Crucial Role of the C-Terminal Domain of Hfq Protein in Genomic Instability
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Media
2.2. Measurement of Tetr Mutation Rates
2.3. Statistical Analysis
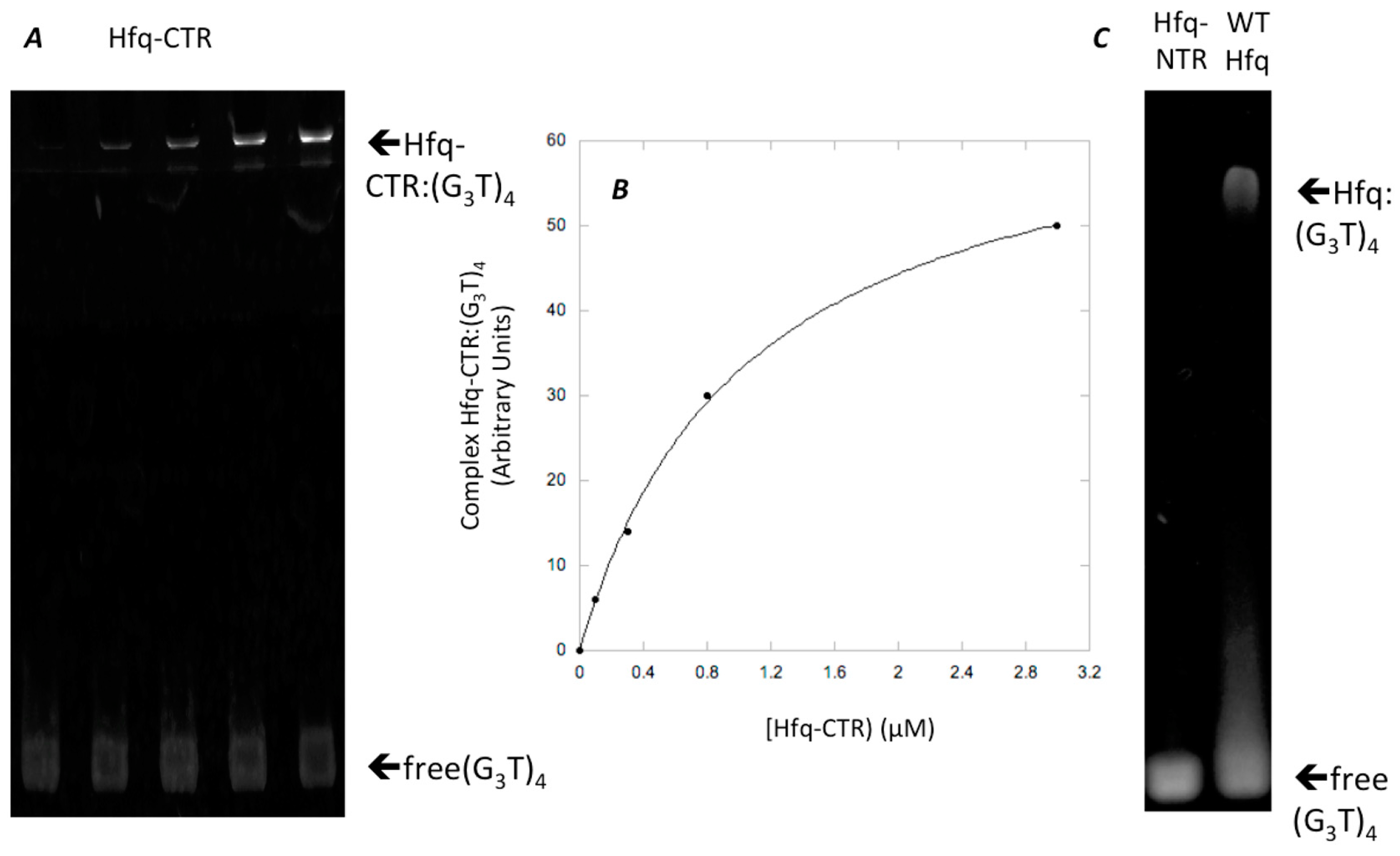
2.4. Binding Assays of Hfq-CTR on d(G3T)4
3. Results
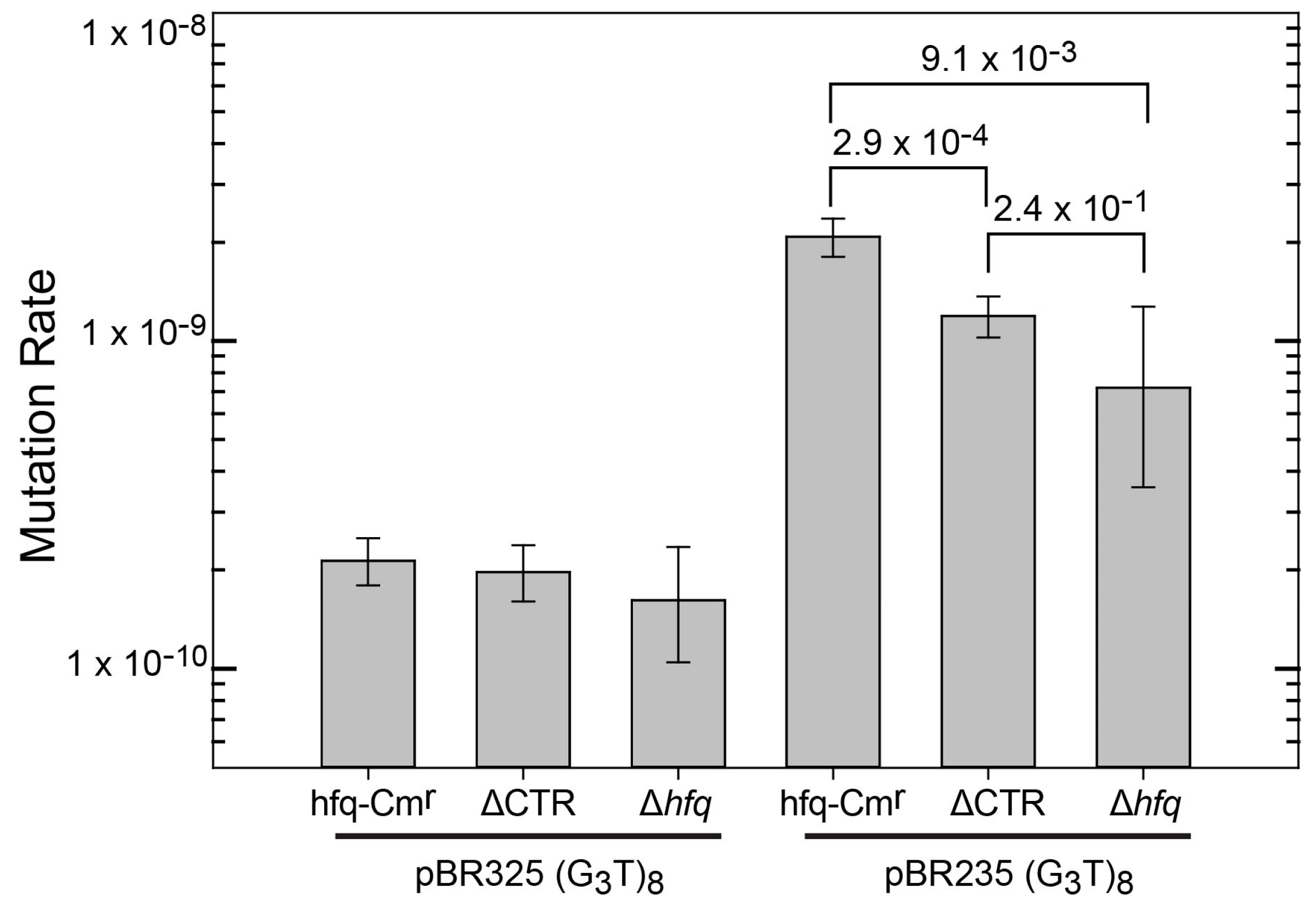
3.1. Influence of Hfq and the Hfq-CTR on the Instability of Quadruplex-Forming Repeats
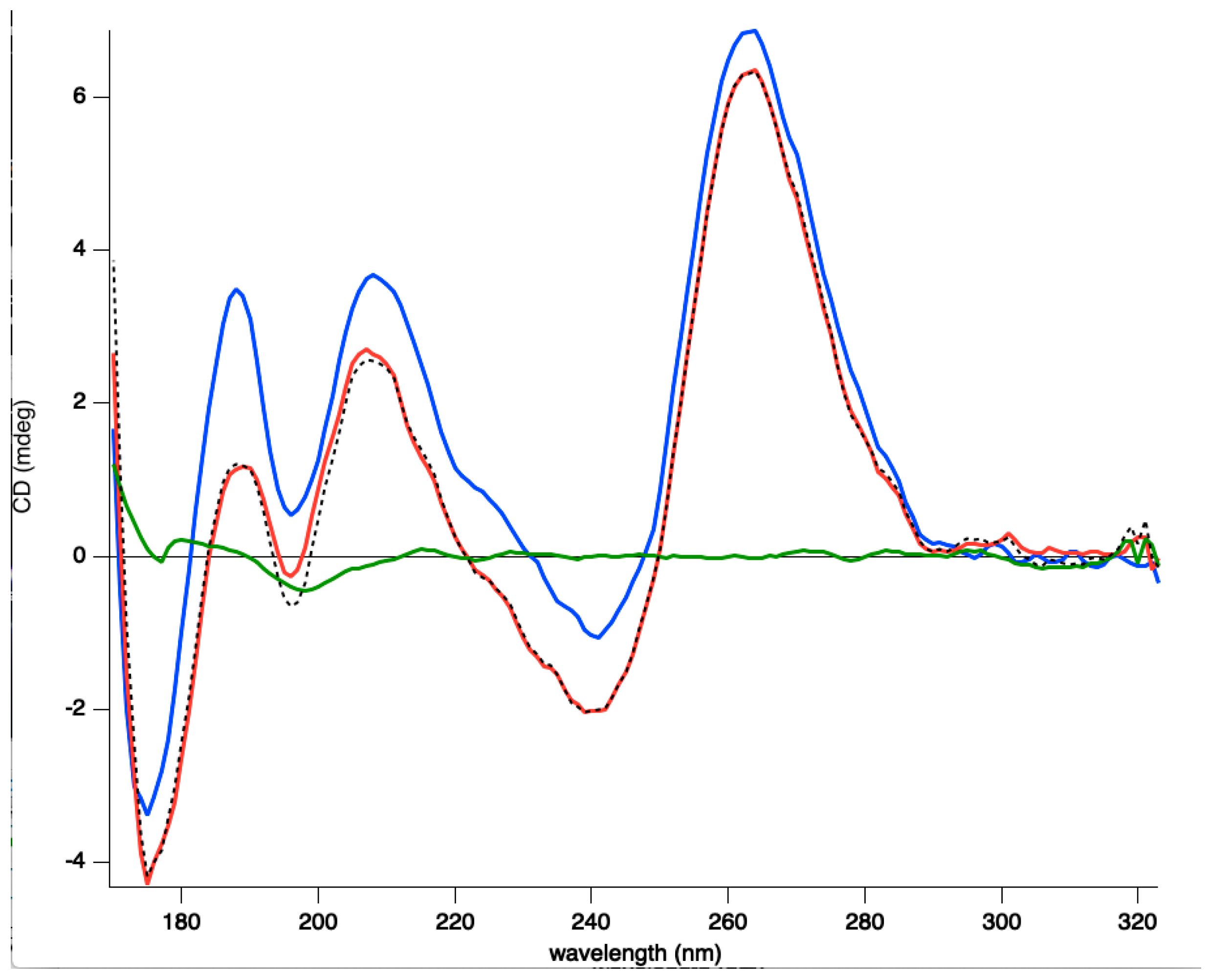
3.2. Interaction of Hfq-CTR with G-Quadruplex DNA
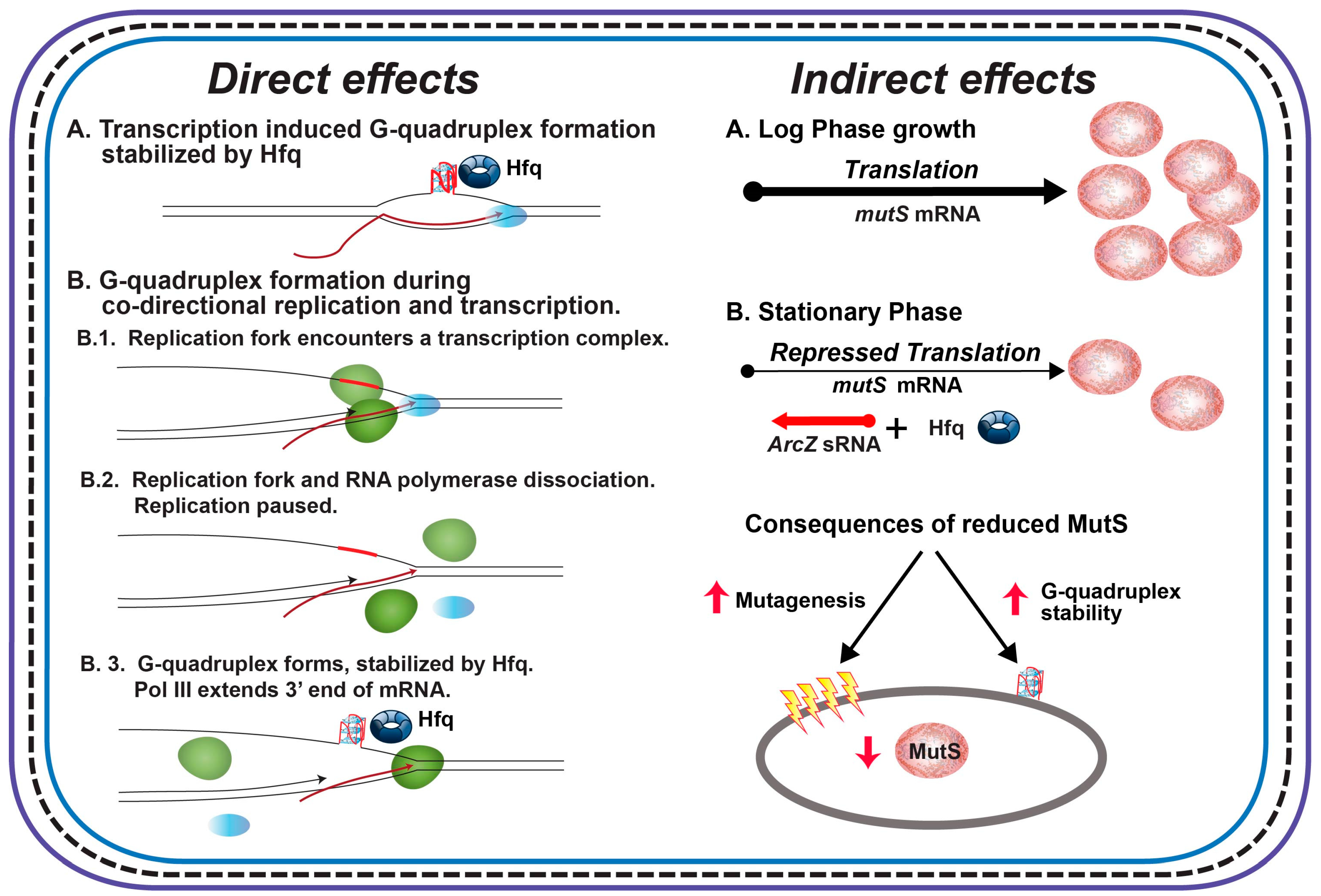
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gellert, M.; Lipsett, M.N.; Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 1962, 48, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Sinden, R.R. DNA Structure and Function; Academic Press: Cambridge, MA, USA, 1994. [Google Scholar]
- Kypr, J.; Kejnovska, I.; Renciuk, D.; Vorlickova, M. Circular dichroism and conformational polymorphism of DNA. Nucleic Acids Res. 2009, 37, 1713–1725. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.J.; Phan, A.T.; Kuryavyi, V. Human telomere, oncogenic promoter and 5’-UTR G-quadruplexes: Diverse higher order DNA and RNA targets for cancer therapeutics. Nucleic Acids Res. 2007, 35, 7429–7455. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Hurley, L.H. Structures, folding patterns, and functions of intramolecular DNA G-quadruplexes found in eukaryotic promoter regions. Biochimie 2008, 90, 1149–1171. [Google Scholar] [CrossRef] [PubMed]
- Gehring, K.; Leroy, J.L.; Gueron, M. A tetrameric DNA structure with protonated cytosine-cytosine base pairs. Nature 1993, 363, 561–565. [Google Scholar] [CrossRef]
- Phan, A.T.; Mergny, J.L. Human telomeric DNA: G-quadruplex, i-motif and Watson-Crick double helix. Nucleic Acids Res. 2002, 30, 4618–4625. [Google Scholar] [CrossRef]
- Kang, C.H.; Berger, I.; Lockshin, C.; Ratliff, R.; Moyzis, R.; Rich, A. Crystal structure of intercalated four-stranded d(C3T) at 1.4 A resolution. Proc. Natl. Acad. Sci. USA 1994, 91, 11636–11640. [Google Scholar] [CrossRef] [PubMed]
- Guo, K.; Pourpak, A.; Beetz-Rogers, K.; Gokhale, V.; Sun, D.; Hurley, L.H. Formation of pseudosymmetrical G-quadruplex and i-motif structures in the proximal promoter region of the RET oncogene. J. Am. Chem. Soc. 2007, 129, 10220–10228. [Google Scholar] [CrossRef]
- Marsico, G.; Chambers, V.S.; Sahakyan, A.B.; McCauley, P.; Boutell, J.M.; Antonio, M.D.; Balasubramanian, S. Whole genome experimental maps of DNA G-quadruplexes in multiple species. Nucleic Acids Res. 2019, 47, 3862–3874. [Google Scholar] [CrossRef] [PubMed]
- Eddy, J.; Maizels, N. Gene function correlates with potential for G4 DNA formation in the human genome. Nucleic Acids Res. 2006, 34, 3887–3896. [Google Scholar] [CrossRef]
- Sen, D.; Gilbert, W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 1988, 334, 364–366. [Google Scholar] [CrossRef]
- Kaplan, O.I.; Berber, B.; Hekim, N.; Doluca, O. G-quadruplex prediction in E. coli genome reveals a conserved putative G-quadruplex-Hairpin-Duplex switch. Nucleic Acids Res. 2016, 44, 9083–9095. [Google Scholar] [CrossRef]
- Du, X.; Wojtowicz, D.; Bowers, A.A.; Levens, D.; Benham, C.J.; Przytycka, T.M. The genome-wide distribution of non-B DNA motifs is shaped by operon structure and suggests the transcriptional importance of non-B DNA structures in Escherichia coli. Nucleic Acids Res. 2013, 41, 5965–5977. [Google Scholar] [CrossRef] [PubMed]
- Bartas, M.; Cutova, M.; Brazda, V.; Kaura, P.; Stastny, J.; Kolomaznik, J.; Coufal, J.; Goswami, P.; Cerven, J.; Pecinka, P. The Presence and Localization of G-Quadruplex Forming Sequences in the Domain of Bacteria. Molecules 2019, 24, 1711. [Google Scholar] [CrossRef] [PubMed]
- Rawal, P.; Kummarasetti, V.B.; Ravindran, J.; Kumar, N.; Halder, K.; Sharma, R.; Mukerji, M.; Das, S.K.; Chowdhury, S. Genome-wide prediction of G4 DNA as regulatory motifs: Role in Escherichia coli global regulation. Genome Res. 2006, 16, 644–655. [Google Scholar] [CrossRef]
- Beaume, N.; Pathak, R.; Yadav, V.K.; Kota, S.; Misra, H.S.; Gautam, H.K.; Chowdhury, S. Genome-wide study predicts promoter-G4 DNA motifs regulate selective functions in bacteria: Radioresistance of D. radiodurans involves G4 DNA-mediated regulation. Nucleic Acids Res. 2013, 41, 76–89. [Google Scholar] [CrossRef] [PubMed]
- Holder, I.T.; Hartig, J.S. A matter of location: Influence of G-quadruplexes on Escherichia coli gene expression. Chem. Biol. 2014, 21, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.Y.; Zheng, K.W.; Zhang, J.Y.; Hao, Y.H.; Tan, Z. Formation of DNA:RNA hybrid G-quadruplex in bacterial cells and its dominance over the intramolecular DNA G-quadruplex in mediating transcription termination. Angew. Chem. Int. Ed. 2015, 54, 2447–2451. [Google Scholar] [CrossRef]
- Shao, X.; Zhang, W.; Umar, M.I.; Wong, H.Y.; Seng, Z.; Xie, Y.; Zhang, Y.; Yang, L.; Kwok, C.K.; Deng, X. RNA G-Quadruplex Structures Mediate Gene Regulation in Bacteria. mBio 2020, 11, e02926-19. [Google Scholar] [CrossRef]
- Bugaut, A.; Balasubramanian, S. 5’-UTR RNA G-quadruplexes: Translation regulation and targeting. Nucleic Acids Res. 2012, 40, 4727–4741. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.M.; Merrick, C.J. G-quadruplexes in pathogens: A common route to virulence control? PLoS Pathog. 2015, 11, e1004562. [Google Scholar] [CrossRef] [PubMed]
- Bhartiya, D.; Chawla, V.; Ghosh, S.; Shankar, R.; Kumar, N. Genome-wide regulatory dynamics of G-quadruplexes in human malaria parasite Plasmodium falciparum. Genomics 2016, 108, 224–231. [Google Scholar] [CrossRef] [PubMed]
- Claessens, A.; Harris, L.M.; Stanojcic, S.; Chappell, L.; Stanton, A.; Kuk, N.; Veneziano-Broccia, P.; Sterkers, Y.; Rayner, J.C.; Merrick, C.J. RecQ helicases in the malaria parasite Plasmodium falciparum affect genome stability, gene expression patterns and DNA replication dynamics. PLoS Genet. 2018, 14, e1007490. [Google Scholar] [CrossRef]
- Harris, L.M.; Monsell, K.R.; Noulin, F.; Famodimu, M.T.; Smargiasso, N.; Damblon, C.; Horrocks, P.; Merrick, C.J. G-Quadruplex DNA Motifs in the Malaria Parasite Plasmodium falciparum and Their Potential as Novel Antimalarial Drug Targets. Antimicrob. Agents Chemother. 2018, 62, e01828-17. [Google Scholar] [CrossRef] [PubMed]
- Stanton, A.; Harris, L.M.; Graham, G.; Merrick, C.J. Recombination events among virulence genes in malaria parasites are associated with G-quadruplex-forming DNA motifs. BMC Genom. 2016, 17, 859. [Google Scholar] [CrossRef]
- Gage, H.L.; Merrick, C.J. Conserved associations between G-quadruplex-forming DNA motifs and virulence gene families in malaria parasites. BMC Genom. 2020, 21, 236. [Google Scholar] [CrossRef]
- Saha, T.; Shukla, K.; Thakur, R.S.; Desingu, A.; Nagaraju, G. Mycobacterium tuberculosis UvrD1 and UvrD2 helicases unwind G-quadruplex DNA. FEBS J. 2019, 286, 2062–2086. [Google Scholar] [CrossRef]
- Shukla, K.; Thakur, R.S.; Ganguli, D.; Rao, D.N.; Nagaraju, G. Escherichia coli and Neisseria gonorrhoeae UvrD helicase unwinds G4 DNA structures. Biochem. J. 2017, 474, 3579–3597. [Google Scholar] [CrossRef]
- Zhou, X.; Ren, W.; Bharath, S.R.; Tang, X.; He, Y.; Chen, C.; Liu, Z.; Li, D.; Song, H. Structural and Functional Insights into the Unwinding Mechanism of Bacteroides sp Pif1. Cell Rep. 2016, 14, 2030–2039. [Google Scholar] [CrossRef] [PubMed]
- Lopes, J.; Piazza, A.; Bermejo, R.; Kriegsman, B.; Colosio, A.; Teulade-Fichou, M.P.; Foiani, M.; Nicolas, A. G-quadruplex-induced instability during leading-strand replication. EMBO J. 2011, 30, 4033–4046. [Google Scholar] [CrossRef]
- Thakur, R.S.; Desingu, A.; Basavaraju, S.; Subramanya, S.; Rao, D.N.; Nagaraju, G. Mycobacterium tuberculosis DinG is a structure-specific helicase that unwinds G4 DNA: Implications for targeting G4 DNA as a novel therapeutic approach. J. Biol. Chem. 2014, 289, 25112–25136. [Google Scholar] [CrossRef] [PubMed]
- Teng, F.Y.; Hou, X.M.; Fan, S.H.; Rety, S.; Dou, S.X.; Xi, X.G. Escherichia coli DNA polymerase I can disrupt G-quadruplex structures during DNA replication. FEBS J. 2017, 284, 4051–4065. [Google Scholar] [CrossRef]
- Ehrat, E.A.; Johnson, B.R.; Williams, J.D.; Borchert, G.M.; Larson, E.D. G-quadruplex recognition activities of E. Coli MutS. BMC Mol. Biol. 2012, 13, 23. [Google Scholar] [CrossRef]
- Takada, A.; Wachi, M.; Kaidow, A.; Takamura, M.; Nagai, K. DNA binding properties of the hfq gene product of Escherichia coli. Biochem. Biophys. Res. Commun. 1997, 236, 576–579. [Google Scholar] [CrossRef]
- Jiang, K.; Zhang, C.; Guttula, D.; Liu, F.; van Kan, J.A.; Lavelle, C.; Kubiak, K.; Malabirade, A.; Lapp, A.; Arluison, V.; et al. Effects of Hfq on the conformation and compaction of DNA. Nucleic Acids Res. 2015, 43, 4332–4341. [Google Scholar] [CrossRef]
- Cech, G.M.; Szalewska-Palasz, A.; Kubiak, K.; Malabirade, A.; Grange, W.; Arluison, V.; Wegrzyn, G. The Escherichia Coli Hfq Protein: An Unattended DNA-Transactions Regulator. Front. Mol. Biosci. 2016, 3, 36. [Google Scholar] [CrossRef]
- Malabirade, A.; Partouche, D.; El Hamoui, O.; Turbant, F.; Geinguenaud, F.; Recouvreux, P.; Bizien, T.; Busi, F.; Wien, F.; Arluison, V. Revised role for Hfq bacterial regulator on DNA topology. Sci. Rep. 2018, 8, 16792. [Google Scholar] [CrossRef]
- Malabirade, A.; Jiang, K.; Kubiak, K.; Diaz-Mendoza, A.; Liu, F.; van Kan, J.A.; Berret, J.F.; Arluison, V.; van der Maarel, J.R.C. Compaction and condensation of DNA mediated by the C-terminal domain of Hfq. Nucleic Acids Res. 2017, 45, 7299–7308. [Google Scholar] [CrossRef]
- Geinguenaud, F.; Calandrini, V.; Teixeira, J.; Mayer, C.; Liquier, J.; Lavelle, C.; Arluison, V. Conformational transition of DNA bound to Hfq probed by infrared spectroscopy. Phys. Chem. Chem. Phys. PCCP 2011, 13, 1222–1229. [Google Scholar] [CrossRef]
- Vogel, J.; Luisi, B.F. Hfq and its constellation of RNA. Nat. Rev. Microbiol. 2011, 9, 578–589. [Google Scholar] [CrossRef]
- Santiago-Frangos, A.; Woodson, S.A. Hfq chaperone brings speed dating to bacterial sRNA. Wiley Interdiscip. Rev. RNA 2018, 9, e1475. [Google Scholar] [CrossRef]
- Parekh, V.J.; Niccum, B.A.; Shah, R.; Rivera, M.A.; Novak, M.J.; Geinguenaud, F.; Wien, F.; Arluison, V.; Sinden, R.R. Role of Hfq in Genome Evolution: Instability of G-Quadruplex Sequences in E. coli. Microorganisms 2019, 8, 28. [Google Scholar] [CrossRef]
- Wilusz, C.J.; Wilusz, J. Lsm proteins and Hfq: Life at the 3’ end. RNA Biol. 2013, 10, 592–601. [Google Scholar] [CrossRef]
- Orans, J.; Kovach, A.R.; Hoff, K.E.; Horstmann, N.M.; Brennan, R.G. Crystal structure of an Escherichia coli Hfq Core (residues 2-69)-DNA complex reveals multifunctional nucleic acid binding sites. Nucleic Acids Res. 2020, 48, 3987–3997. [Google Scholar] [CrossRef]
- Arluison, V.; Folichon, M.; Marco, S.; Derreumaux, P.; Pellegrini, O.; Seguin, J.; Hajnsdorf, E.; Regnier, P. The C-terminal domain of Escherichia coli Hfq increases the stability of the hexamer. Eur. J. Biochem. 2004, 271, 1258–1265. [Google Scholar] [CrossRef]
- Fortas, E.; Piccirilli, F.; Malabirade, A.; Militello, V.; Trepout, S.; Marco, S.; Taghbalout, A.; Arluison, V. New insight into the structure and function of Hfq C-terminus. Biosci. Rep. 2015, 35, e00190. [Google Scholar] [CrossRef]
- Partouche, D.; Militello, V.; Gomez-Zavaglia, A.; Wien, F.; Sandt, C.; Arluison, V. In Situ Characterization of Hfq Bacterial Amyloid: A Fourier-Transform Infrared Spectroscopy Study. Pathogens 2019, 8, 36. [Google Scholar] [CrossRef]
- Santiago-Frangos, A.; Jeliazkov, J.R.; Gray, J.J.; Woodson, S.A. Acidic C-terminal domains autoregulate the RNA chaperone Hfq. eLife 2017, 6, e27049. [Google Scholar] [CrossRef]
- Santiago-Frangos, A.; Kavita, K.; Schu, D.J.; Gottesman, S.; Woodson, S.A. C-terminal domain of the RNA chaperone Hfq drives sRNA competition and release of target RNA. Proc. Natl. Acad. Sci. USA 2016, 113, E6089–E6096. [Google Scholar] [CrossRef]
- Blattner, F.R.; Plunkett, G., III; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The complete genome sequence of Escherichia coli K-12. Science 1997, 277, 1453–1462. [Google Scholar] [CrossRef]
- Hashem, V.I.; Rosche, W.A.; Sinden, R.R. Genetic assays for measuring rates of (CAG)·(CTG) repeat instability in Escherichia coli. Mutat. Res. 2002, 502, 25–37. [Google Scholar] [CrossRef]
- Trinh, T.Q.; Sinden, R.R. Preferential DNA secondary structure mutagenesis in the lagging strand of replication in E. coli. Nature 1991, 352, 544–547. [Google Scholar] [CrossRef]
- Hashem, V.I.; Sinden, R.R. Duplications between direct repeats stabilized by DNA secondary structure occur preferentially in the leading strand during DNA replication. Mutat. Res. 2005, 570, 215–226. [Google Scholar] [CrossRef]
- Sinden, R.R.; Zheng, G.X.; Brankamp, R.G.; Allen, K.N. On the deletion of inverted repeated DNA in Escherichia coli: Effects of length, thermal stability, and cruciform formation in vivo. Genetics 1991, 129, 991–1005. [Google Scholar]
- Edwards, S.F.; Hashem, V.I.; Klysik, E.A.; Sinden, R.R. Genetic instabilities of (CCTG)·(CAGG) and (ATTCT).(AGAAT) disease-associated repeats reveal multiple pathways for repeat deletion. Mol. Carcinog. 2009, 48, 336–349. [Google Scholar] [CrossRef]
- Luria, S.E.; Delbruck, M. Mutations of Bacteria from Virus Sensitivity to Virus Resistance. Genetics 1943, 28, 491–511. [Google Scholar]
- Cech, G.M.; Pakula, B.; Kamrowska, D.; Wegrzyn, G.; Arluison, V.; Szalewska-Palasz, A. Hfq protein deficiency in Escherichia coli affects ColE1-like but not lambda plasmid DNA replication. Plasmid 2014, 73, 10–15. [Google Scholar] [CrossRef]
- Zheng, Q. rSalvador: An R Package for the Fluctuation Experiment. G3 2017, 7, 3849–3856. [Google Scholar] [CrossRef]
- Zheng, Q. Comparing mutation rates under the Luria-Delbrück protocol. Genetica 2016, 144, 351–359. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- MacGregor-Fors, I.; Payton, M.E. Contrasting diversity values: Statistical inferences based on overlapping confidence intervals. PLoS ONE 2013, 8, e56794. [Google Scholar] [CrossRef]
- Malabirade, A.; Morgado-Brajones, J.; Trepout, S.; Wien, F.; Marquez, I.; Seguin, J.; Marco, S.; Velez, M.; Arluison, V. Membrane association of the bacterial riboregulator Hfq and functional perspectives. Sci. Rep. 2017, 7, 10724. [Google Scholar] [CrossRef]
- Réfrégiers, M.; Wien, F.; Ta, H.P.; Premvardhan, L.; Bac, S.; Jamme, F.; Rouam, V.; Lagarde, B.; Polack, F.; Giorgetta, J.L.; et al. DISCO synchrotron-radiation circular-dichroism endstation at SOLEIL. J. Synchrotron Radiat. 2012, 19, 831–835. [Google Scholar] [CrossRef]
- Wien, F.; Wallace, B.A. Calcium fluoride micro cells for synchrotron radiation circular dichroism spectroscopy. Appl. Spectrosc. 2005, 59, 1109–1113. [Google Scholar] [CrossRef]
- Lees, J.G.; Smith, B.R.; Wien, F.; Miles, A.J.; Wallace, B.A. CDtool-an integrated software package for circular dichroism spectroscopic data processing, analysis, and archiving. Anal. Biochem. 2004, 332, 285–289. [Google Scholar] [CrossRef]
- Del Villar-Guerra, R.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angew. Chem. Int. Ed. 2018, 57, 7171–7175. [Google Scholar] [CrossRef]
- Randazzo, A.; Spada, G.P.; da Silva, M.W. Circular dichroism of quadruplex structures. Top. Curr. Chem. 2013, 330, 67–86. [Google Scholar] [CrossRef]
- Holm, A.I.; Kohler, B.; Hoffmann, S.V.; Brøndsted Nielsen, S. Synchrotron radiation circular dichroism of various G-quadruplex structures. Biopolymers 2010, 93, 429–433. [Google Scholar] [CrossRef]
- Hwang, W.; Arluison, V.; Hohng, S. Dynamic competition of DsrA and rpoS fragments for the proximal binding site of Hfq as a means for efficient annealing. Nucleic Acids Res. 2011, 39, 5131–5139. [Google Scholar] [CrossRef]
- Bouloc, P.; Repoila, F. Fresh layers of RNA-mediated regulation in Gram-positive bacteria. Curr. Opin. Microbial. 2016, 30, 30–35. [Google Scholar] [CrossRef]
- Pham, T.M.; Tan, K.W.; Sakumura, Y.; Okumura, K.; Maki, H.; Akiyama, M.T. A single-molecule approach to DNA replication in Escherichia coli cells demonstrated that DNA polymerase III is a major determinant of fork speed. Mol. Microbiol. 2013, 90, 584–596. [Google Scholar] [CrossRef]
- Großmann, P.; Lück, A.; Kaleta, C. Model-based genome-wide determination of RNA chain elongation rates in Escherichia coli. Sci. Rep. 2017, 7, 17213. [Google Scholar] [CrossRef]
- Pomerantz, R.T.; O’Donnell, M. What happens when replication and transcription complexes collide? Cell Cycle 2010, 9, 2537–2543. [Google Scholar] [CrossRef]
- Pomerantz, R.T.; O’Donnell, M. The replisome uses mRNA as a primer after colliding with RNA polymerase. Nature 2008, 456, 762–766. [Google Scholar] [CrossRef]
- Tsui, H.C.; Feng, G.; Winkler, M.E. Negative regulation of mutS and mutH repair gene expression by the Hfq and RpoS global regulators of Escherichia coli K-12. J. Bacterial. 1997, 179, 7476–7487. [Google Scholar] [CrossRef]
- Tsui, H.C.; Feng, G.; Winkler, M.E. Transcription of the mutL repair, miaA tRNA modification, hfq pleiotropic regulator, and hflA region protease genes of Escherichia coli K-12 from clustered Esigma32-specific promoters during heat shock. J. Bacteriol. 1996, 178, 5719–5731. [Google Scholar] [CrossRef]
- Tsui, H.C.; Winkler, M.E. Transcriptional patterns of the mutL-miaA superoperon of Escherichia coli K-12 suggest a model for posttranscriptional regulation. Biochimie 1994, 76, 1168–1177. [Google Scholar] [CrossRef]
- Barreto, B.; Rogers, E.; Xia, J.; Frisch, R.L.; Richters, M.; Fitzgerald, D.M.; Rosenberg, S.M. The Small RNA GcvB Promotes Mutagenic Break Repair by Opposing the Membrane Stress Response. J. Bacteriol. 2016, 198, 3296–3308. [Google Scholar] [CrossRef]
- Bandekar, P.P.; Roopnarine, K.A.; Parekh, V.J.; Mitchell, T.R.; Novak, M.J.; Sinden, R.R. Antimicrobial activity of tryptanthrins in Escherichia coli. J. Med. Chem. 2010, 53, 3558–3565. [Google Scholar] [CrossRef]
- Kim, S.H.; Pytlos, M.J.; Sinden, R.R. Replication restart: A pathway for (CTG)·(CAG) repeat deletion in Escherichia coli. Mutat. Res. 2006, 595, 5–22. [Google Scholar] [CrossRef]
- Rosche, W.A.; Trinh, T.Q.; Sinden, R.R. Leading strand specific spontaneous mutation corrects a quasipalindrome by an intermolecular strand switch mechanism. J. Mol. Biol. 1997, 269, 176–187. [Google Scholar] [CrossRef]
- Trinh, T.Q.; Sinden, R.R. The influence of primary and secondary DNA structure in deletion and duplication between direct repeats in Escherichia coli. Genetics 1993, 134, 409–422. [Google Scholar]
- Chen, J.; Gottesman, S. Hfq links translation repression to stress-induced mutagenesis in E. coli. Genes Dev. 2017, 31, 1382–1395. [Google Scholar] [CrossRef]
- Modrich, P. Mechanisms and biological effects of mismatch repair. Annu. Rev. Genet. 1991, 25, 229–253. [Google Scholar] [CrossRef]
- Majdalani, N.; Cunning, C.; Sledjeski, D.; Elliott, T.; Gottesman, S. DsrA RNA regulates translation of RpoS message by an anti-antisense mechanism, independent of its action as an antisilencer of transcription. Proc. Natl. Acad. Sci. USA 1998, 95, 12462–12467. [Google Scholar] [CrossRef]
- Lease, R.A.; Belfort, M. Riboregulation by DsrA RNA: Trans-actions for global economy. Mol. Microbial. 2000, 38, 667–672. [Google Scholar] [CrossRef]
- Kalra, P.; Mishra, S.K.; Kaur, S.; Kumar, A.; Prasad, H.K.; Sharma, T.K.; Tyagi, J.S. G-Quadruplex-Forming DNA Aptamers Inhibit the DNA-Binding Function of HupB and Mycobacterium tuberculosis Entry into Host Cells. Mol. Ther. Nucleic Acids 2018, 13, 99–109. [Google Scholar] [CrossRef]
- El Hamoui, O.; Yadav, I.; Radiom, M.; Wien, F.; Berret, J.F.; van der Maarel, J.R.C.; Arluison, V. Interactions between DNA and the Hfq Amyloid-like Region Trigger a Viscoelastic Response. Biomacromolecules 2020. [Google Scholar] [CrossRef]
- Wien, F.; Martinez, D.; Le Brun, E.; Jones, N.C.; Vrønning Hoffmann, S.; Waeytens, J.; Berbon, M.; Habenstein, B.; Arluison, V. The Bacterial Amyloid-Like Hfq Promotes In Vitro DNA Alignment. Microorganisms 2019, 7, 639. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parekh, V.J.; Wien, F.; Grange, W.; De Long, T.A.; Arluison, V.; Sinden, R.R. Crucial Role of the C-Terminal Domain of Hfq Protein in Genomic Instability. Microorganisms 2020, 8, 1598. https://doi.org/10.3390/microorganisms8101598
Parekh VJ, Wien F, Grange W, De Long TA, Arluison V, Sinden RR. Crucial Role of the C-Terminal Domain of Hfq Protein in Genomic Instability. Microorganisms. 2020; 8(10):1598. https://doi.org/10.3390/microorganisms8101598
Chicago/Turabian StyleParekh, Virali J., Frank Wien, Wilfried Grange, Thomas A. De Long, Véronique Arluison, and Richard R. Sinden. 2020. "Crucial Role of the C-Terminal Domain of Hfq Protein in Genomic Instability" Microorganisms 8, no. 10: 1598. https://doi.org/10.3390/microorganisms8101598
APA StyleParekh, V. J., Wien, F., Grange, W., De Long, T. A., Arluison, V., & Sinden, R. R. (2020). Crucial Role of the C-Terminal Domain of Hfq Protein in Genomic Instability. Microorganisms, 8(10), 1598. https://doi.org/10.3390/microorganisms8101598





