbas, B.; Witte, H.; Muyzer, G. Genetic diversity of ‘satellite’ bacteria present in cultures of marine diatoms. FEMS Microbiol. Ecol. 2002, 42, 25–35. [Google Scholar] [CrossRef][Green Version]
- Delmont, T.O.; Hammar, K.M.; Ducklow, H.W.; Yager, P.L.; Post, A.F. Phaeocystis antarctica blooms strongly influence bacterial community structures in the Amundsen Sea polynya. Front. Microbiol. 2014, 5, 646. [Google Scholar] [CrossRef]
- Kirchman, D.L.; Meon, B.; Ducklow, H.W.; Carlson, C.A.; Hansell, D.A.; Steward, G.F. Glucose fluxes and concentrations of dissolved combined neutral sugars (polysaccharides) in the Ross Sea and Polar Front Zone, Antarctica. Deep Sea Res. Part II Top. Stud. Oceanogr. 2001, 48, 4179–4197. [Google Scholar] [CrossRef]
- Schriek, R. Effects of light and temperature on the enzymatic antioxidative defense systems in the Antarctic ice diatom Entomoneis kufferathii Manguin. Rep. Polar Res. 2000, 349, 1–130. [Google Scholar]
- Hünken, M.; Harder, J.; Kirst, G.O. Epiphytic bacteria on the Antarctic ice diatom Amphiprora kufferathii Manguin cleave hydrogen peroxide produced during algal photosynthesis. Plant Biol. 2008, 10, 519–526. [Google Scholar] [CrossRef]
- Bertrand, E.M.; McCrow, J.P.; Moustafa, A.; Zheng, H.; McQuaid, J.B.; Delmont, T.O.; Post, A.F.; Sipler, R.E.; Spackeen, J.L.; Xu, K.; et al. Phytoplankton–bacterial interactions mediate micronutrient colimitation at the coastal Antarctic sea ice edge. Proc. Natl. Acad. Sci. USA 2015, 112, 9938–9943. [Google Scholar] [CrossRef]
- Johannessen, O.M.; Miles, M.W. Critical vulnerabilities of marine and sea ice–based ecosystems in the high Arctic. Reg. Environ. Chang. 2011, 11, 239–248. [Google Scholar] [CrossRef]
- Anisimov, O. Potential feedback of thawing permafrost to the global climate system through methane emission. Environ. Res. Lett. 2007, 2, 045016. [Google Scholar] [CrossRef]
- Apprill, A. The Role of symbioses in the adaptation and stress responses of marine organisms. Ann. Rev. Mar. Sci. 2020, 12. [Google Scholar] [CrossRef]
- Burrows, M.T.; Schoeman, D.S.; Buckley, L.B.; Moore, P.; Poloczanska, E.S.; Brander, K.M.; Brown, C.; Bruno, J.F.; Duarte, C.M.; Halpern, B.S.; et al. The pace of shifting climate in marine and terrestrial ecosystems. Science 2011, 334, 652–655. [Google Scholar] [CrossRef]
- Doney, S.C.; Ruckelshaus, M.; Emmett Duffy, J.; Barry, J.P.; Chan, F.; English, C.A.; Galindo, H.M.; Grebmeier, J.M.; Hollowed, A.B.; Knowlton, N.; et al. Climate Change Impacts on Marine Ecosystems. Annu. Rev. Mar. Sci. 2012, 4, 11–37. [Google Scholar] [CrossRef]


| Stress Exposure | Main Effects/Responses | Reference |
|---|---|---|
| Physiological changes | ||
| Low salt concentration | Increase in specific growth rate, biochemical composition shifting towards a lower POC:PON relationship with higher protein content, but reduced fatty acid and carbohydrate content. | [27,44] |
| High salt concentration | Slowing down of cell division and growth rate, reduced cell size and motility, triggering “palmelloid” formation in Chlamydomonas, production of osmoregulatory compounds (e.g., glycerol and proline), increase in ion transmembrane transport and lipids. In the sea-ice diatom, Nitzschia lecointei, small changes in growth rate, effect on cellular metabolite pool sizes | [46,47,48] |
| High levels of ultraviolet radiation (UVR) | Increase in photoprotective pigments | [49,50] |
| Reduction in photosynthetic rate | [51] | |
| Inactivation of specific enzymes | ||
| affecting species diversity and richness. | ||
| Increase in saturated fatty acids, decrease in polyunsaturated fatty acids (PUFAs), small increase in C18 PUFAs | [52] | |
| Low light exposure | Protection from UVR | |
| Limitation in primary production | ||
| Nutrients: Fe limitation | Influence in microalgal growth and composition | [53] |
| Increase in lipid production | [54] | |
| Low pH | Increase in large diatoms, early senescence | [55] |
| Increase in photosynthetic rate and growth rate | ||
| Affecting membrane potential, energy partitioning and enzyme activity | [56] | |
| Low temperature | Maintenance of membrane fluidity thanks to unsaturated fatty acids | [57] |
| Maintenance of sufficient rates of enzyme-catalyzed reactions for key metabolic processes | ||
| Evolution of cold shock and antifreeze proteins | ||
| Photosynthetic electron transport chain adaptations | ||
| Molecular changes | ||
| Genome comparison between cold-adapted and temperate species | In the cold-adapted genome there were highly divergent alleles which were also differentially expressed across various environmental conditions. Main genes identified for cold adaptation were ice-binding proteins IBPs, proton-pumping proteorhodopsins and chlorophyll a/c light-harvesting complex LHC. | [58] |
| Low temperature, high light | Genes encoding proteins of PSII (psbA, psbC) and for carbon fixation (rbcL) were down-regulated | [59] |
| Chaperones (hsp70) and genes for plastid protein synthesis and turnover (elongation factor EfTs, ribosomal protein rpS4, ftsH protease) were up-regulated | ||
| Low temperature, low light | Down-regulation of psbA, psbC, and rbcL | [59] |
| Low temperature, high salinity | Ionic transporters and antiporters, heat shock proteins, genes related to oxidative stress, and three key genes involved in proline synthesis | [60] |
| Low temperature | Expression of various DEAD-box RNA helicase genes, such as CiRH5, CiRH25, CiRH28, and CiRH55, were found significantly up-regulated under freezing treatment | [61] |
| Increase in genes encoding proteins involved in protein translation and transport, including protein transport protein SEC61, signal recognition particle protein, protein involved in vacuolar protein sorting. Increase in Heat shock protein 70, matrix metalloproteinase M11, X-Pro dipeptidyl-peptidase, and protein binding 26S proteasome regulatory complex, nitrate reductase, ferredoxin-nitrite reductase, and nitrate/nitrite transporter. | [62] | |
| Increased temperature | Up-regulation of cytoprotective genes, down-regulation of genes related to photosynthesis, increase in fucoxanthin chlorophyll a/c-binding proteins | [63] |
| High light | Transcripts related to photosynthesis were affected | [64] |
| Identification of an antifreeze protein gene (Cn-AFP) | [65] | |
| High salinity | Increased expression of genes participating in the metabolism of carbohydrates, such as starch, sucrose, soluble sugar, and glucose | [66] |
| Stress Exposure | Main Effects/Responses | Reference |
|---|---|---|
| Physiological changes | ||
| Low temperature | Reduction of primary metabolism | [111] |
| Modulation of PUFA amount for membrane fluidity maintenance, modulation of proteins and carotenoids | [91,112,113,114] | |
| Peptidoglycan thickening | [115,116] | |
| LPS lacking O-chain component | [117] | |
| Increase in short chain and/or unsaturated fatty acids | [117,118] | |
| Increase in enzymes with high catalytic activity | [91,119] | |
| Increase in antifreeze (AFPs) and ice nucleating (INPs) proteins | [120,121] | |
| Increase in cryoprotective compounds, including compatible solutes, extracellular polymeric substances (EPSs) and polyhydroxyalkanoates (PHAs) | [116] | |
| Molecular changes | ||
| Genomic observations | Common genetic traits include those related to oxidative stress, metabolism and energy and nutrient acquisition, cell wall membrane structure and fatty acid biosynthesis, production of cold-shock protein (CSP) and chaperones, production of exopolysaccharides or other extracellular substances, biosynthesis or transport of compatible solutes (i.e., glycine betaine, ectoine, and trehalose), and presence of genes involved in antioxidant activity, such as superoxide dismutase, glutathione peroxidase, glutathione reductase, catalase, aconitase, thioredoxin and ascorbic acid | [122,123] |
| Glycogen synthesis genes | [124] | |
| Presence of gene cluster encoding for glycogen synthase and 4-oxoacyl-ACP reductase, putative secondary metabolite biosynthesis gene clusters for terpene, nonribosomal peptide synthetase (NRPS), and different polyketide synthases (T1PKS and T3PKS) | [125] | |
| Presence of genes related to exopolysaccharide and polyunsaturated fatty acid biosynthesis, or involved in nutrient acquisition, production of proteins associated with ice-binding and light-sensing processes | [122] | |
| Reduced G+C content and peculiar composition for certain aminoacids to increase protein flexibility | [126] | |
| Transcriptome analyses at low temperature | Up-regulation of transcripts involved in oxidative stress, CSP and chaperone production, metabolism and energy management, membrane fluidity assessment | [127] |
| Up-regulation of genes involved in ethanol oxidation (exaA, exaB and ExaC) encoding for a pyrroloquinoline quinone (PQQ)-dependent ethanol dehydrogenase, a cytochrome c550 and an aldehyde dehydrogenase | [128] | |
| Induction of transcripts encoding for antioxidants (suggested by the up-regulation of sodA, bcp and bpoA2), and of genes encoding for key enzymes of the glioxylate cycle (isocitrate lyase and malate synthase) | [129] | |
| Expression for several protein families (i.e., membrane and regulatory proteins, metabolic proteins, especially those involved in NADH and NADPH generation), DNA metabolism and translation apparatus components resulted up-regulated | [130] | |
| Down-regulation of genes involved in transcription, translation, energy production, and most biosynthetic pathways was evidenced, while genes for specific biosynthesis processes (proline, tryptophan, and methionine), chaperones clpB and hsp33, RNases and peptidases were generally up-regulated | [131] | |
| Up-regulation of genes for translation, ribosomal structure and biogenesis and a down-regulation of lipid transport and metabolism | [132] | |
| Proteome analysis at low temperature | Up-regulation was detected for proteins involved in metabolite transport, protein folding, membrane fluidity and aminoacid biosynthesis (protein MetF, ScoB and MmsA). Proteins related to energy production and conversion were down-regulated | [132] |
| Multi-omics (Genomic and Phenomic) analysis | High metabolic versatility, capacity to enhance the uptake of compounds with peculiar role in cryoprotection (i.e., spermine, glutathione, ornithine and other compounds related to glutathione metabolism). Enrichment in genes involved in lipid transport and metabolism and an up-regulation of protein synthesis metabolism | [2] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lauritano, C.; Rizzo, C.; Lo Giudice, A.; Saggiomo, M. Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments. Microorganisms 2020, 8, 1957. https://doi.org/10.3390/microorganisms8121957
Lauritano C, Rizzo C, Lo Giudice A, Saggiomo M. Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments. Microorganisms. 2020; 8(12):1957. https://doi.org/10.3390/microorganisms8121957
Chicago/Turabian StyleLauritano, Chiara, Carmen Rizzo, Angelina Lo Giudice, and Maria Saggiomo. 2020. "Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments" Microorganisms 8, no. 12: 1957. https://doi.org/10.3390/microorganisms8121957
APA StyleLauritano, C., Rizzo, C., Lo Giudice, A., & Saggiomo, M. (2020). Physiological and Molecular Responses to Main Environmental Stressors of Microalgae and Bacteria in Polar Marine Environments. Microorganisms, 8(12), 1957. https://doi.org/10.3390/microorganisms8121957








